Abstract
Purpose
In early stage, ERα-positive breast cancer, concurrent use of endocrine therapy and chemotherapy has not been shown to be superior to sequential use. We hypothesized that genetic biomarkers can aid in selecting patients who would benefit from chemo-endocrine therapy. Our previous studies revealed that ZNF423 is a transcription factor for BRCA1 and an intronic single nucleotide polymorphism (SNP) in ZNF423, rs9940645, determines tamoxifen response. Here, we identified mitosis-related genes that are regulated by ZNF423 which led us to investigate taxane response in a rs9940645 SNP- and tamoxifen-dependent fashion.
Methods
The Cancer Genome Atlas (TCGA) breast cancer dataset was used to identify genes correlated with ZNF423. Quantitative reverse transcription PCR, chromatin immunoprecipitation, and luciferase reporter assays were used to validate the gene regulation. We used CRISPR/Cas9 to engineer paired ZR-75-1 cells which differ only in ZNF423 rs9940645 SNP genotype to test SNP-dependent phenotypes including cell cycle and cell viability. We validated our findings in an additional two breast cancer cell lines, Hs578T-ERα and HCC1500.
Results
Mitosis-related genes VRK1 and PBK, which encode histone H3 kinases, were experimentally validated to be regulated by ZNF423. ZNF423 knockdown decreased VRK1 and PBK expression and activity. Additionally, ZNF423 knockdown enhanced docetaxel-induced G2/M arrest and cytotoxicity through VRK1 or PBK regulation. Lastly, cells carrying the rs9940645 variant genotype had increased G2/M arrest and decreased cell viability when treated with docetaxel in combination with estradiol and 4-OH-TAM.
Conclusions
We identified ZNF423 regulated genes involved in the G2/M phase of the cell cycle. 4-OH-TAM sensitized ERα-positive breast cancer cells to docetaxel in a ZNF423 SNP-dependent manner. Our findings suggest that patients with rs9940645 variant genotype may benefit from concurrent tamoxifen and docetaxel. This would impact a substantial proportion of patients because this SNP has a minor allele frequency of 0.47.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Breast cancer is the most common form of cancer in women both in the USA [1] and worldwide [2]. Endocrine therapy is the most important treatment modality in the majority (about 70%) of women who have estrogen receptor α (ER α)-positive breast cancer. In many women with early stage breast cancer, chemotherapy is considered appropriate based on the risk of recurrence. However, the accepted approach is to utilize the chemotherapy and endocrine therapy sequentially. This approach is based on prospective clinical trials that failed to show an advantage for combining the two modalities over their sequential use [3, 4]. It is of note that both of these trials utilized tamoxifen as the endocrine agent but neither study utilized regimens including taxanes such as paclitaxel and docetaxel, as is commonly used currently. Related to this is the report that ERα mediates paclitaxel resistance in breast cancer cells through inhibition of apoptotic cell death [5]. These findings raise the possibility that concomitant use of endocrine therapy and chemotherapy may be of value in a subset of patients.
In our previous study utilizing patients from the National Surgical Adjuvant Breast and Bowel Project (NSABP) P-1 and P-2 trials, we identified single nucleotide polymorphisms (SNPs) in ZNF423, including rs9940645, that were associated with decreased risk of breast cancer occurrence during selective estrogen receptor modulation (SERM) treatment. We also found that ZNF423 gene expression, and downstream BRCA1, is modulated by 4-hydroxytamoxifen (4-OH-TAM) in an rs9940645 SNP-dependent manner. Specifically, after adding 4-OH-TAM in the presence of 17β-estradiol (E2), ZNF423 and BRCA1 expression was increased in ZNF423 rs9940645 variant, but not wild-type, cells. Because of this striking effect on BRCA1, patients with ZNF423 rs9940645 wild-type genotype could potentially be selected for combination treatment with tamoxifen and olaparib, a PARP inhibitor which has shown significant therapeutic benefit in BRCA1/2 deficient patients on [6, 7]. Importantly, the minor allele frequency for this ZNF423 SNP (rs9940645) is reported to be 0.47 in Ensembl based on all individuals in Phase 3 of the 1000 Genomes Project [8].
In the present study, we used gene correlation analysis using The Cancer Genome Atlas (TCGA) breast cancer dataset to identify candidate genes correlated with ZNF423 and BRCA1 expression. Interestingly, most of the genes correlated with ZNF423 and BRCA1 expression in silico were related to mitosis such as VRK1 (vaccinia-related kinase 1) and PBK (PDZ binding kinase) which encode histone H3 kinases. Based on our previous studies, we hypothesized that 4-OH-TAM might regulate these mitosis-related genes in a ZNF423 rs9940645 SNP genotype-dependent manner and affect drug response to mitosis-targeting agents such as docetaxel. We found that 4-OH-TAM sensitizes cells to docetaxel treatment in ZNF423 rs9940645 variant, but not wild-type, cells. This cytotoxicity phenotype is dependent on VRK1 and PBK. Our data suggest that combined chemo-endocrine therapy may provide additional benefit to breast cancer patients with ZNF423 rs9940645 variant genotype.
Materials and methods
Cell culture
The breast cancer cell lines ZR-75-1 and HCC1500 were purchased from American Type Culture Collection (ATCC). ZR-75-1 cells are homozygous variant for rs9940645, referred to as ZR-75-1 variant. ZR-75-1 cells that are homozygous wild-type for rs9940645, referred to as ZR-75-1 WT, were generated by CRISPR/Cas9 as previously described [7]. Hs578T breast cell line with ERα overexpression, referred to as Hs578T-ERα, was a generous gift from Thomas Spelsberg, Ph.D. (Mayo Clinic, Rochester, MN, USA). ZR-75-1 WT, ZR-75-1 variant, and HCC1500 were maintained in RPMI 1640 medium (Gibco) with 10% fetal bovine serum (FBS) (Atlanta Biologicals), while Hs578T-ERα was maintained in Dulbecco’s modified Eagle medium (DMEM) (Gibco) with 10% FBS. Lymphoblastoid cell lines (Coriell Cell Repository) with known genotypes for the ZNF423 SNP were transfected with ERα and maintained as previously described [6, 7]. Only cell lines that are homozygous for the ZNF423 SNP were chosen.
