Abstract
Thermodynamic analysis of the empirical data for the solvation of cellulose in aqueous NaOH/urea solution was performed in this study; this was achieved by employing the three-dimensional Reference Interaction Site Model theory coupled with the Kovalenko–Hirata closure approximation. The preferential distributions of Na+, OH−, urea, and water that were in a close proximity to the cellulose molecule, enabled the calculation of the solvation energy and the contribution of each solvent species to the solvation energy. By dividing the solvation energy into the solvent potential energy under the consideration of the solvent–solute interaction and the solvent reorganization energy, cellulose solvation in the NaOH/urea solution was observed to be primarily due to reorganization of the water molecules around the cellulose molecule. The solvated structure was suggested to be composed of cellulose as an inclusion in helical clusters of Na+, OH−, urea, and water, wherein the clusters comprised a repeated arrangement of OH− hydrate, water molecules, urea hydrate, and water molecules. Cellulose is suspected to play the role of a water structure maker in the presence of NaOH and urea, and as a water structure breaker in the absence of NaOH and/or urea.
Graphic abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
An alkali solution (e.g., solution of NaOH or LiOH) containing urea or thiourea was developed as a solvent for cellulose (Cai and Zhang 2005; Cai et al. 2004, 2008; Jiang et al. 2014; Jin et al. 2007; Liu et al. 2018; Wang et al. 2017a, b, c; Xiong et al. 2013). For example, 7% (w/w) NaOH/12% (w/w) urea solution successfully dissolved cellulose at − 12 °C, where 7% (w/w) NaOH and 12% (w/w) urea correspond to about 2 M NaOH and 2 M urea, respectively. It has been evaluated as less expensive and toxic than aqueous NaOH with carbon disulfide or N-methylmorpholine N-oxide monohydrate, which have been used in the viscose process and the Lyocell process, respectively (Cai et al. 2004; Jin et al. 2007). Because KOH cannot be used as a substitute for NaOH and LiOH, Na+ hydrate (and Li+ hydrate) are indispensable for cellulose solvation (Cai and Zhang 2005; Xiong et al. 2013; Wang et al. 2017b). The interaction of Na+ and Li+ with oxygen atoms of cellulose was suggested to weaken the intra- and intermolecular hydrogen bonds within cellulose, and to stabilize the electrostatic interaction between OH− and cellulose (Wang et al. 2017a, b). Also, deprotonation of cellulose by OH− was believed to contribute to the cellulose solvation in aqueous solutions of NaOH or LiOH (Bialik et al. 2016; Xiong et al. 2013). Several experimental and simulation studies have been performed to elucidate the solvation mechanism. In Fourier transform infrared spectroscopy (FT-IR) of water cluster in aqueous NaOH/urea solution, the intensity of the peaks associated with hydrogen bonds was found to increase as the temperature decreased from 13 to − 40 °C (Cai et al. 2008). They suggested the formation of a worm-like inclusion complex (IC) encaging cellulose chains in clusters of NaOH, urea, and water. A peak for the above cluster was found in the wide-angle X-ray diffraction analysis (WAXD), and its intensity increased as the temperature decreased from 25 to − 10 °C (Cai et al. 2008). Jiang et al. (2014) applied 13C-, 15N-, and 23Na-NMR spectroscopy to study the interaction between solvent species and cellulose. No chemical shift was found in the 13C-NMR spectrum of cellulose whether it was dissolved in NaOH solution or in NaOH/urea solution. In addition, there was no distinct change in the 15N-NMR spectra of NaOH/urea solution and NaOH/urea/cellulose solution. These experimental results indicated that there was no strong interaction between urea and cellulose. However, the peaks detected in the 23Na-NMR of NaOH solution shifted to upfield when urea or cellulose was added, implying an interaction of Na+ ion with urea or cellulose. The temperature dependence of 23Na relaxation rate was found to be higher in NaOH/urea/cellulose than in NaOH/cellulose, implying that urea contributes to the stability of NaOH/urea/cellulose. Jiang et al. (2014) suggested the formation of a urea–OH− cluster through the interaction between the NH2 groups of urea and OH− ion, and the stabilization of Na+ ion adjacent to the cellulose chain. Molecular dynamics (MD) simulation studies performed by Liu et al. (2018), Wernersson et al. (2015), and Cai et al. (2012) indicated the preferential adsorption of urea on the hydrophobic face of the pyranose rings of cellulose, which mitigated the hydrophobic interactions between neighboring cellulose chains. Liu et al. (2018) showed that NaOH was preferentially positioned near the hydrophilic hydroxyl groups of cellulose and that NaOH hydrates played a crucial role in maintaining the stability of the cellulose IC.
