Abstract
Human chromosomes are arranged in a linear and conserved sequence order that undergoes further spatial folding within the three-dimensional space of the nucleus. Although structural variations in this organization are an important source of natural genetic diversity, cytogenetic aberrations can also underlie a number of human diseases and disorders. Approaches for studying chromosome structure began half a century ago with karyotyping of Giemsa-banded chromosomes and has now evolved to encompass high-resolution fluorescence microscopy, reporter-based assays, and next-generation DNA sequencing technologies. Here, we provide a general overview of experimental methods at different resolution and sensitivity scales and discuss how they can be complemented to provide synergistic insight into the study of human chromosome structural rearrangements. These approaches range from kilobase-level resolution DNA fluorescence in situ hybridization (FISH)-based imaging approaches of individual cells to genome-wide sequencing strategies that can capture nucleotide-level information from diverse sample types. Technological advances coupled to the combinatorial use of multiple methods have resulted in the discovery of new rearrangement classes along with mechanistic insights into the processes that drive structural alterations in the human genome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA copy number alterations and structural variants are a normal part of genetic variation between individuals but can also be a detrimental byproduct of genomic instability. Structural chromosomal alterations—including deletions, insertions, inversions, and translocations—can arise from this instability and underlie the pathology of human diseases and genomic disorders (Lupski 2007). Through the development and use of cytogenetic approaches, some cancers were discovered to be the direct consequence of abnormal structural changes in chromosome sequence order. The first observation was made by Janet Rowley in 1973, who reported that a gross chromosomal rearrangement (GCR) producing a translocation between chromosomes 9 and 21 was consistently present across chronic myelogenous leukemia patients (Rowley 1973). Genomic disorders associated with architectural changes and DNA copy number alterations throughout different chromosomes range from red-green color blindness to hemophilia A, which can be caused by deletion and inversion events, respectively (Lupski 1998).
Cytogenetic analysis of stained mitotic spread preparations comprised one of the earliest methods for the analysis of chromosome structure. Karyotyping is a rapid and cost-effective microscopy-based technique commonly used in diagnostic laboratories for the detection of aneuploidy and visible GCRs from a variety of clinical specimens. With proper cytogenetic training, each chromosome can be identified by a unique banding pattern. The implementation of molecular biology and genomic-based approaches has significantly increased the sensitivity and resolution limits for the detection of GCRs. DNA fluorescence in situ hybridization (FISH) techniques were a major contributor to the field of cytogenetics by highlighting specific genomic regions or chromosomes of interest with user-specified fluorescent DNA probes. Advancements in microscopy have expanded the resolution range for detecting GCRs, although imaging-based approaches remain limited by the inherent challenges of visualizing submicroscopic alterations. More recently, the evolution of second- and third-generation DNA sequencing technologies has accelerated the discovery of GCRs that previously escaped microscopic recognition, enabling the systematic evaluation of structural variants at single-nucleotide resolution (Le Scouarnec and Gribble 2012).
Each method for studying structural chromosomal alterations offers a unique balance between resolution, sensitivity, throughput, time, and cost. For example, karyotyping-based approaches are rapid and require minimal sample processing but can be limited to lower resolution analyses of actively dividing cell types. In contrast, DNA sequencing can detect structural abnormalities at exceptionally high resolution but can be an expensive and lengthy process. In this review, we describe various methods developed over the past few decades (Fig. 1) focusing on the interrogation of human GCRs and discuss their corresponding advantages and limitations for both clinical and experimental use. We further discuss how the combination of cellular and genomic approaches has resulted in the discovery of novel rearrangement types implicated in human disease and the mechanisms that drive their formation.
Seeing is believing: microscopy-based approaches
Classical cytogenetics
Karyotyping is a classic cytogenetic technique used to visualize the complete set of genetic material within individual cells, which can detect abnormalities in chromosome number (aneuploidy) and large-scale structural aberrations (Trask 2002). Initial methods from 1956 utilizing chromosome spreads prepared under hypotonic conditions followed by acetic orcein and Feulgen stains were useful in establishing the normal diploid content of human cells (Tjio and Levan 1956; Ford and Hamerton 1956). However, these stains could not detect structural abnormalities because the uniform staining pattern prevented the identification of distinct chromosomes. Fine structural details were subsequently revealed by banding techniques that were applied to mitotic chromosome spreads. Dating back to 1971, G-banding remains the most commonly used diagnostic technique in clinical cytogenetic laboratories to differentiate individual chromosomes. Mitotic spread preparations typically undergo partial digestion with trypsin and exposure to Giemsa stain followed by conventional light microscopy for visualization (Fig. 2a) (Sumner et al. 1971; Schnedl 1971; Seabright 1971; Yunis et al. 1971). G-banded chromosomes exhibit oscillating light and dark banding patterns caused by the distribution and abundance of AT-rich and GC-rich sequences, producing characteristic patterns unique for each chromosome (Drets and Shaw 1971; Speicher and Carter 2005). Complementary approaches to G-banding include quinacrine staining for AT-rich regions and C-banding for heterochromatic regions of centromeres and the heterochromatic repeats located on the chromosome Yq arm. In addition, silver staining can be used to identify nucleolus organizing regions (NORs) containing ribosomal DNA repeats clustered on the short arm of acrocentric human chromosomes (Goodpasture and Bloom 1975).
Microscopy approaches for the detection of structural chromosomal rearrangements. a Giemsa-banded karyotype derived from an acute lymphoblastic leukemia patient exhibiting two rearrangements. b Rearrangements detected with DNA paint probes targeting chromosome 15 in colorectal cancer cells following exposure to 2 Gy ionizing radiation. c Multi-color chromosome painting of a colorectal cancer cell line carrying a rearrangement between chromosomes 10, 13, and 14. d Schematics of interphase nuclei hybridized to the indicated FISH probes to resolve different types of DNA copy number and structural alterations
DNA fluorescence in situ hybridization
The application of banding-based cytogenetic approaches can be limited by restrictions on cell type and resolution. Proper cytogenetic training and experience are also required to accurately identify each chromosome based on their distinctive banding patterns. Hybridization-based methods, which leverage the ability of DNA strands to denature and re-hybridize, have been a transformative approach for molecular cytogenetics to detect specific DNA sequences of interest with high sensitivity. In 1969, Gall and Pardue used in situ hybridization of radioactive DNA/RNA (Gall and Pardue 1969) and subsequently DNA/DNA (Pardue and Gall 1969) hybrids to map the location of ribosomal DNA sequences in Xenopus oocyte nuclei. The development of DNA FISH (Rudkin and Stollar 1977)—which utilizes fluorescent DNA probes targeting entire chromosomes (paint probes), repetitive genomic sequences (e.g., centromeres, telomeres), or specific loci—enabled the routine visual inspection of chromosome copy number and structure without extensive cytogenetic training.
