Abstract
Understanding population structure and genetic diversity is important for designing effective conservation strategies. As a critically endangered shrub, the six remaining extant populations of spiny daisy (Acanthocladium dockeri) are restricted to country roadsides in the mid-north of South Australia, where the species faces many ongoing abiotic and biotic threats to survival. Currently the spiny daisy is managed by selecting individuals from the extant populations and translocating them to establish insurance populations. However, there is little information available on the genetic differentiation between populations and diversity within source populations, which are essential components of planning translocations. To help fill this knowledge gap, we analysed population structure within and among all six of its known wild populations using 7,742 SNPs generated by a genotyping-by-sequencing approach. Results indicated that each population was strongly differentiated, had low levels of genetic diversity, and there was no evidence of inter-population gene flow. Individuals within each population were generally closely related, however, the Melrose population consisted entirely of clones. Our results suggest genetic rescue should be applied to wild spiny daisy populations to increase genetic diversity that will subsequently lead to greater intra-population fitness and adaptability. As a starting point, we suggest focussing on improving seed viability via inter-population crosses such as through hand pollination experiments to experimentally assess their sexual compatibility with the hope of increasing spiny daisy sexual reproduction and long-term reproductive fitness.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Small, fragmented plant populations often have reduced genetic diversity, which risks elevating inbreeding and genetic drift within populations (Templeton et al. 1990; van Treuren et al. 1991; Heywood 1993; Furlan et al. 2012; Neaves et al. 2015). Inbreeding and genetic drift are particularly concerning for small populations because they can increase the prevalence of disease (O’Brien and Evermann 1988; Hajjar et al. 2008). Genetic erosion is the reduction in effective population size within a population over time; where infrequently occurring alleles are likely to be lost, further hindering population survival by reducing adaptive potential (van Treuren et al. 1991). In addition, populations that are at risk of habitat removal are likely to experience population genetic bottlenecks (Buza et al. 2000). Maintaining or even increasing gene flow between fragmented populations is therefore of high importance as it should help support long-term population fitness and increase evolutionary resilience within a species (Moritz 1999). Mapping genetic diversity within a species of conservation concern provides insights into its genetic risk, which is increasingly being used to guide conservation interventions (i.e. for translocations) (Williams et al. 2014; Rossetto et al. 2021). Such knowledge enables managers to prioritize sites for in situ conservation and facilitates identification of source material for the establishment of ex situ activities such as translocations (Young et al. 2002).
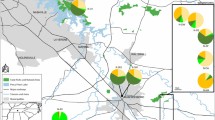
Acanthocladium dockeri (spiny daisy) is listed as critically endangered under the Commonwealth of Australia’s Environment Protection and Biodiversity Conservation Act 1999 and within the IUCN criteria (Sharp et al. 2010; Clarke et al. 2013; Bickerton et al. 2018). It is the only species within the monotypic genus Acanthocladium, meaning it is an important representative of Australian botanical diversity (Jusaitis and Adams 2005; Clarke et al. 2013; Bickerton et al. 2018). Only six small pop Clarke et al. 2013ulations (~ 87–1986 m2) are known to occur within the mid-north of South Australia and are separated by 4-110 km distances. These populations have been named in previous studies as Hart, Telowie, Thornlea, Melrose, Yangya and Rusty Cab (Fig. 1) (Jusaitis and Adams 2005; Clarke et al. 2013; Bickerton et al. 2018). These populations are susceptible to anthropogenic interference as they occur along narrow roadsides that are impacted by agricultural practices. They are exposed to chemical fertilisers and pesticides, herbivory and compete with weeds during the winter months (Clarke et al. 2013). Seed production in the wild is infrequent making recruitment via sexual reproduction rare (Jusaitis 2008). No seedlings have ever been observed in the field despite the presence of insect pollinators on spiny daisy flowers(Jusaitis and Adams 2005). The roughly 50 cm diameter shrubs produce suckers within their woody perennial root system, and this has helped these populations to endure (Clarke et al. 2013). Previous work has shown spiny daisies have low pollen viability and their flowers have a deformed pollen tube possibly contributing to prolonged vegetative reproduction that will eventually cause sterility (Jusaitis and Adams 2005; Jusaitis 2008).
Previous work has assessed the genetic diversity among four populations known at the time (Thornlea, Yangya, Hart and Rusty Cab) using allozymic markers (Jusaitis and Adams 2005). The authors found that each population represented a single, distinct genotype suggesting all individuals within each population were clones and there was no inter-population gene flow (Jusaitis and Adams 2005). Efforts to conserve the spiny daisy included regular site monitoring and maintenance (i.e. pest control and weeding), in addition to the establishment of at least one translocation site for each population (Sharp et al. 2010; Clarke et al. 2013). Subsequent conservation activities further led to the discovery of another two populations (Telowie and Melrose) (Clarke et al. 2013). Although informative, the analysis of allozyme markers only provides a limited representation of the allelic variation in a genome, in comparison to the use of more modern genome-wide techniques (Gaudeul et al. 2004; Narum et al. 2013). It is therefore necessary that population studies use high-resolution genomic methodologies to better inform management and conservation goals.
Next-generation sequencing (NGS) applications have more frequently been used for non-model organisms, improving conservation management strategies (Seeb et al. 2011; Narum et al. 2013). Genotyping by sequencing (GBS) is a popular method that uses NGS platforms to reduce genome complexity and characterise genomic variation across thousands of genome-wide single nucleotide polymorphisms (SNPs). GBS provides a better insight into population structure compared to more traditional methods such as allozymes or microsatellites as there is greater genomic resolution (Beissinger et al. 2013; Narum et al. 2013).
