Abstract
Habitat deterioration and fragmentation increase the risk of wildlife extirpation as they have strong impacts on population size, genetic diversity and gene flow. Small populations are more susceptible to these factors because the loss of genetic diversity by drift and inbreeding occurs at faster rates. Therefore, estimates of genetic diversity and population sizes of threatened and small wildlife populations in deteriorated landscapes are critical for managing and conservation. Here, we used a non-invasive sampling approach in combination with eleven microsatellite loci to evaluate genetic diversity, genetic structure, and demographic parameters of the Neotropical otter (Lontra longicaudis) in three river basins (Actopan, La Antigua and Jamapa), which are priority conservation areas for Veracruz, Mexico. Our results revealed moderate genetic diversity and genetic structure among river basins. However, we detected first-generation migrants among basins, suggesting current gene flow. Effective population size for each basin was considerably lower than the value (Ne < 100) suggested to maintain genetic variation of populations in the short-term. Similarly, census population size was lower than estimates reported for L. longicaudis in a conserved region in Mexico. We did not find evidence of recent genetic bottlenecks for any basin. Our genetic and demographic results suggest that L. longicaudis in the three river basins could be experiencing genetic isolation and erosion, with La Antigua being the most vulnerable basin. Natural fragmentation and habitat deterioration seem to be shaping the observed patterns of genetic variation in the Neotropical otter.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genetic diversity is essential for the adaptation of natural populations to changes in their landscape and environment. Populations that have suffered genetic bottlenecks or that have low genetic diversity are more vulnerable to extirpation as they have less potential to cope with changes resulting from climate change and habitat fragmentation (Crow and Kimura 1970; Frankham et al. 2017). Smaller populations are more susceptible to the erosion of genetic diversity because genetic drift and inbreeding occur at faster rates (Frankham 1996; Franklin and Frankham 1998; Allendorf et al. 2013). Several empirical studies in plants and animals have shown that population size is positively correlated with genetic diversity (e.g., Soulé 1976; Frankham 1996; Hague and Routman 2016).
Habitat loss and fragmentation can have strong impacts on population size, genetic diversity and gene flow (Frankham et al. 2002; Manel et al. 2003; Keyghobadi 2007; Frankham 2010; Cushman et al. 2013). While habitat loss mainly reduces the resources and suitable conditions for the reproduction and growth of individuals (Vergeer et al. 2003; Morris et al. 2008), habitat fragmentation can reduce dispersal and consequently gene flow among remnant populations (Manel et al. 2003; Frankham et al. 2017). The effects of habitat loss and fragmentation are expected to be stronger in threatened species because they are characterized by having several generations of reduced population size and genetic diversity (Spielman et al. 2004; Palstra and Ruzzante 2008; Frankham et al. 2015). Therefore, the estimation of genetic diversity and population sizes in threatened species inhabiting fragmented landscapes are imperative.
The identification of bottlenecks, as well as the estimation of census (Nc) and effective (Ne) population sizes allow generating essential genetic and demographic information, which is key for developing conservation and management plans. Genetic bottlenecks reduce the possibility of survival of individuals from populations that have been subjected to significant declines (Peery et al. 2012). Nc is the number of living individuals in a population, and it is an important demographic parameter for evaluating the conservation status of a species (Knaepkens et al. 2004; Hague and Routman 2016). Ne provides information on the size of an ideal population that would lose genetic diversity as the result of inbreeding or genetic drift, at the same rate as the population under consideration (Wright 1940). The estimation of Ne is essential to warrant the maintenance of genetic diversity and the evolutionary potential of populations (Bouzat 2010).
The Neotropical otter, Lontra longicaudis, is an ecological important species (i.e. top predator and umbrella species) that inhabits freshwater ecosystems. Although L. longicaudis has a wide distribution from northern Mexico to northern Argentina (Larivière 1999), it is classified as a “near threatened” species by the International Union for the Conservation of Nature (IUCN), mainly because of the decrease in its population sizes. Between 1950 and 1970, L. longicaudis was hunted for fur trade (Rheingantz and Trinca 2015), which dramatically reduced its population sizes. A further population reduction of 25% has been estimated to occur in the next 30 years (Pacifici et al. 2013).
In Mexico, L. longicaudis is classified as a threatened species due to its habitat deterioration mainly by pollution, fisheries overexploitation and deforestation (NOM-059 SEMARNAT 2010; Gallo-Reynoso 2007). Despite these threats, population sizes (Ne and Nc) of L. longicaudis have not yet been estimated in Mexico, and there are few studies that have evaluated the genetic diversity and population structure in the species (Ortega et al. 2012; Guerrero et al. 2015; Hernández et al. 2017). Most studies of L. longicaudis in Mexico have estimated abundance employing indirect counting methods (i.e. spraints) (Larivière 1999; Arellano et al. 2012, González-Christen et al. 2013) and more recently molecular markers (Ortega et al. 2012). Only one study has assessed genetic variation using microsatellites, finding moderate levels of genetic diversity (Ho = 0.55), lack of genetic structure, and a maximum displacement of individuals of 17 km in a conserved river system in Chiapas (Ortega et al. 2012). Studies in South American populations of L. longicaudis reported low to high genetic diversity (Ho = 0.322—0.83; Weber et al. 2009, Trinca et al. 2013, 2016). Contemporary genetic structure, gene flow and migration rates have not yet been assessed for L. longicaudis, but for the Eurasian otter (Lutra lutra), studies suggest that gene flow among basins is restricted by physical characteristics of the basins, such as slope and altitude (Janssens et al. 2008; Pagacz 2016).
The main objectives of this study were to estimate genetic diversity and demographic parameters of the Neotropical otter in three river basins in central Veracruz, Mexico: Actopan, La Antigua, and Jamapa. These river basins are classified as highly human perturbed and are priority targets for conservation (Cotler-Ávalos et al. 2010). Specifically, we used non-invasive sampling and eleven microsatellite loci to: (i) evaluate genetic diversity of the Neotropical otter in each basin; (ii) identify patterns of genetic structure within and among basins; (iii) detect contemporary migration events among basins; and (iv) estimate population sizes and test for evidence of genetic bottlenecks. Because the three basins have been subject to a high degree of human disturbance, we expect to find low genetic diversity and small population sizes. Similarly, we anticipate finding strong genetic structure and low contemporary migration rates among basins due to habitat deterioration and to the hierarchical organization of the rivers, which creates a natural fragmented habitat.
Materials and methods
Study area
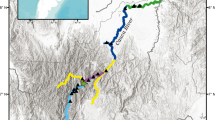
We conducted the study in three river basins of the state of Veracruz in Mexico (Fig. 1). Actopan is the smallest of the river basins, with an area of 1999 km2, followed by La Antigua consisting of 2190 km2 and Jamapa of 3918 km2. In the last decades, human population growth and changes in land use have altered the functional dynamics of the basins (e.g., hydrological filtration processes, evapotranspiration, sediment movement; Cotler et al. 2010). The main economic activities conducted along the river shores of the basins include fishing, livestock, and agriculture (INEGI 2011). Along these basins L. longicaudis is generally found from 0 to 1200 m a.s.l. (Macias-Sánchez and Aranda 1999; Latorre-Cardenas 2013 unpublished data, Hernández-Romero et al. 2017), but it has been reported at up to 2500 m a.s.l., probably due to the presence of aquaculture ponds at these higher altitudes (Hernández-Romero et al. 2018).
Map of the studied area including the rivers and tributaries of the three basins where the sampling of the Neotropical otter was conducted, and the location of the studied area within the state of Veracruz in Mexico (inset in the top right). Color dots in the map indicate the position of the samples of the three individuals that were capture and recapture. The insets on the right show the position of the samples of each capture-recapture individual and the distance between them is indicated
Sample collection
Due to the elusive behavior of Lontra longicaudis, we used a non-invasive approach for sample collection. We collected fresh spraints and anal glands secretions deposited in latrines along the main rivers and third or higher order tributaries of the Actopan, La Antigua and Jamapa basins (Fig. 1). We restricted sampling collection to third-order or higher rivers because they have enough water flow for the otter to inhabit them. We used a hydrological stream layer (hydrological network; INEGI 2016) to identify the rivers and tributaries to be surveyed, employing the spatial tools in ArcGIS v.10.2.1. (ESRI 2013).
