Abstract
Background and Aim
Hepatitis B virus core promoter (CP) mutations can increase risk of hepatocellular carcinoma. The CP region overlaps with the HBV X (HBx) gene, which has been associated with hepatocarcinogenesis. The cyclin kinase inhibitor P53 is an important regulator of cell cycle progression. We determined whether HBx mutants that result from mutations in the CP deregulate P53.
Methods
A HBx combination (combo) mutant with changes in the CP region that corresponded to A1762T/G1764A (TA), T1753A, and T1768A was constructed and expressed in L-02 and Hep3B cells. The effects of CP mutations on expression and degradation of P53, and the effects on cell cycle progression and proliferation were analyzed.
Results
The combo mutant decreased levels of P53 and increased cyclin D1 expression, accelerated P53 degradation in L-02 cells, accelerated cell cycle progression, and increased expression of S-phase kinase-associated protein 2 (Skp2) in L-02 and Hep3B cells. Silencing of Skp2 abrogated the effects of CP mutations on P53 expression. The kinetics of P53 expression correlated with changes in cell cycle distribution.
Conclusions
The HBx mutant with a combination of CP mutations can up-regulate Skp2, which then down-regulates P53 via ubiquitin-mediated proteasomal degradation, increasing the risk of hepatocellular carcinoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is one among the most prevalent and lethal cancers in humans [1] and is the fifth leading cause of cancer globally. Chronic hepatitis B virus (HBV) infection is a significant risk factor for HCC in the developing world, where there are more than 400 million viral carriers [2, 3]. The biology, transmission mode, and epidemiology of HBV continue to be actively investigated and have been recently reviewed. People with chronic HBV infection have approximately a 100-fold increased risk of developing HCC compared with those without chronic HBV infection [4]. The HBV genotype, core promoter (CP) mutations, and high viral load are viral factors associated with an increased risk of HCC [5–7].
The prevalence of the common double mutation A1762T/G1764A (TA) is associated with more aggressive progression of liver disease and development of HCC [8–10]. The truncated pre-S2/S and X HBV genes have been found in hepatoma tissue [11–13]. Other hot spot CP mutations are T1753V and T1768A, which have also been associated with an increased risk of HCC [14, 15]. Those other mutations are almost invariably found in association with the TA mutation, suggesting an important role for the TA double mutation in hepatocarcinogenesis. But the mechanisms by which CP mutations induce hepatocarcinogenesis are not clear.
CP mutations may enhance HBV replication [16, 17]. However, several clinical reports have revealed that the presence of the TA mutation is not always associated with the high HBV-DNA levels [18–20]. HBx protein is a multifunctional regulator that modulates transcription, signal transduction, apoptosis, protein degradation, and cell cycle control and which has been incriminated in hepatocarcinogenesis [21, 22]. The CP region overlaps with the HBX gene, and the mutations in the overlap region may alter the biologic functions of the HBx protein, thereby increasing the risk of HCC [23]. Deregulation of cell cycle-related protein plays an important role in development and progression of HCC [24]. HBx mutants with changes that correspond to a combination of CP mutations up-regulate SKP2, which then down-regulates p21 via ubiquitin-mediated proteasomal degradation [25].
The tumor suppressor gene p53 is also involved in maintenance of the genomic integrity of cells [26]. P53 has many functions, such as transcriptional activation and repression, modulation of factors engaged in DNA repair, cell cycle control, and apoptosis regulation [27–29]. The direct interaction of HBx and P53 both in vitro and in vivo has been investigated recently [30–34]. Increasing evidence supports the view that targeted proteolysis of cell cycle-related proteins by the ubiquitin–proteasome system plays a key role in regulating the growth of cells [35]. HBx inhibits p53 transactivation and p53-dependent apoptosis [31, 32, 36]. However, whether HBx transactivation is affected by p53 is unclear. The relationship between transactivation and the p53-binding functions of HBx has not yet been clearly addressed.
The S-phase kinase-associated protein 2 (Skp2) is an ubiquitin E3 ligase responsible for recognizing and recruiting target proteins for polyubiquitylation. This protein regulates G1–S transition by posttranscriptional regulation of tumor suppressors and may play a crucial role in promoting or preventing cellular transformation and proliferation [35, 37]. We hypothesized that a combination of CP mutations modulates p53 expression through Skp2 and the ubiquitin–proteasome system.
In this study, we constructed a combination (combo) mutant HBX (with mutations corresponding to A1762T/G1764A/T1753A/T1768A) through the gene synthesis and infected both cell L-02 and hepatoma cell lines to compare the effect of the combo mutant versus wild-type (WT) HBx on p53 expression and the pathways that modulate p53 expression.
Materials and Methods
Plasmid Construction
The combo mutant HBx of genotype C corresponding to A1762T/G1764A/T1753A/T1768A, which contains an open reading frame and enhancer/promoter region (nucleotides 1,060–1,838), was constructed through gene synthesis. A FLAG epitope was introduced at the N-terminus of HBx. The HBx-Flag constructs were verified by direct sequencing. HBx protein expression was determined by anti-FLAG antibody (BD Biosciences, San Jose, CA).
