Abstract
Mounting evidences has shown that miRNAs are involved in the development and progression of gastric cancer acts as tumor suppressor genes or oncogenes. In our previous studies, we have found that the up-regulation of miR-106a occurs frequently in human gastric cancer tissues compared with that of normal tissues. Here, we investigate the role of the ectopic expressed miR-106a in the progression and metastasis of gastric cancer in vitro and in vivo. FFPE samples have the priority to be included and qRT-PCR was used to detect the miR-106a expression. Human gastric cancer cells and immortalized gastric epithelial cell were selected and the miR-106a mimic and inhibitor were transfected. Cell growth was determined by MTT method. The flow cytometric analysis for cell apoptosis and transwell assays for evaluating the cell migration and invasion were conducted. Luciferase assay and western blot confirmed the direct binding site of miR-106a and its target. BALB/c nude mice were randomly divided to explore the implantation of gastric cancer cells transfected with miR-106a antagomir. Abnormal over-expression of miR-106a significantly promoted gastric cancer cell proliferation, metastasis, inhibited the cell apoptosis. Functional experiment ascertained that miR-106a interacted with FAS and mediated caspase3 pathway. Knockdown of miR-106a leaded to the attenuation of gastric cancer implantation capacity in vivo. Moreover, expression of TIMP2 was inversely associated with miR-106a in nodule tissues. Apoptotic body was also seen under electron microscope accompanied by silencing of miR-106a. Together, this data indicated that miR-106a may act as an oncogene and contribute to gastric cancer development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastric cancer ranks the fifth most common cancer types and third leading cause of cancer death worldwide [1]. Although a steady decline in gastric cancer incidence and mortality has been observed in the majority of developed countries, the situation is unsatisfactory in developing nations [2]. Gastric cancer continues to present a major challenge due to most cases being diagnosed frequently in advanced stages with various forms of metastasis which means poor prognosis [3]. Thus, understanding the functions of oncogenesis, developmental process of gastric cancer is essential for improving the diagnosis and treatment.
MicroRNAs (miRNAs) are small non-coding RNAs that regulate gene expression at post-transcriptional and translational levels through interaction with their target gene mRNAs in the 3′-untranslated regions [4, 5]. More and more evidences have confirmed that miRNAs are involved in the most important biological processes including proliferation, apoptosis, differentiation, metastasis, angiogenesis which shows its importance in the initiation and development of malignancies [4, 6, 7]. MiR-106a, a potential oncogene, has been reported participating in several malignant tumors progression [8–12]. Our group previously found that the miR-106a was always up-regulated in gastric cancer compared with normal gastric mucosa [13]. Here, we continue to investigate the role of miR-106a and evaluate the mechanism under gastric carcinogenesis and metastasis.
In the present study, we demonstrate that miR-106a is closely associated with gastric cancer cell proliferation, apoptosis and invasion. Moreover, the peritoneal implantation was also affected in vivo experiment. In addition, we provided evidence that FAS and TIMP2 were negatively regulated by miR-106a.
Materials and methods
Tissues and real-time PCR
Human gastric cancer tissues and their corresponding adjacent tissues were collected from Pathological Department of General Hospital of Ningxia Medical University with the approval of local ethics committee and informed consents were also signed, and the exception criterion was the patients with the previous history of radiotherapy and chemotherapy. All these tissues were paraformaldehyde-fixed paraffin-embedded (FFPE) which can guarantee the diagnosis of the carcinoma and adjacent tissues were not infiltrated by any tumor cells. Pathological diagnosis was carried out according to the World Health Organization criteria. RNA from FFPE samples was extracted by RecoverAll™ Total Nucleic Acid Isolation Kit (Ambion) and the isolated RNA was detected under Epoch™ spectrophotometer system (BioTek). cDNA was synthesized using TaqMan MicroRNA Assays (Applied Biosystems) from 10 ng RNA samples. Subsequently, TaqMan Universal Master Mix II (Applied Biosystems) was used to amplify the miR-106a gene with U6 snRNA as its internal reference. The concrete process was described in our previous article [14].
Cell lines
Well-, moderately and poorly differentiated human gastric cancer cells MKN-28, SGC-7901, BGC-823 and MKN-45 as well as immortalized gastric epithelial cell GES-1 were all obtained from the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). Cells were cultured in RPMI-1640 medium (Gibco) with 10 % fetal bovine serum (Gibco) at 37 °C in a 5 % CO2 incubator. RNA from cell lines was isolated by High Pure miRNA Isolation Kit (Roche) and detected using the same method described above.