We conducted sample collection by raft in the main rivers of Actopan and La Antigua, along 45 and 64 km, respectively. In order to detect as many individuals as possible, we carried out surveys on periods of two months from: May to June 2016, November to December 2016 and March to April 2018. Because of the low water level of the Jamapa river, we only performed one survey by raft or walking along the rivers, from November 2017 to February 2018, covering a total of 105 km. We performed all surveys in the morning to obtain fresh otter samples (Lerone et al. 2014), which were collected using sterile sticks, preserved in 1.25 ml of RNA later buffer (Sigma Aldrich), and stored at − 20 °C until genetic analyzes were performed. We registered the geographic coordinates of all collected samples.
We also included blood samples of six captive individuals from the Veracruz aquarium (“Acuario de Veracruz, A.C.”), which were rescued from Jamapa in the same time period of our field work. Veterinarians of the aquarium took blood samples following the ethical protocol “NOM-135-SEMARANT-2004” and the “Code of ethics and welfare animal of the World Association of Zoos and Aquariums (WAZA)”. We preserved blood samples in EDTA at − 20 °C until DNA extractions were performed.
DNA extraction and amplification
We extracted genomic DNA from anal secretions and feces using the ZR Fecal DNA MiniPrep kit (Zymo Research) and from blood samples with the DNeasy Blood & Tissue kit (Qiagen) following the manufacturer instructions. We amplified eleven microsatellite loci: two (RIO 2 and RIO11) previously designed for L. canadensis (Beheler et al. 2004, 2005) and nine (Lolo13, Lolo18, Lolo19, Lolo29, Lolo30, Lolo37, Lolo39, Lolo41 and Lolo48) developed for L. longicaudis (Latorre-Cardenas et al. 2020) using the polymerase chain reaction (PCR). We provide details on DNA extractions and PCR conditions in Latorre-Cardenas et al. (2020) and in Online Resource 1. We pooled PCR products of the eleven microsatellite markers in four multiplex panels, consisting of three loci each (Table S1). Fragment analyses were conducted in a 3730XL sequencer (Applied Biosystems) using 600 LIZ size standard (GeneScan™) at Macrogen, South Korea. We assigned allele size with the aid of the software GeneMapper 4.0 (Thermofisher).
Genotyping quality control
The low amount and quality of the DNA contained in anal secretions and feces could yield low quality of PCR reactions and could increase genotyping errors. Quality control of PCR reactions was assessed by verifying the genotypes of each sample using a multi-tube approach (Taberlet et al. 1996). This approach consists on repeating the PCR reactions of all samples using the same conditions to assure the same alleles are obtained in each replicate (Taberlet et al. 1996). We considered a sample to be heterozygote if the same two alleles were obtained in two identical repetitions of the PCR, and a sample was a homozygote if the same single allele was obtained in three PCR replicates. We assessed genotyping error per locus, based on the number of mismatches between resulting genotypes of the PCR replicates (Pompanon et al. 2005). We also used MICROCHECKER (Van Oosterhout et al. 2004) to determine the presence of large allelic dropout and null alleles.
Individual identification and microsatellite power of discrimination
When employing a non-invasive sampling approach, several samples of the same individual could be collected during the same or in different sampling sessions. We identified unique multilocus genotypes (i.e., individuals) using the ALLELEMATCH package (Galpern 2012) in R v.3.4.2. (R Core Team 2017). This package identifies unique multilocus genotypes in data sets in which the number of individuals is unknown, considering the genotyping error and missing data. We performed all subsequent analyses using the unique multilocus genotypes. To assesses the discriminating power of microsatellites in identifying individuals, we assessed the probability of identity (PID) and the probability of identity between siblings (PIDSIBS) (Waits et al. 2001) for each locus and combining all loci in GIMLET. PID calculates the probability that two unrelated individuals and PIDSIBS that two siblings drawn at random from a population will have the same genotype with a given set of loci.
Genetic diversity
To assess genetic diversity for each basin, we calculated rarefied allelic richness (AR) in FSTAT v.2.9.3.2. (Goudet 2001), rarefied private allelic richness (PAR) in HP-rare (Kalinowski 2004, 2005) and the effective number of alleles (ENA) in GENEALEX v.6.5.02 (Peakall and Smouse 2012). We also calculated observed (HO) and expected (HE) heterozygosity and significant deviations from Hardy–Weinberg equilibrium (HWE) by means of the inbreeding coefficient (FIS), with 10,000 permutations in FSTAT. We tested for significant differences in AR and HO among basins, using 1000 permutations and two-sided tests in FSTAT. To identify which of the basins differed, we performed pairwise comparisons. We also tested for significant differences in HE across basins by means of an ANOVA in R. We used FSTAT to assess linkage disequilibrium (LD) between all pairs of loci. We performed corrections for multiple comparisons with the sequential Bonferroni method (Rice 1989).
Population genetic structure
To assess genetic structure, we identified the most likely number of genetic clusters by employing a Bayesian clustering analysis in STRUCTURE (Pritchard et al. 2000). We used the admixture model with correlated allele frequencies. We ran 10 independent chains for each K (from K = 1 to K = 6). The length of the burn-in was 500,000 followed by 1,000,000 Markov Chain Monte Carlo iterations. We determined the most likely number of genetic clusters in STRUCTURE HARVESTER Web v.0.6.94 (Earl and VonHoldt 2012) using the ΔK statistic (Evano et al. 2005) and the likelihood of the data (ln P(D); Pritchard et al. 2000). We used the software CLUMP v.1.1.2 (Jakobsson and Rosenberg 2007) to average the results of the 10 replicates with the highest ΔK value, and DISTRUCT v.1.1 (Rosenberg 2004) to visualize the genetic clusters. In order to assess genetic substructure, we ran STRUCTURE separately in each of the genetic clusters detected by the initial analysis (see “Results”). We performed these subsequent analyses as described above.
We also performed a Discriminant Analysis of Principal Components (DAPC) in the ADEGENET package v.2.1.1 in R (Jombart et al. 2008) to further evaluate genetic structure. DAPC is a multivariate multilocus analysis designed to maximize genetic variance among groups, while minimizing within group variance (Jombart et al. 2008). DAPC does not assume HWE or LD (Jombart et al. 2008) and it is not sensitive to the presence of null alleles. We assessed the most likely number of clusters from K = 1 to K = 6, with the find.clusters function and selected the K with the lowest BIC. To identify the number of PCs to be retained, we used the dapcCross validation function that selects the optimal number of PCs based on the highest successful assignment with the lowest mean squared error (Jombart et al. 2010).
We calculated genetic differentiation between basins using pairwise FST comparisons and assessed their significance with 1000 permutations in FSTAT. We used the excluding null allele correction method to calculate pairwise FST (Chapuis and Estoup 2007) in FREENA v.1.0. This method prevents overestimation of FST values due to the presence of null alleles. To avoid bias associated to the high polymorphism characteristic of microsatellites, we calculated pairwise DEST (Jost 2008) in the DEMEtics package (Gerlach et al. 2010) implemented in R. We obtained confidence intervals and significance values by performing 1000 bootstrap replicates.
We estimated fine-scale genetic structure among basins using spatial autocorrelation analysis in the program GENEALEX. This analysis allows relating individual pairwise genetic distances, estimated through the autocorrelation coefficient (r) proposed by Smouse and Peakall (1999), with the geographical distances of individuals that fall within a specified distance class (Epperson 2005; Smouse et al. 2008). We used 10 km as the size distance class, 10,000 permutations and 1,000 bootstrap replicates to estimate 95% confidence intervals (CI) around r at each distance class.
Migration estimates
We used a Bayesian assignment method implemented in GENECLASS v.2.0 (Piry et al. 2004) to identify putative first-generation migrants (individuals that were not born in the sampled population), and their most probably population of origin. We used the L = L_home/L_max likelihood criteria and 1000 simulations (Rannala and Mountain 1997). This criterion is the ratio of the likelihood computed for the population where the individual was sampled (L_home) over the highest likelihood value among samples of all populations, including the population where the individual was sampled (L_max). We also employed the USEPOPINFO option and the MIGPRIOR set at 0.05 in STRUCTURE to identify individuals for which the population of origin was different to the one they were sampled from.