The segregation of the cellulose crystal into single molecules followed by solvation of the single molecules in a solvent is employed herein as the concept for cellulose solvation, as illustrated in Scheme 1.
On this basis, it is believed that the preferential solvation of cellulose in an alkali solution with urea can be explained by the thermodynamics for the solvation of the single cellulose molecule. The three-dimensional Reference Interaction Site Model (3D-RISM) theory combined with the Kovalenko-Hirata (KH) closure approximation (Gusarov et al. 2012; Kovalenko 2003, 2013, 2017; Kovalenko and Gusarov 2018) was used in this study to calculate a solvation energy and the distribution of solvent species such as Na+, OH−, urea, and water around the cellulose molecule. 3D-RISM-KH is s molecular theory of solvation based on analytical summation of the free energy diagrams, which yields the 3D site correlation functions in the statistical-mechanical equilibrium ensemble. By solving the RISM integral equations for the correlation functions (Gusarov et al. 2012; Kovalenko 2003, 2013, 2017; Kovalenko and Gusarov 2018), the solvation structure and thermodynamics have been predicted for various chemicals (Kaminski et al. 2010; Huang et al. 2015; Yoshida et al. 2002), bionanomaterials (Silveira et al. 2013; Stoyanov et al. 2014; Silveira et al. 2015) and biomolecular systems (Yoshida et al. 2009; Imai et al. 2007; Omelyan and Kovalenko 2015) in solutions.
The thermodynamics of solvation \( (\Delta {\text{U}}) \), including the solvation energy and the solvent potential energy, were determined based on the above distribution information. The solvation energy \( (\Delta {\text{U}}) \) is defined as the sum of the solvent potential energy (\( \Delta {\text{U}}^{uv} \)) and solvent reorganization energy (\( \Delta {\text{U}}_{R}^{uv} \)) (Ben-Naim 1978; Lazaridis 2000):
where superscripts u and v denote the solute and solvent, respectively. \( \Delta {\text{U}}^{uv} \) embodies the solute–solvent interaction. Once \( \Delta {\text{U}} \) and \( \Delta {\text{U}}^{uv} \) are determined, the above equation enables calculation of \( \Delta {\text{U}}_{R}^{uv} \), which considers the change in the solvent–solvent interaction upon insertion of the solute (cellulose) molecule. \( \Delta {\text{U}} \), \( \Delta {\text{U}}^{uv} \), and \( \Delta {\text{U}}_{R}^{uv} \) correspond to \( \Delta \varepsilon_{KH} \), \( {{\Delta }}\varepsilon^{uv} \), and \( \Delta \varepsilon^{vv} \) in Kovalenko (2017), respectively. In addition to the total values of \( \Delta {\text{U}} \), \( \Delta {\text{U}}^{uv} \), and \( \Delta {\text{U}}_{R}^{uv} \) for the solvation of cellulose, the thermodynamic contribution of each solvent species such as Na+, OH−, urea, and water could be determined. Such thermodynamic information would help in illuminating the structure of cellulose in the NaOH/urea solution (e.g., inclusion complex of cellulose in the NaOH-urea-water clusters).
