Abstract
Dysregulated lncRNAs are proposed to be tightly associated with the progression of various tumors including glioblastoma (GBM). LncRNA Survival Associated Mitochondrial Melanoma-Specific Oncogenic Non-Coding RNA (SAMMSON) has been reported to be an oncogenic lncRNA in several tumors. Nevertheless, the specific role and molecular mechanism of SAMMSON in GBM progression remain unknown. Expression of SAMMSON in GBM tissues and cells was detected by qRT-PCR. CCK-8 and LDH release assays were applied to evaluate cellular viability. Invasion effect was assessed by Transwell invasion assay and western blot analysis of E-cadherin and N-cadherin expression. Apoptosis was detected using flow cytometry analysis and caspase-3 activity assay. The protein levels of phosphatidylinositol-3-kinase (PI3K), phosphorylated (p)-PI3K, protein kinase B (Akt) and p-Akt were estimated by western blot. We found that SAMMSON was highly expressed in GBM tissues and cells. SAMMSON knockdown suppressed cell viability and increased LDH release in GBM cells. Moreover, SAMMSON silencing impeded the invasive ability of GBM cells by regulating epithelial-to-mesenchymal transition (EMT). Furthermore, SAMMSON downregulation increased the apoptotic rate and caspase-3 activity in GBM cells. Additionally, it was demonstrated that the PI3K/Akt pathway was inhibited following SAMMSON silencing in GBM cells. Rescue assays revealed that activation of the PI3K/Akt pathway by 740Y-P abolished SAMMSON knockdown-induced viability reduction, invasion suppression and apoptosis in GBM cells. Taken together, lncRNA SAMMSON knockdown inhibited the malignancy of GBM cells by inactivation of the PI3K/Akt pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glioblastoma (GBM) is the most frequently occurring primary malignant intracranial cancer worldwide, with approximately 13,000 cases diagnosed annually in the USA (Reardon and Mitchell 2017). It is well recognized that most of GBM patients are always diagnosed at a late or metastatic stage, which makes this disease a challenge to be cured (Li et al. 2019). Although the diagnosis and multimodal treatments have been considerably improved, the prognosis of GBM patients still remains unfavorable, with an average survival time of only 12–15 months and a 5-year survival rate of about 3.4% (Davis 2016; Jayachandran et al. 2018). The poor prognosis of GBM patients is mainly attributed to the rapid growth, high invasiveness, resistance to apoptosis and high risk of recurrence (Bai et al. 2011). Up to date, no satisfactory management approaches for GBM exist. Hence, it is of great importance to explore the molecular mechanism underlying the progression of GBM and to search for novel treatment strategies for GBM.
During the last decades, researches on non-coding RNAs, especially long non-coding RNAs (lncRNAs), have evoked considerable interest due to their functional roles in GBM pathogenesis and development (Chen et al. 2017). LncRNAs, a group of RNA transcripts longer than 200 nucleotides, were originally believed as inconsequential transcriptional biological noise that have limited or no protein-coding function (van Bakel and Hughes 2009). Plenty of lncRNAs are proposed to play significantly regulatory roles in diverse aspects of tumor-associated cell events, such as cell growth, differentiation, apoptosis, invasion, and migration (Flynn and Chang 2014; Gonzalez et al. 2015). Recently, extensive studies have demonstrated that lncRNAs are frequently dysregulated in various types of tumors, including GBM, and tightly associated with tumor progression by serving as either oncogenes or cancer suppressors (Huarte 2015). Among these lncRNAs, Survival Associated Mitochondrial Melanoma-Specific Oncogenic Non-Coding RNA (SAMMSON), located on chromosome 3p13–3p14, is a ubiquitously expressed lncRNA that is characterized as an oncogene in cutaneous melanoma (Vendramin et al. 2018). A recent work reported that SAMMSON was upregulated in GBM patients and promoted GBM cell proliferation by downregulating miR-622 (Xie et al. 2019b). However, the detailed functions and mechanism of SAMMSON in GBM still remain to be fully understood.
In our study, we detected the expression pattern of SAMMSON in GBM tissues and cells. Furthermore, we carried out loss-of-function experiments to characterize the biological roles of SAMMSON in GBM cells and its underlying mechanism.
Materials and Methods
Clinical Tissue Collection
Nineteen pairs of GBM tissues and adjacent normal tissues were got from GBM patients undergoing lesion resection at the Second People's Hospital of Huai'an. All the enrolled patients did not receive radiotherapy, chemotherapy, or other anti-tumor therapies prior to surgery. After surgical resection, these collected tissue specimens were rapidly frozen at − 80 °C until RNA isolation. The study protocol was performed with the approval of the Ethics Committee of the Second People's Hospital of Huai’an and signed informed consent was obtained from each individual.
