Abstract
Colorectal cancer (CRC) is a leading cause of cancer-related deaths in both the USA and the world. Recent research has demonstrated the involvement of the gut microbiota in CRC development and progression. Microbial biomarkers of disease have focused primarily on the bacterial component of the microbiome; however, the viral portion of the microbiome, consisting of both bacteriophages and eukaryotic viruses, together known as the virome, has been lesser studied. Here we review the recent advancements in high-throughput sequencing (HTS) technologies and bioinformatics, which have enabled scientists to better understand how viruses might influence the development of colorectal cancer. We discuss the contemporary findings revealing modulations in the virome and their correlation with CRC development and progression. While a variety of challenges still face viral HTS detection in clinical specimens, we consider herein numerous next steps for future basic and clinical research. Clinicians need to move away from a single infectious agent model for disease etiology by grasping new, more encompassing etiological paradigms, in which communities of various microbial components interact with each other and the host. The reporting and indexing of patient health information, socioeconomic data, and other relevant metadata will enable identification of predictive variables and covariates of viral presence and CRC development. Altogether, the virome has a more profound role in carcinogenesis and cancer progression than once thought, and viruses, specific for either human cells or bacteria, are clinically relevant in understanding CRC pathology, patient prognosis, and treatment development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.

Cade Emlet

Mack Ruffin

Regina Lamendella
What Is the Human Virome?
Viruses are the most abundant and widely distributed biological entities on Earth, existing ubiquitously throughout the biosphere in equal or greater distribution than other microbes [1] and thus are influential in nearly every ecosystem [2]. The human body is no exception; within the vast microbial communities that thrive on and within the confines of our anatomy, viruses exist in great abundance. Viruses are capable of regulating bacterial community structure and, subsequently, human health to an extent that has not yet been fully appreciated [3]. It is the subject of recent investigative fervor due to advancements in viral community analyses.
The human virome is a diverse and abundant collection of all viruses found in or on humans and includes both eukaryotic and prokaryotic viruses, encompassing both animal-infecting viruses and bacteriophage. There are, of course, other types of viruses included within the human virome, like archaeal viruses and virophages, which are not as deeply studied due to a limited understanding of their function within the environment of the human body [4]. By either directly affecting host cell behavior and structure or by preying upon certain species of bacteria and subsequently altering bacterial communities, viruses can alter their own environment and hold strong direct or indirect influence over host health and physiology [5, 6]. While much research has focused on the role of the human bacteriome in the etiology and progression of diseases, such as cancer, cardiovascular disease, inflammatory bowel disease, among others, the human virome has been less studied. Recent work is just beginning to shed light on the role the virome may play in human diseases such as periodontitis [7], inflammatory bowel disease [8, 9], cystic fibrosis [10, 11], and cancer [12].
How Can the Virome Induce Carcinogenesis?
In 2012, 15.4% of new cancers in the world were attributable to carcinogenic infections. The following viruses accounted for over 60% of cancers associated with infections: human papillomavirus (640,000), hepatitis B virus (420,000), and Epstein–Barr virus (120,000), among others [13]. The unique aspect of these viruses is that infection does not always lead to cancer; most people infected with the aforementioned viruses do not develop further symptoms, let alone progression to cancer. However, the vast majority of cancerous cohorts have been infected with one of these viruses, or some combination thereof. Viral-linked cancers appear in the setting of persistent infection over years, sometimes decades [14]. This clearly suggests that other issues contribute to the carcinogenic process of the infection that leads to the development of cancer in some patients. The role of other understudied viruses may also be involved in this process.
These potentially oncogenic viruses utilize several different strategies to develop carcinogenic persistence. These strategies include creating conditions for replication (inducing cell cycle progression, altering metabolic reprogramming, inducing angiogenesis, etc.), ensuring correct replication (recruiting or inhibiting DNA damage repair), maximizing viral production (preventing apoptosis until virions mature, evading cellular immune systems), and producing multiple latent episomes or proviruses [15]. There are several different mechanisms by which other viruses may impact carcinogenesis, including genomic alterations, impacting cellular and inflammatory pathways, and shaping bacterial community structure. Changes in human virome composition and diversity have been implicated in periodontal disease [7], HIV [16], cystic fibrosis [10], diseases after antibiotic exposure [17], urinary tract infections [18], and inflammatory bowel disease [19]. In Table 1, we highlight eukaryotic viruses associated with oncogenesis in the gut. These viruses could be conceptualized as drivers in carcinogenesis while the other viral communities facilitate or hinder the chronic infection that leads to cancer. We cannot exclude the other viral communities having a direct impact on the carcinogenesis process.
One such class of viruses to which colorectal cancer can be attributed is the human papillomaviruses (HPV). HPV is a double-stranded, non-enveloped DNA virus, and it is the most common sexually transmitted infectious agent in the USA [20, 21]. HPV has been attributed to a number of cancers, including head and neck squamous cell carcinomas [22], oropharyngeal cancer [23], cervical cancer [24], prostate cancer [25], and colorectal cancer [26], among others. The global increasing incidence of young-onset colorectal cancer has been noted to include a prominent rise in rectal cancer when compared to colon cancer [27]. Persistent HPV high-risk infection is clearly linked to anal and rectal cancer [28].
The reason for the varied effects of HPV can be attributed to the variation in human papillomaviruses themselves, of which over 150 different types have been identified [29]. Only certain types of HPV, including HPV-16, HPV-18, and HPV-45, have been found to be statistically present in colorectal cancer tissue samples, with HPV-16 bearing the strongest prevalence among all HPV types found [30]. It is assumed that oncogenesis begins sometime after integration of the viral genome into the host genome, but how the subsequent induction of cancer occurs remains unknown and contested. The high-risk HPV genes E5, E6, and E7 encode potent oncoproteins that target almost all of the strategies mentioned previously which support replication and persistence [15].
Human polyomaviruses, a class of icosahedral, non-enveloped double-stranded DNA viruses, have also been attributed to cancer [31]. Out of the few human polyomaviruses suspected of having oncogenic properties (including Merkel cell polyomavirus, Trichodysplasia spinulosa polyomavirus, John Cunningham Polyomavirus, Simian Virus 40, and BK polyomavirus), only John Cunningham Polyomavirus (JCV) and BK polyomavirus have been associated with colorectal cancer [32,33,34,35]. Polyomaviruses encode the T-antigen, a non-structural oncogenic protein that is capable of inactivating tumor suppressor proteins p53 and pRB, among other mechanisms of signaling pathway interference [36]. Specific oncogenic mechanisms remain unknown for human polyomaviruses in colorectal cancer, but are implied by their presence in cancerous colorectal tissue samples [37].