Drug treatment
17β estradiol (E2), 4-hydroxytamoxifen (4-OH-TAM), and docetaxel (DCT) were purchased from Sigma-Aldrich. Docetaxel was diluted in dimethyl sulfoxide (DMSO), and E2 and 4-OH-TAM were diluted in ethanol. Prior to all drug treatments, cells were cultured in phenol-free media supplemented with 5% charcoal-stripped serum (CSS) (Thermo Fisher Scientific) for 24 h then in serum-free medium for an additional 24 h to remove all endogenous hormones. Then, cells were cultured in 1% CSS medium with 1 nM E2 for 24 h to stimulate ERα. Afterward, 1 µM 4-OH-TAM was added to the medium. Docetaxel or DMSO control was added 10 min after 4-OH-TAM.
Transient siRNA knockdown and gene overexpression
Control siRNA (D-001206-13), ZNF423 siRNA (D-012907-01, D-012907-02, D-012907-03, D-012907-04), VRK1 siRNA (M-004683-02), PBK siRNA (M-005390-00) were obtained from Dharmacon. Overexpression plasmids pCMV6-XL4 empty vector (EV) and pCMV6-XL4-ZNF423 were generated as previously as described [9]. Cells were reverse transfected in complete media using Lipofectamine RNAiMAX and Lipofectamine 2000 (Life Technologies) for knockdown and overexpression, respectively, as per manufacturer’s recommendations. For the ZNF423 knockdown screen, Hs578T overexpressing ERα cells (Hs578T-ERα cells) were used [6].
Real-time quantitative reverse transcription-PCR
Total RNA was isolated using Qiagen RNeasy kit (QIAGEN). One-step qRT-PCR was performed using 50 ng of RNA template and Power SYBR® Green RNA-to-CT™ kit (Life Technologies) and Step One Plus Real-Time PCR (Applied Biosystems). Primers are provided in Online Resource 1. All assays were performed three times in triplicate.
Luciferase reporter assay
The VRK1 promoter (1018 bp) and PBK promoter (1301 bp) were amplified with Hs578T genomic DNA using primers listed in Online Resource 1. The PCR products were subsequently cloned into pGL4.10 [luc2] plasmid using KpnI and XhoI restriction enzyme sites for VRK1 and XhoI and HindIII restriction enzyme sites for PBK. Plasmid sequences were verified by Sanger sequencing.
24 h after knockdown or overexpression of ZNF423, Hs578T-ERα cells were co-transfected with Firefly and pGL4.74[hRluc/TK] Renilla plasmids at a 10:1 ratio using Lipofectamine 2000 in the presence of 0.1 nM E2 treatment. Luciferase activity was measured with a dual-luciferase reporter assay system (Promega) 48 h after transfection. Firefly luciferase activity was then normalized to the Renilla activity. Primers are provided in Online Resource 1. Each assay was performed in three independent experiments in triplicate.
Chromatin immunoprecipitation assays
ZNF423 protein and DNA complexes were immunoprecipitated and purified using the EpiTect ChIP OneDay kit (QIAGEN). Normal IgG served as negative control. PCR was performed as previously described [6]. PCR products were then loaded on 2% agarose gels for electrophoresed for 2 h in 1 × TAE buffer. PCR primers are provided in Online Resource 1.
Flow cytometry assays
For cell cycle experiments, cells were harvested, fixed in 70% ethanol at 4 °C and stained with propidium iodide (PI) for 30 min. All flow cytometry assays were measured using Attune NxT Flow Cytometer (Thermo Fisher Scientific), and raw data were analyzed with ModFit LT 5.0.
Cell viability assays
For docetaxel assays, ZNF423, VRK1, or PBK were individually knocked down and treated with indicated doses of docetaxel in RPMI 1640 medium with 10% FBS. For rescue experiments, cells were reverse transfected with ZNF423 siRNA or control siRNA and allowed to adhere to the plate. Twelve hours later, ZNF423 knockdown cells were transiently overexpressed with the VRK1 or PBK plasmids (Origene, Rockville, MD). All cells were treated with the indicated docetaxel dose after an additional 12 h. For combination treatment assays, after cells were charcoal stripped for 24 h and serum starved for another 24 h, cells were trypsinized and neutralized with 10% CSS medium. Then, cells were centrifuged and 10,000 cells were seeded into the 96-well plates using 1% CSS medium containing 1 nM E2 for 24 h. Cells were then treated with 1 µM 4-OH-TAM 10 min prior to the indicated docetaxel doses. Cell viability was assessed 72 h after docetaxel treatment using CellTiter 96® AQueous Non-Radioactive Cell Proliferation Assays (Promega) as previously described [10]. All assays were performed in three independent experiments in triplicate.
Western blot
Protein samples were harvested, electrophoresed, and transferred to PVDF membranes using standard wet transfer techniques. Membranes were incubated with either anti-ZNF423 (ABN410, Sigma; ab169096, Abcam), VRK1 (sc-271061, Santa Cruz Biotechnology), PBK (ab75987, Abcam), or β-actin (Sigma) overnight at 4 °C. After washing and incubation of secondary antibody at room temperature for 90 min, membranes were washed and incubated with ECL. Bands were visualized using ChemiDoc Imaging Systems (BioRad).
Statistical analysis
Experimental data were analyzed with GraphPad Prism Software, and statistical methods used are indicated in the figure legends.