Population size and bottleneck estimations
We assessed population sizes by estimating Nc for each basin using the CAPWIRE package (Pennell et al. 2013) implemented in R. This method is based on the capture-recapture of multilocus genotypes collected from non-invasive genetic sampling, during single or multiple sampling sessions and performs well for small populations (< 100 individuals). CAPWIRE considers that individuals may be sampled multiple times per sampling session and uses the assumption of individual replacement. For this analysis, we considered all the performed surveys as a single sampling session. We first obtained the maximum likelihood estimate (MLE) of the population size, using the ‘equal capture model’ (ECM) and then estimated the ‘two-innate rates model’ (TIRM). The later model assumes heterogeneity in the capture probabilities that are related to sex, age and social status of the individuals (Miller et al. 2005). This is particularly important for otters as the scent-marking behavior differs among sexes and age, affecting the number of feces present in the studied area (Arrendal et al. 2007). To select between ECM and TIRM models, we conducted a likelihood ratio test (LRT) and performed 100 parametric bootstraps to estimate 95% confidence intervals for the MLE. We also estimated the density of otters by dividing the number of detected individuals by the total number of kilometers surveyed in each basin (Actopan: 45 km, La Antigua: 64 km and Jamapa: 105 km) and the total studied area (214 km).
We also estimated Ne for each basin employing the linkage disequilibrium information method (LD; Waples and Do 2008) implemented in NeESTIMATOR v.2.0 (Do et al. 2014). NeESTIMATOR and LDne (Waples and Do 2008) are the most robust and accurate single sample genetic estimators of Ne (Gilbert and Whitlock 2015). We chose NeESTIMATOR over LDne as the former allows to set a minimum allele frequency cutoff value to estimate Ne, minimizing the effect of the presence of rare alleles on the estimations (Do et al. 2014). NeESTIMATOR is based on the fact that as Ne decreases, genetic drift and inbreeding generate nonrandom associations among alleles at different loci (Waples 1991; Luikart et al. 2010). We processed the data as a “single sample time” because our sample collection is likely to contain a single generation based on previous reports on Lutra lutra and Lontra canadensis, for which one generation comprises between 2 and 2.5 years (Kruuk 2006; Reed-Smith 2008). We assumed random mating and screened out alleles with frequencies below 0.02.
We inferred the possibility that the Neotropical otter underwent bottlenecks in the software BOTTLENECK v.1.2.02 (Piry et al. 1999), using the two‐phase mutation model (TPM), and the Wilcoxon signed-rank test with 1000 replicates. This algorithm evaluates whether the expected heterozygosity of a population is larger than the expected heterozygosity under drift-migration equilibrium (Luikart et al. 1998).
Results
Genotyping quality control, individual identification and microsatellite power of discrimination
We collected a total of 130 spraints and anal secretion samples along the three river basins. DNA amplification of these samples in at least nine loci, yielded a successful amplification rate of 42%. Genotyping error per locus ranged from 0 to 0.19, with a mean across loci of 0.08 (La Antigua), and 0.09 (Actopan and Jamapa) (Table 1). We did not detect allelic dropout at any loci. We obtained 60 genotypes (54 from non-invasive samples and six from the captive individuals). Among the genotypes of non-invasive samples ALLEMATCH identified 49 unique multilocus genotypes (i.e. individuals). Twelve individuals were from La Antigua, 25 from Actopan, and 18 (including the six captive individuals) from Jamapa. We detected samples of three individuals in different surveys (i.e. recaptures); all recaptures were in the same basin in which individuals were captured. In Jamapa, one individual was recaptured once, and in Actopan one individual was recaptured once and another individual was recaptured in two different occasions. The geographic distance between the position of the capture-recapture individuals was 0.3 km in Jamapa, 4.3 km for one of the individuals in Actopan, and 7.5 and 10.3 km (a total of 17.8 km) for the first and second recaptures of the other individual (Fig. 1).
PID and PIDSIB values were 8.180 × 10–11 and 1.391 × 10–04, respectively, suggesting a high power of these markers to discriminate individuals. Five loci are enough to distinguish between unrelated individuals and 11 to differentiate among siblings with a probability of identity of < 0.0001 (Fig. S1).
Genetic diversity
Genetic diversity estimates per locus and combining the 11 loci are shown in Table 1. All loci were polymorphic, the number of alleles per locus ranged from 3 to 11; the total number of alleles for all loci was 91: 47 were found in La Antigua, 64 in Actopan and 68 in Jamapa. AR per basin varied from 4.04 (La Antigua) to 5.20 (Jamapa), PAR from 0.57 (La Antigua) to 1.22 (Jamapa) and NEA from 2.42 (La Antigua) to 3.61 (Jamapa). HO and HE ranged from 0.47 (Actopan) to 0.55 (La Antigua), and from 0.54 (La Antigua) to 0.69 (Jamapa), respectively. AR (P = 0.007) and HE (P = 0.025) were significantly different between La Antigua and Jamapa, but HO did not differ significantly among basins (P > 0.05; Fig. 2a–c).
FIS values indicated significant deviations from HWE in Actopan and Jamapa, suggesting inbreeding and/or the presence of null alleles. Further inspection of the loci indicated that FIS values were positive and significant for five loci (RIO11, Lolo19 Lolo30, Lolo37, Lolo48) but deficits for a particular locus were not consistently detected across basins (Table 1). MICROCHECKER suggested the presence of null alleles in these five loci and in locus Lolo41, but the frequency of most of them was lower than 0.20 (Table 1), which is considered not to introduce significant biases in the results (Chapuis and Estoup 2007). The exceptions were loci Lolo19 and Lolo37 in Jamapa, for which the frequency of null alleles was slightly higher than 0.20 (Table 1). We included the five loci (RIO11, Lolo19 Lolo30, Lolo37, Lolo48) in the analyses because if null alleles were causing the deviations from HWE we would expect the other two basins (La Antigua and Actopan) to have homozygote deficits at these loci. To evaluate the possible effect of null alleles in the loci with null allele frequencies higher than 0.20 (Lolo19 and Lolo37), we performed estimates of genetic diversity excluding those loci. Excluding loci Lolo19 and Lolo37 did not bias the results, as genetic diversity estimates with and without these two loci were similar (Tables 1, S2). There was no evidence of LD for any of the 55 paired loci comparisons after the Bonferroni correction (P < 0.0009).
Population genetic structure
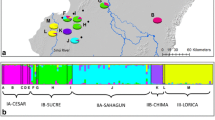
Bayesian clustering analysis in STRUCTURE, including all individuals, showed K = 4 as the most likely number of clusters. This corresponded to the highest value of ΔK (Fig. S2). Individuals from Jamapa and La Antigua formed clusters K1 and K2, respectively; while Actopan individuals were clustered into two additional groups (K3 and K4) (Fig. 3a). Nonetheless, all individuals showed shared ancestry (Fig. 3a). Subsequent analyses for each of the clusters separately did not detect additional substructure in Jamapa and La Antigua (Figs. 3b, S3, S4). However, two genetic clusters were recognized in Actopan (Figs. 3b, S5), supporting the results of the analysis that included all samples. Mixed ancestry was evident in individuals from the two Actopan groups (Fig. 3b). Pie charts of membership scores for each basin suggest that in average the majority of Jamapa individuals were assigned to cluster K1 (blue), while most assignments of La Antigua individuals were to group K2 (green). In Actopan a large proportion of membership assignments were to groups K3 (orange) followed by groups K1 (blue) and K4 (yellow) (Fig. 3c). DAPC identified three genetic clusters (BIC = 3), each one corresponding to a river basin, but some individuals from the three groups overlapped (Fig. 4). To build the plot, ten PCs were retained, constituting 66% of the cumulative variance as predictors of the discriminant analysis.