The aim of our study was to determine levels of genetic diversity within and among six extant populations of the spiny daisy using a genome-wide SNP dataset derived by GBS. We hope to use this information to help conservation management of this species. Based on previous findings, we predicted that each population would be genetically differentiated and contain low levels of genetic diversity. We addressed the following questions: (i) How strong is the genetic differentiation among populations? (ii) How variable is intra-populations genetic diversity? (iii) Is evidence of gene flow present among the extant populations?
Materials and methods
Study sites and sampling
All spiny daisy populations occur along roadsides within close proximity to country townships in the mid-north of South Australia (Fig. 1) (Jusaitis and Adams 2005; Clarke et al. 2013). Some of these sites are surrounded by several species of native vegetation identified in the Spiny Daisy Recovery Guide (Clarke et al. 2013). Thornlea, Yangya and Rusty Cab sites are situated East of Laura and are the closest geographically (4 km) to each other. The Hart site occurs within the Clare Valley, approximately 65 km south of the township Laura, between a main sealed road and an old railway reserve. The sites are located in semi-arid grasslands, adjacent to and sometimes partially on private property. The Telowie site is located approximately 20 km north of Port Pirie and its closest population, Thornlea is 30 km to the south-east of Telowie. Melrose represents the most northerly site, occurring 31 km north-east of Telowie. Plant density of these populations have been monitored over time and was recorded in 2007 by the Department of Environment, Water and Natural Resources (DEWNR) (Appendix Table S1) (Clarke et al. 2013).
Young leaf tissue samples were collected from a total of 90 individuals (15 plants per population) among the six sites. Five leaves from each plant harvested were taken along a 45 m transect at each site. Along the transect, the plants were randomly selected at a three-metre interval and within three metres from the transect. This means that sampled individuals were never less than 3 m apart. In instances where plants were absent at a designated interval, the next closest individual along the transect was sampled and the altered distance was recorded. This occurred at the Hart site where the distance between the last individual sampled and its previous sample was 10 m. Transects were also altered if the distribution of the plant populations was not linear up to 45 m. This was the case with the Telowie population, in which the transect had to be separated into two directions. As most sites contained a small number of individuals, a sample size of 15 individuals was thought to provide an adequate representation of the genetic material within the population.
Young leaves were harvested from the tip of a branch, down to the fourth node as DNA quality is known to improve for younger tissue (Moreira and Oliveira 2011). Leaves were selected from one to three separate branches of the plant. The five leaves from each plant were individually stored and temporarily preserved in liquid nitrogen. Samples were then freeze-dried using the Beta 2–8 LSCplus ice condenser for a 2-day period to allow for the extraction of moisture within the leaf.
DNA extraction and genotyping
DNA was extracted from two to three leaves (6–7 mg) from each individual by Diversity Arrays Technology Pty Ltd (DArT) according to their inhouse protocol (Jaccoud et al. 2001; Kilian et al. 2012). Library preparation involved the restriction enzymes PstI and MseI for complexity reduction which targeted low-copy sequences (Wenzl et al. 2004; Melville et al. 2017; van Deventer et al. 2020). Sequencing was conducted using DArTseq™; a genome complexity reduction sequencing methodology, similar to RADseq (Restriction site Associated DNA sequencing) (Sansaloni et al. 2011; Kilian et al. 2012; Rodger et al. 2021). The procedure was performed using an Illumina short-read platform producing 300,000 reads of 75 base pairs in length which were then filtered and shortened based on sequence quality. The closest relative with a complete genome sequence is the sunflower (Helianthus annuus) with a genome size of 3.6 gigabases (Kane et al. 2011; Badouin et al. 2017). Assuming the spiny daisy genome has a similar size to that of the sunflower, we expected the genome coverage with this methodology to be approximately 0.65% of the spiny daisy genome. SNP calling and initial marker filtering based on sequencing read depth and accuracy were performed using standard procedures developed by DArT (Kilian et al. 2012).
Marker quality control
Filtering of less informative SNPs and SNPs introducing bias was conducted in R v 4.0.2 (2020.06.22) (RStudio Team. 2020) using the DARTR package (Gruber et al. 2018). The sequencing procedure was replicated so that each locus was scored based on genotype repeatability. We applied a reproducibility filter to stringently remove loci which contained inadequate repeatability scores (< 99%). Failure to call SNPs due to reduced DNA quality causes missing genotypes which can severely bias some analyses. Loci containing > 1% missing data were removed. Secondary SNPs occur when multiple SNPs occur within the same sequencing read. These cause linkage disequilibrium, another source of bias for some analysis and were therefore removed. To improve data quality and computational efficiency, a minor allele frequency threshold of 5% was applied (The International HapMap 2005; Würschum et al. 2013).
Population genetic structure and differentiation
Genetic structure within the final dataset was examined in R v 4.0.2 using various approaches. Firstly, a Discriminant Analysis of Principle Components (DAPC) within the R package Adegenet was computed to identify genetic clusters and examine the validity of individual assignment within groups. This analysis employs a k-means clustering algorithm which infers the optimal number of k based on the selection of clusters to maximise between-group variation. The most optimal model was defined upon the value which corresponded to the lowest BIC (Bayesian information criterion). A Principal Coordinates Analysis (PCoA) was then used to visualise genomic variation (Jombart 2008). Lastly, to investigate the extent of gene flow between different geographical locations, individual admixture coefficients were produced in the R package LEA, a Bayesian clustering program similar to that of STRUCTURE (Frichot et al. 2015). Shared ancestry among individuals were examined using models of K ranging between 1 and 10 with each model being repeated ten times. Pairwise Weir et al. (1984) FST estimates were obtained using the R package StAMPP (Pembleton et al. 2013) to estimate population genetic differentiation among the sampling sites. The percentage of loci with fixed allelic differences between populations was calculated and represented as a heat map using the R package Pheatmap. A fixed allele occurs when all individuals in a population contain an allele that is not shared with individuals from any other population. The presence and amount of fixed alleles within a population will make that population more genetically distinct (Huber et al. 2016). We examined isolation by distance (IBD) to determine whether geographic distance or geographic isolation is affecting inter-population gene flow. We used the R package Adegenet to test for IBD with a Mantel test and Pearson’s product-moment correlations using pairwise comparisons of Euclidean geographic distance and FST/(1-FST) between populations (Rousset 1997).
