Abstract
The mechanisms by which breast cancer (BrC) can successfully metastasize are complex and not yet fully understood. Our goal was to identify tumor-induced stromal changes that influence metastatic cell behavior, and may serve as better targets for therapy. To identify stromal changes in cancer-bearing tissue, dual-species gene expression analysis was performed for three different metastatic BrC xenograft models. Results were confirmed by immunohistochemistry, flow cytometry, and protein knockdown. These results were validated in human clinical samples at the mRNA and protein level by retrospective analysis of cohorts of human BrC specimens. In pre-clinical models of BrC, systemic recruitment of S100A8+ myeloid cells—including myeloid-derived suppressor cells (MDSCs)—was promoted by tumor-derived factors. Recruitment of S100A8+ myeloid cells was diminished by inhibition of tumor-derived factors or depletion of MDSCs, resulting in fewer metastases and smaller primary tumors. Importantly, these MDSCs retain their ability to suppress T cell proliferation upon co-culture. Secretion of macrophage inhibitory factor (MIF) activated the recruitment of S100A8+ myeloid cells systemically. Inhibition of MIF, or depletion of MDSCs resulted in delayed tumor growth and lower metastatic burden. In human BrC specimens, increased mRNA and protein levels of S100A8+ infiltrating cells are highly associated with poor overall survival and shorter metastasis free survival of BrC patients, respectively. Furthermore, analysis of nine different human gene expression datasets confirms the association of increased levels of S100A8 transcripts with an increased risk of death. Recruitment of S100A8+ myeloid cells to primary tumors and secondary sites in xenograft models of BrC enhances cancer progression independent of their suppressive activity on T cells. In clinical samples, infiltrating S100A8+ cells are associated with poor overall survival. Targeting these molecules or associated pathways in cells of the tumor microenvironment may translate into novel therapeutic interventions and benefit patient outcome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Although localized breast cancer (BrC) is curable, metastatic disease is overwhelmingly lethal. While a great part of BrC research has focused on the genetic and epigenetic changes that occur in cancer cells themselves [1], the stromal changes which are either induced by BrC cells or are intrinsic to some patients—which may increase the likelihood of malignant spread—are also critical factors being studied for their role in the metastatic process and clinical outcome [2]. It is, therefore, important to identify the causal factors regulating metastasis to reduce cancer mortality. This will ultimately allow the development of new therapies to effectively inhibit metastasis and reduce mortality.
In this study, we compared the differential gene expression of cancer-bearing tissue to that of the corresponding non-tumor-bearing tissue in NOD scid gamma (NSG) null mice—including the mammary fat pad, lymph node, and lung. We had recently shown that orthotopic injection of human BrC cells (both from established cell lines and primary cultures) in the mammary fat pad of NSG mice consistently leads to metastatic spread to distant organs [3]. This mouse model has also been used in studies of neuroblastoma [4] and melanoma [5], where metastasis of human melanoma cells in NSG mice correlates with clinical outcome. We have previously used this model to develop a dual-species microarray analysis method to be able to separately and unambiguously quantify gene expression changes that occur in the host (mouse) tissues as compared with those occurring in the BrC cells (human) [6]. Using this approach we identified a group of genes consistently upregulated in cancer-bearing tissues—both at the primary and secondary sites—that is consistent with an infiltration of myeloid cells.
Bone marrow-derived myeloid immune cells infiltrate malignant tumor sites in large numbers, and are often a prominent feature in the stroma surrounding tumors [7]. Specific attention has been drawn to myeloid-derived suppressor cells (MDSCs)—a heterogeneous mixture of immature myeloid cells playing a role in inflammation, immunosuppression, and metastatic progression [8]. MDSCs accumulate during several pathological conditions such as inflammation, parasitic infections, trauma, sepsis, and cancer [9, 10]. In BrC, the presence of these cells in primary tumors and in the blood stream of patients is associated with poor prognosis [11]. The phenotype of MDSCs in mice is defined by the simultaneous expression of surface markers CD11b+ and Gr1+. Recently, two subpopulations have been further defined based on the expression of the Gr-1 epitopes Ly6C and Ly6G: monocytic MDSCs (M-MDSCs: CD11b+Ly6CHILy6G−) and granulocytic MDSCs (G-MDSCs: CD11b+Ly6G+Ly6C−/low). In addition to the difference in surface marker phenotype, these cells differ in the morphology and the mechanism they utilize to suppress immune functions which can include upregulation of iNOS, arginase and immunosuppressive cytokines; the use of reactive oxygen species [9] or depletion of cystine and cysteine—essential for T cell activation and proliferation—from the microenvironment [12]. Expansion of both M-MDSC and granulocytic MDSC (G-MDSCs) populations has been consistently documented in mouse models in many types of cancer [9].
Expansion, activation, and recruitment of MDSCs can be induced by tumor-derived factors including GM-CSF, VEGF, IL-6 [13], prostaglandin E2 [14], and IL-1 beta [15], which are also elevated in chronic inflammation. Other pro-inflammatory molecules such as members of the S100 family of proteins S100A8 and S100A9 have also been shown to be increased in diseases such as rheumatoid arthritis [16], sepsis [17], and cancers including gastric [18], colon [19], breast [20], and ovarian [21] carcinomas. Initially discovered in cells of myeloid origin, S100A8 and S100A9 are preferentially found as heterodimers, and have been shown to be involved in a variety of intra- and extracellular functions. Importantly, the time course of expression of S100A8 and S100A9 suggests that these proteins are expressed at early stages of monocyte/macrophage differentiation, and their expression is neither detected in resident macrophages nor under acute inflammatory conditions; however, the presence of S100A8+ macrophages is indicative of chronic inflammation [22]. S100A8/S100A9 have been shown to initiate signaling via Toll-like receptor 4 and the receptor for advanced glycation end products (RAGE) [23, 24], both of which are expressed by MDSCs and by BrC cells [23, 25].
In addition to their role as potent immunosuppressors, MDSCs have an important role in driving crucial metastatic processes such as invasion, migration, and angiogenesis by secreting pro-angiogenic factors such as VEGF and MMPs [7, 26] and establishing a pre-metastatic niche [27]. Thus, the full characterization of the phenotype andthe function of MDSCs, and the means by which they facilitate metastasis aside from their immunosuppressive function, may unravel novel biomarkers and therapeutic alternatives that may be translated into the clinical setting.
The genetic instability of cancer cells results in inter- and intratumoral heterogeneity that makes the successful therapeutic targeting of carcinoma specific changes very difficult [28]. Thus, we are interested in studying tumor-induced stromal changes, as the stroma may be more genetically stable providing better targets for therapy. Here we describe the characterization of a myeloid immune cell infiltrate in primary tumors and secondary sites, which enhances BrC progression in a xenograft model of human BrC metastasis. Our studies show that human BrC cells xenografted in NSG mice recruit myeloid cells—including MDSCs—that express S100A8 and are preferentially recruited by metastatic tumors. Moreover, even in the absence of T cells, S100A8+ MDSCs promote BrC primary tumor growth and metastatic spread. Although NSG mice lack T, B, and NK cells; MDSCs isolated from tumors and spleens of tumor-bearing NSG mice retain their suppressive function as confirmed by their ability to suppress T cell proliferation. Additionally, in vivo targeting of these cells (Gr-1 antibody) or of their recruitment—via interruption of tumor-derived cytokine secretion—results in decreased levels of S100A8+ infiltrating cells, delayed tumor growth, and lower metastatic burden.
Importantly, our findings are of clinical relevance, as elevated levels of infiltrating S100A8+ cells are associated with significantly shorter metastasis free survival. Furthermore, analysis of nine independent human gene expression datasets confirms the association of increased levels of S100A8 and S100A9 transcripts with an increased hazard ratio (HR) of death. Thus, targeting of these molecules or associated pathways may ultimately translate into clinically exploitable, novel therapeutic interventions by which targeting cells of the tumor microenvironment, in this case myeloid cells, may greatly benefit patient outcome.
Results
Genomic alterations in host cells of cancer-bearing tissue are consistent with an infiltration of myeloid cells
We performed dual-species gene expression analysis as described previously [6] to analyze the stromal (mouse) gene expression profiles of primary tumors and secondary lesions from MDA-MB-231, MDA-MB-436, and DT28 [29] tumor-bearing NSG mice, and compared them to that of non-tumor-bearing NSG mice. For each model, stromal gene expression at the primary and secondary lesions was compared to that of the corresponding, naïve, non-cancer-bearing tissue. Subsequently, those genes that were commonly and differentially upregulated (p < 0.05; 2-fold) at all sites in each model were compared among models. We focused on the upregulated genes as they may ultimately be translated into potential prognostic or therapeutic targets. These comparisons yielded a core group of 34 stromal (mouse) probes that were consistently upregulated among all cancer-bearing tissue in these models, suggesting that they may be crucial for successful cancer development. Of these 34 probes, 32 correspond to protein coding genes, of which 21 are consistent with an infiltration of myeloid cells in the cancer-bearing tissues. In this study, we focused on the pro-inflammatory genes S100A8 and S100A9 (Fig. 1).
