Abstract
Background
Long non-coding RNAs (lncRNAs) have evolved as a critical regulatory mechanism for almost all biological processes. By dynamically interacting with their molecular partners, lncRNAs regulate gene activity at multiple levels ranging from transcription, pre-mRNA splicing, RNA transporting, RNA decay, and translation of mRNA.
Results and Conclusions
Dysregulation of lncRNAs has been associated with human diseases, including cancer, neurodegenerative, and cardiometabolic diseases. However, as lncRNAs are usually much less conserved than mRNAs at the sequence level, most human lncRNAs are either primate or human specific. The pathophysiological significance of human lncRNAs is still mostly unclear due to the persistent limitations in studying human-specific genes. This review will focus on recent discoveries showing human lncRNAs’ roles in regulating metabolic homeostasis and the potential of targeting this unique group of genes for treatment of cardiometabolic diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
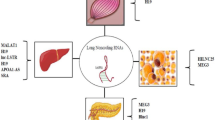
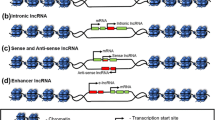
Sensing nutrients and stress to maintain metabolic homeostasis is the most fundamental event in all living organisms. Recent studies have revealed that lncRNAs are dynamically regulated by metabolic clues and constitute an essential layer of regulation contributing to metabolic homeostasis in both normal and cancer cells [1,2,3]. By interacting with transcription factors, epigenetic modifiers, RNA-binding proteins involved in pre-mRNA splicing, RNA modification and transport, and mRNA translation and decay, lncRNAs enable cells to respond to various signaling cues by regulating the expression and activities of genes in a temporal and spatial-sensitive manner (Fig. 1). As such, lncRNAs are able to control the specificity of how transcription factors and RNA-binding proteins recognize their target genes. At the organism level, studies in mouse models have revealed that lncRNAs function through these diverse molecular mechanisms to control the expression of crucial genes in major metabolic pathways, including de novo lipogenesis, fatty acid degradation, glycolysis, gluconeogenesis, lipid uptake and transport, and adipogenesis (Fig. 1). A more comprehensive introduction of regulatory lncRNAs in metabolism can be found in other excellent reviews [3,4,5,6].
Human Data and Experimental System-Driven Discovery of lncRNA Metabolic Regulators
Since mouse model-based studies have demonstrated that lncRNAs perform critical regulatory roles in major metabolic pathways, it is likely that lncRNAs do the same in humans. Then the human lncRNA metabolic regulators could serve as a novel group of therapeutic targets for cardiometabolic diseases, including obesity, diabetes, hyperlipidemia, and fatty liver diseases. However, compared with protein-coding genes, lncRNAs are in general much less conserved at the sequence level among different species [7,8,9,10], which makes applying mouse lncRNAs knowledge to human very challenging. In recent years, efforts have been initiated to define human lncRNA metabolic regulators starting from human samples followed by experimental characterization of their function in physiologically relevant experimental systems including primary human cells, transgenic rodent models, primate models, or humanized mouse models (Fig. 2). These strategies have discovered that a group of human lncRNAs, most of which are non-conserved, function in regulating major metabolic pathways (Table 1). The tight association between these human lncRNAs and disease status suggests their potential to serve as novel drug targets for obesity, diabetes, hyperlipidemia, and fatty liver diseases. A detailed discussion of representative human lncRNA metabolic regulators is included below.
cis-Regulation of APOA1 Expression by APOA1-AS
APOA1 is the major protein component of high-density lipoprotein (HDL) in plasma. Through careful scan of the human APOA1 gene locus, Halley et al. identified a long non-coding natural antisense transcript, APOA1-AS [11]. APOA1-AS shows minimal sequence conservation compared with the mouse genome, suggesting it is a non-conserved human lncRNA. Knockdown of APOA1-AS in cultured hepatocytes resulted in increased transcription of APOA1. The regulatory effect of APOA1-AS is mediated by its ability to recruit histone-modifying enzymes. More importantly, targeting APOA1-AS with short antisense oligonucleotides in African green monkeys resulted in increased hepatic APOA1 RNA and protein levels [11]. Given the critical role of APOA1 in reverse cholesterol transport, targeting APOA1-AS could be an effective therapeutic strategy for cardiovascular diseases.