Population genetic diversity
Observed heterozygosity (HO) and expected heterozygosity (HE) were calculated in the DARTR package (Gruber et al. 2018). An Analysis of Molecular Variance (AMOVA) was computed using GenAlEx v 6.503, implementing 999 permutations, to examine the distribution of genetic diversity within individuals, among individuals and among populations (Peakall and Smouse 2012). To compare levels of relatedness among the populations, pairwise comparisons of relatedness among individuals within one population were performed using the software COANCESTRY v 1.0. Relatedness estimates were obtained using the Wang estimator with 100 bootstraps while accounting for inbreeding (Wang 2011). FIS values were generated within GenAlEx v 6.503. To determine whether samples from each population contained clones, we used COLONY v 2.0.6.5 (Jones and Wang 2010).
Results
Marker quality
From a total of 73,407 SNPs, 7,742 were retained across the 90 individuals after filtering. The majority of SNPs were removed due to low reproducibility (removing 18,578 SNPs) and missing data (removing 45,843 SNPs). Furthermore, the deletion of secondary SNPs and loci below the minor allele frequency threshold removed a further 1,139 and 105 SNPs respectively, bringing the total of retained SNPs within the final dataset to 7,742. This final dataset contained no missing values.
Population structure
Population structure between all sampling localities was supported by several genetic analyses. The DAPC showed that the most likely model of populations structure was K = 6, based on the lowest Bayesian information criterion (Appendix Fig. S1). The PCoA showed a similar result to the DAPC (Fig. 2). The PCoA demonstrated that individuals from each population clustered together, but there was variation between the level of differentiation among populations (Fig. 2). Axis 1 (21.1% of variation) separated the two furthest populations, Melrose (furthest north) and Hart (furthest south), from all the other populations while axis 2 (19.2% of variation) separated Hart and Rusty Cab from all other populations (Fig. 2). Thornlea, Yangya and Telowie were the most genetically similar and Yangya showed the most variation between its individuals.
The proportion of shared ancestry between the six populations produced by the Bayesian clustering program LEA was very small (Fig. 3). Alternative models of K also displayed little admixture between populations (Appendix Fig. S2).
All populations contained similar levels of high differentiation according to pairwise FST estimates (Table 1). The mean pairwise FST estimates between populations was 0.53 ± 0.03 SD (Table 1). Melrose produced slightly higher pairwise FST estimates with all other populations (mean = 0.57 ± 0.007 SD).
AMOVA results displayed significantly greater variation within individuals (63%) than among populations (37%). No variation was detected among individuals (0%) (p = 0.01; Table 2). The proportion of fixed differences between the populations was similar across population pairs (mean = 5.1 ± 0.6%) (Fig. 4). Pairwise comparisons involving Melrose had the highest proportion of fixed differences (mean = 5.8 ± 0.2%) (Fig. 4). The lowest proportion of fixed differences was between Thornlea and Yangya (4.18%).
Genetic diversity and isolation-by-distance
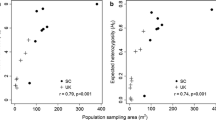
Observed heterozygosity (mean = 0.26 ± 0.03 SD) appeared to be considerably higher than expected heterozygosity (mean = 0.13 ± 0.02 SD) and this pattern was found across all the populations (Fig. 5). Thornlea contained the highest value of expected and observed heterozygosity (0.15; 0.28) and Melrose contained the lowest value of expected heterozygosity and observed heterozygosity (0.09; 0.19). There was no significant trend in isolation by distance despite there being a positive correlation between geographic and genetic distance between pairs of populations (r = 0.48, p = 0.16).
Relatedness within populations
The summary statistics of pairwise relatedness outcomes indicated high relatedness within each population (mean r = 0.89 ± 0.03 SD) (Table 3). Melrose contained the highest relatedness (population mean r = 0.96), and Rusty Cab demonstrated the lowest relatedness (population mean r = 0.85) between individuals (Table 3). Although all FIS results produced a similar range of values (mean = -0.82 ± 0.04 SD) (Table 3) these values were not significant (p = 1). Four of the populations contained only two clones of the 15 individuals sampled. Telowie had slightly more clones (5), but Melrose consisted entirely of clones (Table 3).
Discussion
This study confirms that strong genetic structure is present among all currently known extant populations of spiny daisy with little to no evidence of inter-population gene flow. This indicates that each site represents a unique subset of genetic variation of this species. The inclusion of Telowie and Melrose populations has increased the amount of known genetic diversity that exists within the species by approximately 33%, however the lack in connectivity among these isolated populations reinforces the extinction risk faced by this species (Ellstrand and Elam 1993; Young et al. 1996; Spielman et al. 2004; Frankham and Wilcken 2006). Although relatedness within each population was high, only Melrose represented a single genet. All other populations contained a small number of clones. Our findings therefore differ to that of previous genetic studies that described each population to consist entirely of clones (Jusaitis and Adams 2005). Levels of observed heterozygosity were greater than expected heterozygosity, likely as a result of high levels of clonal reproduction and/or mechanisms preventing sexual reproduction. Our study helps improve the knowledge of the distribution of limited genetic diversity within this critically endangered species and demonstrates that the spiny daisy has likely experienced strong genetic bottlenecks (Rodger et al. 2021), probably due to large-scale habitat disturbance (Jusaitis and Adams 2005; Clarke et al. 2013; Brown and Hodgkin 2015). The varying levels of genetic diversity detected among populations and subtle differences in population structure provide a deeper insight towards prioritization strategies for conservation management (Gardiner et al. 2017).