Transcriptional changes in the cancer-bearing stroma at the primary and secondary cancer sites identify a myeloid cell infiltration signature that is also found in human samples of breast cancer. Venn diagrams summarizing transcriptional changes in each model of cancer-bearing tissues of tumor-bearing mice versus naïve non-tumor-bearing mice (n = 3 mice per group; PT primary tumor, LN met lymph node metastasis). Cut-offs used for gene list generation: p value with FDR <0.05, fold >2. A core group of 34 probes differentially expressed probes in host cells (murine) at all sites in all models. Highlighted genes are expressed in cells of myeloid lineages (blue) and pseudogenes (gray)
Systemic infiltration of S100A8+ and S100A9+ cells is preferentially observed in mouse models of metastatic versus non-metastatic BrC
To confirm the presence of S100A8+ and S100A9+ cells infiltrating the tissue of tumor-bearing mice, immunohistochemical (IHC) staining of both of these proteins was conducted on mammary fat pad and lung tissues from control (non-tumor bearing), MDA-MB-231, MDA-MB-436, and DT28 tumor-bearing NSG mice. Infiltrating S100A8+ and S100A9+ cells were observed in tissue from tumor-bearing mice but not on the naïve tissues (Fig. 2a), confirming our original observation in the transcriptional analysis. We extended this analysis to less aggressive BrC xenograft models available in our lab, including MCF-7, BT474 cells, and two cultures from dissociated primary human breast tumors (DT13 and DT22) [29]. As shown in Fig. 2b, S100A8+ and S100A9+ cells were found at much lower frequencies in these models than in the aggressive ones. Similar results were observed for DT cells in the less immunosuppressed NOD SCID mice (Fig. 2c). Importantly, in immunocompetent BALB/c mice greater numbers of infiltrating S100A8+ cells were observed in the highly aggressive 4T1 tumors compared to the non-metastatic 67NR tumors (Fig. 2d). Moreover, the same BrC cell line used in varyingly immunosuppressed mouse models shows increasing metastatic spread in the more immunocompromised strains as observed for human [29] and murine BrC lines (Fig. S1). Altogether, our results suggest that S100A8+ cells are preferentially—or more efficiently—recruited by metastatic breast tumors and support the role of the host as a critical contributor in mediating the metastatic process.
S100A8+ cells infiltrate tissue of tumor-bearing mice and are found at higher frequency in metastatic tumors in xenograft and syngeneic models of breast cancer. a Micrographs of S100A8 and S100A9 immunohistochemistry staining of mammary fat pad (MFP) and lung from naïve, non-tumor-bearing (NTB), and metastatic tumor-bearing NSG mice (MDA-MB-231, MDA-MB-436, and dissociated culture DT28). Red arrowheads indicate metastatic lesions. Infiltrating S100A8 and S100A9 positive cells are present in tissue from primary and secondary sites of metastatic tumor-bearing mice. b Micrographs of S100A8 and S100A9 immunohistochemistry staining of mammary fat pad (MFP) and lung from NSG mice bearing tumors of low to none metastatic ability (MCF-7, BT474, DT13, and DT22). c Micrographs of S100A8 immunohistochemistry staining of mammary fat pad (MFP) of DT tumor-bearing NOD/SCID mice (dissociated cultures DT13, DT22, and DT28). d Micrographs of S100A8 immunohistochemistry staining of mammary fat pad (MFP) and lung tissues from 4T1 (highly metastatic) and 67NR (non-metastatic) tumor-bearing BALB/c mice. Greater levels of infiltrating S100A8+ cells are found in 4T1 tumors compared to 67NR
S100A8+ MDSCs are recruited by human MDA-MB-231 breast tumors
In order to identify the nature of the infiltrating S100A8+ cells, we performed flow cytometric analysis of dissociated MDA-MB-231 tumors (NSG mice). Tumors were digested and simultaneously stained for the pan-leukocyte marker CD45, myeloid lineage markers CD11b, Ly6C, Ly6G, and F4/80 as well as for S100A8. As shown in Fig. 3a, three subpopulations of CD45+CD11b+ cells are recruited by MDA-MB-231 tumors: Ly6G+Ly6C−, Ly6G−Ly6C+, and Ly6G−Ly6C− cells. These phenotypes are consistent with those observed in G-MDSCs, M-MDSCs, and macrophages, respectively. The high expression of the macrophage marker F4/80 in the Ly6G−Ly6C− population, intermediate expression in the Ly6G−Ly6C+ cells, and lack of expression in the Ly6G+Ly6C− (Fig. 3a), further confirmed these phenotypes. In addition, cellular morphology of these subsets was confirmed by Giemsa–Wright staining (Fig. 3b). Ly6G+Ly6C−F4/80− subset was comprised mainly of PMN cells; Ly6G−Ly6C+F4/80int subset exhibited morphology of immature monocytic cells; and the Ly6G−Ly6C−F4/80hi subset was mainly formed by mature monocytic cells. By flow cytometry, S100A8 expression was found in all of the myeloid subsets, being highest in Ly6G+Ly6C−F4/80− cells (Fig. 3c).
MDA-MB-231 tumor-infiltrating myeloid cells are S100A8+ and include functional MDSCs. a A representative dot-plot of the distribution of tumor-infiltrating myeloid cells (CD45+CD11b+ columns indicate average of results of three independent tumors ± sem) and expression of F4/80 flow cytometric analysis of MDA-MB-231 mammary tumors is shown on the right panels. Cells were gated on CD45+CD11b+ to isolate the myeloid cell populations and analyzed for expression of Ly6C, Ly6G, and F4/80. Bars on the right are averages ± sem of three independent experiments. b Myeloid cell subsets Ly6G+Ly6C−F4/80−, Ly6G−Ly6C+F4/80int, and Ly6G−Ly6C−F4/80hi were sorted by flow cytometry and stained with Giemsa–Wright to evaluate their morphology. c Expression of S100A8 in the subpopulations of myeloid cells (CD45+CD11b+) recruited to MDA-MB-231 mammary tumors. Positive expression of S100A8 was found in 98 % of Ly6G+Ly6C−F4/80− cells, 93 % of Ly6G−Ly6C+F4/80int, and 82 % of Ly6G−Ly6C−F4/80hi. d CellTrace™-labeled splenocytes from non-tumor-bearing BALB/c mice were stimulated to proliferate in vitro and cultured at a 1:3 ratio with the indicated MDSC populations isolated from spleen of naïve NSG mice (left panel), spleens of MDA-MB-231 tumor-bearing NSG mice (middle panel), and from MDA-MB-231 primary tumors (right panel). Proliferation of CD3+CD4+ and CD3+CD8+ T cells was determined after 72 h of co-culture
To confirm that Ly6G−Ly6C+F4/80int and Ly6G+Ly6C−F4/80− are functional MDSCs, splenocytes from non-tumor-bearing BALB/c mice were labeled with CellTrace™ and stimulated to proliferate in co-culture with MDSCs isolated from tumors and spleens from MDA-MB-231 tumor-bearing NSG mice, and as shown in Fig. 3d, splenic and tumor-derived MDSCs retain the ability to suppress T cell proliferation. Importantly, cells sharing the same markers as MDSCs isolated from spleens of non-tumor-bearing NSG female mice did not suppress T cell proliferation under the same co-culture conditions (Fig. 3d). Altogether, our data show that the phenotypes of S100A8+ myeloid cells recruited to the primary site include macrophages and MDSCs.
MDSCs favor primary tumor growth and metastatic spread of MDA-MB-231 cells
Next, we sought to determine the effects of depleting MDSCs on the growth and metastatic potential of MDA-MB-231 tumors. The anti-Gr-1 antibody (RB6-8C5) was administered in vivo to neutralize MDCSs, and thus, induce their depletion. This treatment led to moderately reduced primary tumor growth of MDA-MB-231 cells in the mammary gland of NSG mice when administered at time of cancer cell inoculation (Fig. 4a), as well as when administered at day 21 after cancer cell inoculation (Fig. S2). Depletion of MDSCs significantly reduced the number of metastatic lesions in the lungs, indicating that these cells also promote metastasis of BrC cells (Fig. 4b), consistent with what has been reported on the role of S100A8 in establishing the pre-metastatic niche in 4T1 tumor-bearing immunocompetent mice [30] as well as in other types of cancer [31]. As we had previously confirmed that MDSCs recruited to these tumors express S100A8, we performed IHC staining for this protein and confirmed that treatment of tumor-bearing mice with Gr-1 antibody, and not the isotype control, led to a significant reduction in the number of S100A8+ cells infiltrating primary tumors and lungs (Fig. 4c). Taken together, our data suggest that S100A8+ MDSCs favor BrC tumor growth and metastatic spread.
