Abstract
It is often thought that non-junk or coding DNA is more significant than other cellular elements, including so-called junk DNA. This is for two main reasons: (1) because coding DNA is often targeted by historical or current selection, it is considered functionally special and (2) because its mode of action is uniquely specific amongst the other actual difference makers in the cell, it is considered causally special. Here, we challenge both these presumptions. With respect to function, we argue that there is previously unappreciated reason to think that junk DNA is significant, since it can alter the cellular environment, and those alterations can influence how organism-level selection operates. With respect to causality, we argue that there is again reason to think that junk DNA is significant, since it too (like coding DNA) is remarkably causally specific (in Waters’, in J Philos 104:551–579, 2007 sense). As a result, something is missing from the received view of significance in molecular biology—a view which emphasizes specificity and neglects something we term ‘reach’. With the special case of junk DNA in mind, we explore how to model and understand the causal specificity, reach, and corresponding efficacy of difference makers in biology. The account contains implications for how evolution shapes the genome, as well as advances our understanding of multi-level selection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
How to understand biological function has long been a subject of both philosophical and scientific debate focused on evolutionary biology. Philosophical assessments of biological function have often focused on whether function needs to be understood as a past product of natural selection (Wimsatt 1972; Millikan 1984; Neander 1991) or is aptly described by just what biological causes presently do (Cummins 1975; Hinde 1975; Boorse 1976). The former style of “selected effects” approach is historical—or “etiological,” in philosopher-speak. The latter style of “causal role” approach is often ahistorical but—at least in current biological discourse—nevertheless typically assumes that natural selection should be operating on these entities in the present (Allen and Neal 2020).Footnote 1
In the philosophy of biology, there is also a notion of “actual difference makers” as special targets of biological explanation and investigation (Waters 2007). According to this approach, functional genomic entities like certain stretches of DNA are actual difference makers—since different actual “values” for these causal “variables” (i.e., different sequences of the “same” stretch of DNA among various organisms within a population) can and do generate different actual “values” for intervened-on “variables” (such as RNA and protein) in that population, which can sometimes lead to different actual phenotypic and even fitness “values” for the different organisms in that population.
Here we demonstrate that there is another class of actual difference makers in biology: one which has often been overlooked in discussions of function, role, and import (likely due to the commonly conjoined emphasis on fitness and selection). These are genomic entities whose presence or absence alters the character of the cell and are thus influential—despite that these entities may not have been selected for, and that mutations in these regions do not necessarily affect the fitness of the organism. The presence of these entities generates cellular work, and their complete deletion would likely decrease the fitness of the organism; but their alteration by the typical mutation does not often affect fitness—at least not to the extent that such mutations are detected, eliminated, and/or preserved by organism-level natural selection. One main difference between these genomic regions and typically coding DNAFootnote 2 is not their causal specificity but rather their reach (or lesser extent thereof) and hence their efficacy (both terms described in more detail later in the text). Our aim is to provide a characterization of these overlooked causal entities and relationships, using junk DNA as an illustrative case.
Junk DNA
It is widely agreed (Ponting and Hardison 2011; Rands et al. 2014; Ponting 2017) that approximately 90% of the human genome is not subject to purifying selection and hence—at least according to typical accounts of function in molecular biology—is not functional. Despite this, we argue that paradigmatically “non-functional” DNA, commonly referred to as junk DNA, is still a difference maker (in Waters’ [2007] sense). The fact that there are entities which matter (as they affect cell physiology), yet which are not necessarily sculpted by organism-level natural selection, has profound implications for the evolution of humans, and multicellular life in general.
Junk DNA represents a large fraction of all eukaryotic genomes. Its existence is due to the fact that eukaryotic genomes contain selfish DNA entities (also known as transposable elements), which have the ability to replicate themselves (Doolittle and Sapienza 1980; Orgel and Crick 1980). It is believed that transposable element activity fueled the expansion of the eukaryotic genome. In all eukaryotic genomes examined thus far most transposable elements have accumulated inactivating mutations, with only a minority still having the ability to replicate. These dead transposable elements are a vast graveyard that constitutes much junk DNA.
One critical factor that permitted the expansion of transposable elements, and thus junk DNA, is the nucleus (Martin and Koonin 2006; López-García and Moreira 2006; Palazzo and Gregory 2014). The nucleus effectively divides the cytoplasm into two distinct compartments: the nucleoplasm, where DNA is transcribed into premature messenger RNAs (pre-mRNAs), which are then spliced to form mature messenger RNAs (mRNAs); and the cytoplasm, where these fully processed mRNAs are translated into proteins.Footnote 3 The segregation of mRNA processing machinery away from mRNA translation machinery has two main effects. First, this segregation ensures that for any given mRNA, its splicing is completed before it is allowed to be exported from the nucleus to the cytoplasm (Martin and Koonin 2006). This, in-and-of-itself, has several advantages. It ensures that pre-mRNA is free of translating ribosomes, which would likely collide into—and interfere with—the splicing machinery, and it prevents unprocessed pre-mRNAs from being translated into proteins (many of which are toxic). Hence, this segregation of RNA processing and translation increases the efficiency and fidelity of these two processes. Second, the nucleus/cytoplasmic divide acts as a quality control device. In most eukaryotic systems studied, it has been observed that unprocessed pre-mRNAs and misprocessed mRNAs are not efficiently exported to the cytoplasm, but instead retained in the nucleus and degraded by dedicated RNA decay machinery (Palazzo and Lee 2018). This severely reduces the translation of misprocessed mRNAs, whose protein products are potentially toxic.
So, what does this have to do with junk DNA? Well, DNA in general is a non-specific substrate for RNA polymerase. As a result, intergenic DNA is transcribed at a low level into junk RNA, also known as ‘transcriptional noise’ (Struhl 2007; Palazzo and Gregory 2014; Palazzo and Lee 2015). Like misprocessed mRNAs, most of this junk RNA is retained in the nucleus and eliminated by RNA decay machinery. This makes sense, since if this transcriptional noise was not eliminated and instead allowed to be translated into junk protein, then it would become a significant burden on the organism (Palazzo and Lee 2015). The exposure of the ribosome to misprocessed mRNAs and intergenic RNAs is very toxic in tissue culture cells (Ogami et al. 2017). By retaining junk RNA in the nucleus, and promoting its decay, junk DNA becomes less of a liability. In a sense, the nucleus and all of its associated quality control machinery acts as a global mechanism to suppress the deleterious effects of rampant mRNA mis-processing and transcriptional noise that is inherently present in cells with large junky genomes (Warnecke and Hurst 2011; Rajon and Masel 2011, 2013; Koonin 2016).
Eukaryotic cells have additional mechanisms to reduce the deleteriousness of junk DNA. For example, excess DNA is usually packaged into heterochromatin, which prevents it from being transcribed robustly, thus further limiting the amount of junk RNA that is produced.
Overall, these many processes in eukaryotic cells buffer against the deleteriousness of junk DNA. But these buffering systems are a double-edged sword: by relieving the selection pressure on organisms to get rid of this excess DNA, they also permit junk DNA to accumulate. It is thus possible that the amount of junk and the efficiency of systems that render it less harmful could have arisen in parallel as a sort of evolutionary arms race.
In addition, it is likely that many of these buffering systems are somewhat tuned to the amount of junk DNA they must contend with. Cells with large amounts of junk DNA will likely acquire secondary adaptive changes to help their quality control systems to better deal with this increased load. In addition, these quality control systems have also been co-opted to regulate the functional portion of the genome. For example, RNA decay machinery is also required to properly trim precursor ribosomal RNA to its mature form (Zinder and Lima 2017). Thus, a certain level of junk DNA becomes necessary for the entire system to work properly. In the absence of a certain level of junk RNA, RNA decay machinery will likely start to degrade functional RNAs (Doma and Parker 2007; Garland and Jensen 2020; Wang and Cheng 2020). Similarly, in the absence of junk DNA, proteins that silence this junk by packing it into heterochromatin will likely start to silence functional DNA.
Note that, within this context, junk DNA neither historically evolved for a specific function; nor is it currently being selected for or against for organisms.Footnote 4 According to the predominant accounts of “function” in biology, junk DNA has no function (for the organism). Yet its presence matters for an historical and comparative understanding of evolution. For instance, this type of evolution—one not necessarily directed by natural selection—leads to real differences between diverse branches of the evolutionary tree. Over long stretches of evolutionary time, different organism types have accumulated diverse sets and amounts of these non-functional (yet influential) entities (Gregory et al. 2007). These different entities contribute to features which permit different cellular “life” styles, which can contribute to multi-level selection (Vinogradov 2004; Gregory 2005b). In sum, the existence of junk DNA changes many basic features of the cell. Since these features affect cell physiology, junk DNA is a difference maker. We elaborate on what this means in the next section.