The presence of sporophytic self-incompatibility (SI) is present in many Asteraceae species and serves as an evolutionary mechanism to promote outcrossing within populations and restrict the build-up of deleterious alleles which is often associated with inbreeding in small, isolated populations (Zagorski et al. 1983; Les et al. 1991; Ferrer and Good-Avila 2007). Given the absence of excess homozygosity and sexual reproduction, high relatedness within the spiny daisy populations may reflect increased clonality, whereby the majority of the population represents descendants from a single lineage (Young and Brown 1999). Compatibility of the pollen grain is typically determined by the diploid genotype at the S locus of the paternal individual, inhibiting fertilization between individuals who contain the same S-alleles. The mating system within these spiny daisy populations may be attributed to this mating system (Frankel and Galun 1977; DeMauro 1993). Species which exhibit self-incompatibility often demonstrate limited reproductive performance, particularly via inhibition of the stigma, pollen tube growth and reduced seed set (Lloyd 1968; Oloumi and Rezanejhad 2009).
The co-occurrence between SI and clonality is common among small populations as it helps compensate for limited seed production and reduces selection pressure which favours the breakdown of SI, thus preventing inbreeding (Vallejo-Marín 2007; Franklin-Tong 2008; DeMauro 1993) demonstrated that the partial-SI within the lakeside daisy (Hymenoxys acaulis) can vary depending on the degree of relatedness between mates (DeMauro 1993). She found that although selfing resulted in incompatibility, mating’s between full siblings and between parent and F1 progeny were able to produce seed (DeMauro 1993). A similar incidence may have occurred within the spiny daisy. Jusaitis and Adams (2005) reported that seed collected and artificially raised from Hart contained similar genetic composition to that of their genetically identical parents; however the offspring differed among 2–5 loci and these changes did not contain new alleles (Jusaitis and Adams 2005). The increased resolution of data obtained within this study shows that members within Hart are not all identical, therefore, it is possible that individuals which produce seed may in fact contain different S alleles allowing them to interbreed, although this would be rare due to low genetic diversity (DeMauro 1993). A lack in observed seedling recruitment within the wild due to either SI, inadequate environmental conditions or reduced pollinator activity could each contribute towards an absence of historical sexual reproduction (Jusaitis and Adams 2005).
Although there was no evidence of inter-population gene flow, Thornlea, Rusty Cab and Yangya sites contained lower levels of pairwise genetic differentiation, higher levels of genetic diversity and fewer numbers of clones than Melrose and Telowie. A possible explanation for this is that these populations have been isolated for fewer generations and/or have experienced occasional sexual reproduction in the past, which would reduce genetic drift (Young et al. 2002; Milton et al. 2004). As these three sites occur proximal to each other, and face exposure to similar environmental conditions, increased genetic diversity may reflect historic opportunistic gene flow followed by genetic drift upon being isolated (Milton et al. 2004). Increased spatial separation among populations may have led to increased rates of clonality within populations as a result of limited mate availability. Changes in reproductive strategy from a sexual to vegetative mode in response to patchy geographic distribution has been observed in other Asteraceae species in Australia (Young et al. 2002; Blyth et al. 2021). For example, populations of Rutidosis leiolepis that occur in higher altitudes have been reported to contain greater levels of clonality and population genetic differentiation than those at lower altitudes. Reductions in pollinator activity within these areas, altered length of flowering season, as well as increased disturbance events were considered possible reasons towards increased clonality (Young et al. 2002). Variation in the level of clonality observed among spiny daisy populations may therefore have been influenced by changes in historic connectivity. High intra-population relatedness with strong differentiation, particularly within Melrose and Telowie, indicate effects of genetic drift are likely due to higher rates of cloning (Campbell and Husband 2005).
Clonal reproduction serves as an advantageous strategy within challenging landscapes as it increases population size and maintains fitness within generations due to the repetition of heterozygous genotypes (Young et al. 1999; Stoeckel et al. 2006; Navascués et al. 2010). Clonality can allow for the sharing of resources (such as water, carbohydrates, minerals and photosynthate) among ramets and this competitive advantage can act as a vital tool for population colonisation (Alpert 1996; Pennings and Callaway 2000; Stuefer et al. 2004; Jusaitis and Adams 2005; Donahue and Lee 2008). Our study shows that the extent of clonality and relatedness varies among populations with Melrose consisting entirely of clones. It may be possible that this population has been founded more recently that the other populations, resulting in low genetic diversity in the form of somatic mutations (Stoeckel et al. 2006; de Meeûs et al. 2007; Wang et al. 2018). Asexual reproduction during unfavourable conditions such as when mates or pollinators are limited or during periods of increased environmental stress, can serve as a short-term, cost-effective mechanism to increase population size as it assures reproduction (Wang et al. 2018). The most clonal populations: Telowie and Melrose (respectively containing 33% and 100% clones), also represent the most northern sites. The Melrose population may be at a greatest risk of extinction as it might be at the edge of the climatic limit for this species and does not have the genetic variability to adapt to a changing environment (Dodd and Douhovnikoff 2016). Assisted migration can help retain genetic diversity, and this is important should the original site become further degraded or locally extinct (Clarke et al. 2013; Blyth et al. 2021). The translocation of individuals from all populations (via vegetative cuttings), including Melrose into areas more similar to the expected range of climatic suitability such as at Banrock Station has been carried out (Bickerton et al. 2018) and this serves as a solution towards preventing the loss of species genetic diversity, especially if the original populations become locally extinct. Increased growth rates, flowering and seed set within this population were experienced when plants gained access to improved habitat quality (increased watering and soil tillage), suggesting that the plants may be better suited towards environments which contain greater similarity to that of their historic distribution (Eckert 2002; Sharp et al. 2010; Bickerton and Tourenq 2015; Bickerton et al. 2018). Currently there is a study analysing the genetic diversity within these seedlings to determine whether sexual reproduction has occurred within the species. Further studies which examine the species ecological requirements and assess genetic composition within progeny will lead to opportunities that reintroduce gene flow among populations.