MDSCs favor primary tumor growth and metastatic burden in MDA-MB-231 xenograft model. a Volumes of tumors in mammary fat pads of NSG mice injected with MDA-MB-231 cells and treated with either control (IgG) or anti-Gr-1. Depletion of MDSCs cells by Gr-1 antibody treatment moderately inhibited primary tumor growth. Each data point is the mean estimated volume value (±sem) of six primary tumors. The experiment was performed twice independently. b Number of metastases in lungs of mice bearing MDA-MB-231 tumors and treated with control IgG or anti-Gr-1 antibody. Quantification and representative micrographs of cytokeratin 18 (CK18) metastatic lesions in lung tissue are shown. Depletion of MDSCs by Gr-1 antibody treatment significantly decreased lung metastasis (p < 0.05, Student’s t-test). Each data point is the mean value (±sem) of four–six mice. The experiment was performed twice independently. c Number of S100A8+ cells in lungs of mice bearing MDA-MB-231 tumors and treated with control IgG or anti-Gr-1 antibody. Quantification and representative micrographs of S100A8 immunohistochemical staining of lung tissue are shown. Gr-1 antibody treatment significantly depleted S100A8+ cells (p < 0.05, Student’s t-test). Each data point is the mean value (±sem) of four–six mice. The experiment was performed twice independently
MDA-MB-231 tumor-secreted MIF promotes S100A8+ myeloid cell infiltration
Secretion of tumor-derived cytokines is a known mechanism by which epithelial tumors are able to recruit immune cells [13–15]. As our models utilized BrC cells of human origin in murine hosts, we were able to discern between proteins produced by the individual species. To define the mechanism by which tumor cells with metastatic potential recruit MDSCs to primary and metastatic lesions, we performed a human cytokine-profiling array to analyze the sera of MDA-MB-231 tumor-bearing and naïve NSG mice. There was no significant detection of human proteins in sera of non-tumor-bearing mice, while sera of MDA-MB-231 tumor-bearing mice had higher levels of GM-CSF, IL-8, macrophage inhibitory factor (MIF), and Serpin E1 of human origin, thus, indicating their in vivo production by MDA-MB-231 cells. Using BioPlex assays we confirmed this result and quantified additional tumor-secreted factors—previously reported to be secreted by MDA-MB-231 tumors- including IL-6, IL-8 [32], GM-CSF [33], MIF [34], osteopontin (OPN) [35], and VEGF [36] (Fig. 5a).
Cytokine knockdown does not affect MDA-MB-231 in vitro proliferation or migration abilities. a Profile of 36 human cytokines in serum from MDA-MB-231 tumor-bearing and naïve (non-tumor bearing) NSG mice. Human GM-CSF, IL-8, MIF. and Serpin E1 are present in the serum of MDA-MB-231 tumor-bearing mice. Quantification by BioPlex assay of IL-6, IL-8, GM-CSF, MIF, osteopontin (OPN), and VEGF cytokines in serum of MDA-MB-231 tumor-bearing mice. b Depletion of GM-CSF, IL-8, MIF, and Serpin E1 in MDA-MB-231 cells in vitro was confirmed by immunoblot analysis of conditioned media from stably transduced shRNA cells compared with controls (parental cell line MDA-MB-231 and control shRNA cells). c Cytokine depletion does not affect cell migration. Images of wounds of parental, control shRNA, and cytokine shRNA (GM-CSF, IL-8, MIF, and Serpin E1) cells at 0 and 24 h after wounding. d Cytokine depletion does not affect cell proliferation. Cell number of parental, control shRNA, and cytokine shRNA (GM-CSF, IL-8, MIF, and Serpin E1) cells 1, 3, and 5 days after plating
In order to test the functional consequences of impairing cytokine secretion from tumor cells, stable shRNA-knockdown populations of MDA-MB-231 cells were generated for the four cytokines identified by the serum profiling array. Impaired cytokine production was confirmed by Western blot of conditioned media (Fig. 5b). In vitro, migratory ability and growth rate of cytokine depleted MDA-MB-231 cells was not altered by knockdown of the indicated cytokines (Fig. 5c, d), suggesting that the tumor-derived MIF detected in sera of MDA-MB-231 tumor-bearing mice may be the result of the interaction with the host rather than related to changes in the behavior of MDA-MB-231 cells themselves. We next examined the in vivo effect of the depletion of the above-mentioned cytokines in MDA-MB-231 on tumor growth and metastatic behavior. In our model, depletion of MIF in MDA-MB-231 cells, but not GM-CSF, IL-8, and Serpin E1, moderately decreased primary tumor growth (Fig. 6a) and lowered metastatic burden in the lungs (Fig. 6b). Importantly, a reduction in the number of S100A8+ infiltrating cells was also observed in lung tissue, suggesting that depletion of MIF may inhibit tumor growth and metastasis by inhibiting S100A8+ myeloid cell recruitment in this model (Fig. 6c).
Tumor-derived MIF favors myeloid cell infiltration to primary tumor and metastatic sites, promoting primary tumor growth and metastasis of MDA-MB-231 cells. a Tumor growth curve of MDA-MB-231 shRNA mammary fat pad xenografts in NSG mice. Depletion of MIF secretion by MDA-MB-231 tumors resulted in delayed tumor growth (left panel; p < 0.01, ANOVA) and final primary tumor weight (right panel; p < 0.01, ANOVA). Each data point is the mean value (±sem) of ten primary tumors (NSC non- silencing control). Columns are average of n = 10 mice (±sem). b Number of metastases in lungs of NSG mice bearing MDA-MB-231 shRNA tumor xenografts. Quantification and representative micrographs of cytokeratin 18 (CK18) metastatic lesions in lung tissue are shown. MIF shRNA significantly decreased lung metastasis (p < 0.05, Student’s t-test). Each data point is the mean value (±sem) of ten mice. c Micrographs of S100A8 immunohistochemical staining of lung tissue from NSG mice bearing MDA-MB-231 shRNA tumor xenografts. Decrease in the number of S100A8+ infiltrating cells was observed in lung tissue from mice bearing MIF shRNA tumor xenografts
Validation in clinical samples: increased S100A8 and S100A9 expression associates with increased HR of death
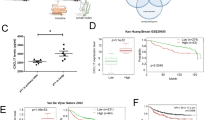
To test whether increased levels of these genes correlate with patient outcome, Oncomine™ (Compendia Bioscience™, part of Life Technologies™, Ann Arbor, MI) was used for analysis and visualization. We performed a meta-analysis to determine the impact of S100A8 expression on short-term overall survival in BrC datasets with S100A8 (n = 10) and S100A9 (n = 9) expression data and overall survival data available from Oncomine™ (Fig. 7a). This meta-analysis revealed that increased S100A8 or S100A9 expression is associated with a significant increase in hazard of a death event within 5 years (random effects model summary statistic HR S100A8: [95 % CI] 1.14 [1.07, 1.21], p = 2.59 e−05; and S100A9 [95 % CI] 1.14 [1.07, 1.21], p = 9.9 e−05). Furthermore, based on the PAM50 classification [37] of the samples in the Curtis dataset, increased levels of S100A8 and S100A9 transcripts are found in the basal and Her2-enriched BrC subtypes (Fig. 7b). Since these microarray data are obtained from the entire tumor, the contribution of each cell type—cancer or cells from the microenvironment—in these samples cannot be separated. Therefore, we examined these transcriptional changes in three available BrC datasets in which stromal cells were isolated and independently analyzed. Indeed, stroma of invasive BrC exhibits greater levels of S100A8 and S100A9 transcripts compared to stroma of normal mammary tissue in these datasets (Fig. 7c) [2, 38]. Similar results were observed for other transcripts identified in our signature (Selplg, Lst1; not shown).