Cell Culture and Treatments
GBM cell lines (U87MG, T98G, U251, LN229, and A172) and normal human astrocyte (NHA) cells were obtained from the American Type Culture Collection (Manassas, VA, USA). These cells were fostered in RPMI-1640 medium (Thermo Fisher Scientific, Inc., Waltham, MA, USA) supplemented with 10% heat-inactivated fetal bovine serum (Wisent, St-Bruno, QC, Canada) and antibiotics (100 U/ml penicillin and 100 μg/ml streptomycin) (Solarbio, Beijing, China) under a water-saturated chamber containing 5% CO2 and 95% air at 37 °C. In the following experiments, siRNA targeting SAMMSON (si-SAMMSON) and siRNA control (si-Ctrl) (RiboBio Co., Ltd., Guangzhou, China) were transfected into U87MG and T98G cells using Lipofectamine 2000 Transfection Reagent (Invitrogen, Carlsbad, CA, USA), followed by treatment for 48 h with 15 μM 740Y-P (Sigma, St. Louis, MO, USA), a phosphatidylinositol-3-kinase (PI3K)/protein kinase B (Akt) activator.
Quantitative Real-Time PCR (qRT-PCR)
Total RNA was isolated from GBM tissues, adjacent normal tissues and cultured cells with TRIzol reagent (Invitrogen). For the detection of SAMMSON expression, isolated RNA was reverse-transcribed into complementary DNA (cDNA) by means of FastKing RT Kit (TIANGEN, Beijing, China). The resultant cDNA was then subjected to qPCR using SYBR Premix DimerEraser (Takara) on a Roche LightCycler480 II instrument (Roche, Basel, Switzerland). The amplification condition was listed as below: 95 °C for 10 min, followed by 35 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 1 min. GAPDH was used as the internal control for SAMMSON. The \(2^{{ - \Delta \Delta C_{{\text{T}}} }}\) method was used to compute the fold changes in SAMMSON expression. The primers used in this research included: SAMMSON, forward 5′-TTCC TCAA CTAT GCAA CTCA A-3′, reverse 5′-TAGA CTAC GGGC TCAT GACT T-3′; GAPDH, forward 5′-GTCT CCTC TGAC TTCA ACAG CG-3′, reverse 5′-ACCA CCCT GTTG CTGT AGCC AA-3′. The experiment was repeated three times.
Cell Viability Assay
Cell counting kit-8 (CCK-8) detection kit (Dojindo, Tokyo, Japan) was taken for the evaluation of cell viability. Following the indicated treatments, U87MG and T98G cells were seeded into 96-well plates at 2 × 103 cells per well and incubated at 37 °C for 48 h. Afterwards, the cells were incubated with 10 µl of CCK-8 reagent for another 2 h, followed by the measurement of optical density at 450 nm by a spectrophotometric plate reader (Bio-Rad Laboratories, Richmond, CA, USA). The experiment was repeated three times in triplicate.
Detection of Lactate Dehydrogenase (LDH) Release
LDH release assay was used to assess the extent of cell death. LDH released from treated U87MG and T98G cells was measured using a Cytotoxicity LDH Assay Kit (JianCheng Bioengineering Institute, Nanjing, China). The experiment was repeated three times in triplicate.
Transwell Invasion Assay
Transwell invasion assay was conducted to determine the invasive ability of treated GBM cells. Briefly, a total of 3 × 104 U87MG and T98G cells suspending in 200 μl serum-free medium were plated into the upper Transwell chamber (8 μm pore size, Millipore, Darmstadt, Germany) covered with matrigel (BD Biosciences, San Jose, CA, USA) and RPMI-1640 medium containing 10% FBS was added into the lower chamber as a chemoattractant. After 48 h, the non-invading cells on the upper chamber were wiped with cotton swabs. The invaded cells on the lower chamber were fixed with 4% polyformaldehyde and stained with 0.1% crystal violet. The invasive ability was evaluated by counting the number of invaded cells in 5 randomly selected fields using an Olympus CKX31 microscope (magnification, × 200; Exsson, Daxing, Beijing, China). The experiment was repeated three times.
Apoptosis Assay by Flow Cytometry
The treated U87MG and T98G cells were collected, rinsed twice with PBS, and resuspended in 1 × binding buffer at a concentration of 6 × 105 cells/ml. These cells were then double-stained with Annexin V-fluorescein isothiocyanate (FITC) and propidium iodine (PI) according to the Annexin V-FITC Apoptosis Detection Kit (Beyotime, Shanghai, China). Finally, a FACSAria III flow cytometer (BD Biosciences) was employed to analyze the stained cells. The experiment was repeated three times.
Caspase-3 Activity Assay
The treated U87MG and T98G cells were harvested by trypsinization, resuspended and lysed in ice-cold cell lysis buffer for 30 min before centrifugation for 5 min. Caspase-3 activity in the supernatant of treated U87MG and T98G cells was measured by a Caspase-3 Colorimetric Assay kit (Promega Corporation, Madison, WI, USA). The experiment was repeated three times in triplicate.