Methods
The 3D-RISM calculations were performed in the canonical ensemble (NVT). The grid spacing was 0.025 Å for 1D RISM and 0.5 Å for 3d RISM. RISM calculations were performed with AMBER16 package. The TIP3P water was used for solvent water. General Amber Force Field (GAFF) was referred for evaluating LJ parameters of the solvent atoms and ions with AM1-BCC charge as shown in Fig. 1 (Case et al. 2016; Jakalian et al. 2000, 2002). A cellulose molecule with the Iβ structure and the degree of polymerization as 8 was chosen in this study as a model cellulose molecule. Its structure was created using cellulose-builder (Gomes and Skaf 2012). Figure 2 shows the structure of glucose, which is the repeating unit of a model cellulose molecule with the Iβ structure. By following the experimental conditions employed for the solvation of cellulose (Cai et al. 2006, 2007, 2008; Cai and Zhang 2005; Qi et al. 2011; Xiong et al. 2014; Yang et al. 2011), 2 M NaOH/2 M urea solution (abbreviated as solvent NU) was used as the solvent for cellulose. NaOH solution (2 M), urea solution (2 M), and pure water were tested for comparison with solvent NU; these solvents are abbreviated as solvent NO, solvent UO, and solvent WA, respectively. The temperature was assigned as 261 K, 280 K or 298 K. The placevent.py program was used to convert the 3D distribution of solvent atoms and ions into their specific positions around cellulose (Sindhikara et al. 2012). The molecular graphics and graphical representation were produced using the Visual Molecular Dynamics (VMD) (Humphrey et al. 1996).
Results and discussion
The probability of finding the solvent atom (or ion) γ at distance r from the cellulose molecule was determined from the 3D-RISM calculations, and is represented as its ratio to the probability of finding γ in the bulk phase. This ratio is expressed as \( g_{\gamma }^{uv} \left( r \right) \), where superscripts u and v denote the solute (i.e., the cellulose molecule) and the specific solvent, respectively. As an example of \( g_{\gamma }^{uv} \left( r \right) \), Fig. 3 graphically shows the regions where \( g_{\gamma }^{uv} \left( r \right) \) is higher than 3 for Na+ and urea. Urea, especially the carbon atom of urea, was preferentially located above and below the pyranose rings (see orange-colored surfaces in Fig. 3), which seems to agree well with the MD simulation studies of Liu et al. (2018), Wernersson et al. (2015), and Cai et al. (2012). Na+ and OH− were mostly distributed along the edges of the pyranose rings (Fig. 3). When the distributions with the higher \( g_{\gamma }^{uv} \left( r \right) \) (e.g., \( g_{\gamma }^{uv} \left( r \right) > 8 \) for Na+ and OH−) were considered, acidic Na+ was found to be positioned approximately 2.5 Å from basic O2 and O3 in the repeating glucose of cellulose. OH−, which should be close to Na+ due to the electrostatic interaction, was positioned approximately 1.6 Å from H6O of cellulose (see Fig. 4a). Such localization of Na+ and OH− suggests their electrostatic interaction with cellulose (Wang et al. 2017b; Liu et al. 2018) and hydrogen bonding between the OH− and H6O of cellulose (Fig. 4a), which leads to deprotonation of the hydroxymethyl groups of cellulose (Bialik et al. 2016; Xiong et al. 2013). The carbon atom of urea with \( g_{\gamma }^{uv} \left( r \right) > 4.2 \) was found to be located above and below O4 or O5 of cellulose and its distance from the pyranose ring was approximately 4.0 Å from O4 (Fig. 4a), which indicates the weak interaction, not likely hydrogen bonding between urea and cellulose (Wang et al. 2017c; Liu et al. 2018). With the use of solvents NU and NO (compare Fig. 4a, b), the positions of Na+ and OH− did not differ significantly, whilst the position of urea did not differ much with the use of solvents NU and UO (compare Fig. 4a, c). This observation implies that the interactions of cellulose with NaOH and urea are not affected by urea and NaOH, respectively.