Regulation of Fatty Acid Degradation by LINC01018
A recent work taking advantage of the rich resources of human genetic data, genome-wide association studies (GWAS), and expression quantitative trait loci (eQTL) defined human lncRNAs whose expression in liver tissues are associated with major metabolic traits and diseases [12]. By using a comprehensive lncRNA annotation, lncRNAKB [18], this GWAS-eQTL integration strategy [19] efficiently identified 726 human lncRNAs associated with at least one cardiometabolic trait. Conservation analysis showed that the average conservation score of these trait-associated lncRNAs is meager, suggesting that many of them are non-conserved and could be primate or human specific. Further characterization of these lncRNAs by liver tissue enrichment analysis, Hi-C and enhancer maker analysis, and lncRNA-mRNA co-expression analysis supports the functional significance of these trait-associated lncRNAs. A primate-specific and liver-enriched lncRNA, LINC01018, was chosen for experimental analyses as its expression was negatively associated with body mass index (BMI) in the human population and positively associated with mRNAs involved in the fatty acid degradation pathway. Loss-of-function experiments were performed in a humanized liver model, in which primary human hepatocytes replaced mouse hepatocytes in immune-deficient mice [20]. Knockdown of LINC01018 led to decreased expression of several genes in the fatty acid degradation pathway, supporting the role of LINC01018 in hepatic catabolism, in line with its negative association with BMI [12]. Notably, the observed regulatory effects of LINC01018 cannot be readily captured in cultured primary hepatocytes, further supporting the necessity of using the humanized mouse model for dissecting the physiological function of human-specific genes. The expression of LINC01018 is dramatically downregulated in nonalcoholic fatty liver disease (NAFLD) and nonalcoholic steatohepatitis (NASH), indicating a pathophysiological role in fatty liver diseases [12].
Transcriptional Regulation of Cholesterol Metabolism by hLMR1
By using a stepwise transcriptome analysis using RNA sequencing data from human liver tissue samples and humanized mouse livers, another group of human lncRNA metabolic regulators whose expressions are sensitive to nutrient and fatty liver disease status was defined [13]. One of these lncRNAs, hLMR1 (human lncRNA metabolic regulator 1), is exclusively expressed in human liver tissue with no mouse homolog identified. The knockdown of hLMR1 led to decreased, while a CRISPR activation [21] of hLMR1 resulted in increased expression of enzymes crucial to cholesterol biosynthesis, supporting a pivotal role of hLMR1 in hepatic cholesterol homeostasis. Remarkably, knockdown of hLMR1 in the humanized liver mouse model resulted in a more than 40% reduction of human low-density lipoprotein/very low-density lipoprotein cholesterol levels. These robust effects of hLMR1 are achieved by its interaction with an RNA-binding protein, PTBP1, which positively regulates the transcription of cholesterol biosynthesis genes [13]. Further experiments in the liver of regular mice found that ectopic expression of hLMR1 could induce the expression of mouse cholesterol biosynthesis genes. Chromatin immunoprecipitation analysis revealed that, similar to what happened in humanized livers, hLMR1 could recruit mouse Ptbp1 to the promoters of cholesterol biosynthesis genes and activate their transcription when expressed in the regular mouse liver [13]. Human genetic analyses showed that several cis eQTL of hLMR1 overlap with GWAS loci for total cholesterol levels. Indeed, this finding is further supported by a very recent large-scale GWAS defining novel loci associated with cholesterol levels in human populations [22]. Considering that both hLMR1 and LINC01018 are primate specific, liver-enriched regulators of lipid metabolism [12, 13], strategies screening for compounds modulating their expression levels could be very attractive for the treatment of fatty liver diseases and dyslipidemia in humans.