Western Blot Analysis
Treated U87MG and T98G cells were lysed in RIPA lysis buffer (Beyotime Institute of Biotechnology, Jiangsu, China) containing 1% phosphatase inhibitor cocktail for 30 min, followed by centrifugation for 10 min to remove cell debris. After detection of protein concentration, equivalent volume of denatured proteins (50 μg/lane) was electrophoresed on 10% SDS-PAGE and electro-transferred onto nitrocellulose membrane. After non-specific binding with 5% non-fat dry milk in TBS with 0.1% tween-20 (TBST) for 2 h at room temperature, the membranes were immuno-probed overnight at 4 °C with the primary antibodies against E-cadherin, N-cadherin, phosphorylated PI3K (p-PI3K), PI3K, phosphorylated Akt (p-Akt), Akt, and β-actin (all from Abcam, Cambridge, MA, USA). After rinsing three times with TBST, the membranes were then incubated with horseradish peroxidase-conjugated secondary antibody (Li-COR Biosciences, Lincoln, NE, USA) at room temperature for 1 h. The objective protein bands were detected with an ECL advanced Western blotting detection kit (Thermo Fisher Scientific, Inc.). The experiment was repeated three times.
Statistics
All results are displayed as the mean ± standard deviation (SD). SPSS 22.0 statistical software package (IBM Corp., Armonk, NY, USA) was taken to perform all statistical analyses. Differences among multiple groups were determined by one-way analysis of variance (ANOVA) with a Dunnett's post hoc test. Values of P < 0.05 were considered to indicate a statistically significant difference.
Results
SAMMSON was Highly Expressed in GBM Tissues and Cells
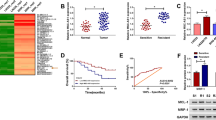
We initially detected the expression pattern of SAMMSON in GBM tissues by qRT-PCR. As presented in Fig. 1a, we observed an elevation of SAMMSON expression in 19 GBM tissues compared with that in adjacent normal tissues. In consistence, we demonstrated that SAMMSON expression was aberrantly upregulated in GBM cell lines including (U87MG, T98G, U251, LN229, and A172) relative to that in normal human astrocyte (NHA) cells (Fig. 1b). In the 5 examined GBM cells, we found that SAMMSON expression was comparatively higher in U87MG and T98G cells than other three GBM cells. Therefore, U87MG and T98G cells were chosen for the next experiments. qRT-PCR analysis was employed to examine the knockdown efficiency of si-SAMMSON in U87MG and T98G cells. As a result, transfection with si-SAMMSON led to a noticeable decrease (65.0% and 76.2%, respectively) of SAMMSON expression in U87MG and T98G cells compared with si-Ctrl group (Fig. 1c).
Expression profile of SAMMSON in GBM tissues and cells. a qRT-PCR analysis of SAMMSON expression in 19 pairs of GBM tissues and adjacent normal tissues. b qRT-PCR analysis of SAMMSON expression in GBM cell lines (U87MG, T98G, U251, LN229, and A172) and normal human astrocyte (NHA) cells. *P < 0.05 compared to NHA, n = 3. c U87MG and T98G cells were transfected with si-SAMMSON or si-Ctrl for 48 h, followed by the detection of SAMMSON expression by qRT-PCR. *P < 0.05 compared to si-Ctrl group, n = 3
SAMMSON Knockdown Repressed the Viability of GBM Cells
To characterize the biological function of SAMMSON in GBM progression, loss-of-function assays were conducted to assess the effects of SAMMSON on cell viability, invasion and apoptosis in GBM cells. CCK-8 assay showed that SAMMSON silencing effectively restricted cell viability in U87MG (Fig. 2a; 32.0% decrease) and T98G (Fig. 2b; 48.6% decrease) cells versus si-Ctrl group. Using LDH release assay, we discovered that LDH release from U87MG (Fig. 2c; 1.55-fold increase) and T98G (Fig. 2d; 2.01-fold increase) cells was increased after depletion of SAMMSON. Overall, these data suggested that SAMMSON knockdown impeded GBM cell viability.
SAMMSON knockdown impeded GBM cell viability. a, b CCK-8 assay was employed to evaluate cell viability in U87MG and T98G cells introduced with si-SAMMSON or si-Ctrl for 48 h. c, d LDH level in si-SAMMSON or si-Ctrl-transfected U87MG and T98G cells was detected by LDH release assay. *P < 0.05 compared to si-Ctrl, n = 3
SAMMSON Knockdown Suppressed the Invasive Ability of GBM Cells
As demonstrated by Transwell invasion assay, we found that SAMMSON silencing in U87MG (Fig. 3a; 65.5% decrease) and T98G (Fig. 3b; 57.9% decrease) cells caused a decrease in the number of invaded cells relative to that in control group. Moreover, we explored the effects of SAMMSON knockdown on the expression of epithelial-to-mesenchymal transition (EMT) markers. Western blot further proved that the expression of epithelial marker E-cadherin was enhanced and the expression of mesenchymal marker N-cadherin was decreased after si-SAMMSON transfection in U87MG (Fig. 3c; 2.21-fold increase and 53.1% decrease for E-cadherin and N-cadherin, respectively) and T98G (Fig. 3d; 1.81-fold increase and 55.3% decrease for E-cadherin and N-cadherin, respectively) cells. Together, these date indicated that SAMMSON knockdown inhibited the invasive ability of GBM cells by regulation of EMT.