Transfection
Hsa-miR-106a mimic, inhibitor, antagomir and their negative controls were commercially synthesized from Ribobio (Guangzhou, China). Transfection was carried out with Lipofectamine 2000 (Invitrogen) method according to the manufacturer’s procedure. The final concentration was 100 nm for mimic or 200 nm for inhibitor and antagomir.
MTT method
MTT assay was used to measure the ability of cell proliferation after transfection with miR-106a mimic. Cells (6 × 103 cells/well) were seeded into 96-well culture plates and transfection was performed the next day at the concentration described above. After reaction for 24, 48, 72 and 96 h, MTT reagent was added to each well and incubated at 37 °C for 4 h. Spectrometric absorbance was read at 490 nm by EnSpire Multimode Plate Reader (PerkinElmer).
Flow cytometric analysis
Flow cytometry was performed to analysis of cell apoptosis after transfection with miR-106a inhibitor. Cells were seeded into 6-well plates at density of 2 × 105 cells/well. After induction for 48 h, cell apoptosis was detected with Annexin V-FITC Apoptosis Detection Kit (Joincare, China) in which 5 μl Annexin V-FITC and 10 μl propidium iodide were added for 15 min and then analyzed by flow cytometry (BD Biosciences).
Luciferase assay
The 3′-UTR of FAS (NM_000043.4) was cloned into the pmiR-RB-REPORT™ luciferase reporter (RiboBio, China). The target gene of FAS’s mutant sequence was designed by mutating the multiple nucleotides complementary to the miR-106a seed region. HEK-293T cell was conducted to detect the luciferase ratio. 24 h after the cells grown to 50–70 % confluence in 96-well plates, a mixture of 100 ng pmiR-RB-REPORT™ h-FAS wild type (WT) or mutant (Mut) reporter plasmid vector together with 50 nM hsa-miR-106a mimic or negative control (RiboBio) were co-transfected using Lipofectamine 2000 (Invitrogen) reagent, and at 48 h lysates were prepared and analyzed by the Dual-Glo luciferase assay (Promage) following the manufacturer’s protocol with Renilla luciferase activity as the reporter gene and firefly luciferase as the reference gene.
Transwell assay
Transwell assay was operated to assess cell migration and invasion after transfection with miR-106a mimic. For migration, cells were starved in serum-free medium and then 2 × 104 cells were seeded into the upper chambers of an insert (8 μm pore size, Merck Millipore), which were soaked into the lower chambers filled with complete growth media. For invasion, the same density of cells was placed into the Matrigel-coated chambers (BD Biosciences). After 24 h of incubation, cells on the upper surfaces of the transwell inserts were scraped with cotton swabs, whereas the cells passed through the membranes were fixed with 4 % paraformaldehyde and stained with 0.1 % crystal violet. The migratory and invasive cells were counted under microscope from five randomly selected fields.
Western blot
Cells were harvested and total protein was lysed with RIPA Buffer (Pierce) contained protease inhibitors PMSF. Proteins were separated by 12 % SDS-PAGE and then electrophoretically transferred to PVDF membranes (Millipore). After blocking with 5 % non-fat milk, the membranes were incubated with rabbit anti-FAS, caspase3 polyclonal antibody (1:1000, Cell Signaling). Samples were incubated with a horseradish peroxidase-conjugated secondary antibody the next day and results were analyzed in the dark room by Immobilon™ Western Chemiluminescent HRP Substrate (Merck Millipore). The expressions of these two proteins were normalized against GAPDH (Santa Cruz Biotechnology).
In vivo tumorigenicity
4-week-old male BALB/c nude mice purchased from Vital River Experimental Animal Technology Co., Ltd (Beijing, China) were housed in sterilized cages under 12 h light/dark cycles with water and food ad libitum. 1 × 107 BGC-823 cells transfected with miR-106a antagomir or negative control were inoculated intraperitoneally using a small incision at cartilago ensiformis to construct the tumor xenograft mouse model. Mice were sacrificed 14 days after inoculation of the cells, and metastatic lesions scattered over the abdomen were visualized after laparotomy and tumor nodules were surgically excised for further analyses.