Modulating of Cholesterol Efflux by CHROME
Hennessy et al. performed gene expression analysis in the plasma and arteries of individuals with coronary artery disease and healthy control subjects to identify heart disease-associated human lncRNAs [14]. They found that a primate-specific lncRNA, which they named CHROME (Cholesterol Homeostasis Regulator of MiRNA Expression), was increased in the setting of human atherosclerotic vascular disease. Functional studies revealed that CHROME is essential to cholesterol efflux in many different cell types, including macrophages and hepatocytes [14]. Mechanistic study found that CHROME positively regulates the expression of genes involved in cholesterol efflux by suppressing microRNAs’ activity. Dietary cholesterol levels regulate the expression of CHROME through an LXR-dependent mechanism in a primate model [14]. The role of CHROME in promoting cholesterol efflux and HDL formation suggests a protective effect in human atherosclerotic vascular disease. However, as most of the functional experiments were performed in cultured cells, the significance of CHROME in regulating cholesterol homeostasis in vivo remains unclear.
Controlling of Adipogenesis by Linc-ADAL
Adipose tissues, including white adipose and thermogenic adipose tissue, are known for their roles in energy storage, secretion of adipokines, and thermogenesis [23]. To explore the role of human lncRNAs in adipose tissue, Zhang et al. performed a deep RNA sequencing analysis using RNA samples from the gluteal subcutaneous adipose tissue of 25 healthy individuals [15]. To increase the power of identification, they implemented de novo transcriptome assembly to determine the expression levels of human lncRNAs in addition to those annotated in GENCODE [24] and the Human BodyMap [25]. Tissue enrichment analysis indicates that 120 out of those human white adipose expressed lncRNAs are highly enriched in adipose tissue, and 53 of those 120 lncRNAs were modulated by bariatric surgery, suggesting their role in energy homeostasis and obesity. Sequence conservation analysis found that 85% of these adipose-enriched and bariatric surgery-regulated human lncRNAs are not conserved in mice [15]. Among all the identified non-conserved human adipose lncRNAs, Linc-ADAL was selected for further functional studies due to its abundant adipose-enriched expression, markedly induced during preadipocyte differentiation and differentially regulated expression between obese and lean humans. Loss-of-function experiments found that Linc-ADAL regulates preadipocyte differentiation and de novo lipogenesis in mature adipocytes [15]. The mechanistic study suggests that Linc-ADAL regulates the expression of genes involved in lipid metabolism at both transcriptional and post-transcriptional levels. These regulatory effects of Linc-ADAL were mediated by its interaction with hnRNPU, an RNA-binding protein previously reported as a transcriptional regulator of lipogenesis [26], and IGF2BP2, an RNA-binding protein regulating mRNA stability [27], respectively. Further studies are needed to determine the pathophysiological significance of Linc-ADAL and other non-conserved human adipose lncRNAs in obesity [15].
Regulation of Lipolysis and Mitochondrial Respiration by LINC00473
In an attempt to study the role of human lncRNAs in adipose tissue, Tran et al. differentiated human adipocytes using stromovascular cells extracted from both abdominal subcutaneous (AbdSQ) and supraclavicular (SClav) depots, representing mostly non-thermogenic and thermogenic adipose tissue depots in human [16]. RNA sequencing was performed in these two groups of adipocytes to determine the gene response to norepinephrine treatment. Among the most strongly induced genes in both groups of cells, LINC00473 was selected for further downstream analysis. The functional significance of LINC00473 was evidenced by the observation of its higher expression in SClav than in participants-matched AbdSQ adipose tissue, its strong co-expression pattern with uncoupling protein 1, and its relative downregulation in SClav samples derived from overweight and obese participants and those with type 2 diabetes mellitus [16]. Characterization of LINC00473 found that it underwent translocation from the nucleus to the cytosol upon treatment with norepinephrine. Antisense oligonucleotide-based RNA pulldown uncovered that when in the cytosol, LINC00473 interacts with lipid droplet protein PLIN1 as well as many mitochondrial proteins, indicating a role in fatty acid metabolism. Knockdown and CRISPR activation experiments confirmed that LINC00473 is critical to induce lipolysis and mitochondrial respiration upon norepinephrine treatment [16]. Overall, the identification of LINC00473 as a primate-specific lncRNA controlling mitochondrial fatty acid oxidation in thermogenic adipose tissue is exciting and with significant implications in human obesity studies. As the functional experiments were largely performed in cultured adipocytes and relied on non-physiological treatment like norepinephrine and forskolin [16], further primate or clinical studies are necessary to determine the physiological role of LINC00473 in human thermogenesis.