SAMMSON knockdown suppressed the invasive ability of GBM cells. U87MG and T98G cells were transfected with si-SAMMSON or si-Ctrl and incubated for 48 h. The number of invaded U87MG (a) and T98G (b) cells was then determined by Transwell invasion assay. *P < 0.05 compared to si-Ctrl, n = 5. The protein levels of E-cadherin and N-cadherin in the treated U87MG (c) and T98G (d) cells were detected by western blot. *P < 0.05 compared to si-Ctrl, n = 3
SAMMSON Knockdown-Induced Apoptosis of GBM Cells
Effects of SAMMSON knockdown on the apoptosis of GBM cells were further evaluated. Using flow cytometry analysis, U87MG (Fig. 4a; 4.83-fold increase) and T98G (Fig. 4b; 5.15-fold increase) cells with si-SAMMSON transfection showed an increased apoptotic rate relative to that in control group. In line with the results of flow cytometry analysis, caspase-3 activity assay revealed that SAMMSON silencing increased caspase-3 activity in U87MG (Fig. 4c; 1.94-fold increase) and T98G (Fig. 4d; 2.17-flod increase) cells in comparison to that in control group. Collectively, these data implied that SAMMSON knockdown promoted apoptosis of GBM cells.
SAMMSON knockdown induced apoptosis of GBM cells. U87MG and T98G cells were received si-SAMMSON or si-Ctrl transfection and incubated for 48 h, followed by the detection of apoptosis (a, b) and caspase-3 activity (c, d) by flow cytometry analysis and caspase-3 activity assay, respectively. *P < 0.05 compared to si-Ctrl, n = 3
Knockdown of SAMMSON Inactivated the PI3K/Akt Pathway in GBM Cells
As is well known, the PI3K/Akt pathway is hyperactivated in different types of human cancers including GBM, and contributes to tumor progression (Li et al. 2016). Therefore, our study tested the phosphorylation status of PI3K and Akt proteins in U87MG and T98G cells after silencing of SAMMSON. Western blot analysis demonstrated that the protein levels of p-PI3K (45.4% and 57.5% decrease for U87MG and T98G cells, respectively) and p-Akt (62.1% and 54.3% decrease for U87MG and T98G cells, respectively) were reduced and no significant changes on the protein levels of PI3K and Akt were observed in si-SAMMSON-transfected U87MG (Fig. 5a) and T98G (Fig. 5b) cells. These findings suggested that SAMMSON knockdown inhibited the activation of the PI3K/Akt pathway in GBM cells.
Activation of the PI3K/Akt Pathway Abolished SAMMSON Knockdown-Induced Viability Reduction of GBM Cells
To further address the role of PI3K/Akt pathway in SAMMSON knockdown-mediated effects, U87MG and T98G cells were transfected with si-SAMMSON or si-Ctrl, followed by treatment with 15 μM 740Y-P, an agonist of PI3K/Akt signaling. CCK-8 assay manifested that treatment with 740Y-P restored SAMMSON knockdown-induced viability reduction in U87MG (Fig. 6a; 1.44-fold increase compared with si-SAMMSON group) and T98G (Fig. 6b; 1.39-fold increase compared with si-SAMMSON group) cells. LDH release assay revealed that SAMMSON silencing-induced increase of LDH release in U87MG (Fig. 6c; 26.6% decrease compared with si-SAMMSON group) and T98G (Fig. 6d; 27.8% decrease compared with si-SAMMSON group) cells was partially attenuated in response to 740Y-P. These results demonstrated that activation of the PI3K/Akt pathway partially rescued SAMMSON knockdown-induced viability reduction of GBM cells.
Activation of the PI3K/Akt pathway abolished SAMMSON knockdown-induced viability reduction of GBM cells. U87MG and T98G cells were transfected with si-SAMMSON or si-Ctrl and then treated with 15 μM 740Y-P for 48 h, followed by evaluation of cell viability (a, b) and LDH release (c, d) by CCK-8 and LDH release assay, respectively. *P < 0.05 compared to si-Ctrl, n = 3. #P < 0.05 compared to si-SAMMSON, n = 3
Activation of the PI3K/Akt Pathway Resisted SAMMSON Knockdown-Induced Invasion Suppression in GBM Cells
Transwell invasion assay further displayed that administration with 740Y-P abolished the inhibitory effect of SAMMSON knockdown on cell invasive ability in U87MG (Fig. 7a; 1.76-fold increase compared with si-SAMMSON group) and T98G (Fig. 7b; 1.82-fold increase compared with si-SAMMSON group) cells. Using western blot analysis, we also discovered that the increase of E-cadherin protein expression and decrease of N-cadherin protein expression in si-SAMMSON-transfected U87MG (Fig. 7c; 21.2% decrease and 1.46-fold increase for E-cadherin and N-cadherin, respectively, compared with si-SAMMSON group) and T98G (Fig. 7d; 23.9% decrease and 1.39-fold increase for E-cadherin and N-cadherin, respectively, compared with si-SAMMSON group) cells were reversed following the addition of 740Y-P. These data suggested that activation of the PI3K/Akt pathway relieved SAMMSON knockdown-induced invasion suppression in GBM cells.