Immunohistochemistry
Immunohistochemical study was conducted to determine the expression of TIMP2 in tumor nodules removed from xenograft mouse model. Nodules were fixed in 10 % formalin and then manufactured into FFPE sections. A rabbit anti-TIMP2 polyclonal antibody (1:200, proteintech) was used in the 2-step plus® Poly-HRP Anti-Mouse/Rabbit IgG Detection System (ZSGB-BIO, China). TIMP2 was stained as yellow in the cytoplasm or nucleus.
In situ hybridization
The expression of miR-106a was examined by in situ hybridization (ISH) method to compare with the TIMP2 in the tumor nodules of nude mice. The miR-106a probe was double-DIG labeled locked nucleic acid (LNA) probe (Exiqon). The probe sequence was as follows: 5′-CTACCTGCACTGTAAGCACTTTT-3′. The ISH was performed in accordance with the company’s instruction. Briefly speaking, the FFPE sections were deparaffinized and rehydration. The miRNA was demasked using 15 μg/ml proteinase K (Exiqon) at 37 °C for 10 min, which allowed the access of probe to hybridize to the miRNA sequence. The hybridization was achieved in a hybridizer (Iris international) with 40 nM of LNA™ probe at 55 °C for 1 h. The DIG was then be recognized by a anti-Digoxigenin-AP antibody (Roche) at 1:800 for 60 min at room temperature. The colorimetric reaction was detected with BCIP/NBT ready-to-use tablets (Roche) at 30 °C for 2 h in a dark oven, and slides were counterstained in Neutral Red Staining Solution (Sigma-Aldrich). The specific miR-106a ISH signal was seen in the cytoplasm or nucleus as dark blue particles.
Transmission electron microscope
The ultra-morphology of gastric cancer cells of nodules was observed by a transmission electron microscope (TEM). Immediately after collection of nodules, the specimens (less than 1 mm3 diameter) were subjected to fixation in 2.5 % glutaraldehyde for 2 h at 4 °C and then post-fixed in 1 % osmic acid for 1 h. After washed several times in phosphate buffer saline, the specimens were dehydrated in increasing concentrations of ethanol. Subsequently, the samples were saturated and embedded in Epon812 epoxy resin and polymerized in an oven. Ultrathin Sects. (50 nm) were cut on an ultramicrotome (LKB, Sweden) followed by double-dyed with uranyl acetate and lead citrate. The cancer cells were characterized under a TEM (HT-7650, HITACHI).
Statistical analysis
Statistical analysis was carried out using SPSS 17.0. Values were shown as mean ± SEM and P < 0.05 was considered to be statistically significant. Paired t test was used to determine the expression of miR-106a in tissues and two sample t-test was used to evaluate the significant difference of in vitro data. One-Way ANOVA was used to detect the results from cell experiments and the relationship between miR-106a and clinicopathologic characteristics of gastric tissues. The bar chart and line chart were chosen to exhibit the difference between the different samples and to describe the absorbency trend over time, respectively.
Results
Relationship of the up-regulation of miR-106a with gastric cancer characteristics
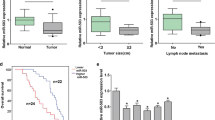
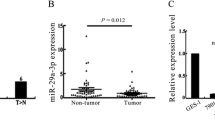
Using real-time PCR assays, we found that miR-106a was frequently up-regulated in gastric cancer tissues even if in FFPE samples. As shown in Fig. 1, the relative expression of miR-106a in gastric cancer tissues was significantly elevated compared with its corresponding non-tumor samples with the fold change expressed by \(2^{{ - \Delta \Delta {\text{C}}_{\text{t}} }}\) at 2.82 ± 1.98 (P = 0.000). Relationship between miR-106a expression and gastric cancer clinical pathological characteristics were unclosed in Table 1. In 50 cases of gastric cancer tissues, there were no significant differences among the miR-106a expression and patient’s gender, age and gastric cancer’s location and dimension that were measured by centimeter. Lauren classification was also be defined including intestinal and diffuse. From our data, the over-expressed miR-106a has the positive correlation with gastric cancer pathological type (P = 0.022), cancer cells infiltration (P = 0.003) and peritoneal implantation (P = 0.017). The result indicated that the high expression of miR-106a in diffuse gastric cancer tissues has linked with malignant degree and therefore could not only promote cancer local invasion but also distant spread.