Cells and Cell Culture
The HCC cell line Hep3B (American Type Culture Collection, Manassas, VA) was cultured in Minimum Essential Medium (MEM; Gibco-BRL, Grand Island, NY) supplemented with penicillin (100 U/mL), streptomycin (100 µg/mL), and 10 % fetal bovine serum (FBS). L-02 cells (obtained from our lab storage) were cultured in RPMI 1640 (Gibco-BRL) supplemented with penicillin (100 U/mL), streptomycin (100 µg/mL), and 10 % FBS. All cells were passaged at a ratio of 1:4 every 4–5 days.
Carboxyfluorescein Diacetate Succinimidyl Ester (CFSE) Labeling
CFSE cell labeling was conducted as previously described. Briefly, cultured transfected L-02 and Hep3B cells were harvested, suspended in phosphate buffered saline (PBS) at a concentration of 107/mL, and labeled with 5 µM CFSE for 15 min at room temperature in the dark. An equal volume of fetal calf serum was added, and the labeled cells were incubated for an additional 5 min. Cells were washed three times in cold PBS and used in the proliferation experiments described below. Typically, the labeling proficiency was over 98 % as analyzed by flow cytometry.
Lentiviral Transfection
The procedure was similar with the previous study (The EMBO Journal (2012) 31, 576–592). In brief, Flag-HBx or its mutations were amplified by PCR and inserted into pENTR/D-TOPO vector using pENTR Directional TOPO Cloning Kits (Invitrogen). The cDNA was then cloned into pLenti4/V5-DEST using Gateway LR recombination reaction following the manufacturer’s protocol (Invitrogen). The viruses were then used to infect cells in the presence of polybrene (6 mg/mL).
Immunofluorescence
Stationary phase cells transfected with virus were washed twice with PBS, added to a sufficient volume of 4 % formaldehyde to cover the cells, fixed at room temperature for 5 min, washed gently with PBS three times for 5 min each, dispensed in cold 100 % methanol to rupture cell membranes, rinsed with PBS for 5 min, and placed at −20 °C for 1 h to reduce nonspecific binding and lower stain background. Cells were incubated with primary antibody (anti-Flag, anti-EGFP, Santa Cruz Biotechnology, Dallas, TX, USA) diluted 1:300 at 4 °C overnight. Unbound antibody was removed by three 5-min PBS washes. Secondary antibody (1:500, Jackson ImmunoResearch, West Grove, PA, USA) was added and incubated at room temperature for 2 h in the dark, followed by addition of 10 μg/mL 4′,6-diamidino-2-phenylindole (DAPI, Santa Cruz Biotechnology) nuclear stain and incubation in the dark at room temperature for 10 min. Excess DAPI was removed by three 5-min PBS washes, and samples were observed by fluorescence microscopy.
Cell Proliferation Assay
Hep3B HCC cells and L-02 cells were plated at a density of 2 × 103 cells in triplicate in wells of 96-well plates (Corning Life Sciences, Lowell, MA). Cell proliferation was measured using the Cell Counting Kit-8 (Dojindo, Rockville, MD) according to the manufacturer’s protocol.
Cell Cycle Analysis
Cell cycle distribution was analyzed using flow cytometry. Briefly, 2 × 106 cells were trypsinized, washed, and fixed in 80 % ethanol overnight at 4 °C in the dark. Fixed cells were washed and resuspended in 50 µg/mL propidium iodide. Cell cycle distribution was analyzed using a Becton–Dickinson FACS Calibur flow cytometer. The percentage of G0/G1 cells versus S versus G2/M was performed using ModFit LT DNA analysis software and the standard diploid model. Linearity was adjusted to match the instrumentation. Although ModFit will automatically identify peaks, the results were also visually checked. For p53 overexpression, cDNA of p53 was amplified and inserted into pcDNA3 vector and these vectors were transfected into cells via Lipofectin (Life Technologies) according to the manufacturer’s protocol.
RNA Interference
Cells were transfected with 100 nmol/L ONTARGET plus SMART pool siRNAs (Dharmacon, Lafayette, CO) directed against human p53, Skp2, or nonspecific control siRNA, respectively. The p53 siRNA sequences were 5′-GAAGAAAATTTCCGCAAAA-3′, and Skip2 siRNA sequence was 5′-ATCAGATCTCTCTACTTTA-3′. siRNA transfections were performed using Lipofectamine RNAiMAX (Invitrogen) according to the manufacturer’s protocol.