Difference makers
In 2007, the philosopher of science C. Kenneth Waters identified the class of “actual difference makers” as the causes that actually make a difference (Waters 2007). Waters offered this contribution to the extensive literature on causality in order to resolve a stubborn puzzle from the historical study of causation. The puzzle is traditionally framed by asking the following question: when someone strikes a match (say, to light a fire), why do we attribute the cause of the fire-lighting-event to the striking of the match, as opposed to the availability of oxygen (or some other, equally crucial element of the causal situation)?
Prior to Waters’ (2007) contribution, the received philosophical view of this type of situation had been—since the 1843 publication of John Stuart Mill’s A System of Logic—one of causal parity. According to Mill, all causes are ontologically equal, or on a par (hence the term ‘parity’ for this sort of view). On Mill’s view, there is no substantial difference between the striking of the match and the availability of oxygen. This position is at odds with our common, intuitive sense that the actual cause of the lit fire is the match-striking-event, and that other elements such as the presence of oxygen are mere background conditions (or some such). Post-Millian philosophical attempts to explain our tendency to pick out one cause among many have tended, as Waters (2007) details, to point to our interests as the cause of our commonsense beliefs in causal priority rather than parity (Hart and Honoré 1959; Mackie 1974). Believing something like “sure, oxygen is in some sense a cause of the fire lighting, but striking the match is the real cause” is a view which attributes causal priority to one cause (striking the match) over others (such as the presence of oxygen). Here, one cause is being thought of as more significant than others (hence the term ‘priority’ for this sort of view).
Waters develops an account designed to pick out commonly emphasized biological causes, such as differences in the structure of DNA (when it varies in ways which generate variation in other variables such as RNA and other molecules). He then explains the perceived significance, or priority, of this DNA relative to other causal elements, such as the structure of accessory proteins (when these do not tend to vary in ways which generate corresponding variation in other variables). Waters accomplishes all this with the help of a trio of distinctions:
1 – potential difference makers (causes whose variables could vary in a population to make a difference in effect) versus actual difference makers (causes whose variables do vary in a population to make a difference in effect). For example: if there was variation within a population among the structures of one of its protein-folding accessory proteins, and that variation generated corresponding variation in protein conformation, then that protein-folding accessory protein would be an actual difference maker in that population. But in the absence of actual variation among the structures of that protein-folding accessory protein generating actual variation in protein expression within a population, that molecule is merely a potential rather than an actual difference maker in that population.
2 – an actual difference maker (working to generate population-based differences in effect, together with other actual difference-making causes) versus the actual difference maker (working to generate population-based differences in effect, without any other actual difference-making causes). Waters points to RNA synthesis in bacterial cells as an example of the latter kind of case—one in which only activated DNA segments act as the actual difference maker in this process, since only that “variable” takes on different “values” for different cells within the relevant population. Although accessory proteins like RNA polymerase are (of course) required for RNA synthesis in bacterial cells, RNA polymerase is not in this case a “variable” which takes on different “values” in this population, in the sense of individual bacterial cells expressing structurally distinct versions of RNA polymerase (or other protein-folding accessory proteins) which then correspondingly generate differences in synthesized RNA. (RNA polymerase is still a potential difference maker, though, since different bacterial cells within a population could presumably contain structurally distinct versions of RNA polymerase which could then presumably affect instances and rate of RNA synthesis.) Alternatively, Waters points to RNA synthesis in eukaryotic cells as an example of the former kind of case—one in which activated DNA segments act as an actual difference maker along with RNA polymerases, since in this more complicated context there are different kinds of RNA polymerases and “presumably, different accessory molecules are also associated with the synthesis of different RNA molecules” (Waters 2007, p. 574).
3 – causally specific difference makers (causal variables whose values can be distinctly many, variation among which corresponds to distinctly many values in effect-variable) versus causally non-specific difference makers (causal variables whose values are not distinctly many, or whose distinctly many values do not correspond to distinctly many values in effect-variable). Woodward (2010) explicates these notions with a contrast between fine-grained (or dial-like) and coarse-grained (or switch-like) control. To listen to a particular station on your radio, you need to both flip the on/off switch and turn the dial to the appropriate frequency. But only turning the dial gives you fine-grained control over which station you might receive, and there are distinctly many positions on that dial which each correspond to distinctly many stations to which you might listen. Obviously, moving the on/off switch to the correct position is required in order to listen, but “the switch is not causally specific with respect to which program is received” (Woodward 2010).
The upshot of all this conceptual machinery—for biology as it is being practiced, and even more specifically for genetics—is that it provides a potential justification for judgments of the causal priority of coding DNA. Therefore, according to Waters, “it makes sense for biologists to say that DNA is not on a causal par with many of the other molecules that play causally necessary roles in the synthesis of RNA and polypeptides” (2007, p. 579). This stance—that of assigning causal significance, or priority, to DNA over other molecules—is indeed a familiar one in biology. And it is explained, according to Waters (2007), by the fact that DNA is an actual as opposed to a merely potential difference maker. Additionally, although DNA is an actual difference maker, among several others (it is not the only actual difference maker), of those assorted actual difference makers, DNA is the causally specific one (its difference-making is not merely causally non-specific.Footnote 5
Junk DNA as an actual difference maker
A surprising thing about this account of causal priority is that it leaves all “activated” DNA on a par with one another—including junk DNA. Waters writes: “An important actual difference in cells is the difference in the nucleotide sequences of RNA molecules synthesized in a cell” (2007, p. 573). As discussed in the “Junk DNA” section, both coding and junk DNA are transcribed in cells; both make for actual differences in the nucleotide sequences of RNA molecules synthesized in cells; both generate causally specific differences in the sequences of RNA molecules synthesized in cells. Any stretch of transcribed DNA which varies among individuals within a population—transcription of which generates corresponding variation in the RNA synthesized among individuals in that population—is going to look, on Waters’ account, like an important actual difference maker in that population. Given what has recently been learned about widespread transcription activity, this includes junk DNA.
Ironically, since junk DNA is likely subject to less (or even zero) purifying selection relative to coding DNA, junk DNA will often tend to vary more among individuals in a population than does coding DNA, and thus junk DNA might even turn out to be more causally specific than coding DNA on Waters’ view. To apply Woodward’s toy example from above, if we think of a stretch of DNA like a radio dial that can be tuned to different stations (where each station is an instance of phenotypic variation existing in the population), then there are likely going to be more stations available for tuning to, on a dial corresponding to a stretch of junk DNA, than on one corresponding to a stretch of coding DNA. A stretch of junk DNA is a causal variable which is likely to take on more distinct values among individuals in a population than is a stretch of coding DNA, and to thereby correspond with more distinct values of the synthesized RNA effect-variable.
But—as discussed above, and contingently so—biologists and philosophers of biology do not typically view causally specific differences generated by actual differences in junk DNA as on a causal par with causally specific differences generated by actual differences in coding DNA. Although Waters’ account of causal specificity goes some way towards explaining why DNA has been causally elevated relative to other actual difference makers (such as RNA polymerase), it does not explain why coding DNA has been causally elevated relative to junk DNA. Junk DNA has the same two features which Waters has picked out as the distinguishing features of coding DNA: actual difference-making and causal specificity. Yet junk DNA is not on a causal par with coding DNA in biology.
Something is missing from Waters’ account; something related to causal specificity but involving additional concepts we hereby term ‘causal reach’ and ‘causal efficacy’.Footnote 6 One obvious, crucial difference between coding and junk DNA is that, although each of these causal variables generate both proximate and distal effects, not just the immediately proximate but also some of the rather more distal effects of coding DNA can be nonetheless quite causally specific. Although junk DNA has both proximate and distal effects, too—and junk DNA often has casually quite specific proximate effects—the more distal effects of junk DNA tend to be significantly reduced in causal specificity, compared with some of those of coding DNA. So, even taking on board much of Waters’ framework for comprehensively understanding the causal priority typically afforded coding DNA, we need to track not just the specificity of actual difference-making causes in biology, but also the extended (or not) reach of these causes, as well as the maintenance (or not) of that specificity throughout the reach of their effects—i.e., the complex combinatorial feature of causal efficacy.Footnote 7
See Fig. 1 for a conceptual depiction of various causes with distinct combinations of specificity, reach, and corresponding efficacy. Causal specificity has already been explained; causal reach has to do with how many effects a single, initiating cause generates.Footnote 8 These effects can: extend straightforwardly out in a linear chain from the initiating cause; scatter out in a burst from the initiating cause, or both scatter and extend in a cascade from the initiating cause. Explained in engineering terms, causal reach can occur via parallel processing (multiple effects radiating out from one causal node), in sequence (a series of proximate and distal effects), or by combination of these two modes.