Isosurface representation of the 3D-distribution function at 261 K, \( g_{\gamma }^{uv} \left( {\text{r}} \right) \) of a solvent atom or ion (γ) when solute u is the single cellulose molecule and solvent v is the aqueous NaOH–urea solution. The isosurface with \( g_{\gamma }^{uv} \left( {\text{r}} \right) \) > 3 is colored yellow for γ of sodium ion and orange for γ of carbon atom of urea
The preferred positioning of Na+, OH−, and urea around the cellulose molecule was suspected to influence the distribution of the water molecules. \( g_{\gamma }^{u12} \left( r \right) \) and \( g_{\gamma }^{u13} \left( r \right) \), were calculated to investigate the effects of urea in the alkali solvent and NaOH in the urea solvent on the distribution of water, respectively:
where superscripts v1, v2, and v3 denote solvents NU, NO, and UO, respectively. The regions where \( g_{\gamma }^{u12} \left( r \right) > 0.3 \) and \( g_{\gamma }^{u12} \left( r \right) < - 0.2 \) are shown in Fig. 5a, b, respectively. Water molecules were found to be positioned closer to both the edges and pyranose rings of the cellulose molecule due to the inclusion of urea in the alkali solvent. The regions where \( g_{\gamma }^{u13} \left( r \right) > 0.3 \) and \( g_{\gamma }^{u13} \left( r \right) < - 0.2 \) are shown in Fig. 5c, d, respectively. Similar to the case with urea, the inclusion of NaOH in the urea solvent enabled positioning of the water molecules closer to the cellulose molecule. Hence, hydrogen bonding between water and cellulose is more probable in solvent NU than in the other solvents. Figure 5a, c show that water molecules surround the cellulose molecule and are clustered, connecting Na+, OH−, and urea through hydrogen bonds, which is correspondent with the clusters of NaOH, urea, and water suggested by Cai et al. (2008).
Graphical representation of difference in the 3D distribution functions of water around the cellulose molecule in two different solvents: \( \Delta g_{\gamma }^{u12} \left( r \right) \) and \( \Delta g_{\gamma }^{u13} \left( r \right). \) The differences are defined as \( g_{\gamma }^{uv1} \left( {\text{r}} \right) - g_{\gamma }^{uv2} \left( {\text{r}} \right) \) and \( g_{\gamma }^{uv1} \left( {\text{r}} \right) - g_{\gamma }^{uv3} \left( {\text{r}} \right) \), respectively, where superscript u denotes the single cellulose molecule. Superscripts v1, v2, and v3 denote solvents NU, NO, and UO, respectively. Subscript γ may be O or H for the oxygen and hydrogen of water, respectively. The isosurfaces with \( \Delta g_{\gamma }^{u12} \left( r \right) > 0.3 \) and \( \Delta g_{\gamma }^{u13} \left( r \right) > 0.3 \) are shown in Fig. 4a, c, respectively, and are colored purple for γ = O and blue for γ = H. The isosurfaces with \( \Delta g_{\gamma }^{u12} \left( r \right) < - 0.2 \) and \( \Delta g_{\gamma }^{u13} \left( r \right) < - 0.2 \) are shown in Fig. 4b, d, respectively, and are colored green for γ = O and pink for γ = H. The very small pink-colored regions correspond to H
The solvation energy (\( \Delta {\text{U}} \)), solvent potential energy (\( \Delta {\text{U}}^{uv} \)), and solvent reorganization energy (\( \Delta {\text{U}}_{R}^{uv} \)), which is equal to the difference between and (see Eq. (9)), were calculated for cellulose in four different solvents at 261 K, 280 K, and 298 K, and are listed in Table 1. The much more negative \( \Delta {\text{U}} \) in solvent NU than in the other solvents indicates that the presence of NaOH and urea is critical in the solvation of cellulose. By comparing \( \Delta {\text{U}}^{uv} \) and \( \Delta {\text{U}}_{R}^{uv} \) in solvent NU, it is deduced that the negative solvation energy originates mostly from \( \Delta {\text{U}}_{R}^{uv} \). As shown in Figs. 4 and 5, the solvent species such as NaOH, urea, and water molecules were clustered around the cellulose molecule. Hence, the hydrogen bonds between these species are believed to be induced by cellulose, which leads to \( \Delta {\text{U}}_{R}^{uv} \) being more negative than \( \Delta {\text{U}}^{uv} \). Because hydrogen bonding is weaker at higher temperature, \( \Delta {\text{U}}_{R}^{uv} \) at higher temperature was smaller (i.e., less negative) than that at lower temperature (i.e., \( \Delta {\text{U}}_{R}^{uv} \left( {261 K} \right) \) < \( \Delta {\text{U}}_{R}^{uv} \left( {280 K} \right) \) < \( \Delta {\text{U}}_{R}^{uv} \left( {298 K} \right) \) < 0). This is consistent with