Activation of the PI3K/Akt pathway abrogated SAMMSON knockdown-induced invasion suppression in GBM cells. U87MG and T98G cells were transfected with si-SAMMSON or si-Ctrl, followed by exposure to 15 μM 740Y-P for 48 h. a, b Cell invasion in the treated U87MG and T98G cells was assessed by Transwell invasion assay. *P < 0.05 compared to si-Ctrl, n = 5. #P < 0.05 compared to si-SAMMSON, n = 5. c, d The protein expression of E-cadherin and N-cadherin in the treated U87MG and T98G cells was examined by western blot analysis. *P < 0.05 compared to si-Ctrl, n = 3. #P < 0.05 compared to si-SAMMSON, n = 3
Activation of the PI3K/Akt Pathway Antagonized SAMMSON Knockdown-Induced Apoptosis in GBM Cells
Flow cytometry analysis hinted that SAMMSON knockdown-induced increase in the apoptotic rate of U87MG (Fig. 8a; 52.6% decrease compared with si-SAMMSON group) and T98G (Fig. 8b; 52.3% decrease compared with si-SAMMSON group) cells was counteracted after the PI3K/Akt pathway was activated by 740Y-P. Meanwhile, we proved that activation of the PI3K/Akt pathway by 740Y-P nullified the increase of caspase-3 activity induced by SAMMSON silencing in U87MG (Fig. 8c; 23.9% decrease compared with si-SAMMSON group) and T98G (Fig. 8d; 28.0% decrease compared with si-SAMMSON group) cells. These findings revealed that activation of the PI3K/Akt pathway antagonized SAMMSON knockdown-induced apoptosis in GBM cells.
Activation of the PI3K/Akt pathway undermined SAMMSON silencing-induced apoptosis in GBM cells. U87MG and T98G cells were transfected with si-SAMMSON or si-Ctrl before exposure to 15 μM 740Y-P for 48 h, followed by the evaluation of apoptosis (a, b) and caspase-3 activity (c, d) by flow cytometry analysis and caspase-3 activity assay, respectively. *P < 0.05 compared to si-Ctrl, n = 3. #P < 0.05 compared to si-SAMMSON, n = 3
Discussion
In the past few years, lncRNAs are a hot topic in the field of oncology and have attracted an increasing research attention. Emerging evidence has demonstrated that a cluster of lncRNAs are deregulated in a wide range of human malignancies including GBM and play essential roles in the pathogenesis and progression of GBM (Xi et al. 2018). For example, it was reported that TRG-AS1 was aberrantly upregulated in GBM tissues and cells, and promoted GBM cell proliferation by acting as a ceRNA of miR-877-5p to regulate SUZ12 expression (Xie et al. 2019a). LncRNA AGAP2-AS1 expression was demonstrated to be upregulated in GBM tissues and cells and predict a poor prognosis in GBM patients, and exert oncogenic functions in GBM by epigenetically silencing TFPI2 expression through binding to EZH2 and LSD1 (Luo et al. 2019). LncRNA AC016405.3 was found to act as an anti-oncogene in GBM cells through modulation of TET2 by sponging miR-19a-5p (Ren and Xu 2019). Therefore, these reports suggest that lncRNAs may be used as biomarker for cancer diagnosis and treatment, as well as potential therapeutic candidates for GBM in the future. SAMMSON is a lineage-specific lncRNA that is overexpressed in several types of tumors including GBM (Xie et al. 2019b). However, up to now, the detailed biological function of SAMMSON in GBM has not been systematically investigated. Herein, we focused on exploring the specific function and underlying mechanism of SAMMSON in GBM cells.