Bacteriophage Shapes Community Structure and Can Indirectly Induce Oncogenesis
Although the effects of eukaryotic viral species have been well documented via functional techniques, the mechanisms by which viral communities induce carcinogenesis in the gut are still being discovered with the advent of next-generation sequencing methods (NGS), including metagenomics and metatranscriptomics techniques. Bacteriophages are indirectly associated with the development of some cancers in humans, particularly in the gastrointestinal system [38,39,40]. Due to the well-established relationship between microbial dysbiosis and the evolution of gastrointestinal malignancy, the relationship between viruses and colorectal cancer is strongly inferred and is currently being investigated [41,42,43,44].
Community-based viral shotgun NGS techniques have revealed that colon virome diversity is altered in individuals with colorectal cancer (CRC) [39], with viral diversity being higher in CRC cohorts [40]. The association between the bacteriophage portion of the enteric virome and CRC is considered to be indirect, as they alter bacterial community structures and bacterial behavior, which can lead to carcinogenesis. Importantly, when asking whether phages in the cancerous gut are primarily lytic or lysogenic, they are not exclusive in this distinction; the vast majority of phages in the gut are temperate, capable of being either lytic or lysogenic [39]. With this in mind, it is difficult to pin carcinogenesis on any specific phage or viral action, as the role of the phage community as a whole is to act as an overarching community modulator for bacteria. In this capacity, phages can serve to reduce some bacterial species, while acting consequently as the indirect impetus for population growth in other, oftentimes more pathogenic, bacterial species. Additionally, horizontal and vertical viral gene transmission is suspected of altering bacterial behavior, particularly with respect to biofilm formation, the alteration of which can also lead to carcinogenesis.
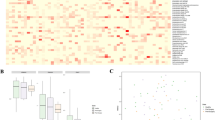
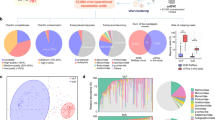
The ability to apply NGS to investigating trans-kingdom microbiomes is enabling a more holistic understanding of the pathogenesis of digestive diseases such as CRC, as tumor initiation and progression are influenced by complex host, environmental, and gut microbial factors. Several bacterial biomarkers have been correlated with CRC and progressional stages of cancer development [45,46,47,48]; however, contributions from viral components of the microbiome remain underexplored. NGS studies in CRC patients have highlighted how opportunistic and persistent viruses have been involved in the course of carcinogenesis [30, 49,50,51,52,53,54,55,56,57,58,59,60,61] and are summarized in Table 2. In addition, phages that infect Gram-negative bacterial hosts, such as enterotoxigenic Bacteroides fragilis and E. coli, as well as Fusobacterium nucleatum, have been associated with CRC development [62,63,64,65]. Bacteriophages also have been shown to have a putative functional role in the regulation of biofilm production (among other virulent functional genes encoded by bacteriophage), which has been implicated in colorectal tumorigenesis [66,67,68,69]. In addition to shaping the bacterial community, bacteriophages have been shown to transfer directly into colonic epithelial cells, promoting tumor growth and invasiveness in CRC [70, 71]. Although the full extent of the enteric virome’s indirect influence on carcinogenesis by means of modulating bacterial populations is in its infancy, a snapshot of how dynamic changes in the virome vary across stages of colorectal cancer is beginning to form (Fig. 1). Shotgun NGS techniques have begun to identify the presence of certain phages as a common factor for early-, mid-, and late-stage colorectal cancer, if not as a driving factor, then potentially as a biomarker (Table 2).
Bacteriophages alter bacterial communities and induce carcinogenesis through varying mechanisms. Panel A: The bacteriophage community serves as a community regulator for bacterial populations, and bacterial communities also have the ability to modulate bacteriophage populations. Dysbiosis in either community can allow carcinogenic bacterial populations to take hold, giving the virome a more indirect role in cancer development. Panel B: This is the hypothetical mechanism by which viruses indirectly induce carcinogenesis. Bacterial and viral communities begin in a healthy and well-regulated state. Variations in the phage community alter bacterial communities, reducing the populations of commensal bacteria while opening a niche for pathogenic (and potentially carcinogenic) bacteria to take home. The deleterious bacterial communities form a biofilm, which spreads due to the biofilm-altering components of some phages. The tight junction between cells is disrupted, allowing bacterial cells to infiltrate the spaces between them; this, in turn, leads to inflammation and creates the perfect environment for opportunistic pathogens, both viral and bacterial. Bacteria then thrive off of peptides secreted by the stressed epithelial cells, which leads them to release carcinogenic reactive oxygen species and polyspermines, subsequently inducing carcinogenesis. Figure 1 is obtained from Hannigan et al. [39]. Reuse is covered under the mBio Creative Commons Attribution 4.0 International license found here https://creativecommons.org/licenses/by/4.0/
Methods for Investigating the Human Virome
Culture-based techniques involve the isolation of human viruses and phages from human-associated environments. The basis for understanding phage ecology and host interactions has been provided by numerous conventional plaque-based assays [72], and these in vitro methods will remain essential in furthering our understanding of these relationships. These cultivation-based techniques are inherently limited by the inability to culture and identify hosts, as well as questions surrounding the ability to develop robust and representative in vitro models [73, 74]. In addition, cultivation-based approaches do not allow for unbiased representation of viral community structure and ecology. While 16S rRNA-based studies have helped revolutionize our understanding of bacterial ecology, absence of conserved molecular marker genes in viruses has complicated the application of marker-specific nucleic acid amplification technologies (NAATs) for viral community profiling [75,76,77,78,79,80]. Targeted sequence capture panels, such as ViroCap, have also been used to enrich nucleic acid from DNA and RNA viruses from viral communities harbored by vertebrate hosts [81].