the study of Cai et al. (2008) that the hydrogen-bonded network structure in aqueous NaOH/urea (i.e., solvent NU) was highly stable at low temperature such as − 12 °C (i.e., 261 K). In solvents other than solvent NU, \( \Delta {\text{U}}_{R}^{uv} \) was positive and not affected by temperature, whereas \( \Delta {\text{U}}^{uv} \) was negative. This suggests that NaOH and urea are both critical for clustering the solvent species and forming hydrogen bonds around the cellulose molecule. The contributions of water, NaOH, and urea to \( \Delta {\text{U}}\), \( \Delta {\text{U}}^{uv} \), and \( \Delta {\text{U}}_{R}^{uv} \) are shown in Tables 1, S1, and S2 (see Supplementary Material for Tables S1 and S2), respectively. The \( \Delta {\text{U}}^{uv} \) values indicate that the attractive interaction of the cellulose molecule with water was stronger than that with NaOH (compare \( \Delta {\text{U}}^{uv} \) for water and NaOH in solvents NU and NO in Table 1 and Table S1) and urea (compare \( \Delta {\text{U}}^{uv} \) for water and urea in solvents NU and UO in Table 1 and Table S2). The contributions of NaOH and urea to the negative solvent reorganization energy in solvent NU were negligible. Hence, NaOH and urea, which are positioned along the edges and above/below the pyranose rings of cellulose (Fig. 3, 4), attract water molecules, facilitating clustering near NaOH and urea around the cellulose molecule (Fig. 5), leading to numerous hydrogen bonds between the clustered water molecules as well as between the cellulose molecule and the nearby water molecules. The significantly negative \( \Delta {\text{U}}_{R}^{uv} \) due to the former hydrogen bonds and the negative \( \Delta {\text{U}}^{uv} \) due to the latter hydrogen bonds enable cellulose solvation in solvent NU. From the 3D RISM data for cellulose solvation, we suggest that the helical cluster of NaOH, urea, and water around the single cellulose molecule allows the solvation of cellulose in the aqueous solution of 2 M NaOH and 2 M urea at temperatures as low as 261 K (see Fig. 6)
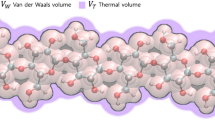
Helical cluster of NaOH, urea, and water around single cellulose molecule. A single cellulose molecule is shown in the ball-and-stick representation within the van der Waals transparent surface (cyan). Sodium ions are represented as yellow van der Waal spheres, whereas blue-colored hydroxide ions are ball-and-stick representations. Orange-colored urea and pink-colored water molecules have the licorice representation. The hydrogen bonds are indicated by the pink dotted line. The top view and side view with the additional distribution of the oxygen of water are shown in Fig. S1 (Supplementary Material)
Conclusions
The solvation of cellulose in aqueous NaOH–urea solution was thermodynamically analyzed using the three-dimensional Reference Interaction Site Model theory coupled with the Kovalenko–Hirata closure approximation. NaOH solution, urea solution, and pure water were tested for comparison with aqueous NaOH–urea solution. The distributions of Na+, OH−, and urea around the cellulose molecule suggests that there is the electrostatic interaction between the charged species (i.e., Na+ and OH−) and cellulose while the weak interaction, not likely hydrogen bonding exists between urea and cellulose. The solvation energy and the contribution of each solvent species to the solvation energy were calculated from the preferential distributions of Na+, OH−, urea, and water in a close proximity to the cellulose molecule. The solvation energy is equal to the sum of the solvent potential energy, which considers the solvent–solute interaction, and the solvent reorganization energy. Cellulose solvation in the NaOH/urea solution was observed to be primarily due to reorganization of the water molecules around the cellulose molecule. The solvated structure was suggested to be cellulose included in helical clusters of Na+, OH−, urea, and water, wherein the clusters comprised a repeated arrangement of OH− hydrate, water molecules, urea hydrate, and water molecules. The aggregates of solvent species connected by hydrogen bonding accounted for the exothermic solvation of cellulose, wherein the solvation energy became less negative with increasing temperature. Cellulose is suspected to play the role of a structural director for water in the presence of NaOH and urea, and as a structure breaker for water in the absence of NaOH and/or urea.