The Pro-Hypertensive Effect of AK098656
To identify human lncRNAs associated with hypertension, Jin et al. performed lncRNA microarray analysis using RNA extracted from the plasma of hypertensive patients and matched healthy controls [17]. This strategy led to the discovery that a non-conserved human lncRNA, AK098656, is highly expressed in the plasma of hypertensive patients, while barely detectable in healthy controls. Further expression analysis found that AK098656 was predominately expressed in human vascular smooth muscle cells (hVSMCs). Overexpression and knockdown experiments in hVSMCs demonstrated that AK098656 promotes hVSMCs synthetic phenotype by positively regulating hVSMCs proliferation and migration [17]. To determine the molecular mechanism of AK098656, chromatin isolation by RNA purification assay followed by mass spectrometry was performed. Two proteins, MYH11 (myosin heavy chain-11), a smooth muscle cell–specific contractile protein and a major pathogenic gene in thoracic aneurysm, and FN1 (fibronectin-1), a necessary factor for VSMCs growth, were found interacting with AK098656. Downstream functional analysis found that AK098656 regulates MYH11/FN1 protein degradation partly through the lysosome pathway [17]. To validate the function of AK098656 in vivo, a transgenic rat model overexpressing AK098656 was generated. Phenotype analysis revealed the development of spontaneous hypertension in AK098656 transgenic rats associated with narrowed resistance arteries [17]. The identification of a human-specific lncRNAs contributing to the development of hypertension is remarkable and could provide novel drug targets for treating hypertension in the human population. However, the lack of homology of AK098656 in non-human primate models makes it very challenging to determine the effects of blocking the expression of AK098656 in hypertension. Further studies aimed at determining the mechanisms controlling the expression of AK098656 would also help to evaluate the significance of this lncRNA in human diseases.
Mouse lncRNA Metabolic Regulators with Reported Human Homologs
Although our understanding of the evolutionary conservation of lncRNAs is largely dependent on primary sequence alignment, the concepts of genomic locus conservation, secondary structure conservation, and small functional element conservation are also emerging [28]. Table 2 summarizes major mouse lncRNA metabolic regulators with reported human homologs. Most of these reported human homologs, however, have not been tested to address whether they perform a similar function in humans as their mouse counterparts. Even the human homolog of Blnc1 was able to rescue the effects of knocking down of cognate Blnc1 in mouse adipocytes, the function of endogenous human Blnc1 was not tested [26]. The only exception to this list is MeXis. Knocking down of its human homolog in human THP-1 macrophages showed similar effects in regulating cholesterol efflux [41], although more evidence is needed to support its in vivo function. Taken together, the pathophysiological significance of these conserved mouse lncRNA metabolic regulators in humans is still awaiting further studies.
Human lncRNAs as Drug Targets for Cardiometabolic Diseases
The recent discovery that human lncRNAs regulate critical metabolic processes and are differentially expressed between healthy and disease individuals suggests that they could serve as therapeutic targets for human diseases [11,12,13,14,15,16,17]. Indeed, compared with protein-coding genes, there are unique features making human lncRNAs attractive drug targets. First, lncRNAs are more tissue specific when compared with mRNAs [42]. Systemic delivery of compounds targeting tissue-specific lncRNAs, like hLMR1, will only affect lipid synthesis in the liver tissue without disrupting the whole-body lipid synthesis. Second, due to the nature of lncRNAs, they function at the RNA level. Thus the fast-growing approaches involving oligonucleotide-based drugs, including small interfering RNA (siRNA) and antisense oligonucleotide (ASO), which induce degradation of target RNAs based on precise complementarity of sequences, are very effective for controlling the activity of lncRNAs [43]. Finally, non-conserved lncRNAs are supposed to serve as regulatory mechanisms fine-tuning the conserved core cellular machinery [44]. Their significance is usually more prominent in disease conditions, making them more effective and safer to serve as targets. Using hLMR1 as an example, Fig. 3 shows potential strategies to target human lncRNA genes in diseases. These strategies include screening for compounds modulating the expression of lncRNAs, siRNA, and ASO targeting specific lncRNAs, and small molecules regulating the interaction between lncRNAs and their protein partners to target human lncRNA genes.