Various causes with distinct combinations of specificity, reach, and corresponding efficacy. Top left, an utterly inefficacious cause with no effects. Bottom right, an extremely efficacious cause—one that has both reach and specificity in terms of its effects. Reach is indicated by number of nodes—where each cause can have either unitary or multiple effects, both proximate and ultimate. Causal specificity is indicated by effect opacity—where each node can either transmit its full pigmentation or produced a faded node with increased transparency. Within rows, causes increase in specificity from left to right; within columns, causes increase in reach from top to bottom. Hence, top right has more specificity though less reach relative to bottom left, which has more reach though less specificity. Both are less efficacious than bottom right, but more efficacious than top left; neither are necessarily insignificant causes. (Prepared by Adam Streed using the matplotlib library for Python 3; all the code is his own)
Causal efficacy tracks interaction of the two prior notions of specificity and reach. This concept is required to monitor the effects a difference maker has—as the effects of that difference maker extend and cascade (or not) and given the causal specificity (or not) of those resulting effects. A causally abrupt, non-specific difference maker is by its nature not very efficacious. A difference maker with extensive reach can nonetheless be rather causally inefficacious if its many “reach” effects are mostly non-specific. Likewise, a highly causally specific difference maker can nonetheless be rather causally inefficacious if it does not have much reach in terms of its effects. A difference maker with both extensive reach and specificity in the effects it causes will be extremely efficacious.
See Fig. 2 for a conceptual depiction of various genetic causes with distinct combinations of specificity, reach, and corresponding efficacy. So, one likely reason coding DNA is considered causally so significant by biologists because it can have quite the causal reach, and not just its proximate but often also its more distal effects can be quite causally specific. Note, however, that not all causally specific changes to coding DNA are on a par with one another—in terms of their causal reach, corresponding efficacy and resulting effect on judgments of causal priority.Footnote 9
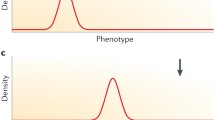
Various genetic causes with distinct combinations of specificity, reach, and corresponding efficacy. a A synonymous, conservative mutation in any transcribed DNA—coding or junk. b A synonymous, semi-conservative mutation in coding DNA. c A non-synonymous, semi-conservative mutation in coding DNA—one that produces a change in protein amino acid sequence but no change in conformation. d A non-synonymous, non-conservative mutation in coding DNA—a genotypic change with phenotypic and other ultimate effects. e Insertion or deletion of a small “repeat” segment of junk DNA. f Insertion or deletion of a large “repeat” segment of junk DNA—one which ultimately produces an uptick in RNA decay machinery. g Insertion or deletion of a large, unique (not a repeat) segment of junk DNA—the size rather than uniqueness of which also produces an uptick in RNA decay machinery. h Insertion or deletion of another large, unique segment of junk DNA—one that produces in virtue of its size and uniqueness not just an uptick in RNA decay machinery but also other downstream, causally non-specific distal effects. Explication of these cases continues in the main text below (Prepared by Adam Streed using the matplotlib library for Python 3; all the code is his own)
A synonymous, conservative mutation in transcribed DNA (e.g., a change from a GGC codon to GGG), although it will produce a causally specific change in synthesized RNA, will not necessarily produce any further changes of note—even if protein synthesis occurs (since GGC and GGG both code for glycine; Fig. 2a).Footnote 10 However, even synonymous mutations in coding DNA can turn out to be not perfectly conservative (i.e., they can be productive of further, causally non-specific effects; Fig. 2b).Footnote 11 Non-synonymous mutations in coding DNA can also vary in terms of the (semi-) conservativeness of their effects. A replacement event which introduces one hydrophobic amino acid for another during protein expression, or one positively charged amino acid for another (Fig. 2c), is likely to be more conservative in its changes than those events which substitute a hydrophobic amino acid for a charged one, or a negatively charged amino acid for a positively charged one. A non-synonymous, non-conservative mutation in coding DNA which introduces a stop codon in a new place will likely affect more than just one amino acid of a protein’s primary sequence (it may cut off several) and could affect not just genotype but also phenotype (Fig. 2d). All these changes (Fig. 2a–d) are proximately causally specific, but they vary in reach, as well as in the specificity of their more distal effects (i.e., their causal efficacy). The fact that these changes also vary in perceived importance (ascending from a to d) shows that not just causal specificity but also causal reach and efficacy are features relevant to judgments of biological importance.Footnote 12
The relevance of causal reach and efficacy to judgments of biological importance becomes even more apparent when considering potential changes to junk DNA. Of course, many changes to junk DNA are unimportant from this point of view. Insertion or deletion of a small, “repeat” segment of junk DNA can fail—even when transcription occurs—to produce any causally specific effects. Such insertions or deletions are likely—if they alter anything at all—to change only the amount or proportion of various segments of junk RNA already synthesized in the cell (since they will neither introduce an entirely new kind of segment nor entirely remove any kind of previously-existing segment; Fig. 2e). To re-apply and extend the radio metaphor from Woodward (2010), this is something like turning up or down the volume on a station to which the cell is already tuned. Insertion or deletion of a larger but still “repeat” segment of junk DNA might produce additional causally non-specific effects further downstream, e.g., an uptick in RNA decay machinery (described in “Junk DNA” section), via sizable alteration of the amount of junk DNA transcribed in the cell (Fig. 2f). Alternatively, insertion or deletion of a large, unique (not a repeat) segment of junk DNA might have causally specific effects on the types of junk RNA that are synthesized in the cell (since its uniqueness means that it will, if that stretch of genome is transcribed, either introduce or eliminate expression of a segmentFootnote 13), but could also have causally non-specific distal effects on, e.g., the amount of RNA decay machinery, via its sizable alteration of the amount of junk RNA being synthesized (Fig. 2g). Finally, insertion or deletion of a large collection of junk DNA segments could have other cascading, causally non-specific distal effects—such as altered patterns of interaction with RNA decay machinery, changes in cellular size, or degradation of functional RNA (introduced in “Junk DNA” section, additional discussion in “Detecting the hidden significance of junk DNA” section; Fig. 2h). As before, these changes tend to vary in perceived biological significance (ascending from e to h).
Our presentation of these cases reflects the fact that the relative significance of junk DNA, when it is significant, does not usually stem from its occasional uniqueness (and corresponding causal specificity). The diagram also helps us to demonstrate, given the reduced causal specificity of this bottom set of cases (Fig. 2e–h) relative to the prior top set (Fig. 2a–d), why coding DNA is considered causally so significant by biologists—because it can have quite the causal reach, and not just its proximate but often also its more distal effects can be quite causally specific. Coding DNA can be a highly efficacious cause, and this reliably leads to judgements of elevated causal importance. But we can further see, given the significant causal reach of some of these cases (Fig. 2d and h) relative to others (Fig. 2a–c and e–g), how not just causal specificity but also reach might play a role in judgements of causal importance. Although junk DNA may rarely be a highly efficacious cause (due to its diminished distal causal specificity), it can still have reach. The conceptual machinery of causal specificity, reach, and corresponding efficacy gives us the tools to understand the utmost causal importance of certain kinds of changes in coding DNA, the relatively reduced in importance but still non-negligible role of certain other kinds of changes in junk DNA, and the common occurrence of trivial sorts of changes in either—even from a relatively restricted, emphasis-on-selection-at-the-level-of-the-organism style of view.
Obviously, we have not exhausted all the diverse ways that either coding or junk DNA can causally affect change (or fail to do so).Footnote 14 We posit that junk DNA, and other non-functional (in the selectionist sense) components of the cell, are a significant part of the evolutionary story of eukaryotes; low-efficacy causes can still be significant. We suspect that the diminished causal efficacy of these components has often made it difficult for either biologists or philosophers to appreciate and characterize their significance, despite their reach.
From the point of view of organism-level selection, junk DNA may have arrived in cells in a non-adaptive manner (at the organismal level), and typical mutations (such as small nucleotide polymorphisms) in a nucleotide sequence of junk DNA may not generally be selected for or against (at the organismal level), but mechanisms within the cell have generally adapted to its presence. In the context of this evolutionary interplay, the presence and potency of the total sum of junk DNA in the cell dictates the need for various cellular machineries of “quality control,” changing the resources required and expended by the cell, and committing the cell to particular strategies. Significant changes to either party (the junk DNA or the cellular clean-up crew) can produce causally significant though non-specific effects. We detail these possibilities, and how to detect them, in the next section.