Results
Candidate genes selection: identifying VRK1 and PBK
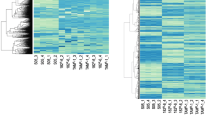
In order to identify ZNF423-regulated genes which might be modulated by 4-OH-TAM in an rs9940645 SNP-dependent manner, we employed the strategy depicted in Fig. 1. First, we performed a BLAST search for the putative ZNF423 binding motif 5′-CCGCCC-3′ [10] in the human genome. We found approximately 16,000 genes which contained one or more ZNF423 binding sequences within the promoter region. Then, we performed gene correlation analysis using The Cancer Genome Atlas (TCGA) ER+ breast cancer dataset (n = 430) to identify genes highly correlated with ZNF423. We also correlated genes with BRCA1, an estrogen-induced and direct target of ZNF423, to further narrow down the gene list. We found 368 genes were correlated with ZNF423 as well as BRCA1 expression with a p value < 10−10. Then, to determine which of these 368 genes might be transcriptionally regulated by ZNF423, we performed a ZNF423 siRNA knockdown screen in Hs578T-ERα cells and measured change in gene expression relative to siRNA control by real-time quantitative reverse transcription-PCR (qRT-PCR). There were 109 genes with at least a two-fold change in gene expression upon ZNF423 knockdown; 99 downregulated and 10 upregulated genes (Online Resource 2). We were also interested in genes that were modulated by 4-OH-TAM in a ZNF423 rs9940645 SNP-dependent manner, similar to our previous BRCA1 discovery [6]. To evaluate this possibility, we used lymphoblastoid cell lines (LCLs) overexpressing ERα with known rs9940645 genotypes with treatments that included control, E2, or E2 plus 4-OH-TAM. We used at least seven different LCLs per genotype for each treatment. We then measured mRNA expression of the 368 genes by qRT-PCR. There were 119 genes that displayed SNP-dependent expression pattern changes in the E2 plus 4-OH-TAM treatment compared to E2 alone (Online Resource 3). Eighty-two of the 119 genes had decreased expression in wild-type cells treated with 4-OH-TAM and increased expression in variant cells treated with 4-OH-TAM, which is the same direction of change that we observed previously with ZNF423 and BRCA1 [6].
Candidate gene selection strategy. The expression of 368 genes was found to be correlated with ZNF423 and BRCA1 expression with a p value < 10−10 in The Cancer Genome Atlas ER+ breast cancer dataset (n = 430). In order to narrow down the candidate gene list, two screens were performed. (Left) ZNF423 was knocked down in Hs578T-ERα cells, and qRT-PCR was performed 48 h later. Expression of the 368 genes was determined relative to β-actin and was normalized to control siRNA. There were 99 genes downregulated and 10 genes upregulated by more than two-fold after ZNF423 knockdown. (Right) Lymphoblastoid cell lines with known ZNF423 SNP genotypes were treated with control, E2, or E2 plus 4-OH-TAM, and gene expression was determined relative to β-actin and normalized to control treatment. Gene expression change was compared in E2 plus 4-OH-TAM treatment relative to E2 alone treatment. 119 genes had a SNP and drug-dependent changes in gene expression. Gene Ontology (GO) analyses were performed to narrow down these two gene lists. The top ranked GO term from each gene list was compared, and VRK1 and PBK were selected for functional validation
To narrow down the candidate gene list further, we performed Gene Ontology (GO) analyses. The gene lists from the two screens, including genes that were significantly changed upon ZNF423 knockdown and genes that showed an rs9940645 SNP and 4-OH-TAM-dependent expression profile, were separately inputted into the ClueGO plugin (version 2.3.5) of the Cytoscape software (3.2.0) using default parameters and a kappa score ≥ 0.4. GO terms were visualized as nodes and grouped based on similar function (Fig. 2a, Online Resource 4). Interestingly, many of the grouped GO terms were related to mitosis (Fig. 2b, c). GO terms were ranked by p value (Online Resource 5–6), and the genes within the most significant GO term from each screen list were compared. There were seven overlapping genes: NCAPH, NEK2, NUSAP1, OIP5, PBK, STIL, and VRK1 (Online Resource 7). We hypothesized that these 7 genes were transcriptionally regulated by ZNF423 and were modulated by 4-OH-TAM in a ZNF423 SNP-dependent manner.
Gene ontology analysis of top genes from the ZNF423 knockdown screen. GO Analysis was performed by inputting genes identified from the ZNF423 knockdown or 4-OH-TAM screen into the Cluego plugin of the Cytoscape software. Depicted is the GO analysis for the ZNF423 knockdown screen. a GO terms are visualized as nodes and grouped into a network based on function (kappa score level ≥ 0.4). The size of each node represents the enrichment significance. b, c Each functional group (GO group) is labeled with a unique color and named using the most significant GO term within the group. Some GO terms were shared by different GO groups. b The pie chart illustrates the proportion by which each GO group was represented among the GO terms. Mitotic nuclear division was the most abundant GO group. c The significance of GO groups is shown ranked by Bonferroni-corrected p value
VRK1 and PBK were chosen for futher functional validation based on literature review. VRK1 is ubiquitously expressed in human tissues and is increased in actively dividing cells. It controls cell cycle progression [11] and is a predictive marker of drug response in multiple tumor types, such as rectal adenocarcinoma [12], hepatocellular carcinoma [13, 14], and ER positive breast cancer [15]. Moreover, VRK1 was identified as a marker for poor prognosis in human breast cancer, particularly in the ER positive subgroup [16, 17]. In addition, increased PBK has been implicated in tumorigenesis [18,19,20]. Its disruption may cause tumor-specific radio-sensitization in several cancer types [21]. Importantly, VRK1 and PBK have both been identified as histone H3 kinases [22, 23]. Because VRK1 and PBK are involved in the same gene ontology of mitosis regulation and because there are agents that target mitosis, we chose these two genes for subsequent functional validation.
Histone H3 Kinases VRK1 and PBK are downstream of ZNF423
First, we evaluated VRK1 and PBK mRNA expression after ZNF423 knockdown or overexpression. We found that ZNF423 knockdown resulted in significantly decreased VRK1 and PBK mRNA expression in cells that were homozygous wild type (CRISPR-Cas9 engineered ZR-75-1 and Hs578T-ERα) and homozygous variant (ZR-75-1 and HCC1500) for the ZNF423 SNP (Fig. 3a, Online Resource 8). Similarly, ZNF423 overexpression significantly increased VRK1 and PBK mRNA expression (Online Resource 8). Luciferase reporter assays were then conducted in ERα stably transfected Hs578T cells (Hs578T-ERα) to measure the transcriptional activity of VRK1 and PBK upon ZNF423 modulation. The luciferase activity of VRK1 and PBK decreased when ZNF423 was knocked down and increased when ZNF423 was overexpressed (Fig. 3b). In order to assess whether ZNF423 can directly bind to VRK1 and PBK promoters, chromatin immunoprecipitation (ChIP) assays were performed in Hs578T-ERα using an anti-ZNF423 antibody. Our results showed that ZNF423 protein binds to the promoter region of both genes (Fig. 3c). Together, these data suggest that ZNF423 acts as a transcription factor by binding to the promoter region and regulating the transcription of histone H3 kinases VRK1 and PBK.