Transcriptional upregulation of S100A8 and S100A9 in human breast tumors is associated with a significant increase in hazard of short-term death event and with aggressive breast cancer subtypes. a Transcriptional upregulation of S100A8 and S100A9 in human breast tumors is associated with a significant increase in hazard of short-term death event. Forest plots of the effect of S100A8 and S100A9 expression on hazard of death event within 5 years of follow-up in independent primary breast cancer datasets. Overall summary statistics are given for the fixed effects model S100A8: [95 % CI] 1.14 [1.07, 1.21], p = 2.59 e−05; and S100A9: [95 % CI] 1.14 [1.07, 1.21], p = 9.9 e−05. Confidence intervals that extend beyond upper range of the forest plot are indicated by arrows. b Transcriptional levels of S100A8 and S100A9 are greater in the basal and Her2 breast cancer subtypes as shown for the Curtis dataset [51] as visualized with Oncomine™. c Transcriptional levels of S100A8 and S100A9 are overexpressed in human stroma of invasive breast carcinoma compared to normal breast stroma as shown for the Finak dataset [2] as visualized with Oncomine™
Higher levels of infiltrating S100A8+ cells correlate with poor clinical outcome
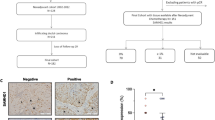
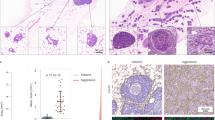
To test the clinical relevance of our findings, we analyzed infiltrating S100A8+ cells in two independent cohorts of invasive BrC specimens. We performed immunohistochemistry (IHC) of S100A8 in a cohort of human breast tumor specimens (n = 244) and counted the S100A8+ positive infiltrating cells. In agreement with the pre-clinical data, high levels of infiltrating S100A8+ cells distant to the tumor correlated with significantly shorter metastasis free survival of BrC patients (Fig. 3a). High levels of infiltrating S100A8+ cells distant to the tumor also correlated with high grade, estrogen-receptor (ER) and progesterone receptor (PR) negativity, epidermal growth factor receptor positivity, basal keratin positivity, and Ki67 (MIB-1) expression (Fig. 8a). Additionally, an independent cohort of invasive BrC specimens was analyzed using AQUA (Automated Quantitative Analysis) staining for assessment of infiltrating S100A8+ cells. In tissue microarrays (TMAs) of BrC specimens with associated clinical parameters and outcome, this analysis showed that infiltrating S100A8+ cells are clearly seen in and adjacent to the tumor (red staining, Fig. 8b). Quantitation of S100A8+ cells in, adjacent to, and at a distance from the tumor and statistical correlation with clinical parameters revealed significant inverse associations of S100A8+ cells with ER and PR and strong associations with Her2 and tumor size and grade (Fig. 8c). These correlations indicate that tumors possessing high levels of infiltrating S100A8+ cells are frequently of the basal subtype, which are clinically more aggressive and have shorter disease free survival [37]. Our data suggest that one of the mechanisms driving this poor prognosis may be the recruitment of S100A8+ myeloid cells.
The presence of infiltrating S100A8+ cells is associated with significantly shorter metastasis free survival in breast cancer patients and with the basal breast cancer subtype. a Kaplan–Meier survival curves of breast cancer patients with tumors that have high numbers of distant infiltrating S100A8+ cells have significantly shorter metastasis free survival (p = 0.043, log-rank test). Patients were categorized into three groups by the number of tumor distant infiltrating S100A8 positive cells present. Fisher’s exact test was used for all statistical analyses except size, which was analyzed using ANOVA. b Representative picture of S100A8 staining of one of two TMAs analyzed by AQUA. c Infiltration of S100A8+ cells (Allred Score) is associated with the basal PAM50 intrinsic breast cancer subtype (n = 62)
Discussion
Much effort has been directed at studying the cell autonomous changes that promote metastatic behavior of BrC cells. While these studies have identified several promising avenues and therapeutic approaches, it is becoming increasingly clear that the genetic instability of cancer cells results in inter- and intratumoral heterogeneity that make the specific targeting of carcinoma very difficult [39]. On the other hand, the changes that occur in the host tissue and recruited cells—including hematopoietic cells, fibroblasts, myofibroblasts, and endothelial cells—are an area under intense investigation. Cells in the tumor microenvironment play a functional role in creating favorable conditions for tumor progression and metastatic spread and are now recognized to be a hallmark of cancer [1], and significant efforts have been aimed at including stromal-targeted therapies in the treatment of BrC including approaches against cancer-associated fibroblasts and immune cells [40].
Our study focused on identifying changes occurring in the cancer-bearing tissues of mice bearing human BrC cells with the goal of identifying novel targets of potential prognostic and therapeutic value for metastatic disease. Using three independent human metastatic BrC models xenografted in NSG mice, we found that a predominant change occurring in tissues of cancer-bearing mice is the infiltration of myeloid cells, as suggested by the function of the majority of genes (21 of 32) consistently identified throughout our models. An accumulating body of evidence has highlighted the role of the immune system in regulating tumor initiation and progression. However, different immune cell populations have different, often opposite, effects on tumor cell behavior. For example, increased infiltration by CD8+ T cells is generally associated with better survival for several types of cancer, whereas infiltration by MDSCs is associated with poorer prognosis [11, 41]. It is, therefore, important to identify which immune cell populations are tumor promoters so that they may be specifically targeted by novel therapeutics. Specifically, MDSCs are believed to promote tumor progression in a number of ways, most notably by induction of T cell tolerance and suppression of other anti-tumor responses by immune cell components and by favoring angiogenesis [42]. In humans, MDSCs are much less well characterized, and it is very important to identify key markers of the specific populations that are tumor promoting in order to avoid the non-specific toxicity associated with targeting the immune system more generally.
In mice, most research on MDSCs has been focused on immunocompetent mouse models of cancer and has allowed the characterization of MDSCs phenotypes and immunosuppressive action. The use of our recently characterized NSG mouse model for human BrC metastasis to study tumor-associated myeloid cells provides an interesting opportunity to address the function of these cells in the absence of T cells. NSG mice are highly prone to metastasis of BrC [3], neuroblastoma [4], and melanoma cells [5]. Interestingly, compared to immunocompetent mice, naïve NSG mice intrinsically have elevated proportions of splenic CD11b+/Gr-1+ cells, and as our own data shows of CD11b+Ly6G+Ly6C−/lo and CD11b+Ly6G−LY6Chi populations (Fig. S4). While the proportion of myeloid cells in spleens is greater in NSG mice than in wild-type mice, we found that in splenic CD11b+ cells expression of S100A8 protein—and likely of other proteins involved in MDSC function—is significantly lower than these same cells isolated from tumor-bearing mice, and they do not show suppressive activity on T cell proliferation. This suggests that expansion and activation of MDSC in this model is triggered by the action of the human BrC cells. In agreement with this, we observed that not all human BrC cell lines are able to neither activate recruitment of S100A8+ myeloid cells nor metastasize in NSG mice, despite them having a pre-existing population of CD11b+/Gr-1+ cells.
In this context, we have determined that human BrC cells recruit populations of S100A8+ myeloid cells, including MDSCs and macrophages, that have strong tumor-promoting effects, which are independent of immune cell tolerance or the suppressive action of MDSCs on T cells, as they are absent in our mouse model. Further confirmation of the importance of MDSCs in our system emerged when animals were treated with a neutralizing Gr-1 antibody, which mainly targets MDSCs exhibited delayed primary tumor growth, lower metastatic spread, and reduced numbers of S100A8+ cells. Furthermore, since we observe differing metastatic potential of the same BrC cell lines in animal models increasing levels of immunosuppression, this strongly supports our contention that a critical element of metastatic progression is host-mediated rather than tumor cell-specific. In addition, we attempted to identify which secreted cytokines may be responsible for activating and recruiting these cells from the bone marrow to the primary tumor. We determined that in our model, MDA-MB-231 cells produce pro-inflammatory cytokines, and other factors previously shown to favor the recruitment of MDSCs including IL-6 [43], GM-CSF [44] and OPN [35], and that tumor-derived MIF played an important role in the recruitment of myeloid cells. MIF is an inflammatory cytokine that is overexpressed in many solid tumors and is associated with poor prognosis [45]. In agreement with our findings, a recent study showed that inhibition of MIF reduced MDSC accumulation in immunocompetent 4T1-tumor-bearing mice, and inhibited primary tumor growth and pulmonary metastasis [30]. Examination of the expression of MIF in a panel of additional human BrC cell lines indicated that the expression of MIF alone might not be solely responsible for the recruitment of S100A8+ MDSC’s and promote BrC metastasis as it is expressed by both metastatic cells lines that recruit S100A8+ MDSCs as well as by non-metastatic cell lines that do not recruit S100A8+ cells (Fig. S3a). This was also confirmed by the quantification of MIF in sera of NSG mice bearing metastatic and non-metastatic breast tumors (Fig. S3b). The latter is in agreement with previous studies showing a dual role for MIF production by metastatic and non-metastatic BrC cell lines [34]. Therefore, while in our model MIF is required for MDA-MB-231 recruitment of S100A8+ cells that favor tumor growth and metastasis in our model, its expression alone is not sufficient to regulate this process. This indicates that additional (known or yet to be identified) mediators of MDSC recruitment and activation must act in concert with MIF to promote BrC progression and metastasis.