Conservation implications and management recommendations
Our study helps improve the knowledge of genetic structure within the critically endangered spiny daisy, demonstrating that it is likely experiencing strong genetic drift probably due to processes that limit sexual reproduction (Sharp et al. 2010; Clarke et al. 2013; Bickerton et al. 2018; Rodger et al. 2021). However, as the evolutionary resilience of a species relies upon adequate levels of genetic diversity which can only be achieved through successful interbreeding, and our study therefore raises concerns about its adaptability and we call for updated conservation management interventions (Bijlsma et al. 2000; Brook et al. 2002; Young et al. 2002; Blambert et al. 2016). We urge the implementation of strategies which enhance reproductive fitness to facilitate sexual reproduction and encourage seed set as the initial step towards species recovery. Although preliminary hand pollination experiments between Hart and Thornlea have been trialled, limited seed viability has been a notable hurdle towards successful germination (Jusaitis and Adams 2005; Sharp et al. 2010; Clarke et al. 2013; Bickerton et al. 2018). Updated crosses among individuals from each population (propagated from vegetative cuttings) would help determine the severity of this problem and explore reproductive compatibility among populations.
Common garden trials enable conservation biologists to measure fitness in parental as well as outcrossed progeny, evaluating the effects and risks of outbreeding depression in a controlled setting (Ottewell et al. 2016; Martín-Forés et al. 2018; Blyth et al. 2020, 2021; Van Rossum and Le Pajolec 2021). These trials have been increasingly used as an effective management approach to re-introduce genetic diversity within clonal, highly related and moderately differentiated in-situ populations as well as guide the design of ex-situ translocation populations (Ellstrand 2014; Ottewell et al. 2016; Blyth et al. 2021; Brunton et al. 2021; Gavin-Smyth et al. 2021). We advise the use of common garden experiments to validate the benefits of introducing novel genotypes within in-situ and ex-situ populations. In addition to habitat suitability, many studies have shown that the composition of genetic diversity of founder individuals acts as a primary driver towards successful early establishment of translocation sites (Schäfer et al. 2020; Van Rossum and Le Pajolec 2021). The increased probability of containing pre-adapted genotypes, as well as the reduced likelihood of bottleneck effects, should improve its evolutionary potential (Gamfeldt and Källström 2007; Breed et al. 2019). The variation of population genetic diversity detected within this study can direct ex-situ conservation efforts by determining sampling intensities within populations for the purpose of genetic rescue (Gardiner et al. 2017; Rodger et al. 2021). As each population represents a subset of unique genetic diversity, representatives from each locality should be used for outcrossing experiments as this will maximise genetic diversity within the progeny. For collection purposes, the intensity of sampling within sites needs to be evaluated depending on the level of genetic diversity within each site (Greenfield et al. 2016; Gardiner et al. 2017; Rossetto et al. 2021). Extensive sampling within clonal sites such as Melrose will help increase population size, however, it may pose an unnecessary use of resources e.g. if sampling is too intense, as it will likely under-represent the genetic diversity of the species (Broadhurst et al. 2008; Gardiner et al. 2017). Instead, pollen from one individual in Melrose could be used to fertilise several individuals in other populations.
Conclusion
Our study has shown that the spiny daisy is at high risk of extinction due to its limited genetic diversity which has likely been influenced by habitat loss. Each population contained (i) strong genetic structure, (ii) low levels of genetic diversity and (iii) little to no evidence of gene flow. Although excessive homozygosity was not detected, high relatedness among individuals suggests this species may be unlikely to increase its genetic diversity without intervention as populations remain isolated. Further studies which examine (i) reproductive compatibility between populations via hand pollination experiments and (ii) species ecological requirements will help provide insight into strategies to improve genetic diversity in addition to facilitating species restoration. Results from common garden trials will also help elucidate the risks and benefits of initiating gene flow among differentiated populations. Our study highlights the urgency of genetic rescue for the spiny daisy and is a guide to develop conservation management strategies which will increase evolutionary resilience within the species.