Chromosome painting has facilitated the accurate identification of chromosomal aberrations (Ried et al. 1998). Initially developed in 1988, chromosome-specific paint probes were made from cloned DNA libraries derived from flow-sorted human chromosomes (Cremer et al. 1988; Lichter et al. 1988). Additionally, chromosome flow-sorting or microdissection followed by PCR amplification has been used to generate whole-chromosome and region-specific paint probes (Telenius et al. 1992). Paint probes are useful for enumerating and confirming inter-chromosomal rearrangements that are challenging to discern by banding-based karyotyping (Fig. 2b). However, chromosome painting is less sensitive for resolving intra-chromosomal rearrangements. Previous studies have leveraged multicolor-banding probes that allow the differentiation of chromosome region (Chudoba et al. 1999; Kosyakova et al. 2013) or paint probes that recognize large, distinct segments of a single chromosome (Ly et al. 2019) to identify intra-chromosomal rearrangements from mitotic spreads.
Repetitive DNA sequences at centromeres and telomeres can also be targeted by FISH probes (Roschke et al. 1996; Genet et al. 2013; Giunta 2018). Individual centromeres harbor a distinct array of repetitive sequence that can be labeled by FISH (Gutiérrez-Mateo et al. 2005; Baumgartner et al. 2006), providing another strategy for identifying chromosomes. Pan-centromere probes can simultaneously label all centromeres by recognizing the α-satellite repeat sequences shared between all human centromeres. This can be useful for detecting acentric or dicentric chromosomes on mitotic spreads. When used in combination with an active centromere marker (e.g., CENP-A), pan-centromere probes can also be leveraged to identify pseudo-dicentric chromosomes carrying an active and inactive centromere or dicentric chromosomes harboring two functional centromeres (Amor et al. 2004). Similarly, telomere probes can be useful for labeling all chromosome ends and detecting dicentric chromosomes caused by chromosome fusions events that produce an interstitial telomeric signal. Subtelomeric probes label specific sequences immediately adjacent to the telomeres of individual chromosomes and can be useful in analyzing GCRs affecting chromosome ends.
Locus-specific probes, originally targeting defined breakpoints (Tkachuk et al. 1990) or derived from chromosome microdissection (Meltzer et al. 1992), can be used to label specific and non-repetitive regions of the genome. Bacterial artificial chromosomes (BACs) harboring cloned genomic segments are now commonly used vectors for producing locus-specific FISH probes (Shizuya et al. 1992; Cheung et al. 2001). These probes carry an average size range of 100–200 kb (Asakawa et al. 1997; Osoegawa et al. 2001) and have been used for diverse applications, ranging from the characterization of chromosomal abnormalities (Bishop 2010) to the determination of three-dimensional nuclear architecture (Steinhaeuser et al. 2002; Nora et al. 2012). Resources for selecting and acquiring defined BAC clones targeting defined regions of the human genome are widely accessible from the UCSC Genome Browser and BACPAC Resources (Children’s Hospital Oakland Research Institute), respectively.
Multi-color chromosome labeling
Chromosome- and region-specific probes require a prior expectation of a rearrangement event of interest, which limits the identification of unexpected GCRs. Spectral karyotyping (SKY) and multiplex FISH (M-FISH)—originally developed in 1996—are similar to painting-based FISH approaches but allows the co-labeling and simultaneous visualization of all 24 human chromosomes from mitotic spread preparations (Schrock et al. 1996; Speicher et al. 1996). Instead of differentiating each chromosome based on light/dark banding patterns or specific FISH probes, SKY and M-FISH probes label each chromosome entirely with unique color combinations. This is achieved by the hybridization of multiple fluorescent probes followed by computational separation on the basis of unique spectral properties for each chromosome. This strategy is particularly useful for identifying subtle inter-chromosomal rearrangements, which are made apparent due to mismatching colors between non-homologous chromosomes (Fig. 2c) (Veldman et al. 1997). Similar to chromosome painting probes, a drawback to this method is the inherent difficulty in exposing intra-chromosomal rearrangements given that individual chromosomes are painted a single and uniform color. This approach, however, provides a comprehensive genomic overview to identify GCRs without a priori knowledge of the affected chromosome(s).
Interphase FISH
Karyotypic analysis requires the preparation of replicated and condensed mitotic chromosomes derived from actively dividing cell populations, which can limit the utility of this method for slow cycling, non-dividing, and/or post-mitotic cells. This can be bypassed by artificially inducing premature chromosome condensation using small molecule phosphatase inhibitors (e.g., calyculin A) (Tosuji et al. 1992) to produce mitotic-like spreads from asynchronous interphase cells (Ly et al. 2017), although the side effects of these inhibitors on chromosome structure are poorly understood. However, molecular cytogenetic approaches can be directly applied to interphase cells to assess decondensed chromatin, which can facilitate the rapid detection of GCRs for screening and diagnostic purposes without extensive experimental manipulation.