Western Blot Analysis
Cell lysates were separated by sodium dodecyl sulfate/polyacrylamide gel electrophoresis, and the resolved proteins were transferred onto nitrocellulose membranes (Bio-Rad, Hercules, CA). The membranes were incubated with primary monoclonal antibody to anti-p53 (Cat#554167, BD Biosciences, San Jose, CA), anti-β-actin (A5316, Sigma-Aldrich, St. Louis, MO), and anti-cyclin D1 (H-295, Santa Cruz Biotechnology). After washing, the membranes were incubated with donkey anti-mouse immunoglobulin G/horseradish peroxidase (Abcam, Cambridge, MA). The bound antibody was visualized by chemiluminescence (Dojindo, Rockville, MD). Relative protein levels were quantified by densitometry analysis using NIH Image J.
Statistical Analysis
The data are presented as the mean ± SEM. Student’s t test was used to compare the difference between groups. P values <0.05 were regarded as statistically significant.
Results
Construction of Combo Mutant HBx and WT HBx Expression Plasmids
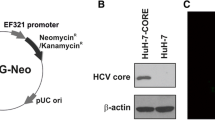
A combo HBx mutant with A1762T/G1764A/T1753A/T1768A and WT HBx of genotype C containing an open reading frame and enhancer/promoter region (nucleotides 1,060–1,838) was constructed through gene synthesis. A FLAG epitope was introduced at the N-terminus of HBx (Fig. 1a). The HBx-Flag constructs were verified by direct sequencing. HBx protein expression was determined by anti-FLAG antibody (BD Biosciences). The lentiviral vector combo mutant HBx and WT HBx can successfully infect L-02 and Hep3B (Fig. 1b). L-02 and Hep3B cells transfected with the combo mutant HBx and WT HBx stably expressed HBX-Flag (Fig. 1c, d).
HBx protein expression. a The HBx combo mutant of genotype C corresponding to A1762T/G1764A/T1753A/T1768A was constructed. The mutant contained an open reading frame and enhancer/promoter region (nucleotides 1,060–1,838). A FLAG epitope was introduced at the N-terminus of HBx. b HBx protein expression was determined by anti-FLAG antibody. The lentiviral vector combo mutant HBx and WT HBx successfully infected L-02 and Hep3B cells. c HBx expression in L-02 cells transfected with WT or combo mutant HBx of C was detected by Western blot. d HBx expression in Hep3B cells transfected with WT or the combo mutant was detected by immunofluorescence
Down-Regulated p53 Expression in the Combo Mutant
p53 expression was detected in L-02 cells transfected with WT and combo mutant HBx. WT HBx had a modest effect on p53 expression in L-02 cells compared with a negative control plasmid. The combo HBx mutant significantly decreased p53 expression compared with a negative control plasmid in hepatoma cell lines (Fig. 2a, b).
Stimulation of Cell Cycle Progression by the Combo Mutant
To determine whether HBx-induced changes in p53 expression could affect cell cycle progression, flow cytometry was performed 48 h after transfection. Consistent with the effect of the combo mutant with respect to decreased p53 expression, the combo mutant significantly increased the proportion of cells in S phase. Transfection with WT HBx did not affect the cell cycle progression (Fig. 3a). To determine whether cell cycle changes observed were truly correlated with p53 expression, p53 overexpression was forced. A significant decrease in the proportion of cells in S phase cotransfected with the combo mutant HBx and p53 compared with cells transfected with combo mutant HBx only was evident (Fig. 3b). These findings indicated that abrogation of p53 expression in cells expressing the combo mutant HBx decreased the proportion of cells that progressed into S phase. Finally, knockdown of p53 by RNA interference increased the proportion of cells in S phase compared with cells transfected with a nontarget siRNA (Fig. 3c). The findings directly indicate that the effect of combo mutant HBx on cell cycle distribution is mainly mediated by p53. To determine whether the combo mutant could influence S-phase progression in L-02 cells, we monitored the expression of cyclin D1, an important regulator of cell cycle progression. The level of cyclin D1 protein was higher in combo mutant-expressing L-02 cells than WT HBx-expressing L-02 cells (Fig. 3d, e). However, cyclin E, another member of cyclin family, did not change much (Fig. 3d, f). A similar result was observed in Hep3B cells (data not shown).
Effect of HBx on cell cycle distribution. a Cell cycle distributions in Hep3B cells expressing WT and HBx combo mutant, with empty vector as a control. Flow cytometry was performed 48 h after transfection. The combo mutant can significantly increase the proportion of cells in S phase, but transfection with WT HBx did not affect cell cycle progression. b Dynamics of p53 expression and cell cycle progression in HBx-transfected Hep3B cells. p53 expression and cell cycle distribution were ascertained at the indicated time points. Overexpression of p53 was associated with a significant decrease in the proportion of cells in S phase cotransfected with the HBx combo mutant and p53 compared with cells transfected with mutant only. c p53 silencing can increase the proportion of S phase in combo mutant HBx-expressing Hep3B cells. p53 siRNA versus nontargeting siRNA. Knockdown p53 by RNA interference can increase the proportion of the cells in S phase compared with the cells transfected with a nontargeting siRNA. d–f Effect of WT and combo mutant HBx on cyclin E, cyclin D1 expression in L-02 cells. Cyclin D1, but not cyclin E, protein level was higher in combo mutant-expressing L-02 than WT HBx-expressing L-02 (*P < 0.05; # P < 0.01)
Combo Mutant Promotes Cellular Proliferation
To detect whether the altered expression of p53 by CP mutations could affect cellular proliferation, growth rate in L-02 cells stably expressing WT and combo mutant HBx was compared. Combo mutant HBx-expressing cells grew significantly more quickly than WT (P < 0.001) HBx-expressing cells (Fig. 4a, b). Similar results were seen in Hep3B cells (Fig. 4c, d).