Conclusion
With more and more lncRNAs functionally characterized in animal models and cultured cells, it is now known that lncRNAs are crucial regulators of gene expression and activity. The lack of sequence conservation raised the fact that most human lncRNAs are human or primate specific. Recent advances in studying non-conserved human lncRNAs have demonstrated that they are functional and differentially regulated in pathophysiological conditions in humans. These findings support the concept that evolutionary conservation should not be the universal criterion for evaluating a gene’s functional significance. Furthermore, human lncRNAs likely account for a large portion of the apparent differences between human and experimental animals like mice [45], thus creating a substantial barrier for their experimental study. Nonetheless, targeting human lncRNAs, especially these non-conserved human lncRNAs, can enable the development of novel therapeutic strategies for treating human diseases.
Future Perspectives
The major limitation of our understanding of primate-specific lncRNAs is still the lack of experimental systems [4]. Especially, this is critical for the study of energy metabolism, an in vivo physiological condition involving organ-organ crosstalk and microenvironment (Fig. 2). The humanized mouse models, which replace mouse cells, tissues, or organs with human ones, provide an excellent tool to understand human-specific regulatory effects [46, 47]. Human organoids are another strategy that allows us to study human-specific genes in a defined experimental system [48]. Of course, all human-like experimental systems have their limitations. It is thus always necessary to lead the functional studies using hypotheses generated from human data.
In addition to limitations in the experimental systems, our current identification of lncRNA metabolic regulators largely relies on determining their expression levels. It is very likely many lncRNAs may undergo regulation by translocation and post-transcriptional modifications with or without obvious changes at the expression levels. For example, the N6-methyladenosine (m6A) methylation of mRNA has emerged as a critical regulatory mechanism controlling the mRNA fate [49]. Recent studies found that many human lncRNAs harbor multiple m6A modification sites and undergo extensive methylation [50, 51]. The m6A modification and other kinds of modifications of lncRNA likely provide another mechanism to control the activity of lncRNA in addition to regulating their expression levels [52,53,54]. As such, compounds precisely modulating the modification of a lncRNA could be attractive for targeting lncRNAs. Another significant challenge is to improve our understanding of the structure-function relationship of human lncRNAs. Recent studies suggest that for some lncRNAs, only a small sequence element is sufficient to account for their function, which suggests that it is not always necessary to determine the conservation of lncRNAs by their full-length sequence alignment [55]. The concept of an RNA structurome driven by transcriptome-wide RNA structure probing coupled with next-generation RNA sequencing is also emerging and may speed up the progress in unveiling the lncRNAs biology [56]. With a more accurate understanding of the structure-function relationship of human lncRNAs, effective screening of small-molecules that modulate the specific interaction between a given lncRNA and its specific protein partners will be a powerful strategy to target lncRNAs for treating human diseases [57]. Recent evidence suggests that human lncRNAs show more variations at both the expression and sequence level among individuals [58, 59]. Each individual may have their unique group of lncRNAs, which contribute to a given individual’s disease susceptibility. Further understanding of the variations of functional lncRNAs in the human population could directly contribute to precision medicine.
Data Availability
Not applicable.
Code Availability
Not applicable.
References
Lu W, Cao F, Wang S, Sheng X, Ma J. LncRNAs: the regulator of glucose and lipid metabolism in tumor cells. Front Oncol. 2019;9:1099.
Lin YH. Crosstalk of lncRNA and cellular metabolism and their regulatory mechanism in cancer. Int J Mol Sci. 2020;21(8):2947.
Zhao XY, Lin JD. Long noncoding RNAs: a new regulatory code in metabolic control. Trends Biochem Sci. 2015;40(10):586–96.