Data Availability
Data will be made available in an open access repository once our paper is published. https://doi.org/10.25451/flinders.20380362
References
Alpert P (1996) Nutrient Sharing in Natural Clonal Fragments of Fragaria Chiloensis. J Ecol 84:395–406
Badouin H, Gouzy J, Grassa CJ, Murat F, Staton SE, Cottret L, Lelandais-Brière C, Owens GL, Carrère S, Mayjonade B, Legrand L, Gill N, Kane NC, Bowers JE, Hubner S, Bellec A, Bérard A, Bergès H, Blanchet N, Boniface MC, Brunel D, Catrice O, Chaidir N, Claudel C, Donnadieu C, Faraut T, Fievet G, Helmstetter N, King M, Knapp SJ, Lai Z, Le Paslier MC, Lippi Y, Lorenzon L, Mandel JR, Marage G, Marchand G, Marquand E, Bret-Mestries E, Morien E, Nambeesan S, Nguyen T, Pegot-Espagnet P, Pouilly N, Raftis F, Sallet E, Schiex T, Thomas J, Vandecasteele C, Varès D, Vear F, Vautrin S, Crespi M, Mangin B, Burke JM, Salse J, Muños S, Vincourt P, Rieseberg LH, Langlade NB (2017) The sunflower genome provides insights into oil metabolism, flowering and Asterid evolution. Nature 546:148–152
Beissinger TM, Hirsch CN, Sekhon RS, Foerster JM, Johnson JM, Muttoni G, Vaillancourt B, Buell CR, Kaeppler SM, de Leon N (2013) Marker Density and Read Depth for Genotyping Populations Using Genotyping-by-Sequencing. Genetics 193:1073
Bickerton D, Field T, Moritz K, Tourenq C (2018) Threatened plant translocation case study: Acanthocladium dockeri (Spiny Daisy), Compositae. Australasian Plant Conservation 26:6–9
Bickerton D, Tourenq C (2015) Translocating the Spiny Daisy: Partnerships with winning outcomes. Australian Network for Plant Conservation Inc
Bijlsma R, Bundgaard J, Boerema A (2000) Does inbreeding affect the extinction risk of small populations?: predictions from Drosophila. J Evol Biol 13:502–514
Blambert L, Mallet B, Humeau L, Pailler T (2016) Reproductive patterns, genetic diversity and inbreeding depression in two closely related Jumellea species with contrasting patterns of commonness and distribution. Ann Bot 118:93–103
Blyth C, Christmas MJ, Bickerton DC, Breed MF, Foster NR, Guerin GR, Mason ARG, Lowe AJ (2021) Genomic, Habitat, and Leaf Shape Analyses Reveal a Possible Cryptic Species and Vulnerability to Climate Change in a Threatened Daisy. Life, p 11
Blyth C, Christmas MJ, Bickerton DC, Faast R, Packer JG, Lowe AJ, Breed MF (2020) Increased Genetic Diversity via Gene Flow Provides Hope for Acacia whibleyana, an Endangered Wattle Facing Extinction. Divers (Basel) 12:299
Breed MF, Harrison PA, Blyth C, Byrne M, Gaget V, Gellie NJC, Groom SVC, Hodgson R, Mills JG, Prowse TAA, Steane DA, Mohr JJ (2019) The potential of genomics for restoring ecosystems and biodiversity. Nat Rev Genet 20:615–628
Broadhurst LM, Lowe A, Coates DJ, Cunningham SA, McDonald M, Vesk PA, Yates C (2008) Seed supply for broadscale restoration: maximizing evolutionary potential. Evol Appl 1:587–597
Brook BW, Tonkyn DW, O’Grady JJ, Frankham R (2002) Contribution of inbreeding to extinction risk in threatened species. Conservation Ecology, p 6
Brown AHD, Hodgkin T (2015) Indicators of Genetic Diversity, Genetic Erosion, and Genetic Vulnerability for Plant Genetic Resources. Cham: Springer International Publishing, Cham. p.^pp. 25–53
Brunton A, Lee DJ, Conroy G, Ogbourne S (2021) Clonal reproduction and low genetic diversity in northern australian santalum lanceolatum (Santalaceae) populations highlights the need for genetic rescue of this commercially significant species. Forests 12:741
Buza L, Young A, Thrall P (2000) Genetic erosion, inbreeding and reduced fitness in fragmented populations of the endangered tetraploid pea Swainsona recta. Biol Conserv 93:177–186
Campbell LG, Husband BC (2005) Impact of clonal growth on effective population size in Hymenoxys herbacea (Asteraceae). Heredity 94:526–532
Clarke A, Robertson MA, Pieck A. (2013) Recovery Plan for Acanthocladium dockeri (Spiny Daisy). (ed. Department of Environment WaNR), Clare, South Australia
de Meeûs T, Prugnolle F, Agnew P (2007) Asexual reproduction: Genetics and evolutionary aspects. Cell Mol Life Sci 64:1355–1372
DeMauro MM (1993) Relationship of Breeding System to Rarity in the Lakeside Daisy (Hymenoxys acaulis var. glabra). Conserv Biol 7:542–550
Dodd RS, Douhovnikoff V (2016) Adjusting to Global Change through Clonal Growth and Epigenetic Variation. Frontiers in Ecology and Evolution, 4
Donahue MJ, Lee CT (2008) Colonization. In: Fath B (ed) Encyclopedia of Ecology (Second Edition). Elsevier, Oxford, pp 42–47
Eckert CG (2002) The loss of sex in clonal plants. Ecology and evolutionary biology of clonal plants.Springer. p.^pp.279–298
Ellstrand N, Elam D (1993) POPULATION GENETIC CONSEQUENCES OF SMALL POPULATION SIZE: Implications for Plant Conservation. Annu Rev Ecol Evol Syst 24:217–242
Ellstrand NC (2014) Is gene flow the most important evolutionary force in plants? Am J Bot 101:737–753
Ferrer MM, Good-Avila SV (2007) Macrophylogenetic Analyses of the Gain and Loss of Self-Incompatibility in the Asteraceae. New Phytol 173:401–414
Frankel R, Galun E (1977) Pollination Mechanisms, Reproduction and Plant Breeding. Springer Berlin / Heidelberg, Berlin, Heidelberg, Berlin, Heidelberg
Frankham R, Wilcken J (2006) Does inbreeding distort sex-ratios? Conservation genetics, 7, 879–893
Franklin-Tong VE (2008) Self-Incompatibility in Flowering Plants: Evolution, Diversity, and Mechanisms, 1. Aufl. edn. Berlin, Heidelberg: Springer-Verlag, Berlin, Heidelberg
Frichot E, François O, O’Meara B (2015) LEA: An R package for landscape and ecological association studies. Methods Ecol Evol 6:925–929
Furlan E, Stoklosa J, Griffiths J, Gust N, Ellis R, Huggins RM, Weeks AR (2012) Small population size and extremely low levels of genetic diversity in island populations of the platypus, Ornithorhynchus anatinus. Ecol Evol 2:844–857