MiR-106a promotes high-grade gastric cancer cell proliferation
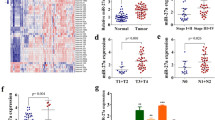
Since we have found that miR-106a was always up-regulated in gastric cancer, we speculated that it might have some potential effects in gastric cancer development. In this regard, the expression of miR-106a was also investigated in gastric cancer cells. As shown in Fig. 2a, the expression of miR-106a varied in four different differentiated gastric cancer cells (P = 0.000). Compared with GES-1 cell, the expression of miR-106a in SGC-7901, BGC-823 and MKN-45 cells were significantly up-regulated with the fold change of 2.50 ± 0.18, 2.45 ± 0.23 and 2.30 ± 0.46 (P = 0.000), unlike the MKN-28 cell with the fold change of 0.96 ± 0.20 (P = 0.823). On the basis of this result, we synthesized the sequence-specific mimic for hsa-miR-106a to increase its expression in representative three different differentiated gastric cancer cells, and then using MTT method to determine the impact of miR-106a on cell proliferation. We found that in contrast to negative group, the proliferations of SGC-7901 and BGC-823 cells, which was moderately and poorly differentiated cells separately, were enhanced greatly from 24 h to 96 h after transduction, whereas there were no changes observed in the wild differentiated gastric cancer MKN-28 cell. GES-1 cell, which was immortalized gastric mucosal epithelial cell, was not express significant difference under the same treatment (Fig. 2b).
MiR-106a in gastric cancer cell lines. a The basic expression level of miR-106a in different differentiated human gastric cancer cells compared with the immortalized GES-1 cell. *P = 0.000. b MTT assays detection of human gastric cancer cell proliferation after transduction with miR-106a mimic or negative control in four typical gastric cancer cells in vitro. *P < 0.05, **P < 0.001
MiR-106a inhibits SGC-7901 cell apoptosis and FAS is its direct downstream target
Since we have found that miR-106a can trigger SGC-7901 and BGC-823 cells proliferation, we want to know whether the miR-106a might induce the cell apoptosis because the acceleration of proliferation is always accompanied by the attenuation of apoptosis. The sequence-specific inhibitor for hsa-miR-106a was conducted in moderately differentiated gastric cancer SGC-7901 cell, and down-regulation of miR-106a in SGC-7901 cell resulted in a higher apoptosis rate than the negative control group (Fig. 3a, b). To further analyze why miR-106a-inhibitor could induce SGC-7901 cell apoptosis, we chosen to examine the apoptosis-associated gene FAS and caspases3 because three kinds of programs jointly predict that FAS is a potential target of miR-106a and caspases3 is considered to be the most important molecule in the process of apoptosis. As shown in Fig. 3c–e, the region of FAS 3′-UTR has sequence complementary with miR-106a. When the wild type of FAS was transfected with miR-106a mimic and negative, the luciferase ratio of WT/miR-106a was significantly decreased compared with WT/NC (P = 0.000). The degree of decreasing is nearly 30 %. When the mutant type of FAS was transfected with miR-106a mimic and negative, no significant difference can be seen between Mut/miR-106a and Mut/NC (P = 0.006). Luciferase assay verified that FAS is a direct target of miR-106a that accepts its regulation. Western blot demonstrated that miR-106a inhibition in SGC-7901 cell can elevate the expression of FAS protein level compared with control group, and subsequently, caspases3 protein level was also increased.
MiR-106a inhibits human gastric cancer SGC-7901 cell apoptosis in vitro. a Flow cytometric analysis of the cells transfected with miR-106a inhibitor or negative control. b The apoptosis rate of miR-106a inhibitor group is higher than control. *P = 0.000. c The FAS 3′-UTR wild type and mutant type sequence are shown with the miR-106a sequence. d The luciferase ratio in HEK-293T cells with wild-type or mutant FAS 3′-UTR vectors and miR-106a mimic or negative control. *P = 0.000. e Apoptosis-related gene protein including FAS and caspase3 were detected by western blot
MiR-106a stimulates the high-grade gastric cancer cell migration and invasion
Since we verified that the ectopic expression of miR-106a has closely associated with gastric cancer cell proliferation and apoptosis, that is to say, miR-106a is involved in many biological processes in the development of gastric cancer. To gain insight into the role of miR-106a, we actualized transwell assays to examine the effect of miR-106a on gastric cancer migration and invasion, the result of which indicated that in SGC-7901 and BGC-823 cells infected with miR-106a inhibitor, migration (49.18 ± 12.12 vs. 104.05 ± 38.90, P = 0.000; 66.09 ± 19.52 vs. 100.27 ± 38.47, P = 0.013) and invasion (46.45 ± 6.70 vs. 102.00 ± 3.54, P = 0.000; 49.45 ± 17.68 vs. 81.45 ± 1.75, P = 0.000) were both accelerated dramatically compared with negative group, nevertheless, the migration (78.10 ± 24.95 vs. 79.38 ± 22.70, P = 0.899) and invasion (51.30 ± 14.95 vs. 52.38 ± 13.70, P = 0.858) of MKN-28 cell was not significantly affected even if under the same concentration of miR-106a inhibitor (Fig. 4). GES-1 is an immortalized cell and did not have any metastatic ability even upon induction by miR-106a.