Having shown in highly immunocompromised NSG mouse model that S100A8+ myeloid cells promote metastasis independently from their immunosuppressive cellular functions, it was important to determine that these cells also play a role in immunocompetent mice. We, and others have shown that 4T1-tumor-bearing BALB/c mice also show infiltration of S100A8+ tumor-associated myeloid cells, similar to our observations in the NSG model. Importantly, we additionally show that this recruitment is not observed in non-metastatic 67NR tumors, confirming our initial observation in xenograft models that metastatic BrC cells preferentially recruit S100A8+ cells. Previous studies have demonstrated that depletion of MDSCs in BALB/c mice suppresses metastasis of 4T1 [30]. Our data suggest that this result may be due to the loss of the metastasis promoting population of S100A8+ myeloid cells that are seen in NSG tumor-bearing mice. In addition, it has been recently shown that mice transplanted with bone marrow from S100A9 knockout mice (which also fail to express S100A8) were much less susceptible to mammary fat pad xenograft tumor growth and metastasis than mice transplanted with wild-type bone marrow. This provides compelling evidence for the role of S100A8/A9+ myeloid cells in driving primary and metastatic tumor growth [46].
A critical part of our study consists in the validation of our observations in mouse models regarding the presence of S100A8+ cells in clinical samples of human BrC. Our data suggest that the infiltrating immune cells contribute a great part of the increased levels of S100A8 transcript and protein found in human breast tumors. In clinical samples of ten independent datasets, we have shown that increased transcript levels of S100A8 and S100A9 correlate with increased hazard of a death event at 5 years and basal BrC. A previous study by Arai et al. investigated the relationship between the expression of S100A8 and S100A9 proteins in the tumor cells with several pathological parameters. They found that protein expression of the S100A8/A9 complex in tumor cells correlated with poor pathological parameters of invasive ductal carcinoma (IDC) as well as with poorly differentiated ductal carcinoma in situ, which has a higher recurrence rate. Additionally, in agreement with our data, strong positive reaction of both proteins in the infiltrating myelomonocytic cells was also detected; however, the correlation of these cells is not addressed in that study [20]. Importantly, in this study we show for the first time that S100A8 protein expression in the host (immune) compartment of clinical samples consistently correlates with poor prognosis, and thus, S100A8+ infiltrating cells should be considered as a prognosis marker in BrC.
Interestingly, a recent study demonstrated that MDSCs in human colon patients are S100A8/A9 positive and that S100A9 staining in combination with anti-CD14 could be used to identify MDSC in whole blood from patients with colon cancer [19]. This suggests that our finding that S100A8+ MDSCs can be used as an additional marker of tumor-promoting myeloid cells in mice, may also apply to humans. Our data are consistent with another recent study that showed that BrC patients with high expression of S100A8/A9 in their lung metastases had a significantly shorter overall survival compared to low S100A8/A9 (p value = 0.01) and [46], supporting our results from primary BrC tumors. These data suggest that it may be possible to inhibit S100A8/A9 myeloid cells in patients to inhibit metastatic progression. Data from pre-clinical models indicate that this strategy may be effective. S100A8/A9 depleting antibodies have been used to inhibit metastasis of Lewis lung cancer cells successfully in mice [27]. In addition, the S100A9 knockout mouse, which also lacks any S100A8 expression, is phenotypically normal, suggesting there should not be major toxicity problems associated with this approach. Moreover, our data suggest that infiltration of S100A8+ myeloid cells occurs most frequently in the basal BrC subtype, which is aggressive and in need of effective targeted therapies. The development of small molecule inhibitors or depleting antibodies targeting S100A8/A9 may, therefore, be an effective therapeutic strategy in these patients.
In summary, our data suggest that during BrC progression, populations of S100A8+ tumor-associated myeloid cells are recruited by tumor-derived factors, including MIF, and promote BrC progression. The use of immunosuppressed mouse models permits the elucidation of novel role(s) of MDSCs aside from their canonical effects on T cells. Analysis of clinical data supports S100A8+ infiltrating cells to be an important prognosis marker in BrC. Our results suggest several new potential therapeutic strategies, including neutralizing antibodies targeting MIF and/or S100A8+ myeloid cells that may be useful for inhibiting BrC progression and metastasis.
Materials and methods
Ethics statement
The Institutional Animal Care and Use Committee (IACUC) at the University of Miami approved the animal experiments described in this study (protocol 11-227). All animals were maintained in accordance with IACUC guidelines.
BrC cell lines and cultures
Human BrC cells (MCF-7, BT474, MDA-MB-231, and MDA-MB-436) were obtained from the American Type Culture Collection and maintained according to the supplier’s instructions. Authentication of the human BrC cell lines was performed by sequencing of the hypervariable regions of the mitochondrial DNA. Human BrC DT cells have been generated and characterized by group as previously described [29]. Murine BrC cells 4T1 and 67NR cells were obtained from the American Type Culture Collection and from the Karmanos Cancer Institute (Detroit, MI), respectively. All cells were harvested at the exponential phase of growth for injection into the mammary fat pads of mice.
Mice
BALB/c, NOD SCID and NOD scid gamma (NSG) mice were purchased from Jackson Laboratory. BrC cells were unilaterally injected by subcutaneous injection at the base of the nipple of the fourth abdominal mammary fat pad of eight-week-old female mice. Human BrC lines (2.5 × 106 cells) and DT cells (1 × 106 cells) were injected in 200 μl of 50:50 Matrigel/Collagen. Murine 4T1 (1 × 105 cells) and 67NR (1 × 105 cells) cells were injected into BALB/c mice. Tumor growth was monitored externally using vernier calipers for up to 30 weeks, and animals were sacrificed when tumors reached 10 % of body weight. Necropsies were performed to identify macro-metastases. Primary tumors and organs were harvested and samples were taken for RNA extraction and for pathological analysis. Pathology samples were fixed in 10 % formalin, paraffin embedded, sectioned and stained with hematoxylin and eosin, or antibodies against cytokeratin 18 (CK18), S100A8, or S100A9. Pathology processing and staining of harvested mouse tissues was performed at the Lombardi Comprehensive Cancer via Science Exchange, Inc. Slides were analyzed by a pathologist to confirm the presence of metastases. RNA was isolated from three independent metastases and primary xenograft tumors for analysis. Normal tissue was harvested from three independent age matched non-tumor-bearing mice.
RNA isolation
RNA was extracted using Trizol Reagent (Invitrogen) according to the manufacturer’s instructions. Concentration and yield of RNA samples were determined using a NanoDrop ND-1000 Spectrophotometer (NanoDrop Technologies). RNA integrity was determined by analysis on an Agilent 2100 Bioanalyzer (Agilent Technologies) following the manufacturer’s recommendations. Only samples with a RIN score >7.0 were used for microarray analysis.
Microarray analysis
Samples were prepared according to the manufacturer’s instructions and analyzed using MouseWG-6_V2 Expression BeadChips (Illumina, Inc.). Non-normalized, non-background subtracted data were generated using the Gene Expression module (v1.9) of GenomeStudio (v2011.1). The datasets were combined before performing log2-transformation and quantile normalization in R (version 2.14.1) using the lumi package at Wayne State University. Differential expression was calculated by performing analysis of variance (ANOVA) using Partek® software (version 6.6). Gene expression alterations were determined by identifying genes differentially expressed in tissues from non-tumor-bearing control mice compared to tissues from mice bearing MDA-MB-231 and MDA-MB-436 mammary fat pad tumor xenografts and metastases. A p value with FDR <0.05 was used as a cut-off for differentially expressed genes. The differentially expressed genes from comparisons were used to create Venn diagrams based on gene symbols. Microarray experiments and analyses of datasets were performed at Wayne State University via Science Exchange, Inc.
S100A8 and S100A9 expression meta-analyses
S100A8 and S100A9 gene expression and clinical follow-up information from the following BrC datasets were obtained from Oncomine™: Bild, Boersma, Curtis, Desmedt, Esserman, Kao, Pawitan, Sorlie, vandeVijver. R statistical software (version 2.15.2; http://www.R-project.org) and the ‘meta’ package (version 3.0-1; http://CRAN.R-project.org/package=meta) were used to perform meta-analysis and forest plot visualization for impact of S100A8 or S100A9 expression on 5-year survival.