Assignment of Neotropical otter individuals to genetic groups (K) based on 11 microsatellite loci performed in STRUCTURE. a Probability of membership of 55 analyzed individuals to belong to an optimal number of genetic clusters K = 4. b Probability of belonging to a group when analyzing separately each of the resulting groups from the initial analysis. Different colors indicate the proportion of membership to a genetic group and the basin where the individuals were collected is indicated at the bottom of the graph. c Geographic location in the basins of each of the 55 individuals identified from the non-invasive samples, indicating with colors their probability of membership to each of the four inferred genetic clusters of the analysis using the entire dataset. Pie charts show averages of cluster assignments for each basin using the complete data set
Scatterplot of discriminant analysis of principal component (DPCA) of the Neotropical otter performed on 11 microsatellites loci. We show genetic clusters in different colors, matching the three basins and the 95% inertia ellipses. Axes correspond to the first two discriminant functions and the circles represent individuals. The lower right inset shows the 10 obtained PCA eigenvalues
The three estimators of genetic differentiation (FST, FST-FREENA and DEST) indicated low to moderate significant genetic differentiation among all basins (Table 2). The comparisons between Actopan and La Antigua showed the highest degree of differentiation, according to the tree statistics; while La Antigua vs. Jamapa comparison showed the lowest, as indicated by DEST and FST values. Pairwise FST and FST-FREENA values were very similar and lower than DEST values, suggesting that the relatively high frequency of null alleles of loci Lolo19 and Lolo37 is not resulting in an overestimation of population differentiation.
Spatial autocorrelation analysis identified a significant relationship between genetic and geographic distances at the first two distance classes (within 20 km; r = 0.068, P = 0.001). The autocorrelation coefficient (r) reached a value of zero at 28 km, suggesting that spatial autocorrelation is lost at such distance (Fig. 5).
Plot of the spatial genetic autocorrelation coefficient (r) of the Neotropical otter individuals from the three river basins using distance classes of 10 km. The upper and lower 95% confidence intervals (CI) are indicated with dashed lines and represent the null hypothesis of a random geographic distribution of otter individuals
Migration estimates
GENECLASS identified 10 individuals (P < 0.001) as putative first-generation migrants (Table 3). A total of five individuals migrated from La Antigua, three of them migrated to Jamapa and two to Actopan. Four individuals migrated from Jamapa, three to Actopan and one to La Antigua; one individual migrated from Actopan to Jamapa. STRUCTURE identified seven first generation migrants (Table 3). Six were individuals collected in Actopan and assigned to genetic clusters K1 (Jamapa) or K4 (one of the Actopan groups), and the other one was collected in La Antigua and assigned to group K3 (one of the Actopan groups). Only two individuals (A140 and L58) were identified as migrants by the two analyses.
Population size and bottleneck estimation
All Nc estimations fitted the TIRM model (P = 0) and calculated 75 individuals in Actopan, 92 in La Antigua and in Jamapa, which sum a total of 259 individuals for the entire studied area (Table 4). According to the 95% confidence intervals NC values did not significantly differ among basins. The density of L. longicaudis was 1.66 individuals km−1 in Actopan, 1.44 individuals km−1 in La Antigua and 0.88 individuals km−1 in Jamapa. The total density of the otter in the three basins was 1.21 individuals km−1, corresponding to 259 individuals in 214 km of rivers. Ne estimates significantly differed among basins, with values being 19.6 for Actopan, 6.4 for La Antigua, and 37.3 for Jamapa (Table 4). According to the Wilcoxon test, we did not find evidence of bottlenecks under the two phase mutation model (Tables 4, S3).
Discussion
Genetic diversity
Genetic diversity of the Neotropical otter was moderate (Mean: HO = 0.50, HE = 0.62, AR = 4.56) and comparable to values reported for the conserved Lacantun river in Mexico (Ortega et al. 2012) and for the human-perturbed Paraná river in Argentina (Trigila et al. 2016). However, it was lower than the genetic diversity of otter populations inhabiting the conserved rivers of the Maquiné Valley in Brazil (Trinca et al. 2013). Similar patterns of lower genetic diversity in Mexican than in South American populations of the Neotropical otter have been detected with mitochondrial makers, and have been attributed to the higher river interconnection and degree of conservation of Amazon basins compared to Mexican basins (Guerrero et al. 2015; Hernández et al. 2017).
Although it is well known that habitat deterioration can have important impacts on genetic diversity (Keyghobadi 2007), this was not clearly evident in our study since we observed moderate genetic diversity levels. It is plausible that genetic diversity of the Neotropical otter has been maintained through time and that not enough generations have passed to detect detrimental effects of habitat deterioration (Goossens et al. 2004; Wozney et al. 2011; Epps and Keyghobadi 2015). The lifetime of L. longicaudis is between 10 and 15 years (Rheingantz and Trinca 2015), which could be enough to generate a time lag that masks the effect of habitat deterioration on genetic diversity. Alternatively, habitat deterioration of the studied river basins may have not reached a threshold value at which drastic reductions of genetic diversity begin, which on average starts when 50% of the habitat has been lost (Pflüger et al. 2019).
Surprisingly, although Jamapa is the most perturbed basin it had the highest values of genetic diversity (AR and HE). This could be because its largest area allows maintaining a larger otter population, which is supported by the largest Ne estimates for Jamapa (see below). The lowest genetic diversity was detected in La Antigua and could be the result of the extreme perturbation reported for the basin (Cotler-Ávalos et al. 2010).
Population genetic structure and migration patterns
The genetic structure observed among basins could reflect the hierarchical organization of the rivers, which can promote higher gene flow within than among basins (Meffe y Vrijenhoek 1988; Hughes et al 2009; Selkoe et al. 2015). Similar patterns of genetic structure have been reported in other semi-aquatic species, such as the capibara (Byrne et al. 2015) and the North American beaver (Crawford et al. 2009). In addition to the hierarchical structure of the basins, other landscape characteristics such as highways, the presence of towns or cities and water pollution could be constraining dispersal of L. longicaudis as it has been reported in other otter species (Janssens et al. 2008; Carranza et al. 2012; Pagacz 2016). A study on the Eurasian otter reported that individuals cross the drainage divides of basins located at lower altitudes (up to 1300 m a.s.l.) and with lower slopes (a maximum of 45°), resulting in a lack of genetic differentiation (Pagacz 2016). The basins included in our study are located at altitudes up to 3180 m a.s.l. and have slopes from 0 to 80° (Latorre-Cardenas et al. pers. obs.), which could be limiting gene flow among basins. The three capture-recapture events that we obtained showed that individuals can move between 0.3 and 18 km (Fig. 1), corresponding to the moderate movement capability of the species reported in other studies (7–17 km; Ortega et al. 2012, Trinca et al. 2013). These distances are similar to the 20–25 km that the spatial autocorrelation analysis suggested (Fig. 4). Similar patterns of fine scale genetic structure were detected for the Eurasian otter, with dispersal distances varying between 21 and 38 km (Quaglieta et al. 2013; Pagacz 2016).
The number and identity of first-generation migrants were different between the two algorithms employed. STRUCTURE identified as migrants the individuals that conformed one of the Actopan genetic groups (K4, yellow group). It is plausible that those individuals are migrating from unsampled basins contiguous (in the North) to Actopan, or that they constitute unsampled genotypes in Jamapa and La Antigua that dispersed to Actopan. Further sampling in basins northern to Actopan as well as in the three studied basins is necessary to distinguish between these possibilities. GENECLASS identified a total of 10 first generation migrants and suggests that La Antigua is the basin with the highest number of emigrants and Actopan with the highest number of immigrants. Several studies have reported that poor habitat quality promotes dispersal of individuals to search for better and optimal habitats (Bowler and Benton 2005; Rémy et al. 2011; Honorato et al. 2015). However, otters can disperse to poor habitats as they are not able to identify the presence of pollutants that can potentially have detrimental effect on their health and fitness (Huang et al. 2018). Based on water quality and prey availability, Actopan basin could constitute a better habitat for the Neotropical otter than Jamapa and La Antigua (Macías-Sánchez and Latorre-Cardenas unpublished data). These differences support the higher migration to Actopan from the other two basins. Further studies evaluating dispersal patterns of the species and habitat quality of the three studied basins are necessary to confirm this possibility.