Zhang X, Li DY, Reilly MP. Long intergenic noncoding RNAs in cardiovascular diseases: challenges and strategies for physiological studies and translation. Atherosclerosis. 2019;281:180–8.
Muret K, Désert C, Lagoutte L, Boutin M, Gondret F, Zerjal T, et al. Long noncoding RNAs in lipid metabolism: literature review and conservation analysis across species. BMC Genomics. 2019;20(1):882.
Zhao Y, Wu J, Liangpunsakul S, Wang L. Long non-coding RNA in liver metabolism and disease: current status. Liver Res. 2017;1(3):163–7.
Necsulea A, Soumillon M, Warnefors M, Liechti A, Daish T, Zeller U, et al. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature. 2014;505(7485):635–40.
Hezroni H, Koppstein D, Schwartz MG, Avrutin A, Bartel DP, Ulitsky I. Principles of long noncoding RNA evolution derived from direct comparison of transcriptomes in 17 species. Cell Rep. 2015;11(7):1110–22.
Chen J, Shishkin AA, Zhu X, Kadri S, Maza I, Guttman M, et al. Evolutionary analysis across mammals reveals distinct classes of long non-coding RNAs. Genome Biol. 2016;17:19.
Washietl S, Kellis M, Garber M. Evolutionary dynamics and tissue specificity of human long noncoding RNAs in six mammals. Genome Res. 2014;24(4):616–28.
Halley P, Kadakkuzha BM, Faghihi MA, Magistri M, Zeier Z, Khorkova O, et al. Regulation of the apolipoprotein gene cluster by a long noncoding RNA. Cell Rep. 2014;6(1):222–30.
Ruan X, Li P, Chen Y, Shi Y, Pirooznia M, Seifuddin F, et al. In vivo functional analysis of non-conserved human lncRNAs associated with cardiometabolic traits. Nat Commun. 2020;11(1):45.
Ruan X, et al. Identification of human long noncoding RNAs associated with nonalcoholic fatty liver disease and metabolic homeostasis. J Clin Invest. 2021;131(1):e136336.
Hennessy EJ, van Solingen C, Scacalossi KR, Ouimet M, Afonso MS, Prins J, et al. The long noncoding RNA CHROME regulates cholesterol homeostasis in primate. Nat Metab. 2019;1(1):98–110.
Zhang X, et al. Interrogation of nonconserved human adipose lincRNAs identifies a regulatory role of linc-ADAL in adipocyte metabolism. Sci Transl Med. 2018;10(446):eaar5987.
Tran KV, Brown EL, DeSouza T, Jespersen NZ, Nandrup-Bus C, Yang Q, et al. Human thermogenic adipocyte regulation by the long noncoding RNA LINC00473. Nat Metab. 2020;2(5):397–412.
Jin L, Lin X, Yang L, Fan X, Wang W, Li S, et al. AK098656, a novel vascular smooth muscle cell-dominant long noncoding RNA, promotes hypertension. Hypertension. 2018;71(2):262–72.
Seifuddin F, et al. lncRNAKB, a knowledgebase of tissue-specific functional annotation and trait association of long noncoding RNA. Sci Data. 2020;7(1):326.
Zhu Z, Zhang F, Hu H, Bakshi A, Robinson MR, Powell JE, et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet. 2016;48(5):481–7.
Hasegawa M, Kawai K, Mitsui T, Taniguchi K, Monnai M, Wakui M, et al. The reconstituted 'humanized liver' in TK-NOG mice is mature and functional. Biochem Biophys Res Commun. 2011;405(3):405–10.
Konermann S, Brigham MD, Trevino AE, Joung J, Abudayyeh OO, Barcena C, et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature. 2015;517(7536):583–8.
Richardson TG, Sanderson E, Palmer TM, Ala-Korpela M, Ference BA, Davey Smith G, et al. Evaluating the relationship between circulating lipoprotein lipids and apolipoproteins with risk of coronary heart disease: a multivariable Mendelian randomisation analysis. PLoS Med. 2020;17(3):e1003062.