References
Ben-Naim A (1978) Standard thermodynamics of transfer. Uses and misuses. J Phys Chem 82:792–803
Bialik E, Stenqvist B, Fang Y, Östlund Å, Furó I, Lindman B, Lund M, Bernin D (2016) Ionization of cellobiose in aqueous alkali and the mechanism of cellulose dissolution. J Phys Chem Lett 7:5044–5048
Cai J, Zhang L (2005) Rapid dissolution of cellulose in LiOH/urea and NaOH/urea aqueous solutions. Macromol Biosci 5:539–548
Cai J, Zhang L, Zhou J, Li H, Chen H, Jin H (2004) Novel fibers prepared from cellulose in NaOH/urea aqueous solution. Macromol Rapid Commun 25:1558–1562
Cai J, Liu Y, Zhang L (2006) Dilute solution properties of cellulose in LiOH/urea aqueous system. J Polym Sci Pol Phys 44:3093–3101
Cai J, Zhang L, Chang C, Cheng G, Chen X, Chu B (2007) Hydrogen-bond-induced inclusion complex in aqueous cellulose/LiOH/urea solution at low temperature. ChemPhysChem 8:1572–1579
Cai J, Zhang L, Liu S, Liu Y, Xu X, Chen X, Chu B, Guo X, Xu J, Cheng H, Han C, Kuga S (2008) Dynamic self-assembly induced rapid dissolution of cellulose at low temperatures. Macromolecules 41:9345–9351
Cai L, Liu Y, Liang H (2012) Impact of hydrogen bonding on inclusion layer of urea to cellulose: study of molecular dynamics simulation. Polymer 53:1124–1130
Case DA, Betz R, Cerutti D, Cheatham TE, Darden T, Duke RE (2016) AMBER 16. University of California, San Fransisco
Gomes TC, Skaf MS (2012) Cellulose-builder: a toolkit for building crystalline structures of cellulose. J Comput Chem 33:1338–1346
Gusarov S, Pujari BS, Kovalenko A (2012) Efficient treatment of solvation shells in 3D molecular theory of solvation. J Comput Chem 33:1478–1494
Huang W, Blinov N, Kovalenko A (2015) Octanol–water partition coefficient from 3D-RISM-KH molecular theory of solvation with partial molar volume correction. J Phys Chem B 17:5588–5597
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Gr 14:33–38
Imai T, Ohyama S, Kovalenko A, Hirata F (2007) Theoretical study of the partial molar volume change associated with the pressure-induced structural transition of ubiquitin. Protein Sci 16(9):1927–1933
Jakalian A, Bush BL, Jack DB, Bayly CI (2000) Fast, efficient generation of high-quality atomic charges. AM1-BCC model: I. Method. J Comput Chem 21:132–146
Jakalian A, Jack DB, Bayly CI (2002) Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II. Parameterization and validation. J Comput Chem 23:1623–1641
Jiang Z, Fang Y, Xiang J, Ma Y, Lu A, Kang H, Huang Y, Guo H, Liu R, Zhang L (2014) Intermolecular interactions and 3D structure in cellulose–NaOH–urea aqueous system. J Phys Chem B 118:10250–10257
Jin H, Zha C, Gu L (2007) Direct dissolution of cellulose in NaOH/thiourea/urea aqueous solution. Carbohyd Res 342:851–858
Kaminski JW, Gusarov S, Wesolowski TA, Kovalenko A (2010) Modeling solvatochromic shifts using the orbital-free embedding potential at statistically mechanically averaged solvent density. J Phys Chem A 114(20):6082–6096
Kovalenko A (2003) Three-dimensional Rism theory for molecular liquids and solid–liquid interfaces. In: Hirata F (ed) Molecular theory of solvation. Springer, Dordrecht, pp 169–275
Kovalenko A (2013) Multiscale modeling of solvation in chemical and biological nanosystems and in nanoporous materials. Pure Appl Chem 85(1):159–199
Kovalenko A (2017) Multiscale modeling of solvation. In: Breitkopf C, Swider-Lyons K (eds) Springer handbook of electrochemical energy. Springer, Heidelberg, pp 95–139
Kovalenko A, Gusarov S (2018) Multiscale methods framework: self-consistent coupling of molecular theory of solvation with quantum chemistry, molecular simulations, and dissipative particle dynamics. Phys Chem Chem Phys 20(5):2947–2969