Tumor immunophenotyping by flow cytometry
Mammary fat pad tumors were collected, minced into small pieces, enzymatically digested with a blend of liberase (Roche) and DNase I (Sigma), and filtered to generate single cell suspensions. Cells were then blocked for 30 min with purified anti-CD16/CD32 (BD Biosciences) and stained with anti-CD45 Brilliant Violet 421 (Biolegend), anti-mouse F4/80 PE (Biolegend), anti-CD11b PerCP-Cy5.5 (Biolegend), anti-Ly6G PE-C7 (BD Biosciences), and anti-Ly6C APC-Cy7 (BD Biosciences). After 30 min incubation, cells were washed in 2 ml of PBS–FBS 1 % buffer. Cell pellets were resuspended in 100 μl of BD Cytofix/Cytoperm solution (BD Bioscience) and incubated on ice for 20 min. Cells were washed twice in 1× PermWash buffer (BD Bioscience) and stained with Alexa Fluor 488-conjugated anti-S100A8 (Novus Biologicals). Cells were washed twice in PermWash buffer and resuspended in 300 μl of PBS 1× and acquired on the same day. Data were acquired using the LSRFortessa instrument (BD Biosciences) and analyzed using FlowJo software.
MDSC depletion
Starting on the day of tumor injection, mice were injected intraperitoneally bi-weekly with 50 μg anti-Gr-1 antibody (eBioscience, clone: RB6-8C5, Functional Grade Purified) or anti-isotype (Rat IgG2b K Isotype Control Functional Grade Purified) control antibody (eBioscience, clone: eB149/10H5).
Cytokine profiling
Following manufacturer’s instructions, serum from naïve (non-tumor-bearing control) and MDA-MB-231 tumor-bearing NSG mice were profiled for the levels of 36 human cytokines using a human cytokine array (Panel A Proteome Profiler, R&D Systems) and BioRad’s BioPlex assays (Cancer Panel 1, 27-plex, and 21-plex cytokine assays).
Preparation of T cells and isolation of tumor-associated MDSCs for proliferation suppression assay
Spleens were removed from non-tumor-bearing BALB/C mice and mechanically disaggregated into single cell suspensions. Cells were washed with PBS and spun at 350 g. RBrCs were lysed by hypotonic shock (RBrC lysis buffer, eBioscience cat. 00-4333). Cells were washed with PBS and labeled with CellTrace™ Violet proliferation reagent (Invitrogen) following manufacturer’s instructions. Cells were resuspended in RPMI supplemented with 10 % FBS, l-glutamax (1×), 50 μM beta-mercaptoethanol, and penicillin/streptomycin 50 U/ml, and plated in 96-well ultra-low attachment plates at 1 × 106 cells/ml and stimulated with ConA (2.5 μg/ml) and recombinant murine IL-2 (10 ng/ml). MDSCs were isolated from spleens and digested tumors using magnetic beads in the MDSC isolation kit mouse (Miltenyi Biotec, cat. 130-094-538) following manufacturer’s instructions.
Cytokine silenced cell lines
Cytokine silenced MDA-MB-231 cells were generated by lentiviral transduction with pools of specific shRNAs from the Open Biosystems pGIPZ shRNA lentiviral collection. Cytokine secretion levels were assessed by immunoblot analysis of conditioned media and cell lysates using anti-IL-8, Serpin E1, MIF, GM-CSF, and anti-actin antibodies (Abcam) to confirm successful silencing. Pooled populations of silenced cells, obtained after 5 days of drug selection (1.0 μg/ml puromycin) without subcloning, were used for experiments. Negative control cells were generated by transducing cells with a pGIPZ non-silencing control (NSC) construct, which did not yield any appreciable knockdown of the cytokines.
Wound-healing assay
Cell migration was assessed using a wound-healing (scratch) assay. Cells were plated in 24 well plates and allowed to proliferate to form a confluent monolayer. A 200 μl pipette tip was used to scratch a single wound through the middle of the cell monolayer. Cells were fixed 0 and 24 h after scratching, stained with crystal violet and imaged. Images from matched time-points were compared to determine differences in the rate of cell migration.
Proliferation assay
5 × 103 cells were plated in 24 well plates and maintained for 5 days in culture. Cells were quantified by trypsinization and counting of the cell number using a Coulter Counter. Assays were performed in triplicate.
Patient data for IHC analysis of infiltrating S100A8+ myeloid cells
A cohort of 245 patients with invasive BrC (186 IDCs, 27 invasive lobular carcinomas, 24 invasive mixed ductal and lobular carcinomas, and 8 invasive breast carcinomas of other special types) was retrospectively retrieved from a prospectively maintained database of patients diagnosed and managed at the Royal Marsden Hospital, London, UK. All patients were treated with therapeutic surgery (69 mastectomy and 156 wide local excision) followed by anthracycline-based chemotherapy. Adjuvant endocrine therapy was prescribed for patients with ER-positive tumors (tamoxifen alone in 96.4 % of the patients for the available follow-up period). Complete follow-up was available for 244 patients, ranging from 0.5 to 125 months (median = 67 months, mean = 67 months). Tumors were graded according to a modified Bloom–Richardson scoring system and size was categorized according to the TNM staging. The details of this cohort of patients have been previously described [47].
Representative 3-μm-thick formalin-fixed paraffin-embedded (FFPE) sections were cut from each tumor. Sections were pre-treated using a pressure cooker for 2.5 min with citrate buffer at pH 6.0. The primary antibody used was a rabbit monoclonal anti-MRP8, clone EPR3554 (Abcam®—Cambridge, UK) and visualization was done using the EnVision™ kit (DAKO®—Copenhagen, Denmark). For developing, the 3,3-diaminobenzidine (DAB), method was used. Positive controls included tonsil tissue, where non-germinal center lymphoid cells expressed MRP8. Negative controls included omission of the primary antibody and substitution of the primary antibody by IgG-matched control.
The semi-quantitative analysis of the distribution of S100A8 expression in the inflammatory component of BrCs was based on the assessment of ten high power fields representative of three compartments of the tumor, namely (i) intratumor compartment (within the tumor cell nests), (ii) distant stroma (defined as >one tumor cell diameter away from the tumor), and (iii) adjacent stroma (within one tumor cell diameter of the tumor), as previously described [41]. Within each compartment of the tumor, cells displaying unequivocal macrophage morphology and expressing membranous MRP8 were counted. The IHC analysis was performed by two of the authors on a multi-headed microscope, without previous knowledge of the expression of other markers and patients’ outcome. Kaplan–Meier curves were generated for metastasis free survival based on numbers of distant infiltrating S100A8+ cells. Significance was determined by log-rank test.
Patient data for AQUA analysis of infiltrating S100A8+ myeloid cells
FFPE tissue blocks of invasive BrCs (n = 62) were obtained from the files of the Department of Pathology, University of Michigan Medical Center, Ann Arbor, MI. This study was conducted with Institutional Review Board approval. After pathological review, a construction of the TMA was constructed from the most representative area using the methodology of Nocito et al. [48]. Normal breast tissue and control tissue was included in the construction of the TMA. Simultaneously, with the construction of the TMA, an addition set of cores, were used for intrinsic subtyping using the PAM50 method [49].