VRK1 and PBK are ZNF423-regulated genes. a ZNF423 was knocked down in the indicated cell lines, and VRK1 and PBK mRNA expression was determined. Gene expression was normalized to actin and is shown relative to siControl transfected cells. b The VRK1 and PBK transcriptional activities were measured by Dual-Luciferase Reporter Assay under ZNF423 knockdown (top) or overexpression (bottom) conditions in ERα stably transfected Hs578T (Hs578T-ERα) cell line which is homozygous wild-type WT for the ZNF423 SNP. The firefly luciferase activity was normalized to the corresponding Renilla activity and is shown as relative luciferase units (RLU). c Chromatin immunoprecipitation assay (ChIP) was performed in Hs578T-ERα and confirmed the binding of ZNF423 protein to the promoter regions of VRK1 (left) and PBK (right). Experiments were repeated in at least three independent experiments. Error bars represent SEM. Student t test was performed relative to control: *p < 0.05, **p < 0.01, and ***p < 0.001
VRK1, PBK, and ZNF423 regulate docetaxel response
Given the known function of VRK1 and PBK in mitosis regulation, we investigated whether ZNF423 could also affect the cell cycle. To test this possibility, we used siRNA to knockdown ZNF423, VRK1, and PBK in ZR-75-1 WT and variant cells and assessed cell cycle phases by propidium iodide staining. We did not observe any significant differences in DMSO-treated cells (data not shown). However, in docetaxel treated cells, knockdown of ZNF423, VRK1, and PBK significantly increased the proportion of cells arrested in the G2/M phase of cell cycle compared to control in both WT and variant ZNF423 genotypes (Fig. 4b). Overexpression of VRK1 and PBK in siZNF423 knockdown cells rescued this phenotype (Fig. 4b). Knockdown and overexpression efficiency was assessed by qRT-PCR (Fig. 4c).
Knockdown of ZNF423, VRK1, and PBK affect docetaxel-induced G2/M arrest. Cell cycle was assessed by propidium iodide staining followed by flow cytometry in CRISPR-Cas9 engineered ZR-75-1 WT and parental ZR-75-1 variant cell lines after the indicated knockdown and overexpression (OE) transfections under DMSO or docetaxel (DCT) treatment. a Representative peaks from an individual experiment are shown. b Quantification of cells in G2/M phase of the cell cycle using ModFit. c Knockdown and overexpression efficiency as measured by qRT-PCR normalized to actin and relative to siControl. Error bars represent SEM. Student t test was performed relative to siZNF423: ****p < 0.0001
Because there was an increase in docetaxel-induced G2/M arrest upon knockdown of ZNF423 and VRK1 and PBK, we questioned whether cytotoxic response to docetaxel was also affected. Knockdown of ZNF423, VRK1, and PBK all resulted in decreased cell viability upon docetaxel treatment in all breast cancer cell lines regardless of ZNF423 SNP genotype (Fig. 5a). Sensitivity to docetaxel was decreased in ZNF423 knockdown cells when VRK1 and PBK were overexpressed (Fig. 5b). Taken together, these results provide evidence that VRK1 and PBK are downstream of ZNF423 and that these three genes regulate mitosis (Fig. 4) and subsequent chemotherapeutic response to docetaxel (Fig. 5).
Docetaxel sensitivity is dependent on VRK1 and PBK. a Knockdown of ZNF423, VRK1, and PBK was performed in the indicated cell lines which were treated with increasing doses of docetaxel (DCT) for 72 h prior to MTS cell viability assay. b ZNF423 knockdown cells were overexpressed (OE) with VRK1 or PBK plasmids and treated with the indicated docetaxel dose. Graphs were plotted, and area under the curve was calculated using GraphPad Prism 7. Data represented as mean ± SEM. For the knockdown experiments shown in (a), AUC in each experiment was compared to siControl (open circle) by one-way ANOVA. For the rescue experiments in (b), AUC for each overexpression condition (closed circle and closed triangle) was compared to siZNF423 (closed square) by one-way ANOVA. ***p < 0.001, ****p < 0.0001
4-OH-TAM sensitizes cells to docetaxel in ZN423 rs9940645 variant genotype
Furthermore, we assessed cell cycle changes in a 4-OH-TAM and docetaxel combination treatment setting. In comparison with docetaxel and E2 alone, addition of 4-hydroxytamoxifen (4-OH-TAM) showed an increase in cells arrested in G2/M phase in cells harboring the variant ZNF423 SNP (Fig. 6a, b). This effect was observed in paired ZR-75-1 cell lines which had been CRISPR-Cas9 engineered to differ only in the rs9940645 genotype.
Combination of 4-OH-TAM and docetaxel treatment increases G2/M arrest in ZNF423 variant breast cancer cells. ZR-75-1 WT and ZR-75-1 variant cells were treated as indicated and subjected to cell cycle analysis by propidium iodide staining. a Representative peaks are shown, and b quantification of cells in G2/M is shown. Error bars represent SEM. Student t test was performed relative to E2 plus docetaxel (DCT) treatment: ***p < 0.001
We then tested whether there was a SNP-dependent cytotoxic effect after combination 4-OH-TAM and docetaxel treatment using a cell viability assay and western blot. ZR-75-1 variant cells were more resistant than ZR-75-1 WT cells when treated with docetaxel in the presence of E2 (Fig. 7a, open circles). However, this resistant phenotype in variant cells can be abolished with the addition of 4-OH-TAM. The same SNP-dependent phenotypes were observed in Hs578T-ERα WT and HCC1500 variant breast cancer cells. At the protein level, ZNF423 and its downstream targets VRK1 and PBK are decreased in the combination 4-OH-TAM and docetaxel setting only in variant cells (Fig. 7b). This is consistent with our previous findings that knockdown of ZNF423, VRK1, or PBK resulted in enhanced docetaxel sensitivity when treated in combination 4-OH-TAM (Fig. 5a).