Impact of HBx mutants on cellular proliferation. a, b Transfected L-02 were labeled by CFSE and cultured for 48 h. The fluorescence intensity of the HBx combo mutant-expressing L-02 cells was reduced more quickly than WT HBx-expressing L-02 cells and the control group. b Flow cytometry was performed to detect the fluorescence intensity of transfected L-02 cells. The proportion of lower fluorescence intensity of the mutant group was higher than that of the WT and control groups. c In transfected Hep3B cells at 24 and 48 h, the average fluorescence intensity of HBx combo mutant-expressing Hep3B cells was lower than that of the WT and control groups. d Growth rates of Hep3B cells stably expressing the mutant were compared with that of cells expressing WT HBx and control. The mutant-expressing Hep3B cells grew significantly more quickly than WT HBx-expressing Hep3B cells and control (*P < 0.05; # P < 0.01)
Accelerated p53 Degradation by the Combo Mutant
To further investigate the potential mechanisms by which HBx regulates p53 expression, the effect of the combo mutant on p53 protein degradation was studied. Protein stability was ascertained in cycloheximide-treated L-02 cells. p53 stability was decreased in cells transfected with the combo mutant, but was unchanged in WT HBx-transfected cells compared with control (Fig. 5a). However, there was no difference among mRNA expressions upon these treatments (Fig. 5b). The results indicated that combo CP mutations accelerated p53 degradation. To unravel the mechanism by which the combo mutant regulated p53 stability, cells were treated with proteasome inhibitor MG132. The effects of mutant HBx on p53 stability were abolished (Fig. 5c, d), indicating HBx regulation of p53 stability via the ubiquitin–proteasomal system.
The combo mutation accelerates p53 protein degradation. a, b Effect of HBx on p53 protein stability/degradation. L-02 cells were transfected transiently with WT or combo mutant HBx, and then treated with cycloheximide, followed by Western blot and Q-PCR analysis. p53 stability was decreased in the mutant-transfected cells and was unchanged in WT HBx-transfected cells compared with control. The average target/β-Actin ratios for each time point were plotted in combo mutation group. There was no difference among mRNA expressions. c, d Immunoblotting for p53 in WT and combo mutant HBx-expressing Hep3B cells with p53 siRNA. Hep3B cells were treated with proteasome inhibitor MG132. The effects of mutant HBx on p53 stability were abolished
CP Mutations Regulate p53 Expression by Skp2
Results from both Hep3B cells (Fig. 6a) and L-02 cells (Fig. 6b) showed that Skp2 expression increased in the combo mutant and was unchanged in WT HBx-expressing cells compared with control. A similar result was observed in Huh7 cells (data not shown). To further investigate the role of Skp2 in regulating p53 expression, Skp2 expression was abrogated by RNA interference to determine the effect on p53 protein expression and cell cycle distribution. Silencing of Skp2 increased the p53 expression in cells transfected with the combo mutant. Skp2 silencing resulted in similar cell cycle distributions regardless of which HBx protein (WT, combo, or control) was expressed. Skp2 knockdown also abolished the enhancement of S-phase entry by the combo mutant (Fig. 6c). The results indicate that HBx can regulate p53 expression mainly through Skp2.
The role of Skp2 on WT and HBx combo mutant modulation of p53 expression and cell cycle distribution. a Investigation of Skp2 protein in WT and mutant HBx-expressing Hep3B cells by Western blot analysis. Skp2 expression increased in the combo mutant and was unchanged in WT HBx-expressing cells compared with control. b Investigation of Skp2 protein in WT and mutant HBx-expressing L-02 cells. Skp2 expression was increased in the combo mutant and unchanged in WT HBx-expressing cells compared with control. c Hep3B cells were transfected with control or Skp2 siRNA for 24 h and then transfected with WT and combo mutant HBx followed by Western blot and determination of cell cycle distribution by fluorescence-activated cell sorting. p53 expression in the Skp2 silenced group was higher than the nontarget siRNA group. The S-phase proportion in Skp2 silencing cells was lower than the nontarget siRNA group. (*P < 0.05)
Discussion
Hepatitis B virus CP resides in the overlapping X gene and plays an important role in HBV replication and hepatocarcinogenesis [38]. The CP is composed of the core upstream regulatory region (URR) and the basal CP. The T1762/A1764 double mutation has been associated with severe forms of liver disease [8, 39–42], although the overall pathogenic significance remains controversial. Clinical observations have suggested that HBV CP mutations generally accumulate in a stepwise fashion before the development of HCC, often beginning with the TA double mutation. T1768A and T1753A are frequent in HCC patients [43–45]. HBx is a multifunctional viral protein that may influence apoptotic and cell cycle regulatory pathways, but the results of these activities are likely affected by the systems in which the studies have been conducted [21]. Several studies reported that HBx can induce apoptosis [46–48], sensitize cells to proapoptotic stimuli [49], prevent apoptosis [50–52], or promote cell proliferation [53, 54]. But the mechanism underlying CP mutation and hepatocarcinogenesis is largely unknown. Since mutations in the CP region in patients with HCC can alter the coding sequence of the overlapping HBx gene, and HBx protein has been previously incriminated in hepatocarcinogenesis, we investigated the impact of combination HBX mutations on the biologic function of HBx protein.