Knockdown of miR-106a suppresses the implantation of gastric cancer cells in vivo
The findings above illustrated that miR-106a expression was correlated with gastric cancer metastatic capacity in vitro. In order to further attest the oncogenic role of miR-106a, we conducted animal experiment to investigate whether miR-106a could promote tumorigenicity in vivo. Hsa-miR-106a-antagomir was transfected into BGC-823 cell to antagonize the high level of miR-106a in this cell line. Treated cells were injected into the abdomen of nude mice to construct the tumor xenograft mouse model (Fig. 5a). As exhibited in Fig. 5b, the xenograft on incision, great omentum, mesenterium, hepatic portal, splenic hilus, and renal hilus in anti-miR-106a group were totally smaller than antagomir control. HE staining showed that gastric cancer cell of anti-miR-106a group had a large necrotic area. Since we had confirmed that TIMP2 was a direct target of miR-106a in our previous study, we stained TIMP2 in these nodules and found that the protein level of TIMP2 in anti-miR-106a group was higher than those of control group, and the high expression of TIMP2 corresponds to the low expression of miR-106a detected by ISH technology (Fig. 5c). In addition, we had observed that SGC-7901 cell apoptosis was induced after using miR-106a inhibitor in vitro, the observation of the apoptotic phenomenon in vivo study was needed to make further investigation of the effect of miR-106a. TEM was chosen to evaluate the ultra-structure of BGC-823 cells induced with miR-106a antagomir. TEM showed tumor cells in anti-miR-106a group developed towards apoptosis morphologically in which apoptotic body was seen in several visual fields under microscope (Fig. 5d). The results in vivo tallied with that of in vitro survey. Thus, in vitro and in vivo study both suggested that miR-106a may probably acted as tumor oncogene to promote gastric cancer progression.
Knockdown of miR-106a decreases the gastric cancer peritoneal implantation ability in vivo. a Animal model were built with cancer cells inoculated intraperitoneally through a small incision at cartilago ensiformis of upper abdomen. b Antagomir-106a-transfected and antagomir-control-transfected BGC-823 cells were injected into nude mice. Xenograft of incision, great omentum, mesenterium, hepatic portal, splenic hilus, and renal hilus were shown. c The inversely correlation of miR-106a and TIMP2 in metastatic nodules. It is represented by the nodules from great omentum. d Representative TEM microphotograph with qualitative observation of the apoptotic bodies on the great omental nodules with anti-miR-106a transfected. Gastric cancer cells exhibited morphological signs of apoptosis, such as chromatin condensation in the nucleus and nuclear fragments adhering to the membrane (upper panel) and an intact apoptotic body with high electron-dense shadow (lower panel) ×1000
Discussion
Increasing studies have demonstrated that miR-106a is always enhanced in many malignancies, especially gastric cancer [13–15]. Up-regulation of miR-106a in gastric cancer is supposed to have some important functions in tumorigenesis and development. As our previous study reported, over-expressed miR-106a has strong correlation with gastric cancer progression [13]. To fully explore the underlying mechanisms, we carried out in vitro and in vivo experiments to inquire into the problem.