Population size and bottlenecks
Franklin (1980) proposed that a Ne = 50 is the minimum size required to avoid inbreeding depression in the short term (five generations), while a Ne = 500 is necessary to maintain the evolutionary potential of a population. More recently Frankham et al. (2014) suggested that Ne should be increase to > 100/1000 to maintain the genetic variability of populations. Ne of Neotropical otter from the three studied basins are therefore too small to maintain the genetic variation of the populations in the short and long term. It is likely that otter populations are subject to the detrimental effects of small population sizes, including inbreeding, genetic drift and mutational meltdown (Franklin 1998; Higgins and Lynch 2001; Frankham et al. 2014).
Although we used the LD method, which is considered the most robust and accurate single sample estimator of Ne for small populations (Gilbert and Whitlock 2015), our estimations should be interpreted with caution. LD assumes random mating, lack of population structure and immigration, some of which could have been violated due to the biology of the Neotropical otter (Wang et al. 2016). For example, in this study, we detected migration among populations. Additionally, in order to obtain Ne estimates with biases lower than 10%, it is recommended to use 25–50 samples per population and 10–20 loci when the actual effective population size is small (Ne ≤ 100; Waples and Yokota 2007; Waples and Do 2010). Population sizes of L. longicaudis are expected to be small due to the few offspring per litter (2 to 3 offspring; Gallo-Reynoso 2007) and our samples sizes were lower than those recommended. Therefore, it is likely that the Ne values are underestimated in a 10–50% (Waples and Do 2010; Wang, 2016). Regardless of the biases of Ne estimations for the Neotropical otter, the values for the three basins provide a good initial approximation and indicate that the studied populations are vulnerable to extirpation. Even if we doubled the obtained Ne values (assuming our values are underestimated in a 50%), the estimates would still be lower than those proposed (Ne > 100/1000) to maintain the viability of the populations.
Despite that the estimation of Nc is considered important to evaluate the conservation status of the species, few studies have estimated it for L. longicaudis using molecular markers (Rheingantz et al. 2017; Aristizabal-Duque et al. 2018). Our estimation of Nc for the entire studied area was 259 (75 in Actopan and 92 in La Antigua and Jamapa), which is equivalent to a density of 1.21 individuals km−1. Compared to other studies, the calculated density is similar to that reported for a population of L. longicaudis in Brazil (1 otter km−1; Trinca et al. 2013) and lower than the reported for the Lacandona jungle (1.95 individuals km−1; Ortega et al. 2012). The lower density of the Neotropical otter in the studied basins compared to the Lacandona population could be associated with the lower anthropogenic activity and the higher fluvial connectivity in the Lacandona that could result in a better habitat quality and thus in larger populations (Ortega et al. 2012).
The Nc estimations of this study should be interpreted with caution as the confidence intervals were large and most individuals were captured only once (singletons), suggesting that otter populations are larger than reported here (Miller et al. 2005). A more intensive sampling of the three basins is necessary to assure more accurate Nc estimates. The use of non-invasive samples, in combination with molecular markers, have limitations as biases in Nc estimations can result from amplification and genotyping errors (Janssens et al. 2008). Nevertheless, studies using this approximation have demonstrated that Nc estimations are similar to those obtained with other methods (Mumma et al. 2015; Biaffi et al. 2017; Ferreira et al. 2018). The genotyping error of our study was relatively low (mean = 0.08) and the amplification success (42%) was comparable to that reported in other studies (Ortega et al. 2012; Trinca et al. 2013, 2016). Nonetheless, some of the collected samples that did not amplify could correspond to recaptures or unsampled individuals, leading to the underestimation of Nc.
Even though we did not find signatures of recent bottlenecks, the low estimates of population sizes and genetic differentiation among basins suggest that the Neotropical otter could be experiencing isolation and genetic erosion. Although some migration is apparent, it could not be enough to counteract the detrimental effects of genetic drift. Further genetic erosion and isolation will occur if population size (both Ne and Nc) do not increase or if gene flow is completely interrupted as the result of habitat fragmentation and deterioration (Bouzat 2010). A previous study using mitochondrial DNA suggested that otter populations in Jamapa and La Antigua underwent recent and sudden population expansions with the Ne increasing approximately 200,000–300,000 years ago (Hernández-Romero et al. 2018). This implies that the small population sizes detected in this study are the result of recent reductions. River pollution (Ramos-Rosas et al. 2012) and hunting (Rheingantz and Trinca 2015) could have promoted decreases in the population size of L. longicaudis, as it has been suggested for the sea and Eurasian otters (Larson et al. 2002, 2012; Tison et al. 2015; Pigneur et al. 2019).
Conservation implications
In this study we estimated genetic and demographic parameters relevant for the management and conservation of the Neotropical otter in Mexico. Genetic diversity was similar to the reported values for other otter species that are endangered according to the UICN, including the sea otter Enhydra lutris (HO = 0.49, HE = 0.47; Gagne et al. 2018) and the giant otter, Pteronura brasiliensis (HO = 0.56, HE = 0.57; Pickles et al. 2012). This highlights the importance of monitoring genetic diversity and population sizes of L. longicaudis in the short and medium-term to detect possible effects of human activities. It is also essential to determine whether levels of genetic diversity in L. longicaudis are enough to maintain the adaptive potential of the species by performing genomic studies to detect loci under selection that are related to traits that allow the adaptation to environmental changes (Beichman et al. 2019; Mable 2019). For instance, Cianfrani et al. (2018), predicted that core areas in the distribution of L. longicaudis will be expose to the negative effects of human impacts, such as illegal poaching, dams, fishing, water contamination, and tourism.
We found differences on genetic diversity among basins with contrasting degrees of habitat deterioration (Cotler-Ávalos et al. 2010). Employing Ecological Niche Modelling, Trigila et al. (2016) suggested that poor habitat suitability, characterized by high human densities, negatively affect the genetic diversity of the Neotropical otter throughout its distributional range. To evaluate those suggested effects, it would be important to conduct a landscape genetic study to explicitly test the effect of different landscape attributes, including natural (slope, altitude, vegetation coverage, density of hydrological networks) and anthropogenic (human density, presence of roads) elements on genetic diversity and connectivity.
The Neotropical otter population of La Antigua is the most threatened of the three basins. Given its small effective population size, it is likely that the genetic diversity cannot be maintained in the short term. In fact, La Antigua showed the lowest genetic diversity and it is under strong anthropogenic pressure that could continue to affect the viability of the population. Reintroduction of individuals from Actopan and Jamapa rivers into La Antigua could help reducing the loss of genetic diversity in the basin. In L. canadensis, reintroductions of a large number of individuals over an extended period of time resulted in the increase of genetic diversity and population sizes of the species (Mowry et al. 2015). The apparent better habitat quality of Actopan and its highest levels of gene flow with the other otter populations, highlights the importance of maintaining its connectivity with the other basins. Even though L. longicaudis population of Jamapa basin exhibited the highest genetic diversity and population sizes, it is important to monitor these population parameters in the short and medium term as habitat deterioration of the basin can have a negative impact. In conclusion, although genetic diversity of the Neotropical otter populations inhabiting the three river basins is moderate and migration events occur among them, it would be important to implement conservation plans for the populations of the three basins as they continue to be under anthropogenic pressure and thus have a negative effect on the maintenance of genetic variation.