Published studies have demonstrated that SAMMSON exerts oncogenic roles in several types of tumors. For example, it was proved that SAMMSON may be an informative biomarker of melanoma malignancy and its silencing reduced melanoma cell growth and survival (SAMMSON Long Noncoding RNA Is Essential for Melanoma Cell Viability 2016). Additionally, SAMMSON was discovered to be upregulated in papillary thyroid carcinoma (PTC) cells, and serve as a novel diagnostic and prognostic biomarker for PTC patients. Moreover, knockdown of SAMMSON inhibited PTC cell proliferation and invasion in vitro as well as tumorigenicity and metastasis in vivo through p300/Specifcity protein 1 (Sp1) axis (Shao et al. 2020). In hepatocellular carcinoma (HCC), SAMMSON was demonstrated to be upregulated in HCC tissues and promote cancer cell migration and invasion (Yang et al. 2019). A study revealed that SAMMSON overexpression induced enhanced self-renewal in liver cancer through EZH2-dependent Wnt/β-catenin activation (Li et al. 2017). In the present study, we confirmed that SAMMSON was highly expressed in GBM tissues and cells. A previous study showed that SAMMSON was upregulated in plasma of glioblastoma patients compared with healthy controls and SAMMSON overexpression promoted the proliferative rate of GBM cells by inhibition of miR-622 expression (Xie et al. 2019b). However, Xie and colleagues did not explored the downstream signaling pathway by which SAMMSON/miR-622 regulated GBM cell proliferation. Our functional investigation further revealed that knockdown of SAMMSON suppressed cell viability and invasion, and induced apoptosis in GBM cells. The present study identified SAMMSON as an oncogenic lncRNA in GBM, which was in accordance with the oncogenic roles of SAMMSON in other cancers.
The PI3K/Akt signaling is a well-known intracellular signaling pathway that modulates various cellular processes, mainly involved in cell proliferation, survival, apoptosis, and invasiveness of tumor cells (Arcaro and Guerreiro 2007; Manning and Cantley 2007). Increasing experimental data have shown that hyper-activation of the PI3K/Akt pathway is frequently detected in multiple cancers including GBM (Wen et al. 2012; Cheng et al. 2009), and therefore contributes to the proliferation and survival of cancer cells during cancer progression (Zhang et al. 2007). The expression levels of p-PI3K and p-Akt in GBM tissues was increased compared with those in normal or lower-grade tumor tissues, while total PI3K and Akt expression levels were not significantly different between GBM and normal or lower-grade tumor tissues (Chakravarti et al. 2004; Wang et al. 2004). The PI3K/Akt pathway has been considered as a critical regulatory pathway in the treatment of malignant GBM (Zhao et al. 2017). Activation of the PI3K/Akt/mTOR signaling pathway was a driving force of EMP1-promoted GBM cell proliferation, migration and invasion potential (Miao et al. 2019). Activation of PI3K/Akt signaling prevented the suppressive effects of RWDD3 downregulation on GBM cell proliferation and migration (Chen et al. 2018). Hence, blocking the PI3K/Akt pathway has been investigated as a potential effective anticancer therapy for GBM (Shingu et al. 2003). To clarify the molecular mechanism underlying the effect of SAMMSON silencing on GBM cells, we analyzed the correlation between SAMMSON and the PI3K/Akt pathway. In this research, we found that the protein expression of p-PI3K and p-Akt was inhibited following knockdown of SAMMSON in GBM cells, suggesting that SAMMSON silencing inactivated the PIK3/Akt pathway in GBM cells. However, in the presence of a PI3K/Akt activator, 740Y-P, SAMMSON silencing-induced cell viability reduction, invasion suppression and apoptosis in GBM cells was partially relieved. Therefore, the PI3K/Akt signaling pathway provides a molecular foundation for the suppression of malignant phenotypes of GBM cells by SAMMSON knockdown. Up to now, there is no evidence that SAMMSON directly regulates the PI3K/Akt pathway. Since we found that the protein levels of p-PI3K and p-Akt were decreased and no significant changes on the protein levels of PI3K and Akt were observed in si-SAMMSON-transfected GBM cells, we deduced that SAMMSON knockdown indirectly regulated the PI3K/Akt pathway. A recent study suggested that SAMMSON overexpression acted as a scaffold to recruit p300 on Sp1 promoter region, resulting in transcriptional activation of Sp1 (Shao et al. 2020). Sp1 can inhibit PTEN expression by binding to the promoter region of PTEN, an upstream negative effector of PI3K/Akt pathway. (Kou et al. 2013; Jia et al. 2014; Xia et al. 2017). Therefore, we inferred that SAMMSON modulated PI3K/Akt pathway may at least by regulating Sp1/PTEN axis.
Conclusion
In summary, we confirmed that SAMMSON was highly expressed in GBM tissues and cells. What’s more, we found that SAMMSON silencing suppressed cell viability and invasion ability, and induced apoptosis in GBM cells partially through inhibition of the PI3K/Akt signaling pathway, contributing to the better understanding of the exact mechanism underlying the roles of SAMMSON in GBM progression. Therefore, our research offers a potential therapeutic target for the treatment of GBM.
