Abstract
To determine the expression of signal transducer and activator of transcription 3 (STAT3) in patients with fragility fractures (FFs) and its effect on the biological function of osteoblasts. The study included 32 patients with FFs who were diagnosed and treated in the research group and 30 concurrent healthy individuals in the control group. We observed STAT3 mRNA expression in the patients with FFs and controls and altered STAT3 mRNA to detect changes in the proliferation, invasion, and apoptosis of osteoblasts. The patients with FFs presented higher serum STAT3 mRNA expression than the controls (P < 0.05). We plotted receiver operating characteristic curves based on the STAT3 mRNA expression and found that the area under the curve for STAT3 mRNA was 0.856 (P < 0.05). Transfection of STAT3 mRNA mimics resulted in increased STAT3 mRNA expression, inhibited cell proliferation as detected by an MTT assay, and increased apoptosis rate, which was determined using flow cytometry with human fetal osteoblastic cell line 1.19 cells. STAT3 mRNA expression was elevated in the serum of patients with FFs and can be used as a biomarker for the diagnosis of the disease. Regulating STAT3 mRNA can inhibit the proliferation and induce the osteoblasts apoptosis.
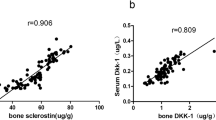
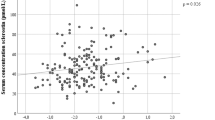
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A fragility fracture (FF) occurs in the absence of trauma and is mostly caused by osteoporosis, thereby frequently affecting elderly patients (Posen et al. 2013; World Health Organization 1998). Osteoporosis decreases bone mass and density; thus, fractures are more likely to occur in everyday life or during minor trauma (Haentjens et al. 2003). Compared with healthy individuals, patients with osteoporosis are at increased risk of fractures, which adversely affects public and individual health (Kanis et al. 2019). Therefore, it is essential to clarify the pathogenesis of FFs and find potential therapeutic targets.
MicroRNA (miR) is a highly conserved endogenous short-chain non-coding RNA that can regulate approximately one third of human genes (Li et al. 2016). Studies on miR function have revealed that they have obvious abnormal expression in various orthopedic diseases and can participate in disease progression. For example, miR-495 is upregulated in osteoarthritis and promotes the development of osteoarthritis by modulating AKT1 (Zhao et al. 2019), whereas miR-106b can inhibit osteoblast differentiation and bone formation by negatively regulating the expression of bone morphogenetic protein-2 (Liu et al. 2017). In addition, miR-142-5p promotes fracture healing in aged mice via the stimulation of osteoblast activity (Tu et al. 2017). Studies have reported the abnormal expression of signal transducer and activator of transcription 3 (STAT3) in osteosarcoma (Li et al. 2019); however, no studies have reported STAT3 expression in FFs. Osteoblasts are the primary cell type in bone formation and crucially contribute to the metabolic balance, growth, and injury repair of bone tissue (Li et al. 2015). Studies have found that miR can interfere with the biological behavior of osteoblasts by regulating the expression of their target genes, thereby participating in the pathogenesis of FFs (Yan et al. 2018; Zhang et al. 2017). We hypothesized that STAT3 could be similarly involved in the pathogenesis of FFs. Therefore, we explored the role and potential mechanisms of STAT3 in FFs by conducting a series of experiments, which might contribute to a better understanding of FF pathogenesis.
Materials and methods
General information
Totally, 32 patients with FFs who were diagnosed and treated in our hospital from March 2018 to November 2019 were included in the research group (RG) (21 men and 11 women; mean age, 50.8 ± 10.6 years). We also included 30 healthy individuals as the control group (CG) who underwent physical examinations at our hospital during the same period (18 men and 12 women; mean age, 51.2 ± 10.3 years). The study was approved by the Medical Ethics Committee of Jinling Hospital, Medicine College, Nanjing University.
Inclusion and exclusion criteria
Inclusion criteria: With complete case data and an informed consent form signed by the patient or their immediate family, all enrolled patients underwent treatment in our hospital after the diagnosis and cooperated with the hospital’s medical staff.
Exclusion criteria: We excluded patients with vital organ damage, other cardiovascular and/or cerebrovascular diseases, physical disabilities, other autoimmune diseases, mental diseases, speech/language disorders, or any disease that might affect the study results. We also excluded referred patients and those with surgical contraindications.