Frayn KN, Karpe F, Fielding BA, Macdonald IA, Coppack SW. Integrative physiology of human adipose tissue. Int J Obes Relat Metab Disord. 2003;27(8):875–88.
Frankish A, Diekhans M, Ferreira AM, Johnson R, Jungreis I, Loveland J, et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 2019;47(D1):D766–73.
Cabili MN, Trapnell C, Goff L, Koziol M, Tazon-Vega B, Regev A, et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 2011;25(18):1915–27.
Mi L, Zhao XY, Li S, Yang G, Lin JD. Conserved function of the long noncoding RNA Blnc1 in brown adipocyte differentiation. Mol Metab. 2017;6(1):101–10.
Bell JL, Wächter K, Mühleck B, Pazaitis N, Köhn M, Lederer M, et al. Insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs): post-transcriptional drivers of cancer progression? Cell Mol Life Sci. 2013;70(15):2657–75.
Diederichs S. The four dimensions of noncoding RNA conservation. Trends Genet. 2014;30(4):121–3.
Zhao XY, Li S, Wang GX, Yu Q, Lin JD. A long noncoding RNA transcriptional regulatory circuit drives thermogenic adipocyte differentiation. Mol Cell. 2014;55(3):372–82.
Zhao XY, Xiong X, Liu T, Mi L, Peng X, Rui C, et al. Long noncoding RNA licensing of obesity-linked hepatic lipogenesis and NAFLD pathogenesis. Nat Commun. 2018;9(1):2986.
Li S, Mi L, Yu L, Yu Q, Liu T, Wang GX, et al. Zbtb7b engages the long noncoding RNA Blnc1 to drive brown and beige fat development and thermogenesis. Proc Natl Acad Sci U S A. 2017;114(34):E7111–20.
Zhao XY, Li S, DelProposto JL, Liu T, Mi L, Porsche C, et al. The long noncoding RNA Blnc1 orchestrates homeostatic adipose tissue remodeling to preserve metabolic health. Mol Metab. 2018;14:60–70.
Yang L, Li P, Yang W, Ruan X, Kiesewetter K, Zhu J, et al. Integrative transcriptome analyses of metabolic responses in mice define pivotal LncRNA metabolic regulators. Cell Metab. 2016;24(4):627–39.
Liu C, Yang Z, Wu J, Zhang L, Lee S, Shin DJ, et al. Long noncoding RNA H19 interacts with polypyrimidine tract-binding protein 1 to reprogram hepatic lipid homeostasis. Hepatology. 2018;67(5):1768–83.
Schmidt E, Dhaouadi I, Gaziano I, Oliverio M, Klemm P, Awazawa M, et al. LincRNA H19 protects from dietary obesity by constraining expression of monoallelic genes in brown fat. Nat Commun. 2018;9(1):3622.
Tao SC, Rui BY, Wang QY, Zhou D, Zhang Y, Guo SC. Extracellular vesicle-mimetic nanovesicles transport LncRNA-H19 as competing endogenous RNA for the treatment of diabetic wounds. Drug Deliv. 2018;25(1):241–55.
Li DY, Busch A, Jin H, Chernogubova E, Pelisek J, Karlsson J, et al. H19 induces abdominal aortic aneurysm development and progression. Circulation. 2018;138(15):1551–68.
Sallam T, Jones MC, Gilliland T, Zhang L, Wu X, Eskin A, et al. Feedback modulation of cholesterol metabolism by the lipid-responsive non-coding RNA LeXis. Nature. 2016;534(7605):124–8.
Sallam T, Jones M, Thomas BJ, Wu X, Gilliland T, Qian K, et al. Transcriptional regulation of macrophage cholesterol efflux and atherogenesis by a long noncoding RNA. Nat Med. 2018;24(3):304–12.
Ding C, Lim YC, Chia SY, Walet ACE, Xu S, Lo KA, et al. De novo reconstruction of human adipose transcriptome reveals conserved lncRNAs as regulators of brown adipogenesis. Nat Commun. 2018;9(1):1329.
Kambarami RA, Chirenje M, Rusakaniko S, Anabwani G. Perinatal mortality rates and associated socio-demographic factors in two rural districts in Zimbabwe. Cent Afr J Med. 1997;43(6):158–62.