In this study, we further examined the expression level of miR-106a in FFPE samples and paid attention to its relationship with gastric histological classification. Its high expression reflected the oncogenic character and also reflected that miRNA from FFPE was always stable and could not be degraded easily throughout formalin treatment. Subsequently, gain-of-function study was implemented in vitro. The result identified that the cell proliferations were different among three gastric cancer cells. In our another paper, we used sequence-specific inhibitor to silence the expression of miR-106a, and observed that the same cells proliferation were depressed, combined with the previous PCR detection that the expression of miR-106a was related to the high-grade Lauren’s type and high-grade cancer cells, the conclusion should be reached that the dysregulation of miR-106a could promote gastric cancer proliferation at least directed against highly malignant cells. Because of the balance between cell proliferation and apoptosis is the most classical mechanism for carcinoma occurrence [16, 17], we wonder whether miR-106a could cause cell apoptosis. FACS assay approved our speculation. Similarly, Wang reported that suppression of miR-106a significantly inhibited gastric cancer cell proliferation and triggered apoptosis [15]. To further clarify which was responsible to the effect elicited by miR-106a in highly malignant gastric cancer cell, miRbase database was selected to retrieval the potential target of miR-106a. Three open access programs—TargetMiner, miRanda and miRTarBase—to predict the target of miR-106a. FAS, which is known to be initiate the extrinsic apoptotic pathway, was authenticated by report gene detection and western blot. From our result, the FAS was direct regulated by miR-106a at posttranscriptional level. Moreover, besides FAS, another apoptosis-related gene caspase3 was also affected by miR-106a and elevated at protein level. FAS is a member of a subset of the TNF receptor super-family known as ‘death receptors’ [18, 19]. By combined with FASL, the death domain of FAS will be exposed, and then recognize and activate the caspase8, followed by caspase3, 7, which eventually lead to apoptosis [20, 21]. Therefore, the results above suggested that miR-106a may promote gastric cancer cell proliferation and inhibit cell apoptosis through FAS-induced extrinsic pathway. Expect for impact on cell growth, the cell motility was also improved in the same cells. Related research also found that a high level of miR-106a could promote cell proliferation, self-renewal and invasion through epithelial-mesenchymal transition [9, 22]. Taken together, our study demonstrated that miR-106a may play an oncogenic role in gastric cancer initiation and progression through promote cell growth, migration and invasion.
It is known that the peritoneal carcinomatosis is the most common metastatic type for advanced gastric cancer and almost account for half of the proportion even after standardized radical gastrectomy [23–25]. The morphological characteristics of gastric cancer are the key factors to determine the ability of peritoneal metastasis. Gastric carcinoma can be categorized into four types. I: tubular differentiation-well, mucus in cytoplasm-poor; II: tubular differentiation-well, mucus in cytoplasm-rich; III: tubular differentiation-poor, mucus in cytoplasm-poor; IV: tubular differentiation-poor, mucus in cytoplasm-rich [26]. Type IV is more inclined to peritoneal dissemination among these four types. BGC-823, which is originated from poorly differentiated gastric cancer tissues, has the strongest ability to implantation than other cells in our experiment since its diffuse signet-ring cell cancer characteristics were observed in HE sections. As reported, the high risk of peritoneal recurrence is particularly in patients with diffuse-mixed tumors and when there is serosal infiltration [27]. Therefore, the peritoneal carcinomatosis is the most important problem that impedes gastric cancer treatment and impels us to elucidate. In vivo study, injection of BGC-823 cells, peritoneal dissemination was very obviously and consistent with another report that peritoneal cancer nodules of various sizes were throughout the whole abdominal cavity and mainly implanted into omentum, intestines were also be obstructed [28]. Beyond that, the metastatic organ observed within the abdomen should including hepatic portal, splenic hilus, and renal hilus because the abdominal wall was incised when injection. The omentum and mesenterium are the most common implanted position because they are membranous structure within the cavity and the area is the largest. The liver, the spleen and kidney were also excised because the carcinoma cells implanted into the gate that blood vessels flow in and out. No matter which kind of organs, the tumor nodules derived from miR-106a group were significantly smaller than those of untreated group, whatever the number or size. TIMP2, which was confirmed as a direct target of miR-106a from the luciferase assays in our previous studies, was detected to verify the influence of miR-106a on cancer dissemination in vivo. As expected, the protein level of TIMP2 in miR-106a antagomir group was higher than control. TIMP2 was inversely correlated with miR-106a on the basis of the ISH proved that miR-106a was silenced. The result suggests that miR-106a may contribute gastric cancer peritoneal carcinomatosis through suppressing TIMP2. It has been reported that TIMP2 has an affinity with MMP2 and this reaction will abrogate the collagenases and gelatinases activity of MMP2, reduce the degradation of extracellular matrix (ECM) and maintain the integrity of basement membrane [29, 30]. MMPs are acknowledged as key player in the regulation of cell-extracellular matrix interactions [31, 32]. When TIMP2 was inhibited, the degradation of ECM was accelerated and tumor metastasis was facilitated. In addition, we observed several apoptotic bodies in miR-106a antagomir group, which supported the result from in vitro experiment that miR-106a suppression could advance the gastric cancer cell death. Thus, in vitro and in vivo study both suggested that miR-106a may exert positive effects in gastric cancer.