Recent improvements in parallelized NGS technologies and bioinformatics have enabled for the first time a deeper and more comprehensive view of human-associated viral communities as compared to cultivation-based and PCR-based methodologies [82]. Since no single methodology can provide an all-inclusive approach, when designing a virome study, one should take into consideration sample source and type of viral particles to be assessed (i.e., DNA and/or RNA genomes, enveloped versus non-enveloped). Due to their small genome size, viral nucleic acids represent a minority of the total nucleic acids recovered from a given sample despite their greater abundance, and thus, isolation of viral DNA/RNA and sample concentration is recommended before shotgun sequencing. It should be noted that the methods used for viral particle purification can have a strong effect on the viral populations recovered and are subject to differential contamination issues [83, 84]. Methods for isolating viral-like particles (VLPs) from human-associated sample matrices often utilize a combination of filtration and centrifugation techniques for concentration of VLPs, followed by the elimination of contaminating cells and free nucleic acids [83, 85]. Viral nucleic acids can then be extracted using a variety of methods including those that utilize phenol–chloroform and Trizol, as well as a variety of commercially available kits, such as DNeasy (Qiagen), or QIAmp Ultrasens Virus kits (QIAGEN, Germantown, MD). Because of the low concentration of viral nucleic acids, prior to shotgun sequencing, an amplification step, such as multiple displacement amplification (MDA), is generally utilized to amplify viral genomes [83, 85, 86]. It should be noted that MDA introduces biases into viral community analysis due to preferential amplification of small circular viruses [87]. To analyze RNA viruses, RNA must first be reverse transcribed into more stable cDNA and then amplified by sequence-independent, single-primer amplification (SISPA) [88] or via amplification-based techniques [89]. Subsequently, library preparation can be performed using Illumina Nextera XT or FLX kits, which require very low nucleic acid inputs, followed by deep sequencing on high-output Illumina sequencers.
Shotgun sequencing approaches have provided unprecedented glimpses into the viral fraction of the human-associated ecosystems, revealing for the first time, in-depth inventories of the composition and functional repertoire of the virome [90,91,92,93]. Microscopic techniques like transmission electron microscopy (TEM) have not only benefited our understanding of viral structure, but they have also aided in the support of novel virus identification from shotgun-based sequencing studies [39, 40, 94, 95]. Perhaps the most comprehensive study characterizing viruses from a variety of sample types associated with the development of various cancers utilized a diverse selection enrichment method targeting all major viral groups followed by high-throughput sequencing [96]. This study investigated total DNA and RNA, mRNA, retroviral [97] and vertebrate viral capture [98], as well as enrichment of small circular DNA capture [99].
Bioinformatics Analysis of Viral Shotgun Data
Transforming shotgun sequencing data into usable outputs for clinicians and biologists requires robust bioinformatics analysis. There are more than 30 different pipelines available to analyze viral shotgun data as reviewed in detail [100]. Among this sea of tools, selecting the appropriate workflow can be challenging [101, 102]. Most workflows include preprocessing and filtering of nontarget sequences, assembling short reads, database searching for taxonomic assignment, and post-processing to detect any potential false positive results. Tool selection should be made based on the type of application, such as simple detection (genus/species) or more specific identification to the subspecies level, the latter of which is generally performed for source-tracking and surveillance purposes. For discovery-based detection projects, finding remote homologs can be accomplished by using alignment-based or composition search algorithms against reference databases that span a wide range of viral taxa; however, these approaches are very computationally intensive and time-consuming [100].
The viral metagenomics field has not yet adopted standard methods for classification of next-generation sequencing (NGS) results. Current tools include a few user-friendly online workflows, including VIROME [103] and Metavir [104]. Several more flexible command line programs are also available and generally require some basic background in linux/unix operating systems. Since methods and study goals vary widely, it is virtually impossible to assess the classification performance of these tools. Generally speaking, if high sensitivity is required, one should minimize the preprocessing steps on sequence data for retention of as many viral reads as possible. If higher specificity is required, such as in a clinical setting, high-specificity workflows such as RIEMS [105] and MetLab [106] can be employed. If the user’s application is more focused on higher precision, such as for variant calling, more aggressive preprocessing/quality filtration and assembly steps should be employed [101]. RINS [107] and Kraken [108] combined with MetLab [106] can perform preprocessing, filtering, and assembly, and each was determined to have high precision [100].
In the era of NGS technologies, viruses that are only represented by sequence data alone will likely need to be formally placed into the classification schemes maintained by the International Committee on the Taxonomy of Viruses [109]. Currently, there are no published guidelines on divergence cutoffs for viruses to be considered new viral taxa. When comparing genomic distances for eukaryotic viruses, there is high consistency between their genetic content and their current family- and genus-level taxonomic assignments, which suggests that eukaryotic viral metagenomic signatures could be classified consistently with the current ICTV taxonomy [110]. However, the taxonomy of prokaryotic viruses is far more divergent as compared with archeal and eukaryotic viruses [111]. Thus, one of the biggest unmet needs of viral metagenomics is the development of a consistent classification of viruses in which assignment at family and finer taxonomic levels is based on genomic relatedness and other evidence-based classification schemes.
Numerous challenges remain in analyzing viral shotgun data. First is the problem of sensitivity and false positive detections. For example, viruses may be missed due to biases associated with wet laboratory procedures, database limitations, and poor sequencing coverage. Contrastingly, viruses that are not present may in fact be detected because of homology to other viruses, incorrect annotation in databases, or sample cross-contamination. The intractability of viruses is also complicated by their high recombination rate and horizontal gene transfer and/or reassortment of genomic segments. Challenges for analyzing viral content in metagenomics datasets have been robustly discussed [75, 102, 112,113,114,115,116,117,118,119,120,121]. In order to make viral NGS data more useful in the clinical realm, we need to move beyond only taxonomic classification. It has been suggested that taxonomy should be associated with confidence scores, as well as potential pathogenicity/cancer associations [122]. Functional gene analysis would enable detection of mechanisms of pathogenicity/carcinogenesis; however, it certainly complicates bioinformatics analysis of shotgun data [123].
Important steps in the much needed standardization of viral metagenomics data [101, 102, 116] are vital to bring this technology into clinical studies, and this will be enabled by comparing and validating results within and between laboratories. In addition, metadata such as sample preparation methodology and sequencing technology should always be reported, as well as the establishment of true and false positive and negative results of synthetic constructs and experimental controls. Benchmarks should also be assessed for bioinformatics workflows, where different filtering, assembly, and annotation methods are tracked to optimize sensitivity and specificity [124,125,126,127,128]. Optimized parameterization will enable tractable and flexible frameworks that enable implementation of different algorithms so that users can utilize the best possible workflow tailored to their application. Updated reference databases, which include newly classified sequences, should be routinely used. While the field of viral metagenomics is young, there is much progress and momentum to advance the field by standardizing and validating methods including the CAMI challenge (http://cami-challenge.org/), OMICtools [129], and COMPARE (http://www.compare-europe.eu/).