References
Allendorf FW, Luikart G, Aitken S (2013) Conservation and the genetics of populations. Blackwell Publishing, Massachusetts
Amos W, Wilmer JW, Fullard K et al (2001) The influence of parental relatedness on reproductive success. Proc Biol Sci 268:2021–2027
Aparicio JM, Ortego J, Cordero PJ (2006) What should we weigh to estimate heterozygosity, alleles or loci? Mol Ecol 15:4659–4665
Arellano E, Sánchez Nuñez E, Mosquera MA (2012) Distribución y abundancia de la nutria neotropical (Lontra longicaudis annectens) en Tlacotalpan, Veracruz, México. Acta Zool Mex 28:270–279
Aristizábal Duque SL, Orozco-Jiménez LY, Zapata-Escobar C, Palacio-Baena JA (2018) Conservation genetics of otters: review about the use of non-invasive samples. Therya 9:85–93
Arrendal J, Vilà C, Björklund M (2007) Reliability of noninvasive genetic census of otters compared to field censuses. Conserv Genet 8:1097–1107
Beichman AC et al (2019) Aquatic adaptation and depleted diversity: a deep dive into the genomes of the sea otter and giant otter. Mol Biol Evol. https://doi.org/10.1093/molbev/msz101
Beheler A, Fike S, Rhodes G, Serfass OE, Thomas L (2004) Development of polymorphic microsatellite loci for North American river otters (Lontra canadensis) and amplification in related Mustelids. Mol Ecol Notes 4:56–58
Beheler A, Fike S, Dharmarajan JA, Rhodes G, Serfass OE, Thomas L (2005) Ten new polymorphic microsatellite loci for North American river otters (Lontra canadensis) and their utility in related mustelids. Mol Ecol Notes 5:602–604
Biaffi D, Williams DA (2017) Use of non-invasive techniques to determine population size of the marine otter in two regions of Peru. Mammal biol 84:12–19
Bouzat JL (2010) Conservation genetics of population bottlenecks: the role of chance, selection, and history. Conserv Genet 11:463–478
Bowler DE, Benton TG (2005) Causes and consequences of animal dispersal strategies: relating individual behaviour to spatial dynamics. Biol Rev 80:205–225
Byrne MS, Quintana RD, Bolkovic MA, Cassini MH, Túnez JI (2015) The role of river drainages in shaping the genetic structure of capybara populations. Genetica 6:645–656
Carranza ML et al (2012) Connectivity providers for semi-aquatic vertebrates: the case of the endangered otter in Italy. Landsc Ecol 27:281–290
Chapman JR, Nakagawa S, Coltman DW, Slate J, Sheldon BC (2009) A quantitative review of heterozygosity–fitness correlations in animal populations. Mol Ecol 18:2746–2765
Chapuis MP, Estoup A (2007) Microsatellite null alleles and estimation of population differentiation. Mol Biol Evol 243:621–631
Cianfrani C, Broennimann O, Loy A, Guisan A (2018) More than range exposure: global otter vulnerability to climate change. Biol Conserv 221:103–113
Cotler-Ávalos H, Garrido A, Bunge V, Cuevas ML (2010) Las cuencas hidrográficas de México: priorización y toma de decisiones. In: Cotler-Ávalos H, et al. (eds) Las cuencas hidrográficas de México. Diagnóstico y priorización. Pluralia Ediciones e Impresiones, Mexico City
Cornuet JM, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144:2001–2014
Crawford JC, Liu Z, Nelson TA, Nielsen CK, Bloomquist CK (2009) genetic population structure within and between beaver (Castor canadensis) populations in Illinois. J Mammal 90:373–379
Crow JF, Kimura M (1970) An introduction in population genetics theory. Harper and Row, New York
Cushman SA, McRae B, Adriaensen F, Beier P, Shirley M, Zeller K (2013) Biological corridors and connectivity. In: Macdonald DW, Willis KJ (eds) Key topics in conservation biology 2. Wiley, Oxford, pp 384–404
Di Fonzo MMI, Pelletier F, Clutton-Brock TH, Pemberton JM, Coulson T (2011) The population growth consequences of variation in individual heterozygosity. PLoS ONE 6:e19667. https://doi.org/10.1371/journal.pone.0019667
Do C, Waples RS, Peel D, Macbeth GM, Tillett BF, Ovenden JR (2014) NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour 14:209–214
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genets Resour 4:359–361
Environmental Systems Research Institute (ESRI) (2013) ArcGIS Desktop: Release 10.2.1. Redlands, CA
Epperson BK (2005) Estimating dispersal from short distance spatial autocorrelation. Heredity 95:7–15
Epps CW, Keyghobadi N (2015) Landscape genetics in a changing world: disentangling historical and contemporary influences and inferring change. Mol Ecol 24:6021–6040
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Ferreira CM, Sabino-Marques H, Barbosa S (2018) Genetic non-invasive sampling (GNIS) as a cost-effective tool for monitoring elusive small mammals. Eur J Wildl Res. https://doi.org/10.1007/s10344-018-1188-8
Frankham R (1996) Relationship of genetic variation to population size in wildlife. Conserv Biol 10:1500–1508
Frankham R (2010) Where are we in conservation genetics and where do we need to go? Conserv Genet 11:661–663
Frankham R (2015) Genetic rescue of small inbred populations: meta-analysis reveals large and consistent benefits of gene flow. Mol Ecol 24:2610–2618
Frankham R, Ballou JD, Briscoe DA (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Frankham R, Bradshaw CJA, Brook BW (2014) Genetics in conservation management: revised recommendations for the 50/500 rules, Red List criteria and population viability analyses. Biol Conserv 170:56–63
Frankham JD, Ralls RK, Eldridge MDB et al (2017) Genetic management of fragmented animal and plant populations. Oxford University Press, Oxford
Franklin IR (1980) Evolutionary change in small populations. In: Soule ME, Wilcox BA (eds) Conservation biology: an evolutionary-ecological perspective. Sinauer Associate, Sunderland, pp 135–149
Franklin IR, Frankham R (1998) How large must populations be to retain evolutionary potential? Anim Conserv 1:69–73
Gagne RB, Tinker MT, Gustafson KD (2018) Measures of effective population size in sea otters reveal special considerations for wide-ranging species. Evol Appl 11:1779–1790
Gallo-Reynoso JP (1997) Situación y distribución de las nutrias en México, en énfasis en Lontra longicaudis annectens MAJOR, 1897. Rev Mex Mastozool 2:10–32
Gallo-Reynoso (2007) Evaluación del riesgo de extinción de Lontra longicaudis de acuerdo al numeral 5–7 de la NOM-059-ECOL-2001. In: Sánchez O et al. Método de Evaluación del Riesgo de Extinción de las Especies Silvestres en México (MER). INE-SEMARNAT CONABIO, Mexico City
Galpern P, Manseau M, Hettinga P, Smith K, Wilson P (2012) Allelematch: an R package for identifying unique multilocus genotypes where genotyping error and missing data may be present. Mol Ecol Resour 12:771–778
Gerlach G, Jueterbock A, Kraemer P, Deppermann J, Harmand P (2010) Calculations of population differentiation based on G(ST) and D: Forget G(ST) but not all of statistics! Mol Ecol 19:3845–3852
Gilbert KJ, Whitlock MC (2015) Evaluating methods for estimating local effective population size with and without migration. Evolution 69:2154–2166
González-Chisten A, Delfín-Alfonso CA, Sosa-Martínez A (2013) Distribución y abundancia de la nutria neotropical (Lontra longicaudis annectens Major, 1897) en el lago de Catemaco Veracruz, México. Therya 4:201–217
Goudet J (2001) FSTAT, version 2.9. 3, A program to estimate and test gene diversities and fixation indices. Lausanne University, Lausanne
Goossens B et al (2005) Patterns of genetic diversity and migration in increasingly fragmented and declining orangutan (Pongo pygmaeus) populations from Sabah, Malaysia. Mol Ecol 14:441–456
Guerrero J, Gallo-reynoso JP, Biek R (2015) Mitochondrial DNA diversity, genetic structure, and demographic history of the Neotropical otter (Lontra longicaudis) in Mexico. J Mammal 96:1162–1173
Hague MT, Routman EJ (2016) Does population size affect genetic diversity? A test with sympatric lizard species. Heredity 116:92–98