Cell sources, reagents, and instruments
The following materials were used: human fetal osteoblastic cell line (hFOB) 1.19 cells (Beijing BeNa Culture Collection, Beijing, China, Resource No. BNCC344663); Lipofectamine™ 2000 (Shanghai Mituo Biotechnology Co., Ltd., Shanghai, China, Cat. No. 11668019); MMT kit and dimethyl sulfoxide reagent (Shanghai Yuanye Biotechnology Co., Ltd., Shanghai, China, Cat. No. S30860, S24295); Roswell Park Memorial Institute-1640 (RPMI-1640; Shanghai Yanjin Biotechnology Co., Ltd., Shanghai, China, Cat. No. 31800-022) medium; phosphate buffer saline and fetal bovine serum (Shanghai Hengfei Biotechnology Co., Ltd., Shanghai, China, Cat. No. P1000, SA133); penicillin–streptomycin dual antibiotic solution (Beijing Biolab Technology Co., Ltd., Beijing, China, Cat. No. MT0104-SEY); TransScript II Green Two-Step qRT-PCR SuperMix (Beijing TransGen Biotech, Beijing, China, AQ301-01); Annexin V/PI apoptosis detection kit (Shanghai Hengfei Biotechnology Co., Ltd., Shanghai, China, Cat. No. CA1020); microplate reader (Beijing Image Trading Co., Ltd., Beijing, China, Cat. No. 21261000); polymerase chain reaction (PCR) instrument (Wuxi Microsep Biotechnology Co., Ltd., Wuxi, China, Cat. No. TC9639); and flow cytometer (Beijing Image Trading Co., Ltd., Beijing, China, Cat. No. AMG0002051). The primer sequence of STAT3 mRNA was designed and synthesized by Shanghai Sangon Bio-tech Co., Ltd. (Table 1).
Cell culture and transfection
We transferred the hFOB 1.19 cellsto RPMI-1640 medium (penicillin–streptomycin dual antibiotic solution, 10% fetal bovine serum) and cultured in an incubator at 37 °C with 5% carbon dioxide. Subsequently, we transfected the cells with STAT3 mRNA mimics and STAT3 mRNA inhibitor using the Lipofectamine™ 2000 kit strictly. All primers were transfected into the cells with the greatest difference in expression.
Quantitative reverse transcriptase (qRT)-PCR detection
Following the manufacturer’s protocol, total RNA was extracted from the collected cells and serum using the TRIzol kit. We determined the purity, concentration, and integrity of the extracted total RNA using an ultraviolet spectrophotometer and agarose gel electrophoresis. We used the 5X TransScript ®II All-in-One SuperMix, TransScript ®miRNA RT Enzyme Mix, and 2 × TS miRNA Reaction Mix kits for reverse transcription strictly according to the manufacturer’s instructions. Next, we performed PCR amplification using a PCR reaction system consisting of 1 μL of cDNA, upstream and downstream primers (0.4 μL each), 10 μL of 2X TransScript® Tip Green qPCR SuperMix, 200 μL of passive reference dye (50×), and ultra-pure sterile water for a final volume of 20 μL. The PCR reaction conditions were as follows: 45 cycles comprising pre-denaturation at 94 °C for 30 s, denaturation at 94 °C for 5 s, and annealing and extension at 60 °C for 30 s. Three replicate wells were used for each sample, and the experiment was performed in triplicates. In this experiment, U6 was used as the internal reference and the delta-delta Ct method was used to analyze the data.
Detection of cell proliferation
We harvested the cells 24 h after transfection, adjusted the concentration to 3 × 104 cells/well, and inoculated the cells in 96-well plates for incubation at 37 °C for 24, 48, 72, and 96 h. We then added 20 μL of MTT solution (5 μg/mL) to each well at each of the aforementioned timepoints and incubated the culture at 37 °C for 4 h before adding 150 μL of dimethyl sulfoxide. We measured the optical density of the cells in each group using a microplate reader at 450 nm.
Detection of apoptosis
After 24 h of transfection, the cells were fully digested using 0.25% trypsin, washed twice with phosphate buffer saline, and added to 100 μL of binding buffer to prepare a suspension of 1 × 106 cells/mL. Next, we added AnnexinV-FITC and PI and incubated the cells for 5 min at room temperature in the dark. We detected apoptosis using the FC500MCL flow cytometry system and repeated the experiment 3 times to obtain the mean value.
Outcome measures
The primary outcome measures were STAT3 mRNA expression in the patients with FFs and healthy controls. We altered the STAT3 mRNA expression to detect the changes in proliferation ability, invasion ability, and apoptosis rate of osteoblasts. The secondary outcome was the diagnostic value of STAT3 mRNA in patients with FFs.