Locus-specific FISH probes are particularly useful for detecting DNA copy number alterations (e.g., deletions, amplifications), aneuploidy using centromere enumeration probes, GCRs, and chromosome breaks in the interphase nucleus (Olsen et al. 2004; Feuk et al. 2006; Vorsanova et al. 2010; Rudin et al. 2012) (Fig. 2d). For example, translocations can be detected by utilizing two-color probes recognizing different chromosome regions at putative rearrangement junctions (Sugita and Hasegawa 2017). The fusion of two normally separated inter-chromosomal signals provides support for the presence of a translocation. In contrast, split signal or breakapart probes, which span or flank a breakpoint of interest, can be used to detect chromosome breaks by generating two separated foci instead of one co-localized signal indicative of an intact chromosome (van der Burg et al. 2004). Panels of locus-specific probes that detect specific gene rearrangement events or oncogene copy number alterations are commercially available and frequently utilized in clinical diagnostic laboratories. Importantly, these approaches require some knowledge of the chromosomal abnormality of interest in order to select and implement specific probe sets targeting these regions.
Interphase FISH can also be quantitative in the assessment of large cell numbers compared to traditional cytogenetic approaches. For example, hundreds to thousands of cells can be readily screened for the presence of a specific GCR within a bulk tumor population. Computational algorithms have also been developed in conjunction with high-throughput imaging strategies to quantitatively measure rare breakage and translocation events at specific loci within the interphase nucleus (Burman et al. 2015). Due to the decondensed nature of DNA during interphase, the resolution of interphase FISH can exceed that of mitotic spreads, ranging from 50 kb to 2 Mb (Speicher and Carter 2005). This resolution can be further improved by using mechanically stretched DNA fibers in conjunction with FISH (fiber FISH) (Wang et al. 1998), increasing the resolution to a general range of 1–500 kb (Raap et al. 1996) and enabling the detection of small abnormalities within a specific region of the genome (Weier 2001; Ju et al. 2015; Nguyen et al. 2017).
Comparative genomic hybridization-based approaches
Hybridization-based approaches suitable for genome-wide analyses include comparative genomic hybridization (CGH) and array CGH, which can infer differences in DNA copy number between two samples (e.g., biopsy from a tumor compared to matched normal tissue). CGH-based approaches combine and fragment two genomic DNA samples for hybridization to a reference mitotic spread or array (Kallioniemi et al. 1992; Solinas-Toldo et al. 1997). Changes in hybridization patterns are indicative of DNA copy number alterations between the two samples, providing information regarding segmental, arm-level, or whole-chromosome copy number changes. Single-nucleotide polymorphism (SNP) arrays are another useful method for genome-wide copy number analysis, which can identify DNA copy number changes on the basis of paternal and maternal SNP ratios (Wang et al. 1998). Although these approaches can offer better resolution than conventional banding or most fluorescent applications, rearrangements that result in copy number neutral events (e.g., balanced translocations) are not detectable.
Recent advances in cytogenetics
Over the past decade, a number of approaches have been developed that has enabled cytogenetics to be performed with higher efficiency, specificity, and resolution. FISH probes can be laborious to prepare from BAC clones, whereas commercially available probes are expensive and limited to pre-existing probe sets. Advancements in DNA synthesis technologies have enabled customizable FISH probes to be generated from synthesized oligonucleotide arrays, including COMBO-FISH (Hausmann et al. 2003) and Oligopaint approaches (Beliveau et al. 2012). Oligo-based FISH probes have emerged as an increasingly popular method to label user-selected sites with higher specificity and resolution. Synthesized oligos are renewable through PCR-based amplification, representing an affordable alternative to commercial probes. Thousands of oligos can be designed bioinformatically (Beliveau et al. 2018) and synthesized to target specific genomic sequences, arrayed to tile across non-repetitive genomic regions, or paint entire chromosomes. This experimental toolkit can expand substantially when combined with methods to amplify weak signals to visualize discrete sequences in the genome (Beliveau et al. 2015; Kishi et al. 2019).
The limitation of cytogenetic analysis on characterizing GCRs is partly due to the resolution power of microscopy-based approaches. Recent advancements in microscopy have greatly improved the resolution and quantitative capabilities of fluorescent imaging. Studies utilizing super-resolution microscopy, including 3D structured illumination microscopy (3D-SIM) (Gustafsson et al. 2008) and stochastic optical reconstruction microscopy (STORM) (Rust et al. 2006), are beginning to emerge for the analysis of interphase and mitotic chromosomes hybridized to FISH probes, significantly increasing the resolution of conventional FISH (Beliveau et al. 2017; Potapova et al. 2019). For example, 3D-STORM has been used to directly image short non-repetitive DNA sequences as short as ~ 2.5 kb in human and mouse cells (Ni et al. 2017).
Live-cell imaging approaches have also recently emerged that allows GCR formation to be visualized in real-time by time-lapse microscopy. Early examples monitored fluorescently labeled bacterial lac repressors bound to two different lac operon arrays integrated into the genome of mammalian cells (Roukos et al. 2013; Roukos et al. 2014). The induction of DNA DSBs at these arrays permitted the tracking of chromosome break ends to undergo aberrant DNA repair. More recently, modifications to the CRISPR system have now allowed endogenous genomic loci to be detected in real-time, leveraging either a nuclease-deficient Cas9 mutant fused to a fluorescent marker (Chen et al. 2013) or through the direct labeling of guide RNAs with a fluorescent molecule (Wang et al. 2019). These approaches have enabled the monitoring of GCR formation in real time, for example, following the induction of multiple DSBs in the genome (Wang et al. 2019).
Quantitative measurements of rare rearrangement events
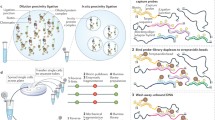
Studying genetic pathways involved in GCR formation can be inherently challenging due to the inefficiencies associated with experimentally generating specific rearrangement events. To systematically interrogate factors that are involved in maintaining genome stability, reporter-based assays can be designed to detect and/or recover specific GCRs formed after DSB induction (Fig. 3). These assays commonly use positive selection to isolate rare events by drug resistance or flow cytometry (Pierce et al. 1999). In an early example, a translocation reporter was developed in which two exons of a neomycin resistance gene are targeted separately to chromosomes 14 and 17 in mouse embryonic stem cells. After inducing DSBs using an I-SceI endonuclease on both chromosomes, an inter-chromosomal translocation resulted in a functional neomycin resistance gene that confers tolerance to antibiotic selection. The frequency of rare translocation events can be extrapolated from the number of survivors following drug selection (Weinstock et al. 2006). Chromosomal inversions and deletions can also be detected following CRISPR/Cas9-induced site-specific DSBs (Li et al. 2015; Mendez-Dorantes et al. 2018; Hu et al. 2019). In these systems, fluorescent protein coding sequences are designed to be non-functional prior to the induction of a specific rearrangement event that activates the fluorescent marker, which can be detected by flow cytometry. These strategies have enabled exceptionally rare GCR events to be detectable and assayed in a quantitative manner, for example, as low as 4 × 10−5 using drug selection approaches (Weinstock et al. 2006).