The combo mutant with changes corresponding to CP mutations T1753A/A1762T/G1764A/T1768A dramatically decreased p53 expression in L-02 cells compared with control. Cell cycle analysis showed the combo mutant markedly increased the proportion of cells in S phase and increased cellular proliferation and growth. The mutant also up-regulated cyclin D1 expression in L-02 cells. Cyclin D1 can promote G1/S-phase transition and acts as a switch for various cellular processes, including initiation of DNA replication [33]. The combo mutant decreased p53 and increased cyclin D1 expression in L-02 cells, providing indirect evidence to support the view that this mutant contributes to DNA replication and cellular proliferation. Tumor suppressor protein p53 can respond to genotoxic and oncogenic stress by inducing cell-growth arrest or by promoting apoptosis [55]. The tumor suppressor function of p53 depends on its activity as a transcription factor.
The F box protein Skp2 is required for substrate recognition of the SCFSkp2 ubiquitin ligase complex. Skp2 targets many proteins for subsequent degradation and ubiquitination, including p21, p27, p57 and tumor suppressors including p130 [56]. Skp2 mediates ubiquitination and degradation of c-Myc, indicating that Skp2 may enhance c-Myc-induced G1/S transition and c-Myc transactivation activity [57]. Skp2 is oncogenic, and its overexpression correlates with the grade of malignancy in many tumors. Skp2 has also been implicated in the regulation of apoptosis. Gastric cancer cells transfected with Skp2 are resistant to induction of apoptosis by actinomycin D [58]. On the other hand, down-regulation of Skp2 by small interfering RNA can decrease cell growth and increase apoptosis [59–61]. However, the exact molecular mechanism by which Skp2 suppresses apoptosis remains unclear.
Our results indicate that Skp2 plays a key role in cellular proliferation. Overexpression of Skp2 significantly reduces p53 expression and promotes cellular proliferation, whereas siRNA-mediated down-regulation of Skp2 measurably increases the level of p53 and abrogates this effect. As overexpression of Skp2 suppresses the transactivation ability of p53, Skp2 may also partially impair p53-mediated cell cycle arrest at the G1 phase. So, Skp2 may targets p53. In this regard, overexpression of Skp2 is expected to show a cooperative effect with the reduction in CDK inhibitors to suppress the p53-mediated cell cycle arrest. Thus, Skp2 is likely to attenuate p53 activation and to trigger cell proliferation, the combination of which influences tumorigenesis.
To determine how CP mutations regulate p53 expression, we detected the effects of WT and the HBx combo mutant on p53 transcription. The mutant markedly decreased p53 expression, and this effect was mediated through the ubiquitin–proteasomal system. Levels of SKP2 were higher in L-02, and Hep3B cells transfected with the mutant than with WT. Thus, SKP2 may be important in HBx regulation of p53. Increased Skp2 expression has been found in many human malignant tumors, including HCC [62, 63]. In addition to p53, SKP2 also targets other cell cycle regulators such as p27, p21, p57, and p130 for polyubiquitination [35]. Although we did not detect these other factors in response to HBx expression, we hypothesize that the ability of combo mutant HBx to promote hepatocarcinogenesis may be mediated by deregulation of additional factors besides p53. Skp2 up-regulation may favor cellular transformation and tumor progression. Therefore, Skp2 may be a therapeutic target for HCC.
Previous studies showed that HBx plays an important role in expression of p53 [64–66]. But these studies did not provide details on the HBx protein sequence. Presently, HBx mutations corresponding to CP T1753A/A1762T/G1764A/T1768A had opposite effects on p53 expression compared with WT HBx. The demonstration that HBx regulates p53 via SKP2 is novel. Finally, the combo CP mutations not only down-regulated p53 expression but also up-regulated Skp2 level in L-02 as well as in cancer cell lines.