Statistical methods
We analyzed the collected data using SPSS 20.0 and visualized the results using GraphPad 7. We used the Kolmogorov–Smirnov test to analyze the dose data distribution, in which the normal distribution data were expressed as mean and standard deviation (mean ± SD). We used the independent sample and paired t-tests for intergroup and intragroup comparisons, respectively. The counting data are presented as percentages and were analyzed using the chi-squared test. We plotted the diagnostic value of STAT3 mRNA in FFs using a receiver operating characteristic curve and the 5-year patient survival using a Kaplan–Meier survival curve, using the log-rank test for analysis. A p-value of < 0.05 was considered significant.
Results
Clinical data
There were no significant differences between the RG and CG with regard to age, sex, body mass index, marital status, ethnicity, residence, smoking, drinking history, and exercise, suggesting comparability (P > 0.05) (Table 2).
STAT3 mRNA expression in RG and CG
We used qRT-PCR to detect STAT3 mRNA in the serum of the RG and CG and found that STAT3 mRNA expression in the RG (4.32 ± 1.45) was higher than that in the CG (1.02 ± 0.34) (P < 0.05).
Diagnostic value of STAT3 mRNA in patients with FFs
The ROC curve revealed an area under the curve for STAT3 mRNA of 0.856 for diagnosing patients with FFs (P < 0.05) (Table 3, Fig. 1).
Effect of STAT3 mRNA on the biological function of osteoblasts
The transfection of STAT3 mRNA mimics increased STAT3 mRNA expression, inhibited cell proliferation, and increased the apoptosis rate in the hFOB 1.19 cells (Fig. 2).
Effect of elevated STAT3 mRNA expression on the proliferation, apoptosis and differentiation of human fetal osteoblastic cell line 1.19 cells. A STAT3 mRNA expression increased in hFOB 1.19 cells after transfection of STAT3 mRNA mimics. B STAT3 mRNA expression in hFOB 1.19 cells in the STAT3 mRNA mimics group and STAT3 mRNA inhibitors group was significantly higher than that in the miR-NC group after transfection of STAT3 mRNA mimics. C hFOB 1.19 cell proliferation was inhibited after transfection of STAT3 mRNA mimics. D The hFOB 1.19 cell apoptosis rate increased after transfection of STAT3 mRNA mimics. E Apoptosis diagram
Discussion
FF is the primary cause of high morbidity and mortality among older adults (Burge et al. 2007). Given the relationship between FF and aging, the number of patients with FFs is increasing with the increasing age of the population, imposing a serious burden on individuals, families, and society (Varahra et al. 2018). Survivors of the first episode of FF are a well-defined high-risk group and are at significantly greater risk of experiencing a second fracture, with the highest incidence within the first 6 months of the first fracture (Munson et al. 2016). Therefore, the pathological processes of FFs are of great social and economic importance and must be explored.
With the expansion of research in recent years, the focus of investigators worldwide has shifted toward genetics. MiRNA can act as endogenous RNA interference to regulate the expression of target genes and participate in the regulation of various physiological and pathological functions (Vitsios et al. 2017). MiRNA can bind to 3′ UTR via base pairing to regulate gene expression (Rouleau et al. 2017) and shows significant expression profiles between different tissues and growth stages, indicating that miRNA has different spatial–temporal expression patterns (Li et al. 2017). Approximately 22 nucleotides in length (Takeda et al. 2015), miRNA can regulate posttranscriptional gene expression by inducing mRNA degradation and inhibiting translation, degrade the target gene, and inhibit the translation of the target gene, thereby completing the posttranscriptional gene silencing (Oxnard et al. 2016). Studies have shown that miRNA can be used as a negative regulator of gene expression to regulate a series of biological functions, thus playing an essential role in various biological processes (Ganju et al. 2017; Rupaimoole et al. 2016).