Challenges and Future Directions
Given the early status of our understanding about the virome and colorectal cancer, there are numerous next steps for future research that could be considered. It may help to divide the potential research strategies into basic science and clinical science approaches. The basic scientific approach would take the current known carcinogenic mechanisms related to sporadic colorectal cancer pathways, hereditary pathways, and nonpolyposis pathways and look for roles that single viruses, such as HPV, may play. However, this may have limited utility given that other microbiota, both bacterial and viral, likely to play a role. As increasing amounts of microbiota are added into the conceptual model of carcinogenesis, complexity increases exponentially. Often the research team lacks expertise in analysis of large complex data sets which impairs the synthesis of the information. Adding experts in managing and analyzing large data sets would improve the synthesis in terms of quality and efficiency. With respect to the clinical aspects of reducing the impact of colorectal cancer suffering, the clinical or public health approach is divided into prevention, early detection, treatment, and surveillance for recurrence. If we knew the large bowel microbiota communities associated with risk of developing adenomatous cancer or which support the expression of a genetic mutation, then we begin looking into interventions to morph the microbial communities into ones associated with less risk. This would lead to a more preventive strategy. The same research could lead to developing an early detection test of large bowel microbiota (alone or in combination with stool and blood testing, or stool genetic markers). Both strategies require access to adults over the age of 40 or 50, an easily collected biological source containing microbiota, and easy/inexpensive assays to evaluate the communities of microbiota. There are plenty of adults undergoing some form of screening for colorectal cancer. There is also plenty of stool to be collected and assayed. This, however, is not always easy or acceptable to patients. As highlighted in this chapter, there are numerous assays that can be used to address the viral communities present. Finally, the reams of data that would be produced require partnership with bioinformaticians from project design through analysis.
Establishing a causative relationship between viruses and colorectal cancer requires a deep investigation the presence of etiologic agents in the diseased state and absence in non-diseased state. There are many challenges associated with fulfilling Koch’s postulates as many viruses cannot be cultured and due to the lack of available models to recapitulate viral pathology. Because of these challenges, alternative means for investigating causality of viruses involved in CRC have been proposed and include detection of viral nucleic acid and antigen in clinical specimens, in addition to visual detection of viral particles. Causation could further be improved with longitudinal sampling at different stages of CRC and demonstration of a host immune response throughout disease course [130].
While the application of NGS to virome profiling is exciting, future studies should incorporate proper quality control measures including negative, positive, and internal controls throughout sampling, extraction, and library preparation, and bioinformatics analysis to minimize false negative and positive detections. For example, several viruses that have been reported as etiological agents have later been shown to be contaminants [131,132,133,134]. In addition, much analytical and clinical validation must be performed for NGS methods to demonstrate test performance is comparable and/or improved compared to nucleic acid amplification techniques (NAATs) [135, 136]. As mandated by the Clinical Laboratory Improvement Amendments (CLIA), NGS viral detection tests will necessitate analytical sensitivity and specificity, reproducibility, and accuracy of both laboratory and bioinformatics processes. These validation procedures should include patient samples with known spike-in standards, as well as positive and negative controls within each testing run. Validation of bioinformatics methods should include in silico controls to test algorithms and databases and perform robust parameter sweeps for optimization of analyses.
NGS-based technologies hold much promise due to the ability to build predictive models that can describe risk of opportunistic and emerging viral infections associated with CRC, which could help personalize treatment options. Integrating the viral diversity and other microorganisms will yield new insights into CRC pathogenesis, by generating reliable diagnostic biomarkers that could inform disease status and treatment strategies. However, the variation in the virome within the gut microbiome of CRC patients is not exclusively explained by clinical factors alone but may also be related to lifestyle factors, such as diet [93, 137, 138]. Thus, careful integration of viral and other microbial components with patient metadata will be imperative in the discovery and generation of reliable CRC biomarkers. Ultra-deep sequencing approaches are also necessary to exhaustively measure viral diversity associated with CRC by circumventing viral enrichment procedures that often lead to biases in viral profiling. Artificial intelligence and machine learning models can then be applied to partition in which metadata information, viral, and other microbial components are most predictive of CRC onset and progression. Thus, as the cost for NGS becomes lower, deeper sequencing approaches will facilitate the integration of trans-kingdom profiling, longitudinally in large CRC cohorts to provide accurate and emerging mechanistic insights into how the virome is contributing to CRC etiology.
There are a variety of challenges that still face the field of utilizing NGS for viral detection in clinical specimens. First, sample collection is challenging, especially obtaining samples that are representative of the true gut ecosystem. Typically, samples are acquired at the time of endoscopy or surgery after the gut has been prepped and thus likely isn’t representative of the normal microenvironment but changes back to usual communities within 2 weeks [139]. In addition, sample preservation should be considered in order to stabilize DNA and RNA from the sample directly at the time of collection. In addition, as previously mentioned, unbiased and comprehensive preparation of viral DNA and RNA should be considered using ultra-high-throughput sequencing, to circumvent enrichment strategies which empart biases on the viral profile.
With respect to bioinformatics analyses, more funding support should be granted for disseminating developed, documented, and tested software distributed through package managers or virtual machines to improve the lifespan of viral informatics software. In addition, attention must be given to the maintenance of updated and well-indexed, centralized sequence databases, with corresponding patient/sample metadata. This will facilitate the discovery of unclassified viral dark matter across several studies. As new software becomes available, benchmarking of software using standardized test datasets will help scientists analyze and optimize viral data analysis across multiple studies. The reporting and indexing of patient health information, socioeconomic data, and other relevant metadata will enable identification of predictive variables and covariates of viral presence and cancer development. In addition, direct communication among software developers and clinicians will help generate analyses that are interpretable and useful for clinicians interested in investigating viral communities and their role in CRC development and progression.
Implications for Clinicians
-
There will be an explosion of information pertaining to the virome and its implications in colorectal cancer over the next 5 years due to commercial advances in high-throughput sequencing.
-
Clinicians need to move away from a single infectious agent model for disease etiology by grasping this new, more encompassing etiological paradigm, in which communities of various microbial components interact with each other and the host.
-
As our understanding of the role of the virome in enteric carcinogenesis expands, new preventive methods, screening/surveillance techniques, or treatment modalities may develop.
-
It may be easy for some people to erroneously associate common infections with cancer. As the concept of viral contributions to carcinogenesis reaches the public, clinicians need to help patients remain calm when they encounter a viral infection.
-
Certain eukaryotic viruses are the driver of gastrointestinal cancers, but most people infected with these viruses do not develop cancer. Thus, it should be noted that infection by these potentially carcinogenic viruses does not necessarily imply future carcinogenesis; rather, their presence should be associated more so with elevated risk.
References
Pan D, Nolan J, Williams KH, et al. Abundance and distribution of microbial cells and viruses in an alluvial aquifer. Front Microbiol. 2017;8:1199.
Rohwer F, Prangishvili D, Lindell D. Roles of viruses in the environment. Environ Microbiol. 2009;11:2771–2774.
Navarro F, Muniesa M. Phages in the Human Body. Front Microbiol. 2017;8:566.
Bekliz M, Colson P, La Scola B. The expanding family of virophages. Viruses. 2016;8:317.