Double immunofluorescence staining and AQUA analysis
Double immunofluorescence staining was performed as previously described [50]. Briefly, after deparaffinization and rehydration, TMA slides were subjected to microwave epitope retrieval in 10 mM citrate buffer, pH 6. After rinsing several times in 10 mM Tris–HCl buffer, pH 8 containing 0.154 M NaCl (TBS), endogenous peroxidase activity was blocked with 2.5 % (v/v) H2O2 in methanol for 15 min. Non-specific binding of the antibodies was extinguished by a 30 min incubation with Background Sniper (BioCare Medical, Concord, CA). The TMA slides were then incubated with the tumor specific antibody, CK (mouse monoclonal antibody, clone KL1, AbD Serotec, Raleigh, NC, cat. MCA144HT, 1:50) overnight at 4 °C and S100A8 [rabbit monoclonal antibody, clone EPR3554, Novus Biologicals, Littleton, CO, cat. NBP1-42076 (1:1,500)] for 60 min at room temperature. Slides are then washed as described above and incubated with a combination of goat anti-mouse IgG conjugated to AF555 (Molecular Probes, Carpinteria, CA, A21424, 1:200) in goat anti-rabbit Envision+ (DAKO) for 60 min at room temperature in a dark humidity tray. The slides are then washed as described above and the target image is developed by a CSA reaction of Cy5 labeled tyramide (PerkinElmer, Waltham, MA, 1:50). The slides are washed with three changes of TBS and stained with the DNA staining dye 4′,6-diaminodo-2-phenylindole (DAPI) in a non-fading mounting media (ProLong Gold, Molecular Probes, Carpinteria, CA). The slides are allowed to dry overnight in a dark dry chamber, and the edges are sealed. The AQUA system (HistoRx, New Haven, Connecticut) was used for the automated image acquisition and analysis. Briefly, images of each TMA core are captured with an Olympus BX51 microscope at three different extinction/emission wavelengths. Within each TMA spot, the area of tumor is distinguished from stromal and necrotic areas by creating a tumor specific mask from the anti-CK protein, which is visualized from the Alexa Fluor 555 signal. The DAPI image is then used to differentiate between the cytoplasmic and nuclear staining within the tumor mask. Finally, the fluorescence pixel intensity of the S100A8 protein/antibody complex is obtained from the Cy5 signal and reported as mean pixel intensity. The stromal area is calculated using the total area of the TMA core minus the tumor specific mask.
References
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674. doi:10.1016/j.cell.2011.02.013
Finak G, Bertos N, Pepin F, Sadekova S, Souleimanova M, Zhao H, Chen H, Omeroglu G, Meterissian S, Omeroglu A, Hallett M, Park M (2008) Stromal gene expression predicts clinical outcome in breast cancer. Nat Med 14(5):518–527. doi:10.1038/nm1764
Iorns E, Drews-Elger K, Ward TM, Dean S, Clarke J, Berry D, El-Ashry D, Lippman M (2012) A new mouse model for the study of human breast cancer metastasis. PLoS ONE 7(10):e47995. doi:10.1371/journal.pone.0047995
Sartelet H, Durrieu L, Fontaine F, Nyalendo C, Haddad E (2012) Description of a new xenograft model of metastatic neuroblastoma using NOD/SCID/Il2rg null (NSG) mice. In Vivo 26(1):19–29
Quintana H, Piskounova E, Shackleton M, Weinberg D, Eskiocak U, Fullen DR, Johnson TM, Morrison SJ (2012) Human melanoma metastasis in NSG mice correlates with clinical outcome in patients. Sci Transl Med 4(159):159ra149. doi:10.1126/scitranslmed.3004599
Iorns E, Clarke J, Ward T, Dean S, Lippman M (2012) Simultaneous analysis of tumor and stromal gene expression profiles from xenograft models. Breast Cancer Res Treat 131(1):321–324. doi:10.1007/s10549-011-1784-8
Murdoch C, Muthana M, Coffelt SB, Lewis CE (2008) The role of myeloid cells in the promotion of tumour angiogenesis. Nat Rev Cancer 8(8):618–631. doi:10.1038/nrc2444
Nagaraj S, Gabrilovich DI (2010) Myeloid-derived suppressor cells in human cancer. Cancer J 16(4):348–353. doi:10.1097/PPO.0b013e3181eb3358
Youn JI, Nagaraj S, Collazo M, Gabrilovich DI (2008) Subsets of myeloid-derived suppressor cells in tumor-bearing mice. J Immunol 181(8):5791–5802
Cuenca AG, Delano MJ, Kelly-Scumpia KM, Moreno C, Scumpia PO, Laface DM, Heyworth PG, Efron PA, Moldawer LL (2011) A paradoxical role for myeloid-derived suppressor cells in sepsis and trauma. Mol Med 17(3–4):281–292. doi:10.2119/molmed.2010.00178
Diaz-Montero CM, Salem ML, Nishimura MI, Garrett-Mayer E, Cole DJ, Montero AJ (2009) Increased circulating myeloid-derived suppressor cells correlate with clinical cancer stage, metastatic tumor burden, and doxorubicin-cyclophosphamide chemotherapy. Cancer Immunol Immunother 58(1):49–59. doi:10.1007/s00262-008-0523-4
Srivastava MK, Sinha P, Clements VK, Rodriguez P, Ostrand-Rosenberg S (2010) Myeloid-derived suppressor cells inhibit T-cell activation by depleting cystine and cysteine. Cancer Res 70(1):68–77. doi:10.1158/0008-5472.CAN-09-2587
Bunt SK, Yang L, Sinha P, Clements VK, Leips J, Ostrand-Rosenberg S (2007) Reduced inflammation in the tumor microenvironment delays the accumulation of myeloid-derived suppressor cells and limits tumor progression. Cancer Res 67(20):10019–10026. doi:10.1158/0008-5472.CAN-07-2354
Sinha P, Clements VK, Bunt SK, Albelda SM, Ostrand-Rosenberg S (2007) Cross-talk between myeloid-derived suppressor cells and macrophages subverts tumor immunity toward a type 2 response. J Immunol 179(2):977–983
Ostrand-Rosenberg S, Sinha P (2009) Myeloid-derived suppressor cells: linking inflammation and cancer. J Immunol 182(8):4499–4506. doi:10.4049/jimmunol.0802740
Odink K, Cerletti N, Bruggen J, Clerc RG, Tarcsay L, Zwadlo G, Gerhards G, Schlegel R, Sorg C (1987) Two calcium-binding proteins in infiltrate macrophages of rheumatoid arthritis. Nature 330(6143):80–82. doi:10.1038/330080a0
Vogl T, Tenbrock K, Ludwig S, Leukert N, Ehrhardt C, van Zoelen MA, Nacken W, Foell D, van der Poll T, Sorg C, Roth J (2007) Mrp8 and Mrp14 are endogenous activators of Toll-like receptor 4, promoting lethal, endotoxin-induced shock. Nat Med 13(9):1042–1049. doi:10.1038/nm1638
Wang L, Chang EW, Wong SC, Ong SM, Chong DQ, Ling KL (2013) Increased myeloid-derived suppressor cells in gastric cancer correlate with cancer stage and plasma S100A8/A9 proinflammatory proteins. J Immunol 190(2):794–804. doi:10.4049/jimmunol.1202088
Zhao F, Hoechst B, Duffy A, Gamrekelashvili J, Fioravanti S, Manns MP, Greten TF, Korangy F (2012) S100A9 a new marker for monocytic human myeloid-derived suppressor cells. Immunology 136(2):176–183. doi:10.1111/j.1365-2567.2012.03566.x
Arai K, Takano S, Teratani T, Ito Y, Yamada T, Nozawa R (2008) S100A8 and S100A9 overexpression is associated with poor pathological parameters in invasive ductal carcinoma of the breast. Curr Cancer Drug Targets 8(4):243–252
Ott HW, Lindner H, Sarg B, Mueller-Holzner E, Abendstein B, Bergant A, Fessler S, Schwaerzler P, Zeimet A, Marth C, Illmensee K (2003) Calgranulins in cystic fluid and serum from patients with ovarian carcinomas. Cancer Res 63(21):7507–7514
Vogl T, Gharibyan AL, Morozova-Roche LA (2012) Pro-inflammatory S100A8 and S100A9 proteins: self-assembly into multifunctional native and amyloid complexes. Int J Mol Sci 13(3):2893–2917. doi:10.3390/ijms13032893
Sinha P, Okoro C, Foell D, Freeze HH, Ostrand-Rosenberg S, Srikrishna G (2008) Proinflammatory S100 proteins regulate the accumulation of myeloid-derived suppressor cells. J Immunol 181(7):4666–4675
Bunt SK, Clements VK, Hanson EM, Sinha P, Ostrand-Rosenberg S (2009) Inflammation enhances myeloid-derived suppressor cell cross-talk by signaling through Toll-like receptor 4. J Leukoc Biol 85(6):996–1004. doi:10.1189/jlb.0708446
Yin C, Li H, Zhang B, Liu Y, Lu G, Lu S, Sun L, Qi Y, Li X, Chen W (2013) RAGE-binding S100A8/A9 promotes the migration and invasion of human breast cancer cells through actin polymerization and epithelial–mesenchymal transition. Breast Cancer Res Treat 142(2):297–309. doi:10.1007/s10549-013-2737-1