References
Arcaro A, Guerreiro AS (2007) The phosphoinositide 3-kinase pathway in human cancer: genetic alterations and therapeutic implications. Curr Genom 8(5):271–306. https://doi.org/10.2174/138920207782446160
Bai RY, Staedtke V, Riggins GJ (2011) Molecular targeting of glioblastoma: drug discovery and therapies. Trends Mol Med 17(6):301–312. https://doi.org/10.1016/j.molmed.2011.01.011
Chakravarti A, Zhai G, Suzuki Y, Sarkesh S, Black PM, Muzikansky A, Loeffler JS (2004) The prognostic significance of phosphatidylinositol 3-kinase pathway activation in human gliomas. J Clin Oncol 22(10):1926–1933. https://doi.org/10.1200/JCO.2004.07.193
Chen G, Cao Y, Zhang L, Ma H, Shen C, Zhao J (2017) Analysis of long non-coding RNA expression profiles identifies novel lncRNA biomarkers in the tumorigenesis and malignant progression of gliomas. Oncotarget 8(40):67744–67753. https://doi.org/10.18632/oncotarget.18832
Chen X, Kuang W, Huang H, Li B, Zhu Y, Zhou B, Yan L (2018) Knockdown of RWD domain containing 3 inhibits the malignant phenotypes of glioblastoma cells via inhibition of phosphoinositide 3-kinase/protein kinase B signaling. Exp Ther Med 16(1):384–393. https://doi.org/10.3892/etm.2018.6135
Cheng CK, Fan QW, Weiss WA (2009) PI3K signaling in glioma–animal models and therapeutic challenges. Brain Pathol 19(1):112–120. https://doi.org/10.1111/j.1750-3639.2008.00233.x
Davis ME (2016) Glioblastoma: overview of disease and treatment. Clin J Oncol Nurs 20(5 Suppl):S2–S8. https://doi.org/10.1188/16.cjon.s1.2-8
Flynn RA, Chang HY (2014) Long noncoding RNAs in cell-fate programming and reprogramming. Cell Stem Cell 14(6):752–761. https://doi.org/10.1016/j.stem.2014.05.014
Gonzalez I, Munita R, Agirre E, Dittmer TA, Gysling K, Misteli T, Luco RF (2015) A lncRNA regulates alternative splicing via establishment of a splicing-specific chromatin signature. Nat Struct Mol Biol 22(5):370–376. https://doi.org/10.1038/nsmb.3005
Huarte M (2015) The emerging role of lncRNAs in cancer. Nat Med 21(11):1253–1261. https://doi.org/10.1038/nm.3981
Jayachandran A, Jonathan GE, Patel B, Prabhu K (2018) Primary spinal cord glioblastoma metastasizing to the cerebellum: a missed entity. Neurol India 66(3):854–857. https://doi.org/10.4103/0028-3886.232347
Jia LF, Huang YP, Zheng YF, Lyu MY, Wei SB, Meng Z, Gan YH (2014) miR-29b suppresses proliferation, migration, and invasion of tongue squamous cell carcinoma through PTEN-AKT signaling pathway by targeting Sp1. Oral Oncol 50(11):1062–1071. https://doi.org/10.1016/j.oraloncology.2014.07.010
Kou XX, Hao T, Meng Z, Zhou YH, Gan YH (2013) Acetylated Sp1 inhibits PTEN expression through binding to PTEN core promoter and recruitment of HDAC1 and promotes cancer cell migration and invasion. Carcinogenesis 34(1):58–67. https://doi.org/10.1093/carcin/bgs336
Li X, Li M, Chen J, Dai H, Wang L, Xiong Y, Zhong Y, Zhang L (2017) SAMMSON drives the self-renewal of liver tumor initiating cells through EZH2-dependent Wnt/beta-catenin activation. Oncotarget 8(61):103785–103796. https://doi.org/10.18632/oncotarget.21792
Li X, Wu C, Chen N, Gu H, Yen A, Cao L, Wang E, Wang L (2016) PI3K/Akt/mTOR signaling pathway and targeted therapy for glioblastoma. Oncotarget 7(22):33440–33450. https://doi.org/10.18632/oncotarget.7961
Li Y, Chen F, Chu J, Wu C, Li Y, Li H, Ma H (2019) miR-148-3p inhibits growth of glioblastoma targeting DNA methyltransferase-1 (DNMT1). Oncol Res 27(8):911–921. https://doi.org/10.3727/096504019x15516966905337
Luo W, Li X, Song Z, Zhu X, Zhao S (2019) Long non-coding RNA AGAP2-AS1 exerts oncogenic properties in glioblastoma by epigenetically silencing TFPI2 through EZH2 and LSD1. Aging (Albany NY) 11(11):3811–3823. https://doi.org/10.18632/aging.102018
Manning BD, Cantley LC (2007) AKT/PKB signaling: navigating downstream. Cell 129(7):1261–1274. https://doi.org/10.1016/j.cell.2007.06.009
Miao L, Jiang Z, Wang J, Yang N, Qi Q, Zhou W, Feng Z, Li W, Zhang Q, Huang B, Chen A, Zhang D, Zhao P, Li X (2019) Epithelial membrane protein 1 promotes glioblastoma progression through the PI3K/AKT/mTOR signaling pathway. Oncol Rep 42(2):605–614. https://doi.org/10.3892/or.2019.7204
Reardon DA, Mitchell DA (2017) The development of dendritic cell vaccine-based immunotherapies for glioblastoma. Semin Immunopathol 39(2):225–239. https://doi.org/10.1007/s00281-016-0616-7