Detection of rare rearrangement events following DNA double-strand break induction. a Schematic to detect intra-chromosomal deletions. A stop cassette flanked by two experimentally induced double-strand break (DSB) sites is inserted between a promoter and GFP coding sequences. After DSB induction, a deletion will result in GFP expression that can be detected by flow cytometry. b Schematic to detect intra-chromosomal inversions. GFP coding sequence is placed in an inverted orientation relative to the promoter. After DSB induction, a correct inversion event will result in GFP expression that can be detected by flow cytometry. c Schematic to detect a translocation. A neomycin resistance gene is split within an intron, each part targeted to two non-homologous chromosomes. DSB induction can result in an inter-chromosomal translocation that brings two separate parts of the neo gene together. The percentage of G418-resistant clones after selection reflects the relative translocation frequency. d Schematic to detect balanced translocations. Two DSBs are induced at different genomic sites, and non-reciprocal or reciprocal translocations generated after DSB repair can be detected by PCR
Another design of the translocation assay is to simultaneously induce DSBs on two chromosomal loci and detect the derivative translocation by nested PCR with primers flanking the putative breakpoint regions (Mateos-Gomez et al. 2015). Altogether, these strategies have been important for elucidating the role of various genetic pathways in facilitating the formation of GCRs. A restriction to these approaches, however, is that these assays are usually designed to detect a specific rearrangement event and therefore requires a prediction of the expected rearrangement outcome. In 2011, translocation sequencing approaches were developed to quantitatively survey all possible rearrangement partners throughout the mouse genome following the induction of a single DSB using an I-SceI endonuclease (Klein et al. 2011; Chiarle et al. 2011).
DNA sequencing-based approaches
Advances in DNA sequencing technology have revolutionized the analyses of human genomes at unprecedented resolution. Nucleotide level information can be captured from a diverse range of sample types, ranging from single cells and bulk tissue to laser microdissected tumor biopsies. Low coverage DNA sequencing (~ 0.1× to 0.5×) from single cells is more frequently being used as a cost-effective method to extrapolate DNA copy number information across the genome. Similarly, RNA sequencing can also be used for DNA copy number analysis since gene expression mostly correlates with the number of gene copies present (Ben-David et al. 2013). As with CGH-based approaches, low coverage sequencing of genomic DNA or RNA libraries typically yields arm-level information of copy number gains and losses but do not provide sufficient sensitivity for GCR analysis and copy number neutral events.
Cancer genome sequencing at high coverage has resulted in the discovery of novel GCRs that were previously undetectable by cytogenetics. Short-read sequencing can be used to identify GCRs through paired-end reads separated by a linker of a known distance (Fig. 4a). Both reads from a pair are aligned to a reference genome, and pairs mapping at the incorrect distance and/or orientation (discordant pairs) or reads that misalign to two locations within the genome (split-end reads) are indicative of potential GCRs (Campbell et al. 2008). Split-end reads, while typically present at a lower frequency depending on the length of the sequencing read, can provide nucleotide-level sequence information. This can be informative at GCR breakpoint junctions to reveal hallmark signatures that are characteristic of specific DNA repair events. Long-read single-molecule sequencing technologies have more recently emerged and permits long, kilobase tracts of sequences to be covered by a single read (Fig. 4b). Higher error rates are frequently associated with current long-read approaches, although repetitive regions, complex GCRs, and unusual sequence structures are more easily resolved compared to short read sequence mapping. Laboratories with access to sequencers can now easily incorporate next-generation DNA sequencing approaches into their experimental workflow. A number of software and computational pipelines are available for sequence mapping and GCR calling, which has greatly accelerated complex sequencing analysis for researchers with limited access to bioinformatic expertise.
Cytogenetics and genomics are commonly used as complementary methods to provide synergistic information regarding chromosome structure, including the mechanisms that drive GCR formation. For example, complex and localized GCRs known as chromothripsis were initially discovered by SNP analysis and paired-end DNA sequencing from a chronic lymphocytic leukemia patient genome (Stephens et al. 2011). These rearrangements, which were hypothesized to arise from the catastrophic fragmentation of one or a few chromosomes, raised mechanistic questions that have been addressed using a number of experimental methods combining cell biology with genomics. Analysis of mitotic spreads by staining and hybridization techniques revealed that chromosome segregation errors resulting in micronuclei formation can trigger the fragmentation (Ly et al. 2017; Crasta et al. 2012) and rearrangement (Ly et al. 2019) of individual chromosomes. Although the complexity of chromothriptic rearrangements are usually difficult to resolve by microscopy, including super-resolution imaging methods such as SIM (Ly et al. 2019), this has been complemented by the use of whole-genome sequencing of both single cells (Zhang et al. 2015) and clonal populations (Ly et al. 2019; Maciejowski et al. 2015) following the induction of chromothripsis.
Another example emerged from initial studies of drug resistance to EGFR inhibitors caused by increased DNA copy numbers of the EGFR locus. Cytogenetic analysis revealed that EGFR amplification can occur on extrachromosomal DNA (ecDNA) (Nathanson et al. 2014), circular DNA structures that can serve as vehicles for oncogene amplification (Verhaak et al. 2019). The relative frequency of ecDNAs across various cancer types and experimental models have more recently been determined by both cytogenetics and genomics (Turner et al. 2017), estimating that these aberrations are present in approximately half of all human tumors. This work has provided evidence that the physical location and arrangement of genes plays an important role in cancer development and therapeutic resistance.