Caputo M, Zoch-Lesniak B, Karch A, et al. Bacterial community structure and effects of picornavirus infection on the anterior nares microbiome in early childhood. BMC Microbiol. 2019;19:1. https://doi.org/10.1186/s12866-018-1372-8.
Yinda CK, Vanhulle E, Conceição-Neto N, et al. Gut virome analysis of cameroonians reveals high diversity of enteric viruses, including potential interspecies transmitted viruses. mSphere. 2019;4:e00585-18. https://doi.org/10.1128/mSphere.00585-18.
Ly M, Abeles SR, Boehm TK, et al. Altered oral viral ecology in association with periodontal disease. mBio. 2014;5:1133. https://doi.org/10.1128/mbio.01133-14.
Ungaro F, Massimino L, Furfaro F, et al. Metagenomic analysis of intestinal mucosa revealed a specific eukaryotic gut virome signature in early-diagnosed inflammatory bowel disease. Gut Microbes. 2019;10:149–158. https://doi.org/10.1080/19490976.2018.1511664.
Fernandes MA, Verstraete SG, Phan TG, et al. Enteric virome and bacterial microbiota in children with ulcerative colitis and crohn disease. J Pediatr Gastroenterol Nutr. 2019;68:30–36. https://doi.org/10.1097/MPG.0000000000002140.
Willner D, Furlan M, Haynes M, et al. Metagenomic analysis of respiratory tract DNA viral communities in cystic fibrosis and non-cystic fibrosis individuals. PLoS ONE. 2009;4:e7370. https://doi.org/10.1371/journal.pone.0007370.
Moran Losada P, Chouvarine P, Dorda M, et al. The cystic fibrosis lower airways microbial metagenome. ERJ Open Res. 2016;2:00096–02015. https://doi.org/10.1183/23120541.00096-2015.
Santiago-Rodriguez TM, Hollister EB. Human virome and disease: high-throughput sequencing for virus discovery, identification of phage-bacteria dysbiosis and development of therapeutic approaches with emphasis on the human gut. Viruses. 2019;11:E656.
Plummer M, de Martel C, Vignat J, et al. Global burden of cancers attributable to infections in 2012: a synthetic analysis. Lancet Glob Health. 2016;4:e609–e616.
Mesri EA, Feitelson MA, Munger K. Human viral oncogenesis: a cancer hallmarks analysis. Cell Host Microbe. 2014;15:266–282.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–674.
Monaco CL, Gootenberg DB, Zhao G, et al. Altered virome and bacterial microbiome in human immunodeficiency Virus-Associated Acquired Immunodeficiency Syndrome. Cell Host Microbe. 2016;19:311–322. https://doi.org/10.1016/j.chom.2016.02.011.
Abeles SR, Ly M, Santiago-Rodriguez TM, et al. Effects of long term antibiotic therapy on human oral and fecal viromes. PLoS ONE. 2015;10:e0134941.
Santiago-Rodriguez TM, Ly M, Bonilla N, et al. The human urine virome in association with urinary tract infections. Front Microbiol. 2015;6:14.
Norman JM, Handley SA, Baldridge MT, et al. Disease-specific alterations in the enteric virome in inflammatory bowel disease. Cell. 2015;160:447–460. https://doi.org/10.1016/j.cell.2015.01.002.
Bzhalava D, Guan P, Franceschi S, et al. A systematic review of the prevalence of mucosal and cutaneous human papillomavirus types. Virology. 2013;445:224–231.
Braaten KP, Laufer MR. Human papillomavirus (HPV), HPV-related disease, and the HPV vaccine. Rev Obstet Gynecol. 2008;1:2.
Kreimer AR, Clifford GM, Boyle P, et al. Human papillomavirus types in head and neck squamous cell carcinomas worldwide: a systematic review. Cancer Epidemiol Biomark Prev. 2005;14:467–475.
D’Souza G, Kreimer AR, Viscidi R, et al. Case-control study of human papillomavirus and oropharyngeal cancer. N Engl J Med. 2007;356:1944–1956.
Muñoz N, Bosch X, de Sanjosé S, et al. Epidemiologic classification of human papillomavirus types associated with cervical cancer. N Engl J Med. 2003;348:518–527.
Yin B, Liu W, Yu P, et al. Association between human papillomavirus and prostate cancer: a meta-analysis. Oncol Lett. 2017;14:1855–1865. https://doi.org/10.3892/ol.2017.6367.
Damin DC, Ziegelmann PK, Damin AP. Human papillomavirus infection and colorectal cancer risk: a meta-analysis. Colorectal Dis. 2013;15:e420–e428.
Araldi RP, Sant’Ana TA, Módolo DG, et al. The human papillomavirus (HPV)-related cancer biology: an overview. Biomed Pharmacother. 2018;106:1537–1556.
Lui RN, Tsoi KKF, Ho JMW, et al. Global increasing incidence of young-onset colorectal cancer across 5 continents: a joinpoint regression analysis of 1,922,167 cases. Cancer Epidemiol Biomark Prev. 2019;28:1275–1282.
Doorbar J, Egawa N, Griffin H, et al. Human papillomavirus molecular biology and disease association. Rev Med Virol.. 2015;25:2–23.
Bodaghi S. Colorectal papillomavirus infection in patients with colorectal cancer. Clin Cancer Res. 2005;11:2862–2867.
De Gascun CF, Carr MJ. Human polyomavirus reactivation: disease pathogenesis and treatment approaches. Clin Dev Immunol. 2013;2013:373579.
Prado JCM, Monezi TA, Amorim AT, et al. Human polyomaviruses and cancer: an overview. Clinics (Sao Paulo). 2018;73:e558s. https://doi.org/10.6061/clinics/2018/e558s.
Dalianis T, Hirsch HH. Human polyomaviruses in disease and cancer. Virology. 2013;437:63–72.
Vilchez RA, Butel JS. Emergent human pathogen simian virus 40 and its role in cancer. Clin Microbiol Rev. 2004;17:495–508.
Khabaz MN, Nedjadi T, Gari MA, et al. Simian virus 40 is not likely involved in the development of colorectal adenocarcinoma. Future Virol. 2016;11:175–180.
Rollison DE. JC virus infection. J Clin Gastroenterol. 2010;. https://doi.org/10.1097/mcg.0b013e3181e0084b.
Khabaz MN, Nedjadi T, Garri MA, et al. BK polyomavirus association with colorectal cancer development. Genet Mol Res. 2016. https://doi.org/10.4238/gmr.15027841.
Cohen LJ. Phages trump bacteria in immune interactions. Sci Transl Med. 2019;11:eaaw5331.