In summary, this study provides a possible mechanism by which HBV CP mutations are associated with an increased risk of HCC. The results suggest that combo CP mutations may promote hepatocarcinogenesis, possibly via down-regulation of p53 expression and, in turn, enhancement of colony formation, cellular proliferation, and acceleration of cell cycle progression. These CP mutations may promote hepatocarcinogenesis through some other mechanisms. These findings will provide further insight into the oncogenic role of combo CP mutations and may lead to the development of novel therapeutic strategies to prevent HBV-related HCC.
References
Farazi PA, Glickman J, Horner J, et al. Cooperative interactions of p53 mutation, telomere dysfunction and chronic liver damage in hepatocellular carcinoma progression. Cancer Res. 2006;66:4766–4773.
Chen JG, Zhang SW. Liver cancer epidemic in China: past present and future. Semin Cancer Biol. 2011;21:59–69.
Yang HI, Lu SN, Liaw YF, Taiwan Community-Based Cancer Screening Project Group, et al. Hepatitis B e antigen and the risk of hepatocellular carcinoma. N Engl J Med. 2002;347:168–174.
el-Deiry WS, Tokino T, Velculescu VE, et al. WAF1, a potential mediator of p53 tumor suppression. Cell. 1993;75:817–825.
Yuen MF, Tanaka Y, Shinkai N, et al. Risk for hepatocellular carcinoma with respect to hepatitis B virus genotypes B/C, specific mutations of enhancer II/core promoter/precore regions and HBV DNA levels. Gut. 2008;57:98–102.
Yang HI, Yeh SH, Chen PJ, et al. Associations between hepatitis B virus genotype and mutants and the risk of hepatocellular carcinoma. J Natl Cancer Inst. 2008;100:1134–1143.
Chen CJ, Yang HI, Su J, et al. Risk of hepatocellular carcinoma across a biological gradient of serum hepatitis B virus DNA level. JAMA. 2006;295:65–73.
Kao JH, Chen PJ, Lai MY, et al. Basal core promoter mutations of hepatitis B virus increase the risk of hepatocellular carcinoma in hepatitis B carrier. Gastroenterology. 2003;124:327–334.
Tsubota A, Arase Y, Ren F, et al. Genotype may correlate with liver carcinogenesis and tumor characteristics in cirrhotic patients infected with hepatitis B virus subtype adw. J Med Virol. 2001;65:257–265.
Yuen MF, Tanaka Y, Mizokami M, et al. Role of hepatitis B virus genotypes Ba and C, core promoter and precore mutations on hepatocellular carcinoma: a case control study. Carcinogenesis. 2004;25:1593–1598.
Hsieh YH, Su IJ, Wang HC, et al. Pre-S mutant surface antigens in chronic hepatitis B virus infection induce oxidative stress and DNA damage. Carcinogenesis. 2004;25:2023–2032.
Hwang GY, Lin CY, Huang LM, et al. Detection of the hepatitis B virus X protein (HBx) antigen and anti-HBx antibodies in cases of human hepatocellular carcinoma. J Clin Microbiol. 2003;41:5598–5603.
Oon CJ, Chen WN, Goh KT, et al. Molecular characterization of hepatitis B virus surface antigen mutant in Singapore patients with hepatocellular carcinoma and hepatitis B virus carrier negative for HBsAg but positive for anti-HBs and anti-HBc. J Gastroenterol Hepatol. 2002;17:S491–S496.
Liu S, Zhang H, Gu C, et al. Associations between hepatitis B virus mutations and the risk of hepatocellular carcinoma: a meta-analysis. J Natl Cancer Inst. 2009;101:1066–1082.
Guo X, Jin Y, Qian G, et al. Sequential accumulation of the mutations in core promoter of hepatitis B virus is associated with the development of hepatocellular carcinoma in Qidong, China. J Hepatol. 2008;49:718–725.
Parekh S, Zoulim F, Ahn SH, et al. Genome replication, virion secretion, and e antigen expression of naturally occurring hepatitis B virus core promoter mutants. J Virol. 2003;77:6601–6612.
Buckwold VE, Xu Z, Chen M, et al. Effects of a naturally occurring mutation in the hepatitis B virus basal core promoter on precore gene expression and viral replication. J Virol. 1996;70:5845–5851.
Chu CJ, Keeffe EB, Han SH, et al. Prevalence of HBV precore/core promoter variants in the United States. Hepatology. 2003;38:619–628.
Fang ZL, Sabin CA, Dong BQ, et al. The association of HBV core promoter double mutations (A1762T and G1764A) with viral load differs between HBeAg positive and anti-HBe positive individuals: a longitudinal analysis. J Hepatol. 2009;50:273–280.
Chun YK, Kim JY, Woo HJ, et al. No significant correlation exists between core promoter mutations, viral replication, and liver damage in chronic hepatitis B infection. Hepatology. 2002;32:1154–1162.
Bouchard MJ, Schneider RJ. The enigmatic X gene of hepatitis B virus. J Virol. 2004;78:12725–12734.
Hu Z, Zhang Z, Doo E, et al. Hepatitis B virus X protein is both a substrate and a potential inhibitor of the proteasome complex. J Virol. 1999;73:7231–7240.