Exploring the 3D genome
Mammalian genomes are spatially organized and compartmentalized in three-dimensional space that can differ across cell types and states. This is critical for the establishment of long-range chromatin contacts to bridge enhancers and promoters. Such non-random and inherent organization of the genome may also be involved in the propensity of some loci or domains to form rearrangements (Roix et al. 2003; Zhang et al. 2012). Our understanding of genome organization has been revolutionized by the development of techniques such as chromosome conformation capture (3C) (Dekker et al. 2002) and Hi-C (Lieberman-Aiden et al. 2009), which provides an overview of the spatial proximity between DNA sequences in the genome. Additionally, combining these strategies with optical mapping has enabled large and complex GCRs to be detected in cancer genomes (Dixon et al. 2018). Although not routinely used for GCR analysis, these approaches can be coupled to other cytogenetic- and genomic-based methods to elucidate how rearrangements in the 2D linear sequence order can disrupt 3D genome organization. For example, the spatial distances between several loci across rearrangement breakpoints can be further measured using locus-specific FISH probes followed by 3D fluorescence microscopy of the interphase nucleus.
Concluding remarks
The past several decades have witnessed an explosion of new and improved approaches for the identification, detection, and study of GCRs—each bearing a unique set of advantages and limitations. Originally encompassing light microscopy of stained chromosomes, we have now transitioned towards visualizing the formation of de novo rearrangements within living cells and resolving rearrangement breakpoints with single-nucleotide accuracy. Future applications will likely center on bypassing the limitations imposed by current techniques, as well as strategies to more seamlessly integrate complementary methods for parallel analyses.
Abbreviations
- BAC:
-
bacterial artificial chromosome
- DNA DSB:
-
DNA double-strand break
- DNA FISH:
-
DNA fluorescence in situ hybridization
- ecDNA:
-
extrachromosomal DNA
- G-banding:
-
giemsa banding
- GCR:
-
gross chromosomal rearrangement
- NOR:
-
nucleolus organizing region
- SIM:
-
structured illumination microscopy
- SNP:
-
single-nucleotide polymorphism
- STORM:
-
stochastic optical reconstruction microscopy
- WGS:
-
whole-genome sequencing
References
Amor DJ, Bentley K, Ryan J, Perry J, Wong L, Slater H, Choo KH (2004) Human centromere repositioning "in progress". Proc Natl Acad Sci U S A 101:6542–6547
Asakawa S, Abe I, Kudoh Y, Kishi N, Wang Y, Kubota R, Kudoh J, Kawasaki K, Minoshima S, Shimizu N (1997) Human BAC library: construction and rapid screening. Gene 191:69–79
Baumgartner A, Weier JF, Weier H-UG (2006) Chromosome-specific DNA repeat probes. J Histochem Cytochem 54:1363–1370
Beliveau BJ, Joyce EF, Apostolopoulos N, Yilmaz F, Fonseka CY, McCole R, Chang Y, Li JB, Senaratne TN, Williams BR, Rouillard JM, Wu CT (2012) Versatile design and synthesis platform for visualizing genomes with Oligopaint FISH probes. Proc Natl Acad Sci U S A 109:21301–21306
Beliveau BJ et al (2015) Single-molecule super-resolution imaging of chromosomes and in situ haplotype visualization using Oligopaint FISH probes. Nat Commun 6:7147
Beliveau BJ, Boettiger AN, Nir G, Bintu B, Yin P, Zhuang X, Wu CT (2017) In situ super-resolution imaging of genomic DNA with OligoSTORM and OligoDNA-PAINT. Methods Mol Biol 1663:231–252
Beliveau BJ et al (2018) OligoMiner provides a rapid, flexible environment for the design of genome-scale oligonucleotide in situ hybridization probes. Proc Natl Acad Sci 115:E2183–E2192
Ben-David U, Mayshar Y, Benvenisty N (2013) Virtual karyotyping of pluripotent stem cells on the basis of their global gene expression profiles. Nat Protoc 8:989–997
Bishop R (2010) Applications of fluorescence in situ hybridization (FISH) in detecting genetic aberrations of medical significance. Biosci Horiz 3:85–95
Burman B, Misteli T, Pegoraro G (2015) Quantitative detection of rare interphase chromosome breaks and translocations by high-throughput imaging. Genome Biol 16:146
Campbell PJ et al (2008) Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nat Genet 40:722–729
Chen B, Gilbert LA, Cimini BA, Schnitzbauer J, Zhang W, Li GW, Park J, Blackburn EH, Weissman JS, Qi LS, Huang B (2013) Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155:1479–1491
Cheung VG, Nowak N, Jang W, Kirsch IR, Zhao S, Chen XN, Furey TS, Kim UJ, Kuo WL, Olivier M, Conroy J, Kasprzyk A, Massa H, Yonescu R, Sait S, Thoreen C, Snijders A, Lemyre E, Bailey JA, Bruzel A, Burrill WD, Clegg SM, Collins S, Dhami P, Friedman C, Han CS, Herrick S, Lee J, Ligon AH, Lowry S, Morley M, Narasimhan S, Osoegawa K, Peng Z, Plajzer-Frick I, Quade BJ, Scott D, Sirotkin K, Thorpe AA, Gray JW, Hudson J, Pinkel D, Ried T, Rowen L, Shen-Ong GL, Strausberg RL, Birney E, Callen DF, Cheng JF, Cox DR, Doggett NA, Carter NP, Eichler EE, Haussler D, Korenberg JR, Morton CC, Albertson D, Schuler G, de Jong PJ, Trask BJ, BAC Resource Consortium (2001) Integration of cytogenetic landmarks into the draft sequence of the human genome. Nature 409:953–958