In this study, we first detected STAT3 mRNA expression in patients using qRT-PCR and found that it was significantly higher in the RG than in the CG, suggesting that STAT3 mRNA may be involved in the pathogenesis of FFs. Patients with FFs have no obvious clinical manifestations before the occurrence of a fracture with no obvious cause and are therefore prone to missing the optimal treatment period (Yang et al. 2018). Studies have found that abnormal miR expression in serum can be used as a potential marker to diagnose various diseases, including FFs (Lin et al. 2019; Moya et al. 2019; Wang et al. 2019). Therefore, we analyzed the diagnostic value of serum STAT3 mRNA in FFs and found an area under the curve of 0.856 for STAT3 mRNA for diagnosing osteoporosis, indicating that STAT3 mRNA has potential as a biological indicator for diagnosing FFs. The decrease in osteoblast proliferation and increase in apoptosis are important factors in the occurrence and development of FFs (Wang et al. 2019). miR is known to regulate many cellular biological processes, such as proliferation, migration, apoptosis, differentiation, and metabolism (Lin et al. 2019). Therefore, in this study, we analyzed the role of STAT3 mRNA in FFs by observing changes in biological behaviors such as cell proliferation and apoptosis after an increase in STAT3 mRNA expression in osteoblasts. Using the MTT assay and flow cytometry, we found that after an increase in STAT3 mRNA expression in osteoblasts, the ability of the osteoblasts to proliferate was inhibited and the rate of apoptosis increased, indicating that STAT3 mRNA can affect the proliferation and apoptosis of osteoblasts and the occurrence and progression of FFs.
This study confirms the preliminary clinical value of STAT3; however, the study has certain limitations. First, the study did not include in vivo experiments, and second, the patients were not followed-up for the prognosis. Therefore, we aim to support our research results by conducting in vivo experiments and prognostic follow-ups in future studies.
In conclusion, STAT3 mRNA expression was elevated in the serum of patients with FFs and can be used as a biomarker to diagnose the disease. STAT3 mRNA regulation can inhibit osteoblast proliferation and induce apoptosis.
Availability of data and materials
The data used to support the findings of this study are available from the corresponding author upon request.
References
Burge R, Dawson-Hughes B, Solomon DH, Wong JB, King A, Tosteson A (2007) Incidence and economic burden of osteoporosis-related fractures in the United States, 2005–2025. J Bone Miner Res 22:465–475. https://doi.org/10.1359/jbmr.061113
Ganju A, Khan S, Hafeez BB, Behrman SW, Yallapu MM, Chauhan SC, Jaggi M (2017) miRNA nanotherapeutics for cancer. Drug Discov Today 22:424–432. https://doi.org/10.1016/j.drudis.2016.10.014
Haentjens P, Autier P, Collins J, Velkeniers B, Vanderschueren D, Boonen S (2003) Colles fracture, spine fracture, and subsequent risk of hip fracture in men and women: a meta-analysis. J Bone Joint Surg Am 85:1936–1943. https://doi.org/10.2106/00004623-200310000-00011
Kanis JA, Cooper C, Rizzoli R, Reginster JY (2019) European guidance for the diagnosis and management of osteoporosis in postmenopausal women. Osteoporos Int 30:3–44. https://doi.org/10.1007/s00198-018-4704-5
Li J, He X, Wei W, Zhou X (2015) MicroRNA-194 promotes osteoblast differentiation via downregulating STAT1. Biochem Biophys Res Commun 460:482–488. https://doi.org/10.1016/j.bbrc.2015.03.059
Li Q et al (2016) miR-139-5p inhibits the epithelial-mesenchymal transition and enhances the chemotherapeutic sensitivity of colorectal cancer cells by downregulating BCL2. Sci Rep 6:27157. https://doi.org/10.1038/srep27157
Li JQ, Rong ZH, Chen X, Yan GY, You ZH (2017) MCMDA: Matrix completion for MiRNA-disease association prediction. Oncotarget 8:21187–21199. https://doi.org/10.18632/oncotarget.15061
Li MH, Wu ZY, Wang Y, Chen FZ, Liu Y (2019) Expression of miR-29 and STAT3 in osteosarcoma and its effect on proliferation regulation of osteosarcoma cells. Eur Rev Med Pharmacol Sci 23:7275–7282. https://doi.org/10.26355/eurrev_201909_18832
Lin C et al (2019) Circulating miR-338 cluster activities on osteoblast differentiation: potential diagnostic and therapeutic targets for postmenopausal osteoporosis. Theranostics 9:3780–3797. https://doi.org/10.7150/thno.34493