Hedrick PW (2005) Genetics of populations. Jones and Bartlett publishers, Massachusetts
Hernández-Romero PC, Gutiérrez-Rodríguez C, Valdespino C, Prieto-Torres DA (2017) The role of geographical and ecological factors on population divergence of the neotropical otter Lontra longicaudis (Carnivora, Mustelidae). Evol Biol 45:37–55
Hernández-Romero PC, Botello-López FJ, Hernández-García N, Espinosa-Rodríguez J (2018) New Altitudinal Record of Neotropical Otter (Lontra longicaudis Olfers, 1818) and Conflict with Fish Farmers in Mexico. Otter Spec Group Bull 35:193–197
Higgins K, Lynch M (2001) Metapopulation extinction caused by mutation accumulation. PNAS 98:2928–2933
Honorato R, Crouzeilles R, Ferreira MS, Grelle CEV (2015) The effects of habitat availability and quality on small mammals abundance in the Brazilian Atlantic Forest. Natureza & Conservação 13:133–138
Huang AC et al (2018) River otters (Lontra canadensis) “trapped” in a coastal environment contaminated with persistent organic pollutants: demographic and physiological consequences. Environ Pollut 238:306–316
Hughes JM, Schmidt DJ, Finn DS (2009) Genes in Streams: using dna to understand the movement of freshwater fauna and their riverine habitat. Bioscience 59:573–583
Instituto Nacional de Estadística, Geografía e Informática - INEGI (2016) Continuo De Elevación Mexicano, CEM 3.0. https://www.inegi.org.mx/app/geo2/elevacionesmex/. Accessed 27 Jan 2016
Instituto Nacional de Estadística y Geografía (INEGI) (2011) Sistema de consultas de las cuencas hidrográficas de México. Disponible en. https://cuencas.ine.gob.mx/cuenca/
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Janssens X, Fontaine MC, Michaux JR, Libois R, De Kermabon J, Defourny P, Baret PV (2008) Genetic pattern of the recent recovery of European otters in southern France. Ecography 31:176–186
Jombart T (2008) Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 94:1471–2156
Jost L (2008) Gst and its relatives do not measure differentiation. Mol Ecol 17:4015–4026
Kalinowski ST (2004) Counting alleles with rarefaction: private alleles and hierarchical sampling designs. Conserv Genet 5:539–543
Kalinowski ST (2005) HP-Rare: a computer program for performing rarefaction on measures of allelic diversity. Mol Ecol Notes 5:187–189
Keyghobadi N (2007) The genetic implications of habitat fragmentation for animals. Can J Zool 85:1049–1064
Knaepkens G, Bervoets L, Verheyen E, Eens M (2004) Relationship between population size and genetic diversity in endangered populations of the European bullhead (Cottus gobio): implications for conservation. Biol Conserv 115:403–410
Kretzmann M, Mentzer L, DiGiovanni R, Leslie MS, Amato G (2006) Microsatellite diversity and fitness in stranded juvenile harp seals (Phoca groenlandica). J Hered 97:555–560
Kruuk H (2006) Otters: ecology, behaviour and conservation. Oxford University Press, Oxford
Lariviére S (1999) Lontra longicaudis. Mamm Species 609:1–5
Larson S, Jameson R, EtnierM FM, Bentzen P (2002) Loss of genetic diversity in sea otters (Enhydra lutris) associated with the fur trade of the 18th and 19th centuries. Mol Ecol 11:1899–1903
Larson S, Jameson R, Etnier M, Jones T, Hall R (2012) Genetic diversity and population parameters of sea otters, Enhydra lutris, before Fur Trade Extirpation from 1741–1911. PLoS ONE 7(3):e32205. https://doi.org/10.1371/journal.pone.0032205
Latorre-Cardenas MC, Gutiérrez Rodríguez C, Lance S (2020) Isolation and characterization of 13 microsatellite loci for the Neotropical otter, Lontra longicaudis, by next generation sequencing. Mol Biol Rep 47:731–736
Lerone L, Mengoni C, Carpaneto GM, Randi E, Loy A (2014) Procedures to genotype problematic non-invasive otter (Lutra lutra) samples. Acta theriol 59:511–520
Luikart G, Sherwin WB, Steele BM, Allendorf FW (1998) Usefulness of molecular markers for detecting population bottlenecks via monitoring genetic change. Mol Ecol 7:963–974
Luikart G et al (2010) Estimation of census and effective population sizes: the increasing usefulness of DNA-based approaches. Conserv Genet 11:355–373
Mable BK (2019) Conservation of adaptive potential and functional diversity: integrating old and new approaches. Conserv Genet 20:89. https://doi.org/10.1007/s10592-018-1129-9
Macías-Sánchez S, Aranda M (1999) Análisis de la alimentación de la nutria Lontra longicaudis (Mammalia: Carnivora) en un sector del Río Los Pescados, Veracruz, México. Acta Zoologica Mexicana 76:49–57. https://doi.org/10.21829/azm.1999.76761699
Manel S, Schwartz MK, Luikart G, Taberlet P (2003) Landscape genetics: combining landscape ecology and population genetics. Trends Ecol Evol 18:189–197
Markert JA, Grant PR, Grant BR, Keller LF, Coombs JL, Petren K (2004) Neutral locus heterozygosity, inbreeding, and survival in Darwin's ground finches (Geospiza fortis and G. scandeus). Heredity 92:306–315
Markert JA, Champlin DM, Gutjahr-Gobell R et al (2010) Population genetic diversity and fitness in multiple environments. BMC Evol Biol. https://doi.org/10.1186/1471-2148-10-205
Meffe GK, Vrijenhoek RC (1988) Conservation genetics in the management of desert fishes. Conserv Biol 2:157–169
Miller CR, Joyce P, Waits LP (2005) A new method for estimating the size of small populations from genetic mark-recapture data. Mol Ecol 14:1991–2005
Morris DW, Clark R, Boyce MS (2008) Habitat and habitat selection: theory, tests, and implications. Isr J Ecol Evol 54:287–294
Mowry RA et al (2015) Genetics and the successful reintroduction of the Missouri river otter. Anim Conserv 18:196–206
Mumma MA, Zieminski C, Fuller TK, Mahoney SP, Waits LP (2015) Evaluating non-invasive genetic sampling techniques to estimate large carnivore abundance. Mol Ecol Resour 15:1133–1144. https://doi.org/10.1111/1755-0998.12390
Ortega J, Navarrete D, Maldonado JE (2012) Non-invasive sampling of endangered neotropical river otters reveals high levels of dispersion in the Lacantun River System of Chiapas, Mexico. Anim Biodiv Conserv 35:59–69
Pacifici M, Santini L, Di Marco M, Baisero D et al (2013) Generation length for mammals. Nat Conserv 5:89–94. https://doi.org/10.3897/natureconservation.5.5734
Pagacz S (2016) The effect of a major drainage divide on the gene flow of a semiaquatic carnivore, the Eurasian otter. J Mamm 97:1164–1176
Palstra FP, Ruzzante DE (2008) Genetic estimates of contemporary effective population size: what can they tell us about the importance of genetic stochasticity for wild population persistence? Mol Ecol 17:3428–3447
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539
Peery MZ, Kirby R, Reid BN, Stoelting R, Doucet-Beer E, Robinson S, Vásquez-Carrillo C, Palsbøll PJN (2012) Reliability of genetic bottleneck tests for detecting recent population declines. Mol Ecol 21:3403–3418
Pennell MW, Stansbury CR, Waits LP, Miller CR (2013) Capwire: a R package for estimating population census size from non-invasive genetic sampling. Mol Ecol Resour 13:154–157
Petit RJ, El Mousadik A, Pons O (2008) Identifying populations for conservation on the basis of genetic markers. Conserv Biol 12:844–855
Pflüger FJ, Signer J, Balkenhol N (2019) Habitat loss causes non-linear genetic erosion in specialist species. Glob Ecol Conserv 17:e00507
Pickles RSA, Groombridge JJ, Rojas VDZ et al (2012) Genetic diversity and population structure in the endangered giant otter, Pteronura brasiliensis. Conserv Genet 13:235–245
Pigneur LM, Caublot G, Fournier-Chambrillon C et al (2019) Current genetic admixture between relictual populations might enhance the recovery of an elusive carnivore. Conserv Genet 20:1133. https://doi.org/10.1007/s10592-019-01199-9
Piry S, Luikart G, Cornuet JM (1999) Computer note. BOTTLENECK: a computer program for detecting recent reductions in the effective size using allele frequency data. J Hered 90:502–503