Lin Y, Nomura T, Yamashita T, et al. The transactivation and p53-interacting functions of hepatitis B virus X protein are mutually interfering but distinct. Cancer Res. 1997;57:5137–5142.
Thorgeirsson SS, Grisham JW. Molecular pathogenesis of human hepatocellular carcinoma. Nat Genet. 2002;31:339–346.
Huang Y, Tong S, Tai AW, et al. Hepatitis B virus core promoter mutations contribute to hepatocarcinogenesis via deregulation of SKP2 and its target p21. Gastroenterology. 2011;141:1412–1421.
Lane DP. Cancer: p53, guardian of the genome. Nature. 1992;358:15–16.
Wu X, Levine AJ. p53 and E2F-l cooperate to mediate apoptosis. Proc Natl Acad Sci USA. 1994;91:23602–23606.
Wang XW, Vermeulen W, Coursen JD, et al. The XPB and XPD DNA helicases are components of the p53-mediated apoptosis pathway. Genes Dev. 1996;10:1219–1231.
Dulić V, Kaufmann WK, Wilson SJ, et al. p53-dependent inhibition of cyclin-dependent kinase activities in human fibroblasts during radiation-induced G1 arrest. Cell. 1994;76:1013–1023.
Feitelson MA, Zhu M, Duan LX, et al. Hepatitis B X antigen and p53 are associated in vitro and in liver tissues from patients with primary hepatocellular carcinoma. Oncogene. 1993;8:1109–1117.
Wang XW, Forrester K, Yeh H, et al. Hepatitis B virus X protein inhibits p53 sequence-specific DNA binding, transcriptional activity, and association with transcription factor ERCC3. Proc Natl Acad Sci USA. 1994;91:2230–2234.
Truant R, Antunovic J, Greenblatt J, et al. Direct interaction of the hepatitis B virus HBx protein with p53 leads to inhibition by HBx of p53 response element-directed transactivation. J Virol. 1995;69:1851–1859.
Ueda H, Ullrich SJ, Gangemi JD, et al. Functional inactivation but not structural mutation of p53 causes liver cancer. Nat Genet. 1995;9:41–47.
Lee H, Lee YH, Huh YS, et al. X-gene product antagonize the p53-mediated inhibition of hepatitis B virus replication through regulation of the pregenomic/core promoter. J Biol Chem. 1995;270:31405–31412.
Frescas D, Pagano M. Deregulated proteolysis by the F-box proteins SKP2 and beta-TrCP: tipping the scales of cancer. Nat Rev Cancer. 2008;8:438–449.
Wang XW, Gibson MK, Vermeulen W, et al. Abrogation of p53-induced apoptosis by the hepatitis B virus X gene. Cancer Res. 1995;52:6012–6016.
Hoeller D, Dikic I. Targeting the ubiquitin system in cancer therapy. Nature. 2009;458:438–444.
Kramvis A, Kew MC. The core promoter of hepatitis B virus. J Viral Hepat. 1999;6:415–427.
Chou YC, Yu MW, Wu CF, et al. Temporal relationship between hepatitis B virus enhancer II/basal core promoter sequence variation and risk of hepatocellular carcinoma. Gut. 2008;57:91–97.
Jang JW, Lee YC, Kim MS, et al. A 13-year longitudinal study of the impact of double mutations in the core promoter region of hepatitis B virus on HBeAg seroconversion and disease progression in patients with genotype C chronic active hepatitis. J Viral Hepat. 2007;14:169–175.
Liu CJ, Chen BF, Chen PJ, et al. Role of hepatitis B virus precore/core promoter mutations and serum viral load on noncirrhotic hepatocellular carcinoma: a case control study. J Infect Dis. 2006;194:594–599.
Sakamoto T, Tanaka Y, Orito E, et al. Novel subtypes (subgenotypes) of hepatitis B virus genotypes B and C among chronic liver disease patients in the Philippines. J Gen Virol. 2006;87:1873–1882.
Tanaka Y, Mukaide M, Orito E, et al. Specific mutations in enhancer II/core promoter of hepatitis B virus subgenotypes C1/C2 increase the risk of Hepatocellular carcinoma. J Hepatol. 2006;45:646–653.
Ito K, Tanaka Y, Orito E, et al. T1653 mutation in the box alpha increases the risk of hepatocellular carcinoma in patients with chronic hepatitis B virus genotype C infection. Clin Infect Dis. 2006;42:1–7.
Takahashi K, Ohta Y, Kanai K, et al. Clinical implications of mutations C-to-T1653 and T-to-C/A/G1753 of hepatitis B virus genotype C genome in chronic liver disease. Arch Virol. 2009;144:1299–1308.
Chirillo P, Pagano S, Natoli G, et al. The hepatitis B virus X gene induces p53-mediated programmed cell death. Proc Natl Acad Sci USA. 1997;94:8162–8167.