Chiarle R, Zhang Y, Frock RL, Lewis SM, Molinie B, Ho YJ, Myers DR, Choi VW, Compagno M, Malkin DJ, Neuberg D, Monti S, Giallourakis CC, Gostissa M, Alt FW (2011) Genome-wide translocation sequencing reveals mechanisms of chromosome breaks and rearrangements in B cells. Cell 147:107–119
Chudoba I, Plesch A, Lörch T, Lemke J, Claussen U, Senger G (1999) High resolution multicolor-banding: a new technique for refined FISH analysis of human chromosomes. Cytogenet Cell Genet 84:156–160
Crasta K, Ganem NJ, Dagher R, Lantermann AB, Ivanova EV, Pan Y, Nezi L, Protopopov A, Chowdhury D, Pellman D (2012) DNA breaks and chromosome pulverization from errors in mitosis. Nature 482:53–58
Cremer T, Lichter P, Borden J, Ward DC, Manuelidis L (1988) Detection of chromosome aberrations in metaphase and interphase tumor cells by in situ hybridization using chromosome-specific library probes. Hum Genet 80:235–246
Dekker J, Rippe K, Dekker M, Kleckner N (2002) Capturing chromosome conformation. Science 295:1306–1311
Dixon JR, Xu J, Dileep V, Zhan Y, Song F, le VT, Yardımcı GG, Chakraborty A, Bann DV, Wang Y, Clark R, Zhang L, Yang H, Liu T, Iyyanki S, An L, Pool C, Sasaki T, Rivera-Mulia JC, Ozadam H, Lajoie BR, Kaul R, Buckley M, Lee K, Diegel M, Pezic D, Ernst C, Hadjur S, Odom DT, Stamatoyannopoulos JA, Broach JR, Hardison RC, Ay F, Noble WS, Dekker J, Gilbert DM, Yue F (2018) Integrative detection and analysis of structural variation in cancer genomes. Nat Genet 50:1388–1398
Drets ME, Shaw MW (1971) Specific banding patterns of human chromosomes. Proc Natl Acad Sci U S A 68:2073–2077
Feuk L, Carson AR, Scherer SW (2006) Structural variation in the human genome. Nat Rev Genet 7:85–97
Ford CE, Hamerton JL (1956) The chromosomes of man. Nature 178:1020–1023
Gall JG, Pardue ML (1969) Formation and detection of RNA-DNA hybrid molecules in cytological preparations. Proc Natl Acad Sci U S A 63:378–383
Genet MD, Cartwright IM, Kato TA (2013) Direct DNA and PNA probe binding to telomeric regions without classical in situ hybridization. Mol Cytogenet 6:42–42
Giunta S (2018) Centromere chromosome orientation fluorescent in situ hybridization (Cen-CO-FISH) detects sister chromatid exchange at the centromere in human cells. Bio-protocol 8:e2792
Goodpasture C, Bloom SE (1975) Visualization of nucleolar organizer regions im mammalian chromosomes using silver staining. Chromosoma 53:37–50
Gustafsson MG et al (2008) Three-dimensional resolution doubling in wide-field fluorescence microscopy by structured illumination. Biophys J 94:4957–4970
Gutiérrez-Mateo C et al (2005) Karyotyping of human oocytes by cenM-FISH, a new 24-colour centromere-specific technique. Hum Reprod 20:3395–3401
Hausmann M et al (2003) COMBO-FISH: specific labeling of nondenatured chromatin targets by computer-selected DNA oligonucleotide probe combinations. Biotechniques 35(564–570):572–567
Hu Q et al (2019) Break-induced replication plays a prominent role in long-range repeat-mediated deletion. EMBO J:e101751
Ju YS et al (2015) Frequent somatic transfer of mitochondrial DNA into the nuclear genome of human cancer cells. Genome Res 25:814–824
Kallioniemi A, Kallioniemi OP, Sudar D, Rutovitz D, Gray JW, Waldman F, Pinkel D (1992) Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors. Science 258:818–821
Kishi JY, Lapan SW, Beliveau BJ, West ER, Zhu A, Sasaki HM, Saka SK, Wang Y, Cepko CL, Yin P (2019) SABER amplifies FISH: enhanced multiplexed imaging of RNA and DNA in cells and tissues. Nat Methods 16:533–544
Klein IA, Resch W, Jankovic M, Oliveira T, Yamane A, Nakahashi H, di Virgilio M, Bothmer A, Nussenzweig A, Robbiani DF, Casellas R, Nussenzweig MC (2011) Translocation-capture sequencing reveals the extent and nature of chromosomal rearrangements in B lymphocytes. Cell 147:95–106
Kosyakova N et al (2013) Generation of multicolor banding probes for chromosomes of different species. Mol Cytogenet 6:6
Le Scouarnec S, Gribble SM (2012) Characterising chromosome rearrangements: recent technical advances in molecular cytogenetics. Heredity (Edinb) 108:75–85
Li Y et al (2015) A versatile reporter system for CRISPR-mediated chromosomal rearrangements. Genome Biol 16:111
Lichter P, Cremer T, Borden J, Manuelidis L, Ward DC (1988) Delineation of individual human chromosomes in metaphase and interphase cells by in situ suppression hybridization using recombinant DNA libraries. Hum Genet 80:224–234
Lieberman-Aiden E, van Berkum N, Williams L, Imakaev M, Ragoczy T, Telling A, Amit I, Lajoie BR, Sabo PJ, Dorschner MO, Sandstrom R, Bernstein B, Bender MA, Groudine M, Gnirke A, Stamatoyannopoulos J, Mirny LA, Lander ES, Dekker J (2009) Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326:289–293
Lupski JR (1998) Genomic disorders: structural features of the genome can lead to DNA rearrangements and human disease traits. Trends Genet 14:417–422
Lupski JR (2007) Genomic rearrangements and sporadic disease. Nat Genet 39:S43–S47
Ly P, Teitz LS, Kim DH, Shoshani O, Skaletsky H, Fachinetti D, Page DC, Cleveland DW (2017) Selective Y centromere inactivation triggers chromosome shattering in micronuclei and repair by non-homologous end joining. Nat Cell Biol 19:68–75