Ye XZ, Yu SC, Bian XW (2010) Contribution of myeloid-derived suppressor cells to tumor-induced immune suppression, angiogenesis, invasion and metastasis. J Genet Genomics = Yi chuan xue bao 37(7):423–430. doi:10.1016/S1673-8527(09)60061-8
Hiratsuka S, Watanabe A, Aburatani H, Maru Y (2006) Tumour-mediated upregulation of chemoattractants and recruitment of myeloid cells predetermines lung metastasis. Nat Cell Biol 8(12):1369–1375. doi:10.1038/ncb1507
Zhang J, Liu J (2013) Tumor stroma as targets for cancer therapy. Pharmacol Ther 137(2):200–215. doi:10.1016/j.pharmthera.2012.10.003
Drews-Elger K, Brinkman JA, Miller P, Shah SH, Harrell JC, da Silva TG, Ao Z, Schlater A, Azzam DJ, Diehl K, Thomas D, Slingerland JM, Perou CM, Lippman ME, El-Ashry D (2014) Primary breast tumor-derived cellular models: characterization of tumorigenic, metastatic, and cancer-associated fibroblasts in dissociated tumor (DT) cultures. Breast Cancer Res Treat. doi:10.1007/s10549-014-2887-9
Simpson KD, Templeton DJ, Cross JV (2012) Macrophage migration inhibitory factor promotes tumor growth and metastasis by inducing myeloid-derived suppressor cells in the tumor microenvironment. J Immunol 189(12):5533–5540. doi:10.4049/jimmunol.1201161
Ichikawa M, Williams R, Wang L, Vogl T, Srikrishna G (2011) S100A8/A9 activate key genes and pathways in colon tumor progression. Mol Cancer Res 9(2):133–148. doi:10.1158/1541-7786.MCR-10-0394
Freund A, Chauveau C, Brouillet JP, Lucas A, Lacroix M, Licznar A, Vignon F, Lazennec G (2003) IL-8 expression and its possible relationship with estrogen-receptor-negative status of breast cancer cells. Oncogene 22(2):256–265. doi:10.1038/sj.onc.1206113
Pederson L, Winding B, Foged NT, Spelsberg TC, Oursler MJ (1999) Identification of breast cancer cell line-derived paracrine factors that stimulate osteoclast activity. Cancer Res 59(22):5849–5855
Verjans E, Noetzel E, Bektas N, Schutz AK, Lue H, Lennartz B, Hartmann A, Dahl E, Bernhagen J (2009) Dual role of macrophage migration inhibitory factor (MIF) in human breast cancer. BMC Cancer 9:230. doi:10.1186/1471-2407-9-230
McAllister SS, Gifford AM, Greiner AL, Kelleher SP, Saelzler MP, Ince TA, Reinhardt F, Harris LN, Hylander BL, Repasky EA, Weinberg RA (2008) Systemic endocrine instigation of indolent tumor growth requires osteopontin. Cell 133(6):994–1005. doi:10.1016/j.cell.2008.04.045
Kurebayashi J, Otsuki T, Kunisue H, Mikami Y, Tanaka K, Yamamoto S, Sonoo H (1999) Expression of vascular endothelial growth factor (VEGF) family members in breast cancer. Jpn J Cancer Res Gann 90(9):977–981
Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS, Thorsen T, Quist H, Matese JC, Brown PO, Botstein D, Lonning PE, Borresen-Dale AL (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98(19):10869–10874. doi:10.1073/pnas.191367098
Karnoub AE, Dash AB, Vo AP, Sullivan A, Brooks MW, Bell GW, Richardson AL, Polyak K, Tubo R, Weinberg RA (2007) Mesenchymal stem cells within tumour stroma promote breast cancer metastasis. Nature 449(7162):557–563. doi:10.1038/nature06188
Diaz-Cano SJ (2012) Tumor heterogeneity: mechanisms and bases for a reliable application of molecular marker design. Int J Mol Sci 13(2):1951–2011. doi:10.3390/ijms13021951
Zardavas D, Baselga J, Piccart M (2013) Emerging targeted agents in metastatic breast cancer. Nat Rev Clin Oncol 10(4):191–210. doi:10.1038/nrclinonc.2013.29
Mahmoud SM, Paish EC, Powe DG, Macmillan RD, Grainge MJ, Lee AH, Ellis IO, Green AR (2011) Tumor-infiltrating CD8+ lymphocytes predict clinical outcome in breast cancer. J Clin Oncol Off J Am Soc Clin Oncol 29(15):1949–1955. doi:10.1200/JCO.2010.30.5037
Condamine T, Gabrilovich DI (2011) Molecular mechanisms regulating myeloid-derived suppressor cell differentiation and function. Trends Immunol 32(1):19–25. doi:10.1016/j.it.2010.10.002
Sumida K, Wakita D, Narita Y, Masuko K, Terada S, Watanabe K, Satoh T, Kitamura H, Nishimura T (2012) Anti-IL-6 receptor mAb eliminates myeloid-derived suppressor cells and inhibits tumor growth by enhancing T-cell responses. Eur J Immunol 42(8):2060–2072. doi:10.1002/eji.201142335
Morales JK, Kmieciak M, Knutson KL, Bear HD, Manjili MH (2010) GM-CSF is one of the main breast tumor-derived soluble factors involved in the differentiation of CD11b−Gr1− bone marrow progenitor cells into myeloid-derived suppressor cells. Breast Cancer Res Treat 123(1):39–49. doi:10.1007/s10549-009-0622-8
Xu X, Wang B, Ye C, Yao C, Lin Y, Huang X, Zhang Y, Wang S (2008) Overexpression of macrophage migration inhibitory factor induces angiogenesis in human breast cancer. Cancer Lett 261(2):147–157. doi:10.1016/j.canlet.2007.11.028
Acharyya S, Oskarsson T, Vanharanta S, Malladi S, Kim J, Morris PG, Manova-Todorova K, Leversha M, Hogg N, Seshan VE, Norton L, Brogi E, Massague J (2012) A CXCL1 paracrine network links cancer chemoresistance and metastasis. Cell 150(1):165–178. doi:10.1016/j.cell.2012.04.042
Resetkova E, Reis-Filho JS, Jain RK, Mehta R, Thorat MA, Nakshatri H, Badve S (2010) Prognostic impact of ALDH1 in breast cancer: a story of stem cells and tumor microenvironment. Breast Cancer Res Treat 123(1):97–108. doi:10.1007/s10549-009-0619-3
Nocito A, Kononen J, Kallioniemi OP, Sauter G (2001) Tissue microarrays (TMAs) for high-throughput molecular pathology research. Int J Cancer 94(1):1–5. doi:10.1002/ijc.1385
Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, Davies S, Fauron C, He X, Hu Z, Quackenbush JF, Stijleman IJ, Palazzo J, Marron JS, Nobel AB, Mardis E, Nielsen TO, Ellis MJ, Perou CM, Bernard PS (2009) Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol Off J Am Soc Clin Oncol 27(8):1160–1167. doi:10.1200/JCO.2008.18.1370
Cooper A, van Doorninck J, Ji L, Russell D, Ladanyi M, Shimada H, Krailo M, Womer RB, Hsu JH, Thomas D, Triche TJ, Sposto R, Lawlor ER (2011) Ewing tumors that do not overexpress BMI-1 are a distinct molecular subclass with variant biology: a report from the Children’s Oncology Group. Clin Cancer Res Off J Am Assoc Cancer Res 17(1):56–66. doi:10.1158/1078-0432.CCR-10-1417
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, Graf S, Ha G, Haffari G, Bashashati A, Russell R, McKinney S, Group M, Langerod A, Green A, Provenzano E, Wishart G, Pinder S, Watson P, Markowetz F, Murphy L, Ellis I, Purushotham A, Borresen-Dale AL, Brenton JD, Tavare S, Caldas C, Aparicio S (2012) The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486(7403):346–352. doi:10.1038/nature10983
Acknowledgments
The authors would like to thank the members of the Oncogenomics Core Facility, Flow Cytometry Core Facility, and the Division of Veterinary Resources at the University of Miami Miller School of Medicine for their assistance during the course of the study. The authors would also like to thank Nanette Bishopric, MD and Barry I. Hudson PhD for helpful discussion of the manuscript. This Project was funded by Breast Cancer Research Foundation (BrCRF) awards to MEL and JMR. Part of these studies was conducted at the Lombardi Comprehensive Cancer Center Histopathology and Tissue Shared Resource which is supported in part by NIH/NCI Grant P30-CA051008. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Cancer Institute or the National Institutes of Health.
Conflict of interest
All authors have no conflicts of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Additional information
Katherine Drews-Elger and Elizabeth Iorns have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Drews-Elger, K., Iorns, E., Dias, A. et al. Infiltrating S100A8+ myeloid cells promote metastatic spread of human breast cancer and predict poor clinical outcome. Breast Cancer Res Treat 148, 41–59 (2014). https://doi.org/10.1007/s10549-014-3122-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-014-3122-4