Liu K et al (2017) Silencing miR-106b accelerates osteogenesis of mesenchymal stem cells and rescues against glucocorticoid-induced osteoporosis by targeting BMP2. Bone 97:130–138. https://doi.org/10.1016/j.bone.2017.01.014
Moya L, Meijer J, Schubert S, Matin F, Batra J (2019) Assessment of miR-98-5p, miR-152-3p, miR-326 and miR-4289 expression as biomarker for prostate cancer diagnosis. Int J Mol Sci. https://doi.org/10.3390/ijms20051154
Munson JC et al (2016) Patterns of prescription drug use before and after fragility fracture. JAMA Intern Med 176:1531–1538. https://doi.org/10.1001/jamainternmed.2016.4814
Oxnard GR et al (2016) Association between plasma genotyping and outcomes of treatment with osimertinib (AZD9291) in advanced non-small-cell lung cancer. J Clin Oncol 34:3375–3382. https://doi.org/10.1200/jco.2016.66.7162
Posen J, Beaton DE, Sale J, Bogoch ER (2013) Bone mineral density testing after fragility fracture: Informative test results likely. Can Fam Physician 59:e564–e571
Rouleau S, Glouzon JS, Brumwell A, Bisaillon M, Perreault JP (2017) 3’ UTR G-quadruplexes regulate miRNA binding. RNA 23:1172–1179. https://doi.org/10.1261/rna.060962.117
Rupaimoole R, Calin GA, Lopez-Berestein G, Sood AK (2016) miRNA deregulation in cancer cells and the tumor microenvironment. Cancer Discov 6:235–246. https://doi.org/10.1158/2159-8290.Cd-15-0893
Takeda M, Okamoto I, Nakagawa K (2015) Pooled safety analysis of EGFR-TKI treatment for EGFR mutation-positive non-small cell lung cancer. Lung Cancer 88:74–79. https://doi.org/10.1016/j.lungcan.2015.01.026
Tu M, Tang J, He H, Cheng P, Chen C (2017) MiR-142-5p promotes bone repair by maintaining osteoblast activity. J Bone Miner Metab 35:255–264. https://doi.org/10.1007/s00774-016-0757-8
Varahra A, Rodrigues IB, MacDermid JC, Bryant D, Birmingham T (2018) Exercise to improve functional outcomes in persons with osteoporosis: a systematic review and meta-analysis. Osteoporos Int 29:265–286. https://doi.org/10.1007/s00198-017-4339-y
Vitsios DM, Davis MP, van Dongen S, Enright AJ (2017) Large-scale analysis of microRNA expression, epi-transcriptomic features and biogenesis. Nucleic Acids Res 45:1079–1090. https://doi.org/10.1093/nar/gkw1031
Wang H, Wei X, Wu B, Su J, Tan W, Yang K (2019) Tumor-educated platelet miR-34c-3p and miR-18a-5p as potential liquid biopsy biomarkers for nasopharyngeal carcinoma diagnosis. Cancer Manag Res 11:3351–3360. https://doi.org/10.2147/cmar.S195654
World Health Organization (1998) Guidelines for preclinical evaluation and clinical trials in osteoporosis. World Health Organization, Geneva
Yan TB, Li C, Jiao GJ, Wu WL, Liu HC (2018) TIMP-1 suppressed by miR-138 participates in endoplasmic reticulum stress-induced osteoblast apoptosis in osteoporosis. Free Radic Res 52:223–231. https://doi.org/10.1080/10715762.2017.1423070
Yang R, Tao Y, Wang C, Shuai Y, Jin L (2018) Circulating microRNA panel as a novel biomarker to diagnose bisphosphonate-related osteonecrosis of the jaw. Int J Med Sci 15:1694–1701. https://doi.org/10.7150/ijms.27593
Zhang Y et al (2017) MicroRNA-221 is involved in the regulation of osteoporosis through regulates RUNX2 protein expression and osteoblast differentiation. Am J Transl Res 9:126–135
Zhao X et al (2019) MicroRNA-495 enhances chondrocyte apoptosis, senescence and promotes the progression of osteoarthritis by targeting AKT1. Am J Transl Res 11:2232–2244
Funding
This work was supported by the project of Jiangsu Health and Family Planning Commission: Role and mechanism of STAT3 regulatory T cell pathway in fracture healing (Project No.: H2018024).
Author information
Authors and Affiliations
Contributions
GS, JL and YT conceived and designed the research and interpreted the results of experiments. GS, JL, ZW, XZ and YT performed experiments, analyzed data. GS and JL wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Ethical approval
The study was approved by the Medical Ethics Committee of Jinling Hospital, Medicine College, Nanjing University.
Consent to participate
All patients and their families agreed to participate in the experiment and signed the informed consent form.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, G., Lu, J., Wu, Z. et al. STAT3 expression in patients with fragility fractures and its effect on the biological function of osteoblasts. Cell Tissue Bank 24, 515–522 (2023). https://doi.org/10.1007/s10561-022-10051-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10561-022-10051-3