Tamoxifen sensitizes ZNF423 variant breast cancer cells to docetaxel treatment. ZR-75-1 WT, ZR-75-1 variant, Hs578T-ERα WT and HCC1500 variant cells treated with docetaxel at the indicated concentrations alone or in combination with 4-OH-TAM. a Sensitivity to docetaxel was assessed by MTS cell viability assay 72 h after treatment. b Western blot was performed after 24 h of combination docetaxel (DCT) and 4-OH-TAM treatment. Experiments were performed in three independent experiments in triplicate. Graphs were plotted and areas under the curve were calculated using GraphPad Prism 7. Data represented as mean ± SEM. Area under the curve was compared to siControl by Student t test. ***p < 0.001, ****p < 0.0001
Discussion
ERα-positive breast cancer is a heterogeneous disease. Using intrinsic subtype classification, ERα-positive tumors can be divided into Luminal A and Luminal B. Compared with luminal A breast cancer, luminal B breast cancer patients have a worse prognosis and typically have a lower response to endocrine therapy or chemotherapy [24] with a large number recurring even after sequential chemo-endocrine adjuvant therapy. Therefore, there is a growing need to develop biomarkers and additional therapeutic strategies to help improve treatment response for these patients [25].
One proposed mechanism of chemoresistance in ER+ breast tumors is activation of ERα-driven cancer survival pathways [26]. It has been theorized that treating with ER blocking agents, such as tamoxifen, concurrently with chemotherapy would help circumvent the ERα-induced chemoresistance mechanism. However, prospective clinical trials failed to show an advantage for combining the two modalities over their sequential use [3, 4]. The results from our current study suggest that a single nucleotide polymorphism, (rs9940645) which is located in ZNF423 intronic region, may help predict which patients would benefit from combined endocrine and chemotherapy.
In our current study, the transcription factor ZNF423 was found to play an important role in regulating the G2/M phase of the cell cycle. This is in accordance with recent reports that ZNF423 regulates cell cycle progression and cell division in Purkinje neuron progenitors [27]. We found that ZNF423 exerts its cell cycle effects partially through the histone H3 kinases VRK1 and PBK. We also observed that VRK1 and PBK expression is directly regulated by ZNF423. Most intriguingly, ZNF423 ChIP-seq (GEO:GSE91794) from the ENCODE project showed binding peaks in the PBK promoter region in HEK293 cells, which is consistent with what we found in breast cancer cells. Although ZNF423 did not showing significant binding peaks in the VRK1 gene in the same dataset, this may be due to cell line-specific differences.
Previous studies have revealed that suppressing the expression of histone H3 kinases, including VRK1 and PBK, causes perturbation of the G2/M phase leading to cell death [22, 23]. It has also been reported that combining histone H3 kinase inhibitors and paclitaxel is synergistic [28, 29]. Similarly, in this study, when VRK1 and PBK were downregulated, it resulted in increased sensitivity to docetaxel. However, this effect was only observed in cell lines harboring the variant ZNF423 rs9940645 SNP genotype when treated in combination with 4-OH-TAM. The underlying mechanism for this SNP-dependent phenotype is mediated through a sensor protein called calmodulin-like protein 3 (CALML3). CALML3 binds to the rs9940645 SNP in the ZNF423 promoter, forms a protein complex with ERα, and cooperates with ERα in estrogen response element (ERE) binding [7].
Whereas our study suggests that concomitant docetaxel and tamoxifen may provide an increase in efficacy in a genotype-determined subset of patients, we have no data on the impact of this combination on toxicity. Multiple attempts were made to integrate tamoxifen into chemotherapy regimens several decades ago [30], however, none of these regimens involved taxanes. Our own experience in adding tamoxifen to a non-taxane chemotherapy regimen in postmenopausal [31] and premenopausal women [32] did not identify any major toxicity signals. However, monitoring for toxicity in clinical trials of a combination of docetaxel plus tamoxifen would be necessary.
There are some limitations in our study. First, the genes selected for the two screens in this study were chosen using an arbitrarily drawn p value of less than 10−10 s, and these results were obtained using cell line models. It is important to validate these findings further using animal models before proceeding in a clinical setting.
Conclusions
In summary, our findings suggest that high risk ERα-positive breast cancer patients with the variant ZNF423 rs9940645 SNP genotype may benefit from the concomitant administration of endocrine therapy (tamoxifen) and chemotherapy (docetaxel). Because the minor allele frequency of rs9940645 is 0.47 [8], there could be a large number of patients who might benefit from this combination treatment.
Data availability
All experimental datasets generated are available in the supplementary materials submitted with this manuscript. Any additional information is available upon reasonable request to the corresponding author.