Gamfeldt L, Källström B (2007) Increasing Intraspecific Diversity Increases Predictability in Population Survival in the Face of Perturbations. Oikos 116:700–705
Gardiner LM, Rakotoarinivo M, Rajaovelona LR, Clubbe C (2017) Population genetics data help to guide the conservation of palm species with small population sizes and fragmented habitats in Madagascar. PeerJ, 2017, e3248-e3248
Gaudeul M, Till-Bottraud I, Barjon F, Manel S (2004) Genetic diversity and differentiation in Eryngium alpinum L. (Apiaceae): comparison of AFLP and microsatellite markers. Heredity 92:508–518
Gavin-Smyth N, Kramer AT, Urbina-Casanova R, Vitt P, Fant JB (2021) Genetic rescue reduces mate limitation in a threatened, clonal, and self-incompatible plant species.Restoration Ecology, 29, e13458
Greenfield A, McPherson H, Auld T, Delaney S, Offord CA, van der Merwe M, Yap J-YS, Rossetto M (2016) Whole-chloroplast analysis as an approach for fine-tuning the preservation of a highly charismatic but critically endangered species, Wollemia nobilis (Araucariaceae). Aust J Bot 64:654
Gruber B, Unmack PJ, Berry OF, Georges A (2018) dartr: An r package to facilitate analysis of SNP data generated from reduced representation genome sequencing. Mol Ecol Resour 18:691–699
Hajjar R, Jarvis DI, Gemmill-Herren B (2008) The utility of crop genetic diversity in maintaining ecosystem services. Agric Ecosyst Environ 123:261–270
Heywood JS (1993) Biparental Inbreeding Depression in the Self-Incompatible Annual Plant Gaillardia pulchella (Asteraceae). Am J Bot 80:545–550
Huber CD, DeGiorgio M, Hellmann I, Nielsen R (2016) Detecting recent selective sweeps while controlling for mutation rate and background selection. Mol Ecol 25:142–156
Jaccoud D, Peng K, Feinstein D, Kilian A (2001) Diversity arrays: a solid state technology for sequence information independent genotyping. Nucleic Acids Res 29:E25–E25
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405
Jones OR, Wang J (2010) COLONY: a program for parentage and sibship inference from multilocus genotype data. Mol Ecol Resour 10:551–555
Jusaitis M (2008) Flowering and seed production in the endangered Spiny Daisy, Acanthocladium dockeri. Australasian Plant Conservation 17:14–15
Jusaitis M, Adams M (2005) Conservation implications of clonality and limited sexual reproduction in the endangered shrub Acanthocladium dockeri (Asteraceae). Aust J Bot 53:535–544
Kane N, Gill N, King M, Bowers J, Bergès H, Gouzy J, Bachlava E, Langlade N, Lai Z, Stewart M, Burke J, Vincourt P, Knapp S, Rieseberg L (2011) Progress towards a reference genome for sunflower. Botany 89:429–437
Kilian A, Wenzl P, Huttner E, Carling J, Xia L, Blois H, Caig V, Heller-Uszynska K, Jaccoud D, Hopper C, Aschenbrenner-Kilian M, Evers M, Peng K, Cayla C, Hok P, Uszynski G (2012) Diversity arrays technology: a generic genome profiling technology on open platforms. Methods Mol Biol 888:67–89
Les DH, Reinartz JA, Esselman EJ (1991) Genetic consequences of rarity in Aster furcatus (Asteraceae), a threatened, self-incompatible plant. Evolution 45:1641–1650
Lloyd DG (1968) Pollen Tube Growth and Seed Set in Self-Incompatible and Self-Compatible Leavenworthia (Cruciferae) Populations. The New Phytologist, 67, 179–195
Martín-Forés I, Acosta-Gallo B, Castro I, de Miguel JM, Del Pozo A, Casado MA (2018) The invasiveness of Hypochaeris glabra (Asteraceae): Responses in morphological and reproductive traits for exotic populations. PLoS ONE 13:e0198849–e0198849
Melville J, Haines ML, Boysen K, Hodkinson L, Kilian A, Date KLS, Potvin DA, Parris KM (2017) Identifying hybridization and admixture using SNPs: application of the DArTseq platform in phylogeographic research on vertebrates. R Soc Open Sci 4:161061–161061
Milton SJ, Dean WRJ, Leuteritz TEJ (2004) Opportunistic and multiple breeding attempts in plants and vertebrates of semi-deserts with unpredictable rainfall events through the year. Trans Royal Soc South Afr 59:43–53
Moreira PA, Oliveira D (2011) Leaf age affects the quality of DNA extracted from Dimorphandra mollis (Fabaceae), a tropical tree species from the Cerrado region of Brazil, vol 10. GMR, Genetics and molecular research, pp 353–358
Moritz C (1999) Conservation units and translocations. Strategies for conserving evolutionary processes. Hereditas 130:217–228
Narum SR, Buerkle CA, Davey JW, Miller MR, Hohenlohe PA (2013) Genotyping-by-sequencing in ecological and conservation genomics. Mol Ecol 22:2841–2847
Neaves LE, Eales J, Whitlock R, Hollingsworth PM, Burke T, Pullin AS (2015) The fitness consequences of inbreeding in natural populations and their implications for species conservation – a systematic map. Environ Evid 4:5
O’Brien SJ, Evermann JF (1988) Interactive influence of infectious disease and genetic diversity in natural populations. Trends Ecol Evol 3:254–259
Oloumi H, Rezanejhad F (2009) Response of pollen tube growth and seed set to controlled pollination and their relation to self-incompatibility in different cultivars of Petunia hybrida. Grana 48:102–108
Ottewell KM, Bickerton DC, Byrne M, Lowe AJ (2016) Bridging the gap: a genetic assessment framework for population-level threatened plant conservation prioritization and decision-making. Divers Distrib 22:174–188
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Pembleton LW, Cogan NOI, Forster JW (2013) StAMPP: an R package for calculation of genetic differentiation and structure of mixed-ploidy level populations. Mol Ecol Resour 13:946–952
Pennings SC, Callaway RM (2000) The Advantages of Clonal Integration under Different Ecological Conditions: A Community-Wide Test. Ecol (Durham) 81:709–716