Hannigan GD, Duhaime MB, Ruffin MT 4th, et al. Diagnostic potential and interactive dynamics of the colorectal cancer virome. mBio. 2018;9:e02248-18.
Nakatsu G, Zhou H, Wu WKK, et al. Alterations in enteric virome are associated with colorectal cancer and survival outcomes. Gastroenterology. 2018;155:e5.
Sobhani I, Tap J, Roudot-Thoraval F, et al. Microbial dysbiosis in colorectal cancer (CRC) patients. PLoS ONE. 2011;6:e16393. https://doi.org/10.1371/journal.pone.0016393.
Tilg H, Adolph TE, Gerner RR, et al. The intestinal microbiota in colorectal cancer. Cancer Cell. 2018;33:954–964.
Schieffer KM, Wright JR, Harris LR, et al. NOD2 genetic variants predispose one of two familial adenomatous polyposis siblings to pouchitis through microbiome dysbiosis. J Crohns Colitis. 2017;11:1393–1397. https://doi.org/10.1093/ecco-jcc/jjx083.
Gogokhia L, Buhrke K, Bell R, et al. Expansion of bacteriophages is linked to aggravated intestinal inflammation and colitis. Cell Host Microbe. 2019;25:e8.
Sze MA, Baxter NT, Ruffin MT 4th, et al. Normalization of the microbiota in patients after treatment for colonic lesions. Microbiome. 2017;5:150.
Baxter NT, Ruffin MT 4th, Rogers MAM, et al. Microbiota-based model improves the sensitivity of fecal immunochemical test for detecting colonic lesions. Genome Med. 2016;8:37.
Baxter NT, Koumpouras CC, Rogers MAM, et al. DNA from fecal immunochemical test can replace stool for detection of colonic lesions using a microbiota-based model. Microbiome. 2016;4:59.
Zackular JP, Rogers MAM, Ruffin MT 4th, et al. The human gut microbiome as a screening tool for colorectal cancer. Cancer Prev Res. 2014;7:1112–1121.
Geuking MB, Weber J, Dewannieux M, et al. Recombination of retrotransposon and exogenous RNA virus results in nonretroviral cDNA integration. Science. 2009;323:393–396.
Horie M, Honda T, Suzuki Y, et al. Endogenous non-retroviral RNA virus elements in mammalian genomes. Nature. 2010;463:84–87.
Klenerman P, Hengartner H, Zinkernagel RM. A non-retroviral RNA virus persists in DNA form. Nature. 1997;390:298–301.
Zhdanov VM. Integration of viral genomes. Nature. 1975;256:471–473.
Enam S, del Valle L, Lara C, et al. Association of human polyomavirus JCV with colon cancer: evidence for interaction of viral T-antigen and beta-catenin. Cancer Res. 2002;62:7093–7101.
Goel A, Li MS, Nagasaka T, et al. Association of JC virus T-antigen expression with the methylator phenotype in sporadic colorectal cancers. Gastroenterology. 2006;130:1950–1961.
Hori R, Murai Y, Tsuneyama K, et al. Detection of JC virus DNA sequences in colorectal cancers in Japan. Virchows Arch. 2005;447:723–730.
Karpinski P, Myszka A, Ramsey D, et al. Detection of viral DNA sequences in sporadic colorectal cancers in relation to CpG island methylation and methylator phenotype. Tumour Biol.. 2011;32:653–659.
Mou X, Chen L, Liu F, et al. Prevalence of JC virus in Chinese patients with colorectal cancer. PLoS ONE. 2012;7:e35900. https://doi.org/10.1371/journal.pone.0035900.
Ouaïssi M, Studer AS, Mege D, et al. Characteristics and natural history of patients with colorectal cancer complicated by infectious endocarditis. Case control study of 25 patients. Anticancer Res. 2014;34:349–353.
Theodoropoulos G, Panoussopoulos D, Papaconstantinou I, et al. Assessment of JC polyoma virus in colon neoplasms. Dis Colon Rectum. 2005;48:86–91.
Vilkin A, Ronen Z, Levi Z, et al. Presence of JC virus DNA in the tumor tissue and normal mucosa of patients with sporadic colorectal cancer (CRC) or with positive family history and Bethesda criteria. Dig Dis Sci. 2012;57:79–84. https://doi.org/10.1007/s10620-011-1855-z.
zur Hausen H. Red meat consumption and cancer: reasons to suspect involvement of bovine infectious factors in colorectal cancer. Int J Cancer. 2012;130:2475–2483.
Boleij A, Hechenbleikner EM, Goodwin AC, et al. The Bacteroides fragilis toxin gene is prevalent in the colon mucosa of colorectal cancer patients. Clin Infect Dis. 2015;60:208–215. https://doi.org/10.1093/cid/ciu787.
Castellarin M, Warren RL, Freeman JD, et al. Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res. 2012;22:299–306. https://doi.org/10.1101/gr.126516.111.
Cuevas-Ramos G, Petit CR, Marcq I, et al. Escherichia coli induces DNA damage in vivo and triggers genomic instability in mammalian cells. Proc Natl Acad Sci USA. 2010;107:11537–11542. https://doi.org/10.1073/pnas.1001261107.
Niu YD, McAllister TA, Nash JHE, et al. Four Escherichia coli O157:H7 phages: a new bacteriophage genus and taxonomic classification of T1-like phages. PLoS ONE. 2014;9:e100426.
Dejea CM, Wick EC, Hechenbleikner EM, et al. Microbiota organization is a distinct feature of proximal colorectal cancers. Proc Natl Acad Sci USA. 2014;111:18321–18326. https://doi.org/10.1073/pnas.1406199111.
Johnson CH, Dejea CM, Edler D, et al. Metabolism links bacterial biofilms and colon carcinogenesis. Cell Metab. 2015;21:891–897. https://doi.org/10.1016/j.cmet.2015.04.011.
Secor PR, Sweere JM, Michaels LA, et al. Filamentous bacteriophage promote biofilm assembly and function. Cell Host Microbe. 2015;18:549–559. https://doi.org/10.1016/j.chom.2015.10.013.
Waldor MK, Mekalanos JJ. Lysogenic conversion by a filamentous phage encoding cholera toxin. Science. 1996;272:1910–1914.
Nguyen S, Baker K, Padman BS, et al. Bacteriophage transcytosis provides a mechanism to cross epithelial cell layers. mBio. 2017;8:e01874-17.
Lehti TA, Pajunen MI, Skog MS, et al. Internalization of a polysialic acid-binding Escherichia coli bacteriophage into eukaryotic neuroblastoma cells. Nat Commun. 2017;8:1915.