Abbreviations
- 4-OH-TAM:
-
4-Hydroxytamoxifene
- CALML3:
-
Calmodulin-like protein 3
- CRISPR:
-
Clustered, regularly interspaced short palindromic repeats
- DCT:
-
Docetaxel
- E2:
-
17β-Estradiol
- ER:
-
Estrogen receptor
- ERE:
-
Estrogen response element
- GO:
-
Gene ontology
- GWAS:
-
Genome-wide association study
- LCL:
-
Lymphoblastoid cell line
- NSABP:
-
National surgical adjuvant breast and bowel project
- OE:
-
Overexpress
- PARP:
-
Poly(ADP-ribose) polymerase
- PBK:
-
PDZ binding kinas
- qRT-PCR:
-
Quantitative reverse transcription polymerase chain reaction
- SERM:
-
Selective estrogen receptor modulator
- SNP:
-
Single nucleotide polymorphism
- TCGA:
-
The Cancer Genome Atlas
- WT:
-
Wild type
- VRK1:
-
Vaccinia-related kinase 1
- ZNF423:
-
Zinc finger protein 423
References
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA: a cancer journal for clinicians 68 (1):7–30. https://doi.org/10.3322/caac.21442
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, Parkin DM, Forman D, Bray F (2015) Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer 136(5):E359–E386. https://doi.org/10.1002/ijc.29210 doi
Albain KS, Barlow WE, Ravdin PM, Farrar WB, Burton GV, Ketchel SJ, Cobau CD, Levine EG, Ingle JN, Pritchard KI, Lichter AS, Schneider DJ, Abeloff MD, Henderson IC, Muss HB, Green SJ, Lew D, Livingston RB, Martino S, Osborne CK (2009) Adjuvant chemotherapy and timing of tamoxifen in postmenopausal patients with endocrine-responsive, node-positive breast cancer: a phase 3, open-label, randomised controlled trial. Lancet 374(9707):2055–2063. https://doi.org/10.1016/s0140-6736(09)61523-3
Bedognetti D, Sertoli MR, Pronzato P, Del Mastro L, Venturini M, Taveggia P, Zanardi E, Siffredi G, Pastorino S, Queirolo P, Gardin G, Wang E, Monzeglio C, Boccardo F, Bruzzi P (2011) Concurrent vs sequential adjuvant chemotherapy and hormone therapy in breast cancer: a multicenter randomized phase III trial. Journal of the National Cancer Institute 103(20):1529–1539. https://doi.org/10.1093/jnci/djr351
Sui M, Huang Y, Park BH, Davidson NE, Fan W (2007) Estrogen receptor alpha mediates breast cancer cell resistance to paclitaxel through inhibition of apoptotic cell death. Cancer Res 67(11):5337–5344. https://doi.org/10.1158/0008-5472.can-06-4582
Ingle JN, Liu M, Wickerham DL, Schaid DJ, Wang L, Mushiroda T, Kubo M, Costantino JP, Vogel VG, Paik S, Goetz MP, Ames MM, Jenkins GD, Batzler A, Carlson EE, Flockhart DA, Wolmark N, Nakamura Y, Weinshilboum RM (2013) Selective estrogen receptor modulators and pharmacogenomic variation in ZNF423 regulation of BRCA1 expression: individualized breast cancer prevention. Cancer Discov 3(7):812–825. https://doi.org/10.1158/2159-8290.CD-13-0038
Qin S, Ingle JN, Liu M, Yu J, Wickerham DL, Kubo M, Weinshilboum RM, Wang L (2017) Calmodulin-like protein 3 is an estrogen receptor alpha coregulator for gene expression and drug response in a SNP, estrogen, and SERM-dependent fashion. Breast Cancer Res 19(1):95. https://doi.org/10.1186/s13058-017-0890-x
Zerbino DR, Achuthan P, Akanni W, Amode MR, Barrell D, Bhai J, Billis K, Cummins C, Gall A, Girón CG, Gil L, Gordon L, Haggerty L, Haskell E, Hourlier T, Izuogu OG, Janacek SH, Juettemann T, To JK, Laird MR, Lavidas I, Liu Z, Loveland JE, Maurel T, McLaren W, Moore B, Mudge J, Murphy DN, Newman V, Nuhn M, Ogeh D, Ong CK, Parker A, Patricio M, Riat HS, Schuilenburg H, Sheppard D, Sparrow H, Taylor K, Thormann A, Vullo A, Walts B, Zadissa A, Frankish A, Hunt SE, Kostadima M, Langridge N, Martin FJ, Muffato M, Perry E, Ruffier M, Staines DM, Trevanion SJ, Aken BL, Cunningham F, Yates A, Flicek P (2018) Ensembl 2018. Nucl Acids Res 46(D1):D754–D761. https://doi.org/10.1093/nar/gkx1098
Liu D, Ho MF, Schaid DJ, Scherer SE, Kalari K, Liu M, Biernacka J, Yee V, Evans J, Carlson E, Goetz MP, Kubo M, Wickerham DL, Wang L, Ingle JN, Weinshilboum RM (2017) Breast cancer chemoprevention pharmacogenomics: deep sequencing and functional genomics of the ZNF423 and CTSO genes. NPJ Breast Cancer 3:30. https://doi.org/10.1038/s41523-017-0036-4
Niu N, Qin Y, Fridley BL, Hou J, Kalari KR, Zhu M, Wu TY, Jenkins GD, Batzler A, Wang L (2010) Radiation pharmacogenomics: a genome-wide association approach to identify radiation response biomarkers using human lymphoblastoid cell lines. Genome Res 20(11):1482–1492. https://doi.org/10.1101/gr.107672.110
Valbuena A, Sanz-García M, López-Sánchez I, Vega FM, Lazo PA (2011) Roles of VRK1 as a new player in the control of biological processes required for cell division. Cell Signal 23(8):1267–1272. https://doi.org/10.1016/j.cellsig.2011.04.002
del Puerto-Nevado L, Marin-Arango JP, Fernandez-Aceñero MJ, Arroyo-Manzano D, Martinez-Useros J, Borrero-Palacios A, Rodriguez-Remirez M, Cebrian A, Gomez del Pulgar T, Cruz-Ramos M, Carames C, Lopez-Botet B, Garcia-Foncillas J (2016) Predictive value of vrk 1 and 2 for rectal adenocarcinoma response to neoadjuvant chemoradiation therapy: a retrospective observational cohort study. BMC Cancer 16:519. https://doi.org/10.1186/s12885-016-2574-9
Huang W, Cui X, Chen Y, Shao M, Shao X, Shen Y, Liu Q, Wu M, Liu J, Ni W, Lu C, Wan C (2016) High VRK1 expression contributes to cell proliferation and survival in hepatocellular carcinoma. Pathol Res Pract 212(3):171–178. https://doi.org/10.1016/j.prp.2015.11.015
Lee N, Kwon JH, Kim YB, Kim SH, Park SJ, Xu W, Jung HY, Kim KT, Wang HJ, Choi KY (2015) Vaccinia-related kinase 1 promotes hepatocellular carcinoma by controlling the levels of cell cycle regulators associated with G1/S transition. Oncotarget 6(30):30130–30148. https://doi.org/10.18632/oncotarget.4967