Detecting the hidden significance of junk DNA
One way to illustrate the difference between significant but not functional (in the selections sense) and functional biological entities is to explore how these entities as a whole might affect the fitness of an organism: to explore their hypothetical, collective effect. In theory, this could be done. In practice, the amount of resources that would be required to carry out the full experiment may not be feasible. First, we would need to pick a genome. Next, we would need to unambiguously identify non-functional DNA by deleting small segments of the genome. If we chose to investigate the human genome, the deletion size that we would use would be similar to mutations that naturally occur in human populations, since it is precisely these types of mutations that are seen by natural selection. Any deletions that did not affect the number of offspring could be tagged for further investigation. Then, we would combine these deletions in increasing amounts and evaluate how these affect the number of offspring of the resulting organism.
Cataloguing functional versus non-functional roles
If given unlimited resources and time, we could in principle test whether each segment of the chosen genome is functional. Any region of DNA can play a causally specific, sequence-dependent role or a causally non-specific, sequence-independent role.
To test whether any given region of DNA has a sequence-dependent role, we could introduce nucleotide substitutions. However, this would not test whether the region in question has more non-specific roles that contribute to its reach. For example, it is known that 5′ untranslated regions have to be a minimal length—if they are any shorter than this minimum, the translational start site will not be reliably recognized and the mRNA will not be properly translated (Kozak 1991). Thus the 5′ untranslated region acts as a spacer, one that can be quite important—since even from a perspective that emphasizes organismal-level selection, it is a physical entity that contributes to the cellular environment.
To test whether any given region DNA has a sequence-independent role, we could systematically delete these regions.Footnote 15 Assessing the rate of insertions or deletions (indels) between organisms has been used to determine that most annotated non-coding transcripts produced in mammalian cells are non-functional, although these analyses are statistical in nature (Ponting 2017). To precisely assess each individual region, we would have to empirically test it. We would begin by deleting as much DNA as what occurs in natural mutations, since these alterations are the ones on which selection operates. The size of the average deletion can vary quite considerably, typically because those regions of the genome which contain repeat elements are quite prone to large alterations. In non-repeat regions it has been estimated that 96% of all deletions are smaller than 16 base pairs in length (Mullaney et al. 2010). In repeat regions, spontaneous deletions are more common and can be larger. Although very long deletions are rarer, they can accumulate in lineages over generations, and as a result the size of any given repeat region varies considerably in the human population, with many of these deletions being tens of thousands of base pairs in length (Jeffreys et al. 1985; Redon et al. 2006). With this in mind, we would begin our experiment by deleting ~ 100 base pair segments in parts of the genome that show no sequence conservation between humans and other mammals, which represents about 95% of the human genome. It is likely that some of this non-conserved DNA plays a lineage specific-role in humans (i.e., has been under selection in the human lineage but is not conserved between humans and related species, such as chimpanzees), although most estimates place this at less than 5% of the genome (Ward and Kellis 2012; Rands et al. 2014; Gulko et al. 2015; Ponting 2017).
To perform this experiment in an informative manner we would need to track the reproductive success of thousands of individuals having the same mutation under very controlled conditions to infer whether a given deletion was slightly deleterious. Thus, in practice, the effect of slight changes to fitness would be almost impossible to assess. Nevertheless, in theory, this experiment could be done.
Testing the boundary between non-functional and functional roles
With all the functional and non-functional DNA cataloged (though only from the organism-level-selection point of view), we could then move on, in a rather interesting way, to determining how consequential each region of so-called junk DNA might be. We could combine previously tested deletions of junk DNA together into unnatural mutations (i.e., mutations not typically seen in nature, and thus not subjected to selection pressures) and determine their effect on reproductive success.
To reiterate, this experiment would not test whether these sequences are functional (in the purifying selectionist sense), as selection would never encounter these types of combinatorial mutations. Instead, this would be one means of testing whether given regions of junk DNA are influential (i.e., whether they are difference makers). The degree to which deletion mutations would have to be combined before they affect fitness (i.e., reproductive success) might indicate how (in)efficacious these regions are as difference makers.
We expect that quite a large number of deletions must be combined before we would be able to detect any effect on fitness. Indeed, it has been seen that deleting 1.5 Megabases of non-coding DNA from the mouse genome (approximately 0.06% of the entire genome) had no measurable effect on the fitness of the animals (Nóbrega et al. 2004). How much of the junk DNA genome would need to be eliminated before measurable effects can be detected is hard to estimate at this time.
Nevertheless, there would likely come a point where the diffuse effects of deleting many junk DNA regions, when combined, would become detectable and significant—and that this would show the cumulative “difference making” effect of the presence of junk DNA (relative to its absence), along with the corresponding cellular adjustment to its usual presence (relative to its absence). We might expect that quite a sizable amount of junk DNA would need to be removed before we would be able to measure any effects on fitness.
Of course, in practice there would be additional complications. Different genomic regions may play redundant roles, thus elimination of one or the other may have little to no fitness effect, while deletion of both will have a large impact on fitness. This combination would not test the diffuse effects of junk DNA, but rather uncover much more specific effects that were previously hidden. Untangling these two possibilities may be almost impossible.
Non-functional yet significant roles for junk DNA
So, what would the massive elimination of junk DNA change? Alternatively, what non-functional roles does it normally play? Below we list some of the causally non-specific yet nonetheless significant effects which can make junk DNA consequential for the cell.
Cell size
It is well known that DNA content correlates with the size of the cell nucleus, which in turn correlates with the total cell size (Cavalier-Smith, 1978; Gregory, 2001). It has thus been inferred that DNA content affects cell size. Thus, massive reductions in the amount of junk DNA would lead to a reduction in cell size. This would fundamentally change many aspects of cell physiology. First, as cell size changes, surface areas and volumes do not scale with one another in a linear fashion. As cells become smaller, the ratio of surface area/volume increases. An increase in cell membrane/volume ratio would allow molecules to diffuse more rapidly into and out of cells, and thus allow for greater rates of metabolism. It is widely believed that the small genome size of both birds and bats, and the accompanying increase in metabolic rate, was a necessary prerequisite for the evolution of flight (Hughes and Hughes 1995; Gregory 2002). Indeed, even within birds, those which have the smallest genomes and have the most rapid metabolism are the hummingbirds and those with the largest genomes are flightless birds (Gregory et al. 2009). Interestingly, in the avian lineage it appears that genome size reduction happened before the evolution of flight as cell size estimates for closely related non-avian dinosaurs were found to be quite small (Organ 2007).This suggests that there was no initial selection pressure to reduce genomes due to metabolic constraints associated with flight, but rather that the organisms that happened to have small genomes had the metabolic prerequisites that were necessary to evolve flight. Subsequently, it appears that bird genomes have stayed small due to occasional large genomic deletions, which are possibly driven by positive selection (Kapusta et al. 2017).
The increase of surface area/volume as the genome contracts is true not only for the cell (cell plasma membrane/cell volume) but also for some of the organelles, for example the nucleus. Other organelles, such as mitochondria, would most probably remain the same. These alterations will have their own consequences. For example, in the nucleus, newly made mRNAs are known to randomly diffuse from their site of production to the nuclear pore (Shav-Tal et al. 2004), thus the larger the nucleus, the more time it will take for newly made mRNAs to reach the cytoplasm where they are translated into proteins. If we compare the nuclear export rate (t1/2; time it takes for half of the pool of mRNA to be exported from the nucleus to the cytosol) for the exact same mRNA (in this case the ftz reporter mRNA), it is 2 h in frog oocytes (Luo and Reed 1999), whose nuclei have a radius of 200 µm; but only 15 min in mammalian cells (Palazzo et al. 2007), whose nuclei have a radius of about 4 µm. Nuclear export rates (t1/2) are 3–5 times faster in yeast, whose nuclei have a radius that is smaller than a micrometer (Oeffinger and Zenklusen 2012). The nuclear export rate of mRNA is one of the main determinants of the lag time between gene activation and protein production, and can affect biological processes such as the timing of circadian rhythm as well as how cells respond to their environment (Hoyle and Ish-Horowicz 2013).
Besides gene expression timing, it is likely that other cellular processes would also be affected by cell size (Cavalier-Smith 1978). It is not entirely clear how drastic changes in cell size would affect the development and the proper function of tissues and organs, but it is reasonable to expect that many of these may become dysregulated. Note that we are not claiming that cell size could never undergo a drastic change due to these constraints. It is likely that if an organism gradually lost DNA over long periods of time, this would be accompanied by other adaptive changes which would compensate for these alterations in cell physiology. But it is likely that a drastic reduction of cell size, without all the accompanying adaptive changes, would lead to a fair number of problems.