Ogilvie LA, Jones BV. The human gut virome: a multifaceted majority. Front Microbiol. 2015;6:918.
Amann RI, Ludwig W, Schleifer KH. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev. 1995;59:143–169.
Rappé MS, Giovannoni SJ. The uncultured microbial majority. Annu Rev Microbiol. 2003;57:369–394.
Edwards RA, Rohwer F. Viral metagenomics. Nat Rev Microbiol. 2005;3:504–510.
García-Arroyo L, Prim N, Martí N, et al. Benefits and drawbacks of molecular techniques for diagnosis of viral respiratory infections. Experience with two multiplex PCR assays. J Med Virol. 2016;88:45–50.
Reijans M, Dingemans G, Klaassen CH, et al. RespiFinder: a new multiparameter test to differentially identify fifteen respiratory viruses. J Clin Microbiol. 2008;46:1232–1240. https://doi.org/10.1128/JCM.02294-07.
Afshar RM, Mollaie HR. Detection of HBV resistance to lamivudine in patients with chronic hepatitis B using Zip nucleic acid probes in Kerman, southeast of Iran. Asian Pac J Cancer Prev. 2012;13:3657–3661.
Mercier-Delarue S, Vray M, Plantier JC, et al. Higher specificity of nucleic acid sequence-based amplification isothermal technology than of real-time PCR for quantification of HIV-1 RNA on dried blood spots. J Clin Microbiol. 2014;52:52–56. https://doi.org/10.1128/JCM.01848-13.
Wu D, Liu F, Liu H, et al. Detection of serum HCV RNA in patients with chronic hepatitis C by transcription mediated amplification and real-time reverse transcription polymerase chain reaction. Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2014;39:664–672.
Wylie TN, Wylie KM, Herter BN, et al. Enhanced virome sequencing using targeted sequence capture. Genome Res. 2015;25:1910–1920.
Moustafa A, Xie C, Kirkness E, et al. The blood DNA virome in 8,000 humans. PLoS Pathog. 2017;13:e1006292. https://doi.org/10.1371/journal.ppat.1006292.
Kleiner M, Hooper LV, Duerkop BA. Evaluation of methods to purify virus-like particles for metagenomic sequencing of intestinal viromes. BMC Genom. 2015;16:7.
Qin J, Li R, Raes J, et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. 2010;464:59–65.
Thurber RV, Haynes M, Breitbart M, et al. Laboratory procedures to generate viral metagenomes. Nat Protoc. 2009;4:470–483.
Reyes A, Semenkovich NP, Whiteson K, et al. Going viral: next-generation sequencing applied to phage populations in the human gut. Nat Rev Microbiol. 2012;10:607–617.
Kim K-H, Chang H-W, Nam Y-D, et al. Amplification of uncultured single-stranded DNA viruses from rice paddy soil. Appl Environ Microbiol. 2008;74:5975–5985.
Wang D, Urisman A, Liu YT, et al. Viral discovery and sequence recovery using DNA microarrays. PLoS Biol. 2003;1:E2. https://doi.org/10.1371/journal.pbio.0000002.
Abbas AA, Diamond JM, Chehoud C, et al. The perioperative lung transplant virome: torque teno viruses are elevated in donor lungs and show divergent dynamics in primary graft dysfunction. Am J Transplant. 2017;17:1313–1324. https://doi.org/10.1111/ajt.14076.
Breitbart M, Hewson I, Felts B, et al. Metagenomic analyses of an uncultured viral community from human feces. J Bacteriol. 2003;185:6220–6223. https://doi.org/10.1128/jb.185.20.6220-6223.2003.
Reyes A, Haynes M, Hanson N, et al. Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature. 2010;466:334–338. https://doi.org/10.1038/nature09199.
Kim M-S, Park E-J, Roh SW, et al. Diversity and abundance of single-stranded DNA viruses in human feces. Appl Environ Microbiol. 2011;77:8062–8070.
Minot S, Sinha R, Chen J, et al. The human gut virome: inter-individual variation and dynamic response to diet. Genome Res. 2011;21:1616–1625. https://doi.org/10.1101/gr.122705.111.
Handley SA, Devkota S. Going viral: a novel role for bacteriophage in colorectal cancer. mBio. 2019;10:e02626-18.
Zheng T, Li J, Ni Y, et al. Mining, analyzing, and integrating viral signals from metagenomic data. Microbiome. 2019;7:42. https://doi.org/10.1186/s40168-019-0657-y.
Mollerup S, Asplund M, Friis-Nielsen J, et al. High-throughput sequencing-based investigation of viruses in human cancers by multienrichment approach. J Infect Dis. 2019;220:1312–1324. https://doi.org/10.1093/infdis/jiz318.
Vinner L, Mourier T, Friis-Nielsen J, et al. Investigation of human cancers for retrovirus by low-stringency target enrichment and high-throughput sequencing. Sci Rep. 2015;5:13201. https://doi.org/10.1038/srep13201.
Mühlemann B, Jones TC, de Barros Damgaard P, et al. Ancient hepatitis B viruses from the Bronze Age to the Medieval period. Nature. 2018;557:418–423.
Hansen TA, Fridholm H, Frøslev TG, et al. New type of papillomavirus and novel circular single stranded DNA virus discovered in urban Rattus norvegicus using circular DNA enrichment and metagenomics. PLoS ONE. 2015;10:e0141952. https://doi.org/10.1371/journal.pone.0141952.
Nooij S, Schmitz D, Vennema H, et al. Overview of virus metagenomic classification methods and their biological applications. Front. Microbiol. 2018;9:749.
Posada-Cespedes S, Seifert D, Beerenwinkel N. Recent advances in inferring viral diversity from high-throughput sequencing data. Virus Res. 2017;239:17–32.
Rose R, Constantinides B, Tapinos A, et al. Challenges in the analysis of viral metagenomes. Virus Evol. 2016;2:vew022.
Wommack KE, Bhavsar J, Polson SW, et al. VIROME: a standard operating procedure for analysis of viral metagenome sequences. Stand Genom Sci. 2012;6:427–439. https://doi.org/10.4056/sigs.2945050.
Roux S, Tournayre J, Mahul A, et al. Metavir 2: new tools for viral metagenome comparison and assembled virome analysis. BMC Bioinform. 2014;15:76.
Scheuch M, Höper D, Beer M. RIEMS: a software pipeline for sensitive and comprehensive taxonomic classification of reads from metagenomics datasets. BMC Bioinform. 2015;16:69.
Norling M, Karlsson-Lindsjö OE, Gourlé H, et al. MetLab: an in silico experimental design, simulation and analysis tool for viral metagenomics studies. PLoS ONE. 2016;11:e0160334.