In conclusion, we demonstrated that miR-106a functioned as an oncogene through potentially targeting FAS and TIMP2 in gastric cancer involving in promotion of cell growth, invasion and peritoneal implantation. These findings provide new insights into the biological functions of miR-106a contributing to gastric cancer proliferation and metastasis.
References
GLOBOCAN 2012: Estimated Cancer Incidence, Mortality and Prevalence Worldwide in 2012 (2014) http://globocan.iarc.fr/Pages/fact_sheets_cancer.aspx?cancer=all. Accessed 22 July 2014
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A (2015) Global cancer statistics, 2012. CA Cancer J Clin 65(2):87–108. doi:10.3322/caac.21262
Piazuelo MB, Correa P (2013) Gastric cancer: overview. Colomb Med 44(3):192–201
Lujambio A, Lowe SW (2012) The microcosmos of cancer. Nature 482(7385):347–355. doi:10.1038/nature10888
Schirle NT, Sheu-Gruttadauria J, MacRae IJ (2014) Structural basis for microRNA targeting. Science 346(6209):608–613. doi:10.1126/science.1258040
Kloosterman WP, Plasterk RH (2006) The diverse functions of microRNAs in animal development and disease. Dev Cell 11(4):441–450. doi:10.1016/j.devcel.2006.09.009
Bushati N, Cohen SM (2007) microRNA functions. Annu Rev Cell Dev Biol 23:175–205. doi:10.1146/annurev.cellbio.23.090506.123406
Landais S, Landry S, Legault P, Rassart E (2007) Oncogenic potential of the miR-106-363 cluster and its implication in human T-cell leukemia. Cancer Res 67(12):5699–5707. doi:10.1158/0008-5472.can-06-4478
Li P, Xu Q, Zhang D, Li X, Han L, Lei J, Duan W, Ma Q, Wu Z, Wang Z (2014) Upregulated miR-106a plays an oncogenic role in pancreatic cancer. FEBS Lett 588(5):705–712. doi:10.1016/j.febslet.2014.01.007
Wang Z, Wang B, Shi Y, Xu C, Xiao HL, Ma LN, Xu SL, Yang L, Wang QL, Dang WQ, Cui W, Yu SC, Ping YF, Cui YH, Kung HF, Qian C, Zhang X, Bian XW (2014) Oncogenic miR-20a and miR-106a enhance the invasiveness of human glioma stem cells by directly targeting TIMP-2. Oncogene. doi:10.1038/onc.2014.75
Hong L, Lai M, Chen M, Xie C, Liao R, Kang YJ, Xiao C, Hu WY, Han J, Sun P (2010) The miR-17-92 cluster of microRNAs confers tumorigenicity by inhibiting oncogene-induced senescence. Cancer Res 70(21):8547–8557. doi:10.1158/0008-5472.CAN-10-1938
Yue B, Sun B, Liu C, Zhao S, Zhang D, Yu F, Yan D (2015) Long non-coding RNA Fer-1-like protein 4 suppresses oncogenesis and exhibits prognostic value by associating with miR-106a-5p in colon cancer. Cancer Sci 106(10):1323–1332. doi:10.1111/cas.12759
Zhu M, Zhang N, He S, Lui Y, Lu G, Zhao L (2014) MicroRNA-106a targets TIMP2 to regulate invasion and metastasis of gastric cancer. FEBS Lett 588(4):600–607. doi:10.1016/j.febslet.2013.12.028
Zhu M, Zhang N, He S (2014) Similarly up-regulated microRNA-106a in matched formalin-fixed paraffin-embedded and fresh frozen samples and the dynamic changes during gastric carcinogenesis and development. Pathol Res Pract 210(12):909–915. doi:10.1016/j.prp.2014.07.008
Wang Z, Liu M, Zhu H, Zhang W, He S, Hu C, Quan L, Bai J, Xu N (2013) miR-106a is frequently upregulated in gastric cancer and inhibits the extrinsic apoptotic pathway by targeting FAS. Mol Carcinog 52(8):634–646. doi:10.1002/mc.21899
Fox JG, Wang TC (2007) Inflammation, atrophy, and gastric cancer. J Clin Investig 117(1):60–69. doi:10.1172/JCI30111
Goubran HA, Kotb RR, Stakiw J, Emara ME, Burnouf T (2014) Regulation of tumor growth and metastasis: the role of tumor microenvironment. Cancer Growth Metastasis 7:9–18. doi:10.4137/CGM.S11285