Lazaridis T (2000) Solvent reorganization energy and entropy in hydrophobic hydration. J. Phys Chem B 104:4964–4979
Liu G, Sun H, Liu G, Zhang H, Yuan S, Zhu Q (2018) A molecular dynamics study of cellulose inclusion complexes in NaOH/urea aqueous solution. Carbohyd Polym 185:12–18
Omelyan I, Kovalenko A (2015) MTS-MD of biomolecules steered with 3D-RISM-KH mean solvation forces accelerated with generalized solvation force extrapolation. J Chem Theory Comput 11(4):1875–1895
Qi H, Yang Q, Zhang L, Liebert T, Heinze T (2011) The dissolution of cellulose in NaOH-based aqueous system by two-step process. Cellulose 18:237–245
Silveira RL, Stoyanov SR, Gusarov S, Skaf MS, Kovalenko A (2013) Plant biomass recalcitrance: effect of hemicellulose composition on nanoscale forces that control cell wall strength. J Am Chem Soc 135(51):19048–19051
Silveira RL, Stoyanov SR, Gusarov S, Skaf MS, Kovalenko A (2015) Supramolecular interactions in secondary plant cell walls: effect of lignin chemical composition revealed with the molecular theory of solvation. J Phys Chem Lett 6(1):206–211
Sindhikara DJ, Yoshida N, Hirata F (2012) Placevent: an algorithm for prediction of explicit solvent atom distribution—application to HIV-1 protease and F-ATP synthase. J Comput Chem 33:1536–1543
Stoyanov SR, Lyubimova O, Gusarov S, Kovalenko A (2014) Computational modeling of the structure relaxation and dispersion thermodynamics of pristine and modified cellulose nanocrystals in solution. Nord Pulp Pap Res J 29(1):144–155
Wang S, Sun P, Zhang R, Lu A, Liu M, Zhang L (2017a) Cation/macromolecule interaction in alkaline cellulose solution characterized with pulsed field-gradient spin-echo NMR spectroscopy. Phys Chem Chem Phys 19:7486–7490
Wang S, Lyu K, Sun P, Lu A, Liu M, Zhuang L, Zhang L (2017b) Influence of cation on the cellulose dissolution investigated by MD simulation and experiments. Cellulose 24:4641–4651
Wang S, Sun P, Liu M, Lu A, Zhang L (2017c) Weak interactions and their impact on cellulose dissolution in an alkali/urea aqueous system. Phys Chem Chem Phys 19:17909–17917
Wernersson E, Stenqvist B, Lund M (2015) The mechanism of cellulose solubilization by urea studied by molecular simulation. Cellulose 22:991–1001
Xiong B, Zhao P, Cai P, Zhang L, Hu K, Cheng G (2013) NMR spectroscopic studies on the mechanism of cellulose dissolution in alkali solutions. Cellulose 20:613–621
Xiong B, Zhao P, Hu K, Zhang L, Cheng G (2014) Dissolution of cellulose in aqueous NaOH/urea solution: role of urea. Cellulose 21:1183–1192
Yang Q, Qi H, Lue A, Hu K, Cheng G, Zhang L (2011) Role of sodium zincate on cellulose dissolution in NaOH/urea aqueous solution at low temperature. Carbohyd Polym 83:1185–1191
Yoshida K, Yamaguchi T, Kovalenko A, Hirata F (2002) Structure of tert-butyl alcohol − water mixtures studied by the RISM theory. J Phys Chem B 106(19):5042–5049
Yoshida N, Imai T, Phongphanphanee S, Kovalenko A, Hirata F (2009) Molecular recognition in biomolecules studied by statistical-mechanical integral-equation theory of liquids. J Phys Chem B 113(4):873–886
Acknowledgments
This work was supported by Yonsei University, and we would like to thank Editage (www.editage.co.kr) for English language editing.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Huh, E., Yang, JH., Lee, CH. et al. Thermodynamic analysis of cellulose complex in NaOH–urea solution using reference interaction site model. Cellulose 27, 6767–6775 (2020). https://doi.org/10.1007/s10570-020-03202-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10570-020-03202-w