As is the case with birds and bats, it is likely that drastic changes in cell size—and the accompanying adaptive changes to compensate for changes in cell physiology—could alter the organism in such a way that it could open up or limit the ability of the organism to occupy certain niches. In insects and vertebrates it has been documented that certain aspects of development and cell physiology are restricted to animals with a limited range of genome sizes (Gregory 2005b). It is also worth noting that in comparison to animals, plants—whose development and organ function appears to be more plastic—more often experience whole genome duplication events which effectively double their cell size. In animals, whole genome duplication events are rarer and seen most in amphibians (frogs, axolotls)—a class of animals that tend to have large genomes, and thus may have extensive buffering capacities. Again, this may indicate that drastic changes in cell size may only be compatible with certain types of cell physiology. The idea that genome size is consequential due to its effects on cell size has been advanced previously; however, unlike previous commentators (e.g. Cavalier-Smith 1978), we believe that the typical indel mutations that alter cell size are not subject to selection, as these indels are so small (see “Cataloguing functional versus non-functional roles” section) that each contributes negligibly to the overall dimensions of the cell. There may be some species that have enough variation in genome size for it to be under selection, but these appear to be the exceptions rather than the rule (Blommaert 2020).
Timing of cell division
It has been noted that cells with larger genomes take longer to divide. Superficially this makes sense as one would expect it would take more time to replicate the increased amount of DNA. However, this can easily be remedied by increasing the number of DNA polymerases in the cell, and also increasing the origins of replication which would allow all the extra polymerases to have access to the DNA. It thus remains unclear how genome size impacts cell division timing.
Regardless of the reasons, the timing of cell division will have drastic effects on the organism’s ability to grow, generate new tissue, and heal after injuries. It is not hard to see that changing this timing would have a major effect on how the organism develops and its overall fitness.
Providing excess substrates for cellular machineries
As we discussed in “Junk DNA” section, junk DNA also provides additional, non-specific substrates for enzymes in the cell to engage with. All enzymes have a preferred substrate, but they will act on non-optimal substrates as well, especially when subjected to non-adaptive evolutionary pressures (Khersonsky and Tawfik 2010; Bar-Even et al. 2011; Tawfik 2020; Copley 2020). Normally the amount of activity associated with these non-optimal substrates is negligible; however, when the amount of non-optimal substrate is in vast excess of the preferred substrate, non-optimal reactions can become substantial. This is true for RNA polymerase enzymes (Struhl 2007), which transcribe DNA into RNA. It is also true of DNA binding proteins (Villar et al. 2014), both those that recognize certain motifs to regulate gene activation, and those that package DNA non-specifically. The easiest solution to deal with an excess of non-optimal substrates is to increase the number of proteins so the excess substrate binding does not interfere with its normal selected activity. In other words, the amounts and activities of all these enzymes are under selection to be able to accomplish their job in a cellular environment filled with excess non-specific substrates: junk DNA and junk RNA.
For other enzymes, their preferred substrates are junk DNA and junk RNA. This is especially true for RNA decay enzymes, since these generally operate with a certain load of junk RNA that they must constantly eliminate. A drastic drop in junk RNA would mean that these enzymes would work more-and-more on their non-optimal substrates, in this case functional RNAs. This “kinetic competition” in RNA quality control systems has been well documented (Doma and Parker 2007; Garland and Jensen 2020; Wang and Cheng 2020). Other proteins that fall in this class are DNA binding proteins that package and silence junk DNA into heterochromatin.
It is possible that many of these enzymes are regulated by feedback loops that “sense” how much excess substrates there are. A large decrease in the number of excess substrates could lead to a decrease in the levels or activities of these enzymes. This, however, may severely impair the proper functioning of enzymes that have both junk RNA, and functional RNAs, as their optimal substrates. For example many RNA decay enzymes also help to trim ribosomal RNAs and other functional non-coding RNAs (Zinder and Lima 2017). Thus, a drastic reduction in junk RNA, if it activated a feedback loop to decrease total RNA decay capacity, may inadvertently diminish the cell’s ability to properly process functional RNAs.
Other (even) more speculative effects
There are possibly other general aspects of the cellular environment that would be altered if a substantial amount of junk DNA were removed.
In eukaryotic cells, long stretches of DNA that contain numerous genes adopt complicated loops, commonly referred to as “topological associated domains” (TADs). There has been much speculation about the importance of these structures (Szabo et al. 2019; Ghavi-Helm et al. 2019; Beagan and Phillips-Cremins 2020). In some cases, these are thought to bring together distant DNA regions, which allows one DNA regulatory element to influence how a distant gene is either turned on or off. In other cases, it is believed that the genes found in one of these loops are co-regulated. It is possible that elimination of large fractions of junk DNA could perturb these local architectural arrangements of the genome. It should be noted that the relative importance of TADs remains controversial. Moreover, it is unclear whether any given TAD would be disrupted if all of the junk within the TAD were to be removed. We understand even less about larger scale architectural features of the genome, how these are affected by junk DNA, and how these structures ultimately contribute to fitness. Nevertheless, effects on DNA architecture could arise, both at the local and global levels, if a genome were massively compacted.
Another aspect of cell physiology that could change is some of its biophysical properties. Recently, there has been a renewed interest in how certain biopolymers tend to form molecular condensates which phase separate from the bulk solution (Banani et al. 2017). Several recent reports suggest that large portions of the genome which are silenced—and normally referred to as heterochromatin—form these condensates (Larson and Narlikar 2018). Similarly, excess amounts of RNA can also form different condensates and these may constitute the matrix of many membrane-less organelles (Jain and Vale 2017; Treeck et al. 2018; Quinodoz et al. 2021). Removal of junk DNA and RNA may alter the amount of these condensates in the nucleus, and this could ultimately impact the biophysical properties of various subcellular regions. Again, it is unclear whether the presence or absence of these condensates would alter the cellular environment enough to impact how functional parts of the genome operate, but it remains possible.
It is likely that many other cell biological processes would be disrupted after large fractions of the junk DNA were to be eliminated. Our goal is not to provide a complete list, but simply to point out that there are many general aspects of the cellular environment that are likely affected by junk DNA.
Summary
Even from a selection-at-the-level-of-the-organism point of view, junk DNA impacts the environment of the cell in which the more traditionally “functional” parts of the genome operate. Normally these impacts are distal, non-specific, and difficult to detect. We can help ourselves conceive of and understand these impacts by imagining how the cell would react to or be changed by the sudden and drastic removal of large swaths of junk DNA. Note that we are not suggesting that these bits of junk DNA are necessarily functional in the traditional sense. Rather, we are suggesting that junk DNA has causal reach; that its presence more than its specificity affects the cell and by extension the organism. There would be significant effects on cell size, timing of cell division, cellular enzymatic activity—to say the least—if it were drastically altered. These are all features and activities of the cell for which junk DNA is, in a normal context, quite important—even though said junk is not subject to the normal processes of natural selection as paradigmatically understood.
Our analysis of junk DNA thus differs from some notable others: those who solely focus on how junk DNA lacks function at the level of the organism (Palazzo and Gregory 2014) as well as those who argue that junk DNA is functional after all (Cavalier-Smith 1978; ENCODE Project Consortium et al. 2012; Mattick and Dinger 2013; Freeling et al. 2015). Similarly, a more nuanced view could also extend to other biological difference makers which do not typically act as organism-level targets of natural selection. Carving out an intermediary role—not necessarily a functional one, but still significant—allows for the recognition of more complex and diffuse elements of cause and effect in biology.
Potential objections and replies
Here we briefly consider two potential objections to the view we have articulated, and offer a pair of pre-emptive replies.
Objection: junk DNA can be eliminated by natural selection—as evidenced in eukaryotic lineages that have undergone genome reduction—and thus its presence must have adaptive or maladaptive effects on cell physiology
Reply: We do not argue that the long-term elimination of junk DNA cannot happen. In the case of some eukaryotes, such as in certain species of Arabidopsis mustard greens (Hu et al. 2011) and Arachis peanuts (Ren et al. 2018), this has occurred within a relatively very short time span. As stated above, flying birds and bats have undergone numerous large deletions that have prevented their genomes from growing due to the periodic invasion of transposable elements in their genomes (Kapusta et al. 2017). Having said that, it is not clear that these genomic reductions were due to positive selection for smaller genomes. It is also possible that genomes could get smaller by non-adaptive mechanisms; for example, when the rate of DNA deletion surpasses the rate of insertion (Petrov 2002), as is likely the case in Fugu pufferfishes (Neafsey and Palumbi 2003). If these non-adaptive processes were ultimately responsible for reducing genome size, we would expect that said organisms also acquired secondary adaptive mutations—so that their cells could operate once their basic properties (such as cell size, timing of cell division, altered rate of molecule diffusion) have significantly changed. It remains possible that in certain organisms, size reductions (or increases) in the genome could be adaptive for a particular reason (Blommaert 2020). For example, it has been suggested that as mammalian lineages expanded after the extinction of the non-avian dinosaurs, mammalian genomes may have undergone a reduction in size due to an increase in the effective population sizes and an increased selection against transposable elements (Lynch et al. 2011). We would expect this to result in the streamlining of the genome only if the advantage that it conferred outweighed all of the negative side effects associated with smaller genomes. Moreover, these effects would not be expected to be linear. We anticipate that quite a sizeable fraction of the genome would have to be eliminated before any adverse effects would be felt. Once that level of depletion is reached, we would expect that these organisms would acquire secondary adaptive changes to counter any negative effects.