Cullen SP, Martin SJ (2015) Fas and TRAIL ‘death receptors’ as initiators of inflammation: implications for cancer. Semin Cell Dev Biol 39:26–34. doi:10.1016/j.semcdb.2015.01.012
Vilmont V, Filhol O, Hesse AM, Coute Y, Hue C, Remy-Tourneur L, Mistou S, Cochet C, Chiocchia G (2015) Modulatory role of the anti-apoptotic protein kinase CK2 in the sub-cellular localization of Fas associated death domain protein (FADD). Biochim Biophys Acta 1853(11 Pt A):2885–2896. doi:10.1016/j.bbamcr.2015.08.001
Walczak H, Krammer PH (2000) The CD95 (APO-1/Fas) and the TRAIL (APO-2L) apoptosis systems. Exp Cell Res 256(1):58–66. doi:10.1006/excr.2000.4840
Lavrik IN (2014) Systems biology of death receptor networks: live and let die. Cell Death Dis 5:e1259. doi:10.1038/cddis.2014.160
Yu D, Shin HS, Lee YS, Lee YC (2014) miR-106b modulates cancer stem cell characteristics through TGF-beta/Smad signaling in CD44-positive gastric cancer cells. Lab Investig 94(12):1370–1381. doi:10.1038/labinvest.2014.125
Moon YW, Jeung HC, Rha SY, Yoo NC, Roh JK, Noh SH, Kim BS, Chung HC (2007) Changing patterns of prognosticators during 15-year follow-up of advanced gastric cancer after radical gastrectomy and adjuvant chemotherapy: a 15-year follow-up study at a single korean institute. Ann Surg Oncol 14(10):2730–2737. doi:10.1245/s10434-007-9479-4
Glockzin G, Piso P (2012) Current status and future directions in gastric cancer with peritoneal dissemination. Surg Oncol Clin N Am 21(4):625–633. doi:10.1016/j.soc.2012.07.002
Coccolini F, Gheza F, Lotti M, Virzi S, Iusco D, Ghermandi C, Melotti R, Baiocchi G, Giulini SM, Ansaloni L, Catena F (2013) Peritoneal carcinomatosis. World J Gastroenterol 19(41):6979–6994. doi:10.3748/wjg.v19.i41.6979
Goseki N, Takizawa T, Koike M (1992) Differences in the mode of the extension of gastric cancer classified by histological type: new histological classification of gastric carcinoma. Gut 33(5):606–612
Roviello F, Marrelli D, de Manzoni G, Morgagni P, Di Leo A, Saragoni L, De Stefano A (2003) Prospective study of peritoneal recurrence after curative surgery for gastric cancer. Br J Surg 90(9):1113–1119. doi:10.1002/bjs.4164
Mei LJ, Yang XJ, Tang L, Hassan AH, Yonemura Y, Li Y (2010) Establishment and identification of a rabbit model of peritoneal carcinomatosis from gastric cancer. BMC Cancer 10:124. doi:10.1186/1471-2407-10-124
Liotta LA, Stetler-Stevenson WG (1991) Tumor invasion and metastasis: an imbalance of positive and negative regulation. Cancer Res 51(18 Suppl):5054s–5059s
Murphy G (2011) Tissue inhibitors of metalloproteinases. Genome Biol 12(11):233. doi:10.1186/gb-2011-12-11-233
Murphy G, Nagase H (2008) Progress in matrix metalloproteinase research. Mol Aspects Med 29(5):290–308. doi:10.1016/j.mam.2008.05.002
Kessenbrock K, Plaks V, Werb Z (2010) Matrix metalloproteinases: regulators of the tumor microenvironment. Cell 141(1):52–67. doi:10.1016/j.cell.2010.03.015
Acknowledgments
This work was supported by Grants from Ningxia Natural Science Foundation of China (No. NZ1230).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no financial interest to disclose.
Rights and permissions
About this article
Cite this article
Zhu, M., Zhang, N., He, S. et al. MicroRNA-106a functions as an oncogene in human gastric cancer and contributes to proliferation and metastasis in vitro and in vivo. Clin Exp Metastasis 33, 509–519 (2016). https://doi.org/10.1007/s10585-016-9795-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10585-016-9795-9