Bhaduri A, Qu K, Lee CS, et al. Rapid identification of non-human sequences in high-throughput sequencing datasets. Bioinformatics. 2012;28:1174–1175.
Wood DE, Salzberg SL. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 2014;15:R46.
Lefkowitz EJ, Dempsey DM, Hendrickson RC, et al. Virus taxonomy: the database of the International Committee on Taxonomy of Viruses (ICTV). Nucl Acids Res. 2018;46:D708–D717. https://doi.org/10.1093/nar/gkx932.
Aiewsakun P, Adriaenssens EM, Lavigne R, et al. Evaluation of the genomic diversity of viruses infecting bacteria, archaea and eukaryotes using a common bioinformatic platform: steps towards a unified taxonomy. J Gen Virol. 2018;99:1331–1343.
Aiewsakun P, Simmonds P. The genomic underpinnings of eukaryotic virus taxonomy: creating a sequence-based framework for family-level virus classification. Microbiome. 2018;6:38.
Wommack KE, Bhavsar J, Ravel J. Metagenomics: read length matters. Appl Environ Microbiol. 2008;74:1453–1463.
Wooley JC, Ye Y. Metagenomics: facts and artifacts, and computational challenges*. J Comput Sci Technol. 2009;25:71–81.
Tang P, Chiu C. Metagenomics for the discovery of novel human viruses. Future Microbiol. 2010;5:177–189.
Wooley JC, Godzik A, Friedberg I. A primer on metagenomics. PLoS Comput Biol. 2010;6:e1000667.
Fancello L, Raoult D, Desnues C. Computational tools for viral metagenomics and their application in clinical research. Virology. 2012;434:162–174.
Thomas T, Gilbert J, Meyer F. Metagenomics—A guide from sampling to data analysis. Microb Inform Exp. 2012;2:3.
Pallen MJ. Diagnostic metagenomics: potential applications to bacterial, viral and parasitic infections. Parasitology. 2014;141:1856–1862.
Hall RJ, Draper JL, Nielsen FGG, et al. Beyond research: a primer for considerations on using viral metagenomics in the field and clinic. Front Microbiol. 2015;6:224.
McIntyre ABR, Ounit R, Afshinnekoo E, et al. Comprehensive benchmarking and ensemble approaches for metagenomic classifiers. Genome Biol. 2017;18:182.
Nieuwenhuijse DF, Koopmans MPG. Metagenomic sequencing for surveillance of food- and waterborne viral diseases. Front Microbiol. 2017;8:230.
Randle-Boggis RJ, Helgason T, Sapp M, et al. Evaluating techniques for metagenome annotation using simulated sequence data. FEMS Microbiol Ecol. 2016;92:fiw095.
Lindgreen S, Adair KL, Gardner PP. An evaluation of the accuracy and speed of metagenome analysis tools. Sci Rep. 2016;6:19233.
Treangen TJ, Koren S, Sommer DD, et al. MetAMOS: a modular and open source metagenomic assembly and analysis pipeline. Genome Biol. 2013;14:R2.
Scholz M, Lo C-C, Chain PSG. Improved assemblies using a source-agnostic pipeline for MetaGenomic Assembly by Merging (MeGAMerge) of contigs. Sci Rep. 2015;4:6480.
Smits SL, Bodewes R, Ruiz-Gonzalez A, et al. Assembly of viral genomes from metagenomes. Front Microbiol. 2014;5:714. https://doi.org/10.3389/fmicb.2014.00714.
Vázquez-Castellanos JF, García-López R, Pérez-Brocal V, et al. Comparison of different assembly and annotation tools on analysis of simulated viral metagenomic communities in the gut. BMC Genom. 2014;15:37.
Deng X, Naccache SN, Ng T, et al. An ensemble strategy that significantly improves de novo assembly of microbial genomes from metagenomic next-generation sequencing data. Nucleic Acids Res. 2015;43:e46. https://doi.org/10.1093/nar/gkv002.
Henry VJ, Bandrowski AE, Pepin A-S, Gonzalez BJ, Desfeux A. OMICtools: an informative directory for multi-omic data analysis. Database. 2014.
Fredericks DN, Relman DA. Sequence-based identification of microbial pathogens: a reconsideration of Koch’s postulates. Clin Microbiol Rev. 1996;9:18–33.
Arias M, Fan H. The saga of XMRV: a virus that infects human cells but is not a human virus. Emerg Microbes Infect. 2014;3:1–6.
Alter HJ, Mikovits JA, Switzer WM, et al. A multicenter blinded analysis indicates no association between chronic fatigue syndrome/myalgic encephalomyelitis and either xenotropic murine leukemia virus-related virus or polytropic murine leukemia virus. mBio. 2012;3:e00266-12. https://doi.org/10.1128/mbio.00266-12.
Erlwein O, et al. DNA extraction columns contaminated with murine sequences. PLoS ONE. 2011;6:e23484.
Naccache SN, Greninger AL, Lee D, et al. The perils of pathogen discovery: origin of a novel parvovirus-like hybrid genome traced to nucleic acid extraction spin columns. J Virol. 2013;87:11966–11977. https://doi.org/10.1128/JVI.02323-13.
Lewandowska DW, Zagordi O, Abinden A, et al. Unbiased metagenomic sequencing complements specific routine diagnostic methods and increases chances to detect rare viral strains. Diagn Microbiol Infect Dis. 2015;83:133–138.
Schlaberg R, Chiu CY, Miller S, et al. Validation of metagenomic next-generation sequencing tests for universal pathogen detection. Arch Pathol Lab Med. 2017;141:776–786.
Kim M-S, Bae J-W. Spatial disturbances in altered mucosal and luminal gut viromes of diet-induced obese mice. Environ Microbiol. 2016;18:1498–1510.
Minot S, Bryson A, Chehoud C, et al. Rapid evolution of the human gut virome. Proc Natl Acad Sci USA. 2013;110:12450–12455. https://doi.org/10.1073/pnas.1300833110.
Nagata N, Tohya M, Fukuda S, et al. Effects of bowel preparation on the human gut microbiome and metabolome. Sci Rep. 2019;9:4042. https://doi.org/10.1038/s41598-019-40182-9.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Regina Lamendella receives funding from the National Science Foundation (NSF1805549) and the National Institute of Health - NIA (1R15AG052933-01). She is also the CEO of Wright Labs, LLC and the VP of research and development for Contamination Source Identification.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Emlet, C., Ruffin, M. & Lamendella, R. Enteric Virome and Carcinogenesis in the Gut. Dig Dis Sci 65, 852–864 (2020). https://doi.org/10.1007/s10620-020-06126-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-020-06126-4