Objection: a lack of negative selection against junk DNA shows that it is a reservoir for future function
Reply: This teleological argument is commonly asserted by biologists (Makalowski 2000, 2003; Muotri et al. 2007)—despite the fact that it has been thoroughly rejected by philosophers of biology and evolutionary theorists (e.g., Hull, 1965). The increase or elimination of junk DNA by positive selection within a lineage is strictly subject to how this alteration impacts an organism’s immediate fitness level (not some future potential fitness level). Lineages that do gain properties due to a steady increase or decrease in junk after many generations (after all junk DNA is a difference maker, albeit a relatively inefficacious one) may outcompete other lineages. So, there may be selection on a higher level between lineages, as suggested elsewhere (Jablonski 2008; Brunet and Doolittle 2015). But this type of selection will not change the amount of junk DNA within a lineage; it will only impact whether one lineage is selected over another that differs in this difference maker. Junk DNA elements may also be co-opted by natural selection to produce new functional units. In some cases, new functions may also arise by constructive neutral evolution (i.e., non-adaptive processes). For example: the conversion of junk RNA to functional lncRNAs (Palazzo and Lee 2018; Palazzo and Koonin 2020). While the efficacy of a particular difference maker remains low, it will be subject to non-adaptive evolution.
From this standpoint, a fair question to ask is whether the future exaptation of junk DNA can ever make it (more or less) influential. In line with previous commentaries (Linquist et al. 2020), we think it is important to describe in what time frame we are speaking. Our arguments evaluate whether non-functional (traditionally understood) elements such as junk DNA are influential even from an organismic-level view and in the present. This entails an examination of their current effects, and how efficacious they are in affecting the organism’s phenotype. Of course, this could change in the future. Certain elements of junk could become more efficacious, especially if they acquire new properties, to the point that they transition from non-functional to functional, even on typically selectionist accounts of function. Other transitions are likely (see Graur et al., 2015).
Conclusions
Throughout molecular biology, as well as in the philosophy of biology, function has often been seen as binary in terms of both type and influence: entities are either functional, or not; and only the functional ones are significant. But there are non-functional (traditionally understood) yet significant entities in biology—entities such as junk DNA. According to the dominant accounts of function in biology, most junk DNA is a non-functional entity—since it is and never was a target of selection at the level of the organism. Nonetheless, junk DNA is often a significant entity in the cellular context, and we have adapted Waters’ (2007) account of DNA as an actual difference maker to account for the significance of junk DNA. Namely, junk DNA typically has causal specificity but only for a limited extent of its reach, and thereby it is a relatively inefficacious actual difference maker in cells. Our identification of this option within the conceptual landscape—that of causal reach with proximate, but not distal, causal specificity—allows us to characterize the importance of junk DNA, while maintaining and explaining why it might be assessed as of lesser significance when compared to (for instance) enduringly causally specific stretches of coding DNA.
On both our and Waters’ (2007) accounts, junk DNA is an actual difference maker, and a causally specific one (at least proximally), precisely because it tends to be involved in actual biological process (such as that of making defined RNAs), and because specific changes in it as a cause correspond with specific changes in its effects (e.g., changes in the nucleotide sequences of junk DNA generate specific changes in resultingly transcribed RNA). However, junk DNA is often quite causally inefficacious: despite that it can produce causally specific proximate effects, its cascading ultimate effects tend towards the non-specific. The constrained significance (for the organism) of junk DNA is not due to a total absence of causal specificity, but rather a limited reach for its causal specificity, and resulting limitations in its causal efficacy.
Junk DNA nonetheless contributes to the environment of the cell in which other, more traditionally functional entities exist. Total DNA levels control cell size, the amount of transcriptional noise, the number of transcription factor binding sites, the timing of cell division and likely many other aspects of the cellular environment. More paradigmatically functional entities of the cell have been tuned by natural selection to operate within this environment. This is especially true for quality control processes, which specifically evolve to deal with junk DNA and all its associated biochemical activity. On the one hand, and despite being an actual difference maker, junk DNA is not typically subject to organism-level natural selection. When non-functional DNA is mutated by natural processes, this does not alter the fitness of the organism sufficiently for natural selection to eliminate such mutations. For these reasons and given the way ‘function’ is predominantly understood in biology and philosophy of biology, junk DNA is typically conceived of as a non-functional entity. On the other hand, various components of cellular machinery exist precisely in order to deal with the effects of junk DNA, and the cellular workload is significantly affected by the need to (for instance) perform routine clean-up of the products of junk DNA transcription. If large swaths of junk DNA were eliminated, this would likely affect the fitness of the organism. Although such instances of widespread elimination of junk DNA are highly unlikely to naturally occur, such alterations could in principle could be performed in the lab. For these alternative reasons, and in our proposed terms, junk DNA is significant for the cell—despite its paradigmatically non-functional role.
Ultimately—since there are actual, causally-specific difference makers which shape and impact the cellular environment, but which are not typically subject to organism-level natural selection—our discussion demonstrates that for certain biological organisms, there can be influential features which nonetheless are non-functional in the selectionist sense, i.e., they are not shaped by natural selection. Indeed, in organisms for which drift dominates, it is possible for non-functional-yet-influential biological components to proliferate—especially when the production of these components is fueled by additional processes. The activity of transposable elements appears, in eukaryotes at least, to have driven the proliferation of junk DNA in just such a manner.
Notes
Linquist et al. (2020) argue that it is conceptually incoherent for molecular biologists to adopt “causal role” approaches to function in biology, while also inferring that said functions imply a contribution to fitness. The fact that such a position is inherently confused does not prohibit its presence or prevalence in the literature.
Although we primarily focus on coding DNA, our arguments apply equally to other unambiguously functional genomic elements, such as non-coding RNA genes and promoter regions.
It remains unclear whether the extensive RNA processing found in eukaryotic mRNAs pre-dated the evolution of the nucleus/cytosolic divide, or whether the evolution of the nucleus permitted RNA processing to evolve and proliferate (López-García and Moreira 2015). Despite this uncertainty, it does appear that the last eukaryotic common ancestor was intron-rich (Rogozin et al. 2003).
Although transposable elements are not being selected for within the species (and in many cases are selected against, as they are a mutational hazard), they are under selection pressure within the genome to become better at copying themselves. For more on how selection at different levels operates on transposable elements, see (Brunet and Doolittle 2015). In any case, active transposable elements make up only a small fraction of all junk DNA in most eukaryotes. Most junk DNA consists of either inactive TEs, small simple repeats, or other heterogeneous sequences of uncertain origin (Gregory 2005a).
Though see (Griffiths et al. 2015) for a persuasive challenge—one different from the trajectory followed in this manuscript—to the notion that causal specificity in combination with actual difference making can uniquely and consistently pick out DNA as the most special and important cause when it comes to transcription and translation across different genes and organisms.
Here we follow Woodward (2010) in the sense that we do not offer these concepts as strict criteria for causation, but rather as features which—although they might not be necessary conditions which must pertain for a relationship to be causal—can nonetheless “play important roles in particular scientific contexts” (p. 288).
If causal specificity and reach are first-order properties, then causal efficacy is a second-order property of reach in combination with specificity.
Our use of the term ‘causal specificity’ refers to what Woodward (2010) calls the fine-grained influence conception of causal specificity, a conception inspired by Lewis’s notion of influence (Lewis 2000). But there is another candidate conception of causal specificity, one Woodward (2010) calls the one to one conception of causal specificity. Whereas the fine-grained influence conception of causal specificity picks out those causal variables which take on many distinct values corresponding to many distinct values in effect, the one to one conception of causal specificity picks out those causal variables which have solitary effects—in an only-one-cause, only-one-effect manner. As Woodward (2010) stresses, causes which are specific in this one to one sense can be powerful targets for intervention, precisely because of how clean-cut and controlled the effects of intervening on these causes can be. Our concept of causal reach acts as something of a counterpart to this one-to-one sense of causal specificity, since causal reach picks out those causal variables with extended and possibly even cascading effects—in an only-one-cause, yet-many-effects manner. Causes which have reach in this one-to-many sense can also be powerful targets for intervention, because of how sweeping and far-reaching the effects of intervening on these causes can be. Just as there is more than one sense of causal specificity, so too is there more than one sense of causal power (where power is here understood as tracking a cause’s prospects for acting as a target for intervention).