Kim H, Lee H, Yun Y. X-gene product of hepatitis B virus induces apoptosis in liver cells. J Biol Chem. 1998;73:381–385.
Terradillos O, Pollicino T, Lecoeur H, et al. p53-independent apoptotic effects of the hepatitis B virus HBx protein in vivo and in vitro. Oncogene. 1998;17:2115–2123.
Su F, Schneider RJ. Hepatitis B virus HBx protein sensitizes cells to apoptotic killing by tumor necrosis factor alpha. Proc Natl Acad Sci USA. 1997;94:8744–8749.
Elmore LW, Hancock AR, Chang SF, et al. Hepatitis B virus X protein and p53 tumor suppressor interactions in the modulation of apoptosis. Proc Natl Acad Sci USA. 1997;94:14707–14712.
Marusawa H, Matsuzawa S, Welsh K, et al. HBXIP functions as a cofactor of survivin in apoptosis suppression. EMBO J. 2003;22:2729–2740.
Shih WL, Kuo ML, Chuang SE, et al. Hepatitis B virus X protein inhibits transforming growth factor-beta-induced apoptosis through the activation of phosphatidylinositol 3-kinase pathway. J Biol Chem. 2000;275:25858–25864.
Koike K, Moriya K, Yotsuyanagi H, et al. Induction of cell cycle progression by hepatitis B virus HBx gene expression in quiescent mouse fibroblasts. J Clin Invest. 1994;94:44–49.
Madden CR, Finegold MJ, Slagle BL. Hepatitis B virus X protein acts as a tumor promoter in development of diethylnitrosamine induced preneoplastic lesions. J Virol. 2001;75:3851–3858.
Vousden KH, Lu X. Live or let die: the cell’s response to p53. Nat Rev Cancer. 2002;2:594–604.
Nakayama KI, Nakayama K. Ubiquitin ligase: cell-cycle control and cancer. Nat Rev Cancer. 2006;6:369–381.
Kim SY, Herbst A, Tworkowski KA, et al. Skp2 regulates Myc protein stability and activity. Mol Cell. 2003;11:1177–1188.
Masuda TA, Inoue H, Sonoda H, et al. Clinical and biological significance of S-phase kinase-associated protein 2 (Skp2) gene expression in gastric carcinoma: modulation of malignant phenotype by Skp2 overexpression, possibly via p27 proteolysis. Cancer Res. 2003;62:3819–3825.
Jiang F, Caraway NP, Li R, et al. RNA silencing of S-phase kinase-interacting protein 2 inhibits proliferation and centrosome amplification in lung cancer cells. Oncogene. 2005;24:3409–3418.
Lee SH, McCormick F. Downregulation of Skp2 and p27/Kip1 synergistically induces apoptosis in T98G glioblastoma cells. J Mol Med. 2005;83:296–307.
Harada K, Supriatno Kawashima Y, et al. Down-regulation of S-phase kinase associated protein 2 (Skp2) induces apoptosis in oral cancer cells. Oral Oncol. 2005;41:623–630.
Gao D, Inuzuka H, Tseng A, et al. Phosphorylation by Akt1 promotes cytoplasmic localization of Skp2 and impairs APCCdh1-mediated Skp2 destruction. Nat Cell Biol. 2009;11:397–408.
Calvisi DF, Ladu S, Pinna F, et al. SKP2 and CKS1 promote degradation of cell cycle regulators and are associated with hepatocellular carcinoma prognosis. Gastroenterology. 2009;137:1816–1826.
Qiao L, Leach K, McKinstry R, et al. Hepatitis B virus X protein increases expression of p21(Cip-1/WAF1/MDA6) and p27(Kip-1) in primary mouse hepatocytes, leading to reduced cell cycle progression. Hepatology. 2001;34:906–917.
Park US, Park SK, Lee YI, et al. Hepatitis B virus-X protein upregulates the expression of p21waf1/cip1 and prolongs G1¡S transition via a p53-independent pathway in human hepatoma cells. Oncogene. 2000;19:3384–3394.
Kwun HJ, Jang KL. Natural variants of hepatitis B virus X protein have differential effects on the expression of cyclin-dependent kinase inhibitor p21 gene. Nucleic Acids Res. 2004;32:2202–2213.
Acknowledgments
This work was supported by National Natural Science Foundation of Guangdong Province (S2013010016015, Meihai Deng), The National Natural Science Fund for Young Scholars of China (81000177, Yuesi Zhong) and The Projects on Science and Technology plan of Guangzhou City (2013J410061-1955, Meihai Deng).
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
Zhicheng Yao and Kunpeng Hu have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Yan, J., Yao, Z., Hu, K. et al. Hepatitis B Virus Core Promoter A1762T/G1764A (TA)/T1753A/T1768A Mutations Contribute to Hepatocarcinogenesis by Deregulating Skp2 and P53. Dig Dis Sci 60, 1315–1324 (2015). https://doi.org/10.1007/s10620-014-3492-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-014-3492-9