Piry S, Alapetite A, Cornuet JM, Paetkau D, Baudouin L, Estoup A (2004) GENECLASS2: a software for genetic assignment and first-generation migrant detection. J Hered 95:536–539
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Pompanon F, Bonin A, Bellemain E, Taberlet P (2005) Genotyping errors: causes, consequences and solutions. Nat Rev 6:847–859
Quaglietta L, Fonseca VC, Hájková P, Mira A, Boitani L (2013) Fine-scale population genetic structure and short-range sex-biased dispersal in a solitary carnivore, Lutra lutra. J Mammal 94:561–571
R Core Team (2017) R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing. https://www.R-project.org/
Ramos-Rosas NN, Valdespino C, García-Hernández J, Gallo-Reynoso JP, Olguín EJ (2012) Heavy metals in the habitat and throughout the food chain of the Neotropical otter, Lontra longicaudis, in protected Mexican wetlands. Environ Monit Assess 185:1163–1173
Rannala B, Mountain JL (1997) Detecting immigration by using microsatellite genotypes. Proc Natl Acad Sci USA 94:9197–9201
Reed-Smith J (2008) North American river otter husbandry notebook. John Ball Zoological Garden, Grand Rapids
Reed-Smith J (2012) North American river otter husbandry notebook. Grand Rapids Press, Michigan
Rémy A, Le Galliard J, Gundersen G, Steen H, Andreassen HP (2011) Effects of individual condition and habitat quality on natal dispersal behaviour in a small rodent. J Anim Ecol 80:929–937
Rheingantz ML, Trinca CS (2015) Lontra longicaudis. The IUCN Red List of Threatened Species. https://doi.org/10.2305/IUCN.UK.2015-2.RLTS.T12304A21937379.en
Rheingantz ML, Trinca S-P (2017) The Neotropical otter Lontra longicaudis: a comprehensive update on the current knowledge and conservation status of this semiaquatic carnivore. Mamm Rev 47:291–305
Rheingantz ML, de Menezes JFS, de Thoisy B (2014) Defining neotropical Otter Lontra Longicaudis distribution, conservation priorities and ecological frontiers. Trop Conserv Sci 7:214–229
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
SEMARNAT (2010) NORMA Oficial Mexicana NOM-059-SEMARNAT-2010, Protección ambiental-Especies nativas de México de flora y fauna silvestres-Categorías de riesgo y especificaciones para su inclusión, exclusión o cambio-Lista de especies en riesgo. https://www.biodiversidad.gob.mx/especies/pdf/NOM_059_SEMARNAT_2010.pdf. Accessed 28 oct 2017
Selkoe KA, Scribner KT, Galindo HM (2015) Waterscape genetics——applications of landscape genetics to Rivers, Lakes, and Seas. In: Balkenhol N, Cushman S, Storfer AT, Waits L (eds) Landscape genetics. Wiley, Chichester, pp 220–246
Smouse P, Peakall R (1999) Spatial autocorrelation analysis of individual multiallele and multilocus genetic structure. Heredity 82:561–573
Smouse PE, Peakall R, Gonzales E (2008) A heterogeneity test for fine-scale genetic structure. Mol Ecol 17:3389–3400
Soulé M (1976) Allozyme variation: its determinants in space and time. In: Ayala F (ed) Molecular evolution. Sinauer Associates, Sunderland, pp 60–77
Spielman D, Brook BW, Frankham R (2004) Most species are not driven to extinction before genetic factors impact them. Proc Natl Acad Sci USA 101:15261–15264
Taberlet P, Griffin S, Goossens B, Questiau S et al (1996) Reliable genotyping of samples with very low DNA quantities using PCR. Nucleic Acids Res 24:3189–3194
Tallmon DA, Koyuk A, Luikart G, Beaumont MA (2008) ONeSAMP: a program to estimate effective population size using approximate Bayesian computation. Mol Ecol Res 8:299–301
Tison JL, Blennow V, Palkopoulou E et al (2015) Population structure and recent temporal changes in genetic variation in Eurasian otters from Sweden. Conserv Genet 16:371–384
Townsend AK, Taffs CC, Wheeler SS et al (2018) Low heterozygosity is associated with vector-borne disease in crows. Ecosphere 9:e02407. https://doi.org/10.1002/ecs2.2407
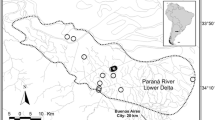
Trigila AP, Gómez JJ, Cassini MH, Túnez JI (2016) Genetic diversity in the Neotropical river otter, Lontra longicaudis (Mammalia, Mustelidae), in the Lower Delta of Parana River, Argentina and its relation with habitat suitability. Hydrobiologia 768:287–298
Trinca CS, Jaeger CF, Eizirik E (2013) Molecular ecology of the Neotropical otter (Lontra longicaudis): non-invasive sampling yields insights into local population dynamics. Biol J Linn Soc 109:932–948
Valière N (2002) GIMLET: a computer program for analyzing genetic individual identification data. Mol Ecol Notes 2:377–379
van Oosterhout C, Hutchinson WF, Willis DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Vergeer P, Rengelink R, Copal A, Ouborg NJ (2003) The interacting effects of genetic variation, habitat quality and population size on performance of Succisa pratensis. J Ecol 91:18–26
Waits LP, Luikart G, Taberlet P (2001) Estimating the probability of identity among genotypes in natural populations: cautions and guidelines. Mol Ecol 10:249–256
Wang J (2009) A new method for estimating effective population sizes from a single sample of multilocus genotypes. Mol Ecol 18:2148–2164
Wang J (2016) A comparison of single-sample estimators of effective population sizes from genetic marker data. Mol Ecol 25:4692–4711
Waples RS (1991) Genetic methods for estimating the effective size of cetacean populations. Rep Int Whal Commn 13:279–300
Waples RS (2008) ldne: a program for estimating effective population size from data on linkage disequilibrium. Mol Ecol Resour 4:753–756
Waples RS, Do C (2010) Linkage disequilibrium estimates of contemporary Ne using highly variable genetic markers: a largely untapped resource for applied conservation and evolution. Evol Appl 3:244–262
Waples RS, Yokota M (2007) Temporal estimates of effective population size in species with overlapping generations. Genetics 175:219–233
Weber L, Hildebrand C, Ferreira A, Pedarass G, Levy J, Colares EP (2009) Microsatellite genotyping from faeces of Lontra longicaudis from southern Brazil. Iheringia Sér Zool 99:5–11
Willoughby JR, Sundaram M, Wijayawardena BK, Kimble SJA et al (2015) The reduction of genetic diversity in threatened vertebrates and new recommendations regarding IUCN conservation rankings. Biol Conserv 191:495–503
Wozney KM, Haxton TJ, Kjartanson S et al (2011) Genetic assessment of lake sturgeon (Acipenser fulvescens) population structure in the Ottawa River. Environ Biol Fish 90:183
Wright S (1940) Breeding Structure of Populations in Relation to Speciation. Am Nat 74:232–248
Acknowledgements
This work was partially supported by the National Geographic Society Early Career Grant (Grant No. # WW-185ER-17), by the Rufford Small Grants Foundation (Grant No. ID-19592-2) and by research funds from the Instituto de Ecología, A.C. (Grant No. 20012-11-080). María Camila Latorre-Cardenas is grateful with the Posgrado en Ciencias Biológicas of the Universidad Nacional Autónoma de México for the academic support provided during her doctoral studies and with the Consejo Nacional de Ciencia y Tecnología (CONACyT) for the Doctoral scholarship (#414864). This research constitutes a requirement for obtaining the doctoral degree of Latorre-Cardenas. The “Acuario de Veracruz, A.C.” donated blood from six individuals. Pablo C. Hernández-Romero, Tarcisio Solis and Luz Magali Sánchez Méndez provided field assistance; and Luz Magali Sánchez Méndez, Denisse Maldonado Sánchez and Cristina Bárcenas laboratory assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.





Rights and permissions
About this article
Cite this article
Latorre-Cardenas, M.C., Gutiérrez-Rodríguez, C. & Rico, Y. Estimating genetic and demographic parameters relevant for the conservation of the Neotropical otter, Lontra longicaudis, in Mexico. Conserv Genet 21, 719–734 (2020). https://doi.org/10.1007/s10592-020-01283-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-020-01283-5