This does not necessarily mean that such causes have absolutely zero further, downstream effects—that the causal reach of such a change is nil. Woodward (2010) discusses the issue of causal proportionality, which pertains to choices of explanatory level and extent. All causal representations offered here are necessarily, selectively representative; they depict some details and ignore others.
In some cases, common codons are more rapidly decoded than rare codons, and thus the presence of certain codons can affect the rate at which a particular mRNA is translated. In bacteria, codon choice can even be selected for, in order to optimize the rate of production for certain proteins (Ermolaeva 2001). Even in mammalian cells, the use of different synonymous codons in certain mRNAs can have profound effects. For example, in mammalian actin mRNAs, the presence of particular rare codons can slow down protein synthesis, leading to the modification of the nascent actin polypeptide at a particular amino acid (Zhang et al. 2010). In the absence of such rare codons, the actin protein is synthesized very quickly, allowing it to fold and bury the amino acid before it can be modified.
This is not to say that now we have completely explained judgements of biological significance. For instance, this critique is being offered from within the selectionist framework. If we stepped outside of that framework, effects on fitness might become markedly less interesting and important. If we cared more about molecular differentiation and diversity, we might prioritize different casual or even non-causal features over specificity, reach, and efficacy. We thank our reviewers for drawing attention to this limitation of the paper’s framing.
This is like adding or losing a station at the end of the tuning dial.
There are of course other functional and non-functional DNA elements, not discussed here, with distinct combinations of causal specificity, reach, and efficacy. For example: promoters, telomeres, origins of replication, and genes for RNA molecules that directly impact a cell biological process (e.g., tRNA, rRNA). Although these DNA elements do not specify proteins directly, they nevertheless have precise downstream causal effects that are specific and efficacious. Some, such as tRNA genes, are very analogous, as they are transcribed into tRNAs which have specific folds based on their sequence and have roles in decoding mRNAs into proteins. A base change in this gene would lead to a transcriptional product that had a corresponding base change. This altered tRNA would likely have a profound change in its ability to decode mRNA into proteins. Other DNA elements, such as promoters, are not themselves copied into RNA, and hence do not appear to have specific causal effects, but they nevertheless influence how often and under what conditions nearby DNA sequences are transcribed into RNA. Base changes in promoters may alter how much and under what conditions the nearby genes are transcribed, thus their effects are quite efficacious. Considerations of space necessitate some limit to the molecular variety we showcase here.
In principle the mutational analysis of putative junk DNA has already been done. Given the number of humans alive today (7 × 109), the size of the diploid genome (6 × 109), and the fact that each person carries on the order of 100 de novo mutations each generation (Ségurel et al. 2014; Sung et al. 2016), this means that every base in the human genome has been altered in approximately 100 humans (with the exception of those alterations that are lethal). About 80–90% of these mutations are single nucleotide polymorphisms, with the rest (about 10–20%) being indels (Mills et al. 2006, 2011; Mullaney et al. 2010; Sung et al. 2016). Thus, every base in the human genome has been deleted de novo in 5–10 humans. In addition, individuals will also have indels inherited from their parents, so the overall number of mutations that each individual harbors is quite high, numbering over a thousand (Agrawal and Whitlock 2012). Suffice to say that by sequencing the genomes of all humans alive, we could determine which parts of the human genome can allow for alterations without compromising viability. Having said that, humans are diploid, so compromising a putative functional element in one copy of the genome is often alleviated by the presence of a second wildtype copy of the same element. This likely explains why most individuals carry many inactivating mutations in essential genes yet do not display any problems, likely because they have a second fully functional version of these defective genes (MacArthur et al. 2012).
References
Agrawal AF, Whitlock MC (2012) Mutation load: the fitness of individuals in populations where deleterious alleles are abundant. Annu Rev Ecol Evol Syst 43:115–135. https://doi.org/10.1146/annurev-ecolsys-110411-160257
Allen C, Neal J (2020) Teleological notions in biology. In: Zalta EN (ed) The Stanford encyclopedia of philosophy, Spring 2020. Metaphysics Research Lab, Stanford University, Stanford
Banani SF, Lee HO, Hyman AA, Rosen MK (2017) Biomolecular condensates: organizers of cellular biochemistry. Nat Rev Mol Cell Biol 18:285–298. https://doi.org/10.1038/nrm.2017.7
Bar-Even A, Noor E, Savir Y et al (2011) The moderately efficient enzyme: evolutionary and physicochemical trends shaping enzyme parameters. Biochemistry 50:4402–4410. https://doi.org/10.1021/bi2002289
Beagan JA, Phillips-Cremins JE (2020) On the existence and functionality of topologically associating domains. Nat Genet 52:8–16. https://doi.org/10.1038/s41588-019-0561-1
Blommaert J (2020) Genome size evolution: towards new model systems for old questions. Proc R Soc B Biol Sci 287:20201441. https://doi.org/10.1098/rspb.2020.1441
Boorse C (1976) Wright on functions. Philos Rev 85:70–86. https://doi.org/10.2307/2184255
Brunet TDP, Doolittle WF (2015) Multilevel selection theory and the evolutionary functions of transposable elements. Genome Biol Evol 7:2445–2457. https://doi.org/10.1093/gbe/evv152
Cavalier-Smith T (1978) Nuclear volume control by nucleoskeletal DNA, selection for cell volume and cell growth rate, and the solution of the DNA C-value paradox. J Cell Sci 34:247–278
Copley SD (2020) The physical basis and practical consequences of biological promiscuity. Phys Biol. https://doi.org/10.1088/1478-3975/ab8697
Cummins R (1975) Functional analysis. J Philos 72:741–765. https://doi.org/10.2307/2024640
Dayhoff MO (1972) Atlas of Protein Sequence and Structure, 5th edn. National Biomedical Research Foundation., Washington D.C
Doma MK, Parker R (2007) RNA quality control in eukaryotes. Cell 131:660–668. https://doi.org/10.1016/j.cell.2007.10.041
Doolittle WF, Sapienza C (1980) Selfish genes, the phenotype paradigm and genome evolution. Nature 284:601–603
ENCODE Project Consortium, Bernstein BE, Birney E et al (2012) An integrated encyclopedia of DNA elements in the human genome. Nature 489:57–74. https://doi.org/10.1038/nature11247
Ermolaeva MD (2001) Synonymous codon usage in bacteria. Curr Issues Mol Biol 3:91–97
Freeling M, Xu J, Woodhouse M, Lisch D (2015) A Solution to the C-value paradox and the function of junk DNA: the genome balance hypothesis. Mol Plant 8:899–910. https://doi.org/10.1016/j.molp.2015.02.009
Garland W, Jensen TH (2020) Nuclear sorting of RNA. Wires RNA 11:e1572. https://doi.org/10.1002/wrna.1572
Ghavi-Helm Y, Jankowski A, Meiers S et al (2019) Highly rearranged chromosomes reveal uncoupling between genome topology and gene expression. Nat Genet 51:1272–1282. https://doi.org/10.1038/s41588-019-0462-3
Grantham R (1974) Amino Acid Difference Formula to Help Explain Protein Evolution. Science 185:862–864. https://doi.org/10.1126/science.185.4154.862
Graur D, Zheng Y, Azevedo RBR (2015) An evolutionary classification of genomic function. Genome Biol Evol 7:642–645. https://doi.org/10.1093/gbe/evv021
Gregory TR (2001) The bigger the C-value, the larger the cell: genome size and red blood cell size in vertebrates. Blood Cells Mol Dis 27:830–843. https://doi.org/10.1006/bcmd.2001.0457
Gregory TR (2002) A bird’s-eye view of the C-value enigma: genome size, cell size, and metabolic rate in the class Aves. Evolution 56:121–130. https://doi.org/10.1111/j.0014-3820.2002.tb00854.x