Salzano M, Vazquez-Cedeira M, Sanz-Garcia M, Valbuena A, Blanco S, Fernandez IF, Lazo PA (2014) Vaccinia-related kinase 1 (VRK1) confers resistance to DNA-damaging agents in human breast cancer by affecting DNA damage response. Oncotarget 5(7):1770–1778. https://doi.org/10.18632/oncotarget.1678
Fournier MV, Martin KJ, Kenny PA, Xhaja K, Bosch I, Yaswen P, Bissell MJ (2006) Gene expression signature in organized and growth-arrested mammary acini predicts good outcome in breast cancer. Cancer Res 66(14):7095–7102. https://doi.org/10.1158/0008-5472.can-06-0515
Martin KJ, Patrick DR, Bissell MJ, Fournier MV (2008) Prognostic breast cancer signature identified from 3D culture model accurately predicts clinical outcome across independent datasets. PLoS One 3(8):e2994. https://doi.org/10.1371/journal.pone.0002994
Zykova TA, Zhu F, Wang L, Li H, Bai R, Lim DY, Yao K, Bode AM, Dong Z (2017) The T-LAK cell-originated protein kinase signal pathway promotes colorectal cancer metastasis. EBioMedicine 18:73–82. https://doi.org/10.1016/j.ebiom.2017.04.003
Ohashi T, Komatsu S, Ichikawa D, Miyamae M, Okajima W, Imamura T, Kiuchi J, Kosuga T, Konishi H, Shiozaki A, Fujiwara H, Okamoto K, Tsuda H, Otsuji E (2017) Overexpression of PBK/TOPK relates to tumour malignant potential and poor outcome of gastric carcinoma. Br J Cancer 116(2):218–226. https://doi.org/10.1038/bjc.2016.394
Ohashi T, Komatsu S, Ichikawa D, Miyamae M, Okajima W, Imamura T, Kiuchi J, Nishibeppu K, Kosuga T, Konishi H, Shiozaki A, Fujiwara H, Okamoto K, Tsuda H, Otsuji E (2016) Overexpression of PBK/TOPK contributes to tumor development and poor outcome of esophageal squamous cell carcinoma. Anticancer Res 36(12):6457–6466. https://doi.org/10.21873/anticanres.11244
Pirovano G, Ashton TM, Herbert KJ, Bryant RJ, Verrill CL, Cerundolo L, Buffa FM, Prevo R, Harrap I, Ryan AJ, Macaulay V, McKenna WG, Higgins GS (2017) TOPK modulates tumour-specific radiosensitivity and correlates with recurrence after prostate radiotherapy. Br J Cancer 117(4):503–512. https://doi.org/10.1038/bjc.2017.197
Kang TH, Park DY, Choi YH, Kim KJ, Yoon HS, Kim KT (2007) Mitotic histone H3 phosphorylation by vaccinia-related kinase 1 in mammalian cells. Mol Cell Biol 27(24):8533–8546. https://doi.org/10.1128/MCB.00018-07
Park JH, Lin ML, Nishidate T, Nakamura Y, Katagiri T (2006) PDZ-binding kinase/T-LAK cell-originated protein kinase, a putative cancer/testis antigen with an oncogenic activity in breast cancer. Cancer Res 66(18):9186–9195. https://doi.org/10.1158/0008-5472.CAN-06-1601
Nguyen PL, Taghian AG, Katz MS, Niemierko A, Abi Raad RF, Boon WL, Bellon JR, Wong JS, Smith BL, Harris JR (2008) Breast cancer subtype approximated by estrogen receptor, progesterone receptor, and HER-2 is associated with local and distant recurrence after breast-conserving therapy. J Clin Oncol 26(14):2373–2378. https://doi.org/10.1200/JCO.2007.14.4287
Gyorffy B, Hatzis C, Sanft T, Hofstatter E, Aktas B, Pusztai L (2015) Multigene prognostic tests in breast cancer: past, present, future. Breast Cancer Res 17:11. https://doi.org/10.1186/s13058-015-0514-2
Ades F, Zardavas D, Bozovic-Spasojevic I, Pugliano L, Fumagalli D, de Azambuja E, Viale G, Sotiriou C, Piccart M (2014) Luminal B breast cancer: molecular characterization, clinical management, and future perspectives. J Clin Oncol 32(25):2794–2803. https://doi.org/10.1200/JCO.2013.54.1870
Casoni F, Croci L, Bosone C, D’Ambrosio R, Badaloni A, Gaudesi D, Barili V, Sarna JR, Tessarollo L, Cremona O, Hawkes R, Warming S, Consalez GG (2017) Zfp423/ZNF423 regulates cell cycle progression, the mode of cell division and the DNA-damage response in Purkinje neuron progenitors. Development 144(20):3686–3697. https://doi.org/10.1242/dev.155077
Scharer CD, Laycock N, Osunkoya AO, Logani S, McDonald JF, Benigno BB, Moreno CS (2008) Aurora kinase inhibitors synergize with paclitaxel to induce apoptosis in ovarian cancer cells. J Transl Med 6:79. https://doi.org/10.1186/1479-5876-6-79
Payton M, Bush TL, Chung G, Ziegler B, Eden P, McElroy P, Ross S, Cee VJ, Deak HL, Hodous BL, Nguyen HN, Olivieri PR, Romero K, Schenkel LB, Bak A, Stanton M, Dussault I, Patel VF, Geuns-Meyer S, Radinsky R, Kendall RL (2010) Preclinical evaluation of AMG 900, a novel potent and highly selective pan-aurora kinase inhibitor with activity in taxane-resistant tumor cell lines. Cancer Res 70(23):9846–9854. https://doi.org/10.1158/0008-5472.CAN-10-3001
Ingle JN (1984) Integration of hormonal agents and chemotherapy for the treatment of women with advanced breast cancer. Mayo Clin Proc 59(4):232–238
Ingle JN, Everson LK, Wieand HS, Martin JK, Votava HJ, Wold LE, Krook JE, Cullinan SA, Paulsen JK, Twito DI et al (1988) Randomized trial of observation versus adjuvant therapy with cyclophosphamide, fluorouracil, prednisone with or without tamoxifen following mastectomy in postmenopausal women with node-positive breast cancer. J Clin Oncol 6(9):1388–1396. https://doi.org/10.1200/jco.1988.6.9.1388
Ingle JN, Everson LK, Wieand HS, Cullinan SA, Wold LE, Hagen JB, Martin JK, Krook JE, Fitzgibbons RG, Foley JF et al (1989) Randomized trial to evaluate the addition of tamoxifen to cyclophosphamide, 5-fluorouracil, prednisone adjuvant therapy in premenopausal women with node-positive breast cancer. Cancer 63(7):1257–1264
Acknowledgements
We would like to acknowledge Thomas Spelsberg, Ph.D. for providing the Hs578T-ERα cell line.