Ly P, Brunner SF, Shoshani O, Kim DH, Lan W, Pyntikova T, Flanagan AM, Behjati S, Page DC, Campbell PJ, Cleveland DW (2019) Chromosome segregation errors generate a diverse spectrum of simple and complex genomic rearrangements. Nat Genet 51:705–715
Maciejowski J, Li Y, Bosco N, Campbell PJ, de Lange T (2015) Chromothripsis and Kataegis induced by telomere crisis. Cell 163:1641–1654
Mateos-Gomez PA et al (2015) Mammalian polymerase θ promotes alternative NHEJ and suppresses recombination. Nature 518:254
Meltzer PS, Guan XY, Burgess A, Trent JM (1992) Rapid generation of region specific probes by chromosome microdissection and their application. Nat Genet 1:24–28
Mendez-Dorantes C, Bhargava R, Stark JM (2018) Repeat-mediated deletions can be induced by a chromosomal break far from a repeat, but multiple pathways suppress such rearrangements. Genes Dev
Nathanson DA, Gini B, Mottahedeh J, Visnyei K, Koga T, Gomez G, Eskin A, Hwang K, Wang J, Masui K, Paucar A, Yang H, Ohashi M, Zhu S, Wykosky J, Reed R, Nelson SF, Cloughesy TF, James CD, Rao PN, Kornblum HI, Heath JR, Cavenee WK, Furnari FB, Mischel PS (2014) Targeted therapy resistance mediated by dynamic regulation of extrachromosomal mutant EGFR DNA. Science 343:72–76
Nguyen K, Puppo F, Roche S, Gaillard MC, Chaix C, Lagarde A, Pierret M, Vovan C, Olschwang S, Salort-Campana E, Attarian S, Bartoli M, Bernard R, Magdinier F, Levy N (2017) Molecular combing reveals complex 4q35 rearrangements in Facioscapulohumeral dystrophy. Hum Mutat 38:1432–1441
Ni Y et al (2017) Super-resolution imaging of a 2.5 kb non-repetitive DNA in situ in the nuclear genome using molecular beacon probes. eLife 6:e21660
Nora EP et al (2012) Spatial partitioning of the regulatory landscape of the X-inactivation Centre. Nature 485:381–385
Olsen KE, Knudsen H, Rasmussen BB, Balslev E, Knoop A, Ejlertsen B, Nielsen KV, Schönau A, Overgaard J, Danish Breast Cancer Co-operative Group (2004) Amplification of HER2 and TOP2A and deletion of TOP2A genes in breast cancer investigated by new FISH probes. Acta Oncol 43:35–42
Osoegawa K, Mammoser AG, Wu C, Frengen E, Zeng C, Catanese JJ, de Jong PJ (2001) A bacterial artificial chromosome library for sequencing the complete human genome. Genome Res 11:483–496
Pardue ML, Gall JG (1969) Molecular hybridization of radioactive DNA to the DNA of cytological preparations. Proc Natl Acad Sci U S A 64:600–604
Pierce AJ, Johnson RD, Thompson LH, Jasin M (1999) XRCC3 promotes homology-directed repair of DNA damage in mammalian cells. Genes Dev 13:2633–2638
Potapova TA, Unruh JR, Yu Z, Rancati G, Li H, Stampfer MR, Gerton JL (2019) Superresolution microscopy reveals linkages between ribosomal DNA on heterologous chromosomes. J Cell Biol 218:2492–2513
Raap AK, Florijn RJ, Blonden LAJ, Wiegant J, Vaandrager JW, Vrolijk H, den Dunnen J, Tanke HJ, van Ommen G (1996) Fiber FISH as a DNA mapping tool. Methods 9:67–73
Ried T, Schröck E, Ning Y, Wienberg J (1998) Chromosome painting: a useful art. Hum Mol Genet 7:1619–1626
Roix JJ, McQueen PG, Munson PJ, Parada LA, Misteli T (2003) Spatial proximity of translocation-prone gene loci in human lymphomas. Nat Genet 34:287–291
Roschke A, Ning Y, Smith ACM, Macha M, Precht K, Riethman H, Ledbetter DH, Flint J, Horsley S, Regan R, Kearney L, Knight S, Kvaloy K, Brown WRA (1996) A complete set of human telomeric probes and their clinical application. National Institutes of Health and Institute of Molecular Medicine collaboration. Nat Genet 14:86–89
Roukos V, Voss TC, Schmidt CK, Lee S, Wangsa D, Misteli T (2013) Spatial dynamics of chromosome translocations in living cells. Science 341:660–664
Roukos V, Burgess RC, Misteli T (2014) Generation of cell-based systems to visualize chromosome damage and translocations in living cells. Nat Protoc 9:2476–2492
Rowley JD (1973) A new consistent chromosomal abnormality in chronic myelogenous leukaemia identified by quinacrine fluorescence and Giemsa staining. Nature 243:290–293
Rudin CM, Durinck S, Stawiski EW, Poirier JT, Modrusan Z, Shames DS, Bergbower EA, Guan Y, Shin J, Guillory J, Rivers CS, Foo CK, Bhatt D, Stinson J, Gnad F, Haverty PM, Gentleman R, Chaudhuri S, Janakiraman V, Jaiswal BS, Parikh C, Yuan W, Zhang Z, Koeppen H, Wu TD, Stern HM, Yauch RL, Huffman KE, Paskulin DD, Illei PB, Varella-Garcia M, Gazdar AF, de Sauvage FJ, Bourgon R, Minna JD, Brock MV, Seshagiri S (2012) Comprehensive genomic analysis identifies SOX2 as a frequently amplified gene in small-cell lung cancer. Nat Genet 44:1111–1116
Rudkin GT, Stollar BD (1977) High resolution detection of DNA-RNA hybrids in situ by indirect immunofluorescence. Nature 265:472–473
Rust MJ, Bates M, Zhuang X (2006) Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat Methods 3:793–795
Schnedl W (1971) Banding pattern of human chromosomes. Nat New Biol 233:93–94
Schrock E et al (1996) Multicolor spectral karyotyping of human chromosomes. Science 273:494–497
Seabright M (1971) A rapid banding technique for human chromosomes. Lancet 2:971–972
Shizuya H, Birren B, Kim UJ, Mancino V, Slepak T, Tachiiri Y, Simon M (1992) Cloning and stable maintenance of 300-kilobase-pair fragments of human DNA in Escherichia coli using an F-factor-based vector. Proc Natl Acad Sci U S A 89:8794–8797