Ruan X, et al. Identification of human long non-coding RNAs associated with nonalcoholic fatty liver disease and metabolic homeostasis. J Clin Invest. 2020.
Matsui M, Corey DR. Non-coding RNAs as drug targets. Nat Rev Drug Discov. 2017;16(3):167–79.
Ransohoff JD, Wei Y, Khavari PA. The functions and unique features of long intergenic non-coding RNA. Nat Rev Mol Cell Biol. 2018;19(3):143–57.
Esguerra JL, Eliasson L. Functional implications of long non-coding RNAs in the pancreatic islets of Langerhans. Front Genet. 2014;5:209.
Ito R, Takahashi T, Ito M. Humanized mouse models: application to human diseases. J Cell Physiol. 2018;233(5):3723–8.
Walsh NC, Kenney LL, Jangalwe S, Aryee KE, Greiner DL, Brehm MA, et al. Humanized mouse models of clinical disease. Annu Rev Pathol. 2017;12:187–215.
Kim J, Koo BK, Knoblich JA. Human organoids: model systems for human biology and medicine. Nat Rev Mol Cell Biol. 2020;21(10):571–84.
Kadumuri RV, Janga SC. Epitranscriptomic code and its alterations in human disease. Trends Mol Med. 2018;24(10):886–903.
Zhang H, Shi X, Huang T, Zhao X, Chen W, Gu N, et al. Dynamic landscape and evolution of m6A methylation in human. Nucleic Acids Res. 2020;48(11):6251–64.
Liu J, et al. Landscape and regulation of m(6)A and m(6)Am methylome across human and mouse tissues. Mol Cell. 2020;77(2):426–40 e6.
Liu N, Zhou KI, Parisien M, Dai Q, Diatchenko L, Pan T. N6-methyladenosine alters RNA structure to regulate binding of a low-complexity protein. Nucleic Acids Res. 2017;45(10):6051–63.
Wu Y, et al. m(6)A-induced lncRNA RP11 triggers the dissemination of colorectal cancer cells via upregulation of Zeb1. Mol Cancer. 2019;18(1):87.
Ni W, Yao S, Zhou Y, Liu Y, Huang P, Zhou A, et al. Long noncoding RNA GAS5 inhibits progression of colorectal cancer by interacting with and triggering YAP phosphorylation and degradation and is negatively regulated by the m(6)A reader YTHDF3. Mol Cancer. 2019;18(1):143.
Zampetaki A, Albrecht A, Steinhofel K. Long non-coding RNA structure and function: is there a link? Front Physiol. 2018;9:1201.
Kwok CK, Tang Y, Assmann SM, Bevilacqua PC. The RNA structurome: transcriptome-wide structure probing with next-generation sequencing. Trends Biochem Sci. 2015;40(4):221–32.
Donlic A, Hargrove AE. Targeting RNA in mammalian systems with small molecules. Wiley Interdiscip Rev RNA. 2018;9(4):e1477.
Kornienko AE, Dotter CP, Guenzl PM, Gisslinger H, Gisslinger B, Cleary C, et al. Long non-coding RNAs display higher natural expression variation than protein-coding genes in healthy humans. Genome Biol. 2016;17:14.
Bhartiya D, Scaria V. Genomic variations in non-coding RNAs: structure, function and regulation. Genomics. 2016;107(2-3):59–68.
Funding
This study was funded by NHLBI Division of Intramural Research funds to HC (1ZIAHL006103, 1ZIAHL006159).
Author information
Authors and Affiliations
Contributions
Xiangbo Ruan initiated the idea for the article, performed the literature search, and drafted the manuscript.
Corresponding author
Ethics declarations
Ethics Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The author declares no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article belongs to the Topical Collection: Translating genome medicine to treatments
Rights and permissions
About this article
Cite this article
Ruan, X. Targeting Human lncRNAs for Treating Cardiometabolic Diseases. Cardiovasc Drugs Ther 35, 655–662 (2021). https://doi.org/10.1007/s10557-021-07158-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10557-021-07158-2