Funding
This work was supported by The Breast Cancer Research Foundation (BCRF-18-076) and Eisenberg Foundation. GW was supported by ChuYing Charity Foundation. JZ was supported by the Mayo Clinic Medical Scientist Training Program (T32 GM065841) and Initiative for Maximizing Student Development (R25 GM055252).
Author information
Authors and Affiliations
Contributions
GW, SQ, and JZ participated in data acquisition, data analysis, and manuscript writing. GW designed the study and drafted the manuscript. SQ generated the ZR75-1 CRISPR Cas9 genome edited cell line. ML carried out the ZNF423 and 4-OH-TAM screens. JNI provided invaluable clinical expertise and assisted in manuscript preparation. RMW conceived the study, participated in the study design, and provided guidance in data interpretation. KS provided clinical advice and helped apply for the ChuYing Charity Foundation support. LW conceived the study, participated in the study design, coordinated the study, and is responsible for all data as described. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
ML is currently affiliated with AbbVie and, however, was not at the time of involvement in this study. LW and RMW are co-founders and stockholders in OneOme, LLC, a pharmacogenomic decision support company.
Ethical approval
All experiments performed in this publication comply with US laws. There were no human participants or animals used in this study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10549_2019_5194_MOESM1_ESM.xlsx
Supplementary material 1. Online Resource 1: Table S1 Primers sequences for luciferase and ChIP assay are provided 5’ to 3’. Catalog numbers for predesigned qRT-PCR primers are also listed. (XLSX 9 KB)
10549_2019_5194_MOESM2_ESM.xlsx
Supplementary material 2. Online Resource 2: Table S2 Hs578T-ERα cells were transiently transfected with siRNA targeting each of the 368 candidate genes and an siRNA control (siControl). Gene expression was measured by by real-time quantitative reverse transcription-PCR (qRT-PCR) and normalized to housekeeping genes. There were 109 genes with at least a twofold change in gene expression upon ZNF423 knockdown; 99 downregulated and 10 upregulated genes. (XLSX 12 KB)
10549_2019_5194_MOESM3_ESM.xlsx
Supplementary material 3. Online Resource 3: Table S3 Lymphoblastoid cell lines (LCLs) overexpressing ERα with known rs9940645 genotypes were treated with vehicle control, E2, or E2 plus 4-OH-TAM. We used at least 7 different LCLs per genotype for each treatment. Gene expression of the 368 genes was measured by quantitative reverse transcription-PCR (qRT-PCR). There were 119 genes that displayed SNP-dependent expression pattern changes in the E2 plus 4-OH-TAM treatment compared to E2 alone. Eighty-two of the 119 genes had decreased expression in wild-type cells treated with 4-OH-TAM and increased expression in variant cells treated with 4-OH-TAM, which is the same direction of change that we observed previously with ZNF423 and BRCA1. (XLSX 20 KB)
10549_2019_5194_MOESM4_ESM.pptx
Supplementary material 4. Online Resource 4: Figure S1 GO Analysis was performed by inputting genes identified from the ZNF423 knockdown or 4-OH-TAM screen into the Cluego plugin of the Cytoscape software. Depicted is the GO analysis for the 4-OH-TAM screen. (A) GO terms are visualized as nodes and grouped into a network based on function (kappa score level ≥ 0.4). The size of each node represents the enrichment significance. (B-C) Each functional group (GO group) is labeled with a unique color and named using the most significant GO term within the group. Some GO terms were shared by different GO groups. (B) The pie chart illustrates the proportion by which each GO group was represented among the GO terms. Mitotic nuclear division was the most abundant GO group. (C) The significance of GO groups is shown ranked by Bonferroni-corrected p value. (PPTX 981 KB)
10549_2019_5194_MOESM5_ESM.xlsx
Supplementary material 5. Online Resource 5: Table S4 Results from gene ontology analysis of the 109 genes which had at least a twofold change in gene expression upon ZNF423 knockdown. (XLSX 20 KB)
10549_2019_5194_MOESM6_ESM.xls
Supplementary material 6. Online Resource 6: Table S5 Results from gene ontology analysis of the 119 genes that displayed SNP-dependent expression pattern changes in the E2 plus 4-OH-TAM treatment compared to E2 alone. (XLS 64 KB)
10549_2019_5194_MOESM7_ESM.xls
Supplementary material 7. Online Resource 7: Table S6 Comparison of genes within the most significant GO term from each screen. There were 7 overlapping genes from top GO terms, including VRK1 and PBK. (XLS 28 KB)
10549_2019_5194_MOESM8_ESM.pptx
Supplementary material 8. Online Resource 8: Figure S2 ZR-75-1 cells with WT or variant SNP genotype were transiently (A) knocked down with the indicated siRNA or (B) overexpressed with the indicated plasmids. Then, qRT-PCR was performed to determine the gene expression of ZNF423, VRK1, and PBK relative to control. Data shown as mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001. (PPTX 359 KB)
Rights and permissions
About this article
Cite this article
Wang, G., Qin, S., Zayas, J. et al. 4-Hydroxytamoxifen enhances sensitivity of estrogen receptor α-positive breast cancer to docetaxel in an estrogen and ZNF423 SNP-dependent fashion. Breast Cancer Res Treat 175, 567–578 (2019). https://doi.org/10.1007/s10549-019-05194-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-019-05194-z