Solinas-Toldo S, Lampel S, Stilgenbauer S, Nickolenko J, Benner A, Döhner H, Cremer T, Lichter P (1997) Matrix-based comparative genomic hybridization: biochips to screen for genomic imbalances. Genes Chromosomes Cancer 20:399–407
Speicher MR, Carter NP (2005) The new cytogenetics: blurring the boundaries with molecular biology. Nat Rev Genet 6:782–792
Speicher MR, Gwyn Ballard S, Ward DC (1996) Karyotyping human chromosomes by combinatorial multi-fluor FISH. Nat Genet 12:368–375
Steinhaeuser U, Starke H, Nietzel A, Lindenau J, Ullmann P, Claussen U, Liehr T (2002) Suspension (S)-FISH, a new technique for interphase nuclei. J Histochem Cytochem 50:1697–1698
Stephens PJ, Greenman CD, Fu B, Yang F, Bignell GR, Mudie LJ, Pleasance ED, Lau KW, Beare D, Stebbings LA, McLaren S, Lin ML, McBride D, Varela I, Nik-Zainal S, Leroy C, Jia M, Menzies A, Butler AP, Teague JW, Quail MA, Burton J, Swerdlow H, Carter NP, Morsberger LA, Iacobuzio-Donahue C, Follows GA, Green AR, Flanagan AM, Stratton MR, Futreal PA, Campbell PJ (2011) Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144:27–40
Sugita S, Hasegawa T (2017) Practical use and utility of fluorescence in situ hybridization in the pathological diagnosis of soft tissue and bone tumors. J Orthop Sci 22:601–612
Sumner AT, Evans HJ, Buckland RA (1971) New technique for distinguishing between human chromosomes. Nat New Biol 232:31–32
Telenius H, Carter NP, Bebb CE, Nordenskjöld M, Ponder BA, Tunnacliffe A (1992) Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics 13:718–725
Tjio JH, Levan A (1956) The Chromosome Number of Man. Hereditas 42:1–6
Tkachuk DC, Westbrook CA, Andreeff M, Donlon TA, Cleary ML, Suryanarayan K, Homge M, Redner A, Gray J, Pinkel D (1990) Detection of bcr-abl fusion in chronic myelogeneous leukemia by in situ hybridization. Science 250:559–562
Tosuji H, Mabuchi I, Fusetani N, Nakazawa T (1992) Calyculin a induces contractile ring-like apparatus formation and condensation of chromosomes in unfertilized sea urchin eggs. Proc Natl Acad Sci U S A 89:10613–10617
Trask BJ (2002) Human cytogenetics: 46 chromosomes, 46 years and counting. Nat Rev Genet 3:769–778
Turner KM, Deshpande V, Beyter D, Koga T, Rusert J, Lee C, Li B, Arden K, Ren B, Nathanson DA, Kornblum HI, Taylor MD, Kaushal S, Cavenee WK, Wechsler-Reya R, Furnari FB, Vandenberg SR, Rao PN, Wahl GM, Bafna V, Mischel PS (2017) Extrachromosomal oncogene amplification drives tumour evolution and genetic heterogeneity. Nature 543:122–125
van der Burg M, Poulsen TS, Hunger SP, Beverloo HB, Smit EM, Vang-Nielsen K, Langerak AW, van Dongen J (2004) Split-signal FISH for detection of chromosome aberrations in acute lymphoblastic leukemia. Leukemia 18:895–908
Veldman T, Vignon C, Schrock E, Rowley JD, Ried T (1997) Hidden chromosome abnormalities in haematological malignancies detected by multicolour spectral karyotyping. Nat Genet 15:406–410
Verhaak RGW, Bafna V, Mischel PS (2019) Extrachromosomal oncogene amplification in tumour pathogenesis and evolution. Nat Rev Cancer 19:283–288
Vorsanova SG, Yurov YB, Iourov IY (2010) Human interphase chromosomes: a review of available molecular cytogenetic technologies. Mol Cytogenet 3:1
Wang DG, Fan JB, Siao CJ, Berno A, Young P, Sapolsky R, Ghandour G, Perkins N, Winchester E, Spencer J, Kruglyak L, Stein L, Hsie L, Topaloglou T, Hubbell E, Robinson E, Mittmann M, Morris MS, Shen N, Kilburn D, Rioux J, Nusbaum C, Rozen S, Hudson TJ, Lipshutz R, Chee M, Lander ES (1998) Large-scale identification, mapping, and genotyping of single-nucleotide polymorphisms in the human genome. Science 280:1077–1082
Wang H et al. (2019) CRISPR-mediated live imaging of genome editing and transcription. Science
Weier H-UG (2001) DNA Fiber mapping techniques for the assembly of high-resolution physical maps. J Histochem Cytochem 49:939–948
Weinstock DM, Elliott B, Jasin M (2006) A model of oncogenic rearrangements: differences between chromosomal translocation mechanisms and simple double-strand break repair. Blood 107:777–780
Yunis JJ, Roldan L, Yasmineh WG, Lee JC (1971) Staining of satellite DNA in metaphase chromosomes. Nature 231:532–533
Zhang Y, McCord R, Ho YJ, Lajoie BR, Hildebrand DG, Simon AC, Becker MS, Alt FW, Dekker J (2012) Spatial organization of the mouse genome and its role in recurrent chromosomal translocations. Cell 148:908–921
Zhang CZ, Spektor A, Cornils H, Francis JM, Jackson EK, Liu S, Meyerson M, Pellman D (2015) Chromothripsis from DNA damage in micronuclei. Nature 522:179–184
Acknowledgments
We thank Kathleen Wilson and Sangeeta Patel (UT Southwestern Medical Center) for providing figures of G-banded karyotypes. This work was supported by the US National Institutes of Health (R00CA218871 to P.L.) and the Cancer Prevention and Research Institute of Texas (RR180050 to P.L.).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Beth Sullivan
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hu, Q., Maurais, E.G. & Ly, P. Cellular and genomic approaches for exploring structural chromosomal rearrangements. Chromosome Res 28, 19–30 (2020). https://doi.org/10.1007/s10577-020-09626-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-020-09626-1