Ren S, Xu Y (2019) AC016405.3, a novel long noncoding RNA, acts as a tumor suppressor through modulation of TET2 by microRNA-19a-5p sponging in glioblastoma. Cancer Sci 110(5):1621–1632. https://doi.org/10.1111/cas.14002
SAMMSON (2016) Long noncoding RNA is essential for melanoma cell viability. Cancer Discov 6(5):470. https://doi.org/10.1158/2159-8290.cd-rw2016-062
Shao L, Sun W, Wang Z, Dong W, Qin Y (2020) Long noncoding RNA SAMMSON promotes papillary thyroid carcinoma progression through p300/Sp1 axis and serves as a novel diagnostic and prognostic biomarker. IUBMB Life 72(2):237–246. https://doi.org/10.1002/iub.2158
Shingu T, Yamada K, Hara N, Moritake K, Osago H, Terashima M, Uemura T, Yamasaki T, Tsuchiya M (2003) Synergistic augmentation of antimicrotubule agent-induced cytotoxicity by a phosphoinositide 3-kinase inhibitor in human malignant glioma cells. Cancer Res 63(14):4044–4047
van Bakel H, Hughes TR (2009) Establishing legitimacy and function in the new transcriptome. Brief Funct Genom Proteom 8(6):424–436. https://doi.org/10.1093/bfgp/elp037
Vendramin R, Verheyden Y, Ishikawa H, Goedert L, Nicolas E, Saraf K, Armaos A, Delli Ponti R, Izumikawa K, Mestdagh P, Lafontaine DLJ, Tartaglia GG, Takahashi N, Marine JC, Leucci E (2018) SAMMSON fosters cancer cell fitness by concertedly enhancing mitochondrial and cytosolic translation. Nat Struct Mol Biol 25(11):1035–1046. https://doi.org/10.1038/s41594-018-0143-4
Wang H, Wang H, Zhang W, Huang HJ, Liao WS, Fuller GN (2004) Analysis of the activation status of Akt, NFκB, and Stat3 in human diffuse gliomas. Lab Investig 84(8):941–951. https://doi.org/10.1038/labinvest.3700123
Wen PY, Lee EQ, Reardon DA, Ligon KL, Alfred Yung WK (2012) Current clinical development of PI3K pathway inhibitors in glioblastoma. Neuro Oncol 14(7):819–829. https://doi.org/10.1093/neuonc/nos117
Xi J, Sun Q, Ma L, Kang J (2018) Long non-coding RNAs in glioma progression. Cancer Lett 419:203–209. https://doi.org/10.1016/j.canlet.2018.01.041
Xia SS, Zhang GJ, Liu ZL, Tian HP, He Y, Meng CY, Li LF, Wang ZW, Zhou T (2017) MicroRNA-22 suppresses the growth, migration and invasion of colorectal cancer cells through a Sp1 negative feedback loop. Oncotarget 8(22):36266–36278. https://doi.org/10.18632/oncotarget.16742
Xie H, Shi S, Chen Q, Chen Z (2019a) LncRNA TRG-AS1 promotes glioblastoma cell proliferation by competitively binding with miR-877-5p to regulate SUZ12 expression. Pathol Res Pract 215(8):152476. https://doi.org/10.1016/j.prp.2019.152476
Xie J, Wang X, Liu S, Chen C, Jiang F, Mao K, Zeng F (2019b) LncRNA SAMMSON overexpression distinguished glioblastoma patients from patients with diffuse neurosarcoidosis. NeuroReport 30(12):817–821. https://doi.org/10.1097/wnr.0000000000001278
Yang S, Cai H, Hu B, Tu J (2019) LncRNA SAMMSON negatively regulates miR-9–3p in hepatocellular carcinoma cells and has prognostic values. Biosci Rep. https://doi.org/10.1042/bsr20190615
Zhang X, Jin B, Huang C (2007) The PI3K/Akt pathway and its downstream transcriptional factors as targets for chemoprevention. Curr Cancer Drug Targets 7(4):305–316. https://doi.org/10.2174/156800907780809741
Zhao HF, Wang J, Shao W, Wu CP, Chen ZP, To ST, Li WP (2017) Recent advances in the use of PI3K inhibitors for glioblastoma multiforme: current preclinical and clinical development. Mol Cancer 16(1):100. https://doi.org/10.1186/s12943-017-0670-3
Funding
None.
Author information
Authors and Affiliations
Contributions
CL conceived and designed the study. HN and KW conducted the experiments. PX, JZ, and WL analyzed the data. HN and CL drafted the manuscript. All authors discussed the results.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical Approval
All procedures performed in studies involving human participants were in accordance with the Ethics Committee of the Second People's Hospital of Huai'an.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ni, H., Wang, K., Xie, P. et al. LncRNA SAMMSON Knockdown Inhibits the Malignancy of Glioblastoma Cells by Inactivation of the PI3K/Akt Pathway. Cell Mol Neurobiol 41, 79–90 (2021). https://doi.org/10.1007/s10571-020-00833-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-020-00833-2