Rodger YS, Pavlova A, Sinclair S, Pickup M, Sunnucks P (2021) Evolutionary history and genetic connectivity across highly fragmented populations of an endangered daisy. Heredity 126:846–858
Rossetto M, Yap J-YS, Lemmon J, Bain D, Bragg J, Hogbin P, Gallagher R, Rutherford S, Summerell B, Wilson TC (2021) A conservation genomics workflow to guide practical management actions.Global ecology and conservation, 26, e01492
Rousset F (1997) Genetic Differentiation and Estimation of Gene Flow from F-Statistics Under Isolation by Distance. Genetics 145:1219–1228
Sansaloni C, Petroli C, Jaccoud D, Carling J, Detering F, Grattapaglia D, Kilian A (2011) Diversity Arrays Technology (DArT) and next-generation sequencing combined: genome-wide, high throughput, highly informative genotyping for molecular breeding of Eucalyptus. BMC Proceedings, 5, P54
Schäfer D, Vincent H, Fischer M, Kempel A (2020) The importance of genetic diversity for the translocation of eight threatened plant species into the wild.Global Ecology and Conservation, 24, e01240
Seeb JE, Carvalho G, Hauser L, Naish K, Roberts S, Seeb LW (2011) Single-nucleotide polymorphism (SNP) discovery and applications of SNP genotyping in nonmodel organisms. Mol Ecol Resour 11(Suppl 1):1–8
Sharp A, Clarke A, Jusaitis M, Pieck A, Slattery P, Potter D (2010) Translocations of the critically endangered spiny daisy in the Mid-north of South Australia. p.^pp.340–344
Spielman D, Brook BW, Frankham R (2004) Most species are not driven to extinction before genetic factors impact them. Proc Natl Acad Sci USA 101:15261–15264
Stoeckel S, Grange J, FernÁNdez-Manjarres JF, Bilger I, Frascaria-Lacoste N, Mariette S (2006) Heterozygote excess in a self-incompatible and partially clonal forest tree species -Prunus avium L. Mol Ecol 15:2109–2118
Stuefer JF, Gomez Garcia SE, Molken MARv (2004) Clonal integration beyond resource sharing: implications for defence signalling and disease transmission in clonal plant networks. Evol Ecol 18:647–667
Templeton AR, Shaw K, Routman E, Davis SK (1990) The genetic consequences of habitat fragmentation. Ann Mo Bot Gard 77:13–27
The International HapMap (2005) A haplotype map of the human genome. Nature 437:1299–1320
Vallejo-Marín M (2007) The paradox of clonality and the evolution of self-incompatibility. Plant Signal Behav 2:265–266
van Deventer R, Rhode C, Marx M, Roodt-Wilding R (2020) The development of genome-wide single nucleotide polymorphisms in blue wildebeest using the DArTseq platform. Genomics 112:3455–3464
Van Rossum F, Le Pajolec S (2021) Mixing gene pools to prevent inbreeding issues in translocated populations of clonal species. Mol Ecol 30:2756–2771
van Treuren R, Bijlsma R, van Delden W, Ouborg NJ (1991) The significance of genetic erosion in the process of extinction. I. Genetic differentiation in Salvia pratensis and Scabiosa columbaria in relation to population size. Heredity 66:181–189
Wang J (2011) coancestry: a program for simulating, estimating and analysing relatedness and inbreeding coefficients. Mol Ecol Resour 11:141–145
Wang Z, Xie L, Prather CM, Guo H, Han G, Ma C (2018) What drives the shift between sexual and clonal reproduction of Caragana stenophylla along a climatic aridity gradient? BMC Plant Biol 18:91
Weir B, Cockerham C, Weir BS, Cockerham CC (1984) Estimating F-Statistics for the Analysis of Population-Structure. Evolution 38: 1358–1370. Evolution, 38, 1358–1370
Wenzl P, Carling J, Kudrna D, Jaccoud D, Huttner E, Kleinhofs A, Kilian A (2004) Diversity Arrays Technology (DArT) for whole-genome profiling of barley. Proceedings of the National Academy of Sciences, 101, 9915–9920
Williams AV, Nevill PG, Krauss SL (2014) Next generation restoration genetics: applications and opportunities. Trends Plant Sci 19:529–537
Würschum T, Langer SM, Longin CFH, Korzun V, Akhunov E, Ebmeyer E, Schachschneider R, Schacht J, Kazman E, Reif JC (2013) Population structure, genetic diversity and linkage disequilibrium in elite winter wheat assessed with SNP and SSR markers. Theor Appl Genet 126:1477–1486
Young A, Boyle T, Brown T (1996) The population genetic consequences of habitat fragmentation for plants. Trends Ecol Evol 11:413–418
Young A, Brown A (1999) Paternal bottleneck in fragmented populations of the grassland daisy Rutidosis leptorrhynchoides. Genet Res 73:117–117
Young AG, Hill JH, Murray BG, Peakall R, Muell (2002)(Asteraceae). Biological Conservation, 106,71–78
Zagorski JS, Ascher PD, Widmer RE (1983) Multigenic self incompatibility in hexaploid Chrysanthemum. Euphytica 32:1–7
Acknowledgements
We would like to thank Trees for Life for their assistance with this study.
Funding
This study was partially funded by a Flinders University Honours scholarship.
Open Access funding enabled and organized by CAUL and its Member Institutions
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors do not have any conflicts of interest to declare.
>Author declaration
We declare that the content of this article is being submitted only to Conservation Genetics and has not been previously published elsewhere. Furthermore, this manuscript does not contain material which has been published by or plagiarised from other sources, using references to accredit known sources for where the information was obtained.
Permit to conduct research
Permission to conduct research was given by the Government of South Australia, Department for Environment and Water, permit number E26945-1.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Brooks, E., Slender, A.L., Cu, S. et al. A range-wide analysis of population structure and genomic variation within the critically endangered spiny daisy (Acanthocladium dockeri). Conserv Genet 23, 1027–1037 (2022). https://doi.org/10.1007/s10592-022-01468-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-022-01468-0