Gregory TR (2005a) Synergy between sequence and size in large-scale genomics. Nat Rev Genet 6:699–708. https://doi.org/10.1038/nrg1674
Gregory TR (2005b) Genome size evolution in animals. In: The Evolution of the Genome. Elsevier, San Diego, pp 3–87
Gregory TR, Nicol JA, Tamm H et al (2007) Eukaryotic genome size databases. Nucleic Acids Res 35:D332-338. https://doi.org/10.1093/nar/gkl828
Gregory TR, Andrews CB, McGuire JA, Witt CC (2009) The smallest avian genomes are found in hummingbirds. Proc R Soc B Biol Sci 276:3753–3757. https://doi.org/10.1098/rspb.2009.1004
Griffiths PE, Pocheville A, Calcott B et al (2015) Measuring causal specificity. Philos Sci 82:529–555. https://doi.org/10.1086/682914
Gulko B, Hubisz MJ, Gronau I, Siepel A (2015) A method for calculating probabilities of fitness consequences for point mutations across the human genome. Nat Genet 47:276–283. https://doi.org/10.1038/ng.3196
Hart HLA, Honoré AM (1959) Causation in the law. Clarendon Press, Oxford
Hinde RA (1975) The concept of function. In: Function and evolution in behavior. Clarendon Press, Oxford, pp 3–15
Hoyle NP, Ish-Horowicz D (2013) Transcript processing and export kinetics are rate-limiting steps in expressing vertebrate segmentation clock genes. Proc Natl Acad Sci USA 110:E4316–E4324. https://doi.org/10.1073/pnas.1308811110
Hu TT, Pattyn P, Bakker EG et al (2011) The Arabidopsis lyrata genome sequence and the basis of rapid genome size change. Nat Genet 43:476–481. https://doi.org/10.1038/ng.807
Hughes AL, Hughes MK (1995) Small genomes for better flyers. Nature 377:391. https://doi.org/10.1038/377391a0
Hull DL (1965) The effect of essentialism on taxonomy—two thousand years of stasis (I). Br J Philos Sci 15:314–326
Jablonski D (2008) Species selection: theory and data. Annu Rev Ecol Evol Syst 39:501–524
Jain A, Vale RD (2017) RNA phase transitions in repeat expansion disorders. Nature 546:243. https://doi.org/10.1038/nature22386
Jeffreys AJ, Wilson V, Thein SL (1985) Hypervariable ‘minisatellite’ regions in human DNA. Nature 314:67–73. https://doi.org/10.1038/314067a0
Kapusta A, Suh A, Feschotte C (2017) Dynamics of genome size evolution in birds and mammals. PNAS 114:E1460–E1469. https://doi.org/10.1073/pnas.1616702114
Khersonsky O, Tawfik DS (2010) Enzyme promiscuity: a mechanistic and evolutionary perspective. Annu Rev Biochem 79:471–505. https://doi.org/10.1146/annurev-biochem-030409-143718
Koonin EV (2016) Splendor and misery of adaptation, or the importance of neutral null for understanding evolution. BMC Biol 14:114. https://doi.org/10.1186/s12915-016-0338-2
Kozak M (1991) A short leader sequence impairs the fidelity of initiation by eukaryotic ribosomes. Gene Expr 1:111–115
Larson AG, Narlikar GJ (2018) The role of phase separation in heterochromatin formation, function, and regulation. Biochemistry 57:2540–2548. https://doi.org/10.1021/acs.biochem.8b00401
Lewis D (2000) Causation as influence. J Philos 97:182–197. https://doi.org/10.2307/2678389
Linquist S, Doolittle WF, Palazzo AF (2020) Getting clear about the F-word in genomics. PLoS Genet 16:e1008702. https://doi.org/10.1371/journal.pgen.1008702
López-García P, Moreira D (2006) Selective forces for the origin of the eukaryotic nucleus. BioEssays 28:525–533. https://doi.org/10.1002/bies.20413
López-García P, Moreira D (2015) Open questions on the origin of eukaryotes. Trends Ecol Evol 30:697–708. https://doi.org/10.1016/j.tree.2015.09.005
Luo MJ, Reed R (1999) Splicing is required for rapid and efficient mRNA export in metazoans. Proc Natl Acad Sci USA 96:14937–14942
Lynch M, Bobay L-M, Catania F et al (2011) The repatterning of eukaryotic genomes by random genetic drift. Annu Rev Genomics Hum Genet 12:347–366. https://doi.org/10.1146/annurev-genom-082410-101412
MacArthur DG, Balasubramanian S, Frankish A et al (2012) A systematic survey of loss-of-function variants in human protein-coding genes. Science 335:823–828. https://doi.org/10.1126/science.1215040
Mackie JL (1974) The cement of the universe: a study of causation. Clarendon Press, Oxford
Makalowski W (2000) Genomic scrap yard: how genomes utilize all that junk. Gene 259:61–67. https://doi.org/10.1016/S0378-1119(00)00436-4
Makalowski W (2003) Not junk after all. Science 300:1246–1247. https://doi.org/10.1126/science.1085690
Martin W, Koonin EV (2006) Introns and the origin of nucleus-cytosol compartmentalization. Nature 440:41–45. https://doi.org/10.1038/nature04531
Mattick JS, Dinger ME (2013) The extent of functionality in the human genome. HUGO J 7:2. https://doi.org/10.1186/1877-6566-7-2
Millikan RG (1984) Language, thought, and other biological categories: new foundations for realism. MIT Press, Cambridge
Mills RE, Luttig CT, Larkins CE et al (2006) An initial map of insertion and deletion (INDEL) variation in the human genome. Genome Res 16:1182–1190. https://doi.org/10.1101/gr.4565806
Mills RE, Pittard WS, Mullaney JM et al (2011) Natural genetic variation caused by small insertions and deletions in the human genome. Genome Res 21:830–839. https://doi.org/10.1101/gr.115907.110
Miyata T, Miyazawa S, Yasunaga T (1979) Two types of amino acid substitutions in protein evolution. J Molecular Evol 12:219–236. https://doi.org/10.1007/BF01732340
Mullaney JM, Mills RE, Pittard WS, Devine SE (2010) Small insertions and deletions (INDELs) in human genomes. Hum Mol Genet 19:R131–R136. https://doi.org/10.1093/hmg/ddq400
Muotri AR, Marchetto MCN, Coufal NG, Gage FH (2007) The necessary junk: new functions for transposable elements. Hum Mol Genet 16:R159–R167. https://doi.org/10.1093/hmg/ddm196
Neafsey DE, Palumbi SR (2003) Genome size evolution in pufferfish: a comparative analysis of diodontid and tetraodontid pufferfish genomes. Genome Res 13:821–830. https://doi.org/10.1101/gr.841703
Neander K (1991) The teleological notion of ‘function.’ Australas J Philos 69:454–468. https://doi.org/10.1080/00048409112344881
Nóbrega MA, Zhu Y, Plajzer-Frick I et al (2004) Megabase deletions of gene deserts result in viable mice. Nature 431:988–993. https://doi.org/10.1038/nature03022
Oeffinger M, Zenklusen D (2012) To the pore and through the pore: a story of mRNA export kinetics. Biochim Biophys Acta 1819:494–506. https://doi.org/10.1016/j.bbagrm.2012.02.011
Ogami K, Richard P, Chen Y et al (2017) An Mtr4/ZFC3H1 complex facilitates turnover of unstable nuclear RNAs to prevent their cytoplasmic transport and global translational repression. Genes Dev 31:1257–1271. https://doi.org/10.1101/gad.302604.117
Organ CL, Shedlock AM, Meade A, Pagel M, Edwards SV (2007) Origin of avian genome size and structure in non-avian dinosaurs. Nature 446:180–184. https://doi.org/10.1038/nature05621
Orgel LE, Crick FH (1980) Selfish DNA: the ultimate parasite. Nature 284:604–607
Palazzo AF, Gregory TR (2014) The case for junk DNA. PLoS Genet 10:e1004351. https://doi.org/10.1371/journal.pgen.1004351
Palazzo AF, Koonin EV (2020) Functional long non-coding RNAs evolve from junk transcripts. Cell 183:1–12. https://doi.org/10.1016/j.cell.2020.09.047
Palazzo AF, Lee ES (2015) Non-coding RNA: what is functional and what is junk? Front Genet 6:2. https://doi.org/10.3389/fgene.2015.00002
Palazzo AF, Lee ES (2018) Sequence determinants for nuclear retention and cytoplasmic export of mRNAs and lncRNAs. Front Genet 9:440. https://doi.org/10.3389/fgene.2018.00440
Palazzo AF, Springer M, Shibata Y et al (2007) The signal sequence coding region promotes nuclear export of mRNA. PLoS Biol 5:e322. https://doi.org/10.1371/journal.pbio.0050322
Petrov DA (2002) Mutational equilibrium model of genome size evolution. Theor Popul Biol 61:531–544. https://doi.org/10.1006/tpbi.2002.1605
Ponting C (2017) Biological function in the twilight zone of sequence conservation. BMC Biol 15:71. https://doi.org/10.1186/s12915-017-0411-5
Ponting CP, Hardison RC (2011) What fraction of the human genome is functional? Genome Res 21:1769–1776. https://doi.org/10.1101/gr.116814.110
Quinodoz SA, Jachowicz JW, Bhat P et al (2021) RNA promotes the formation of spatial compartments in the nucleus. Cell 184:5775-5790.e30. https://doi.org/10.1016/j.cell.2021.10.014
Rajon E, Masel J (2011) Evolution of molecular error rates and the consequences for evolvability. PNAS 108:1082–1087. https://doi.org/10.1073/pnas.1012918108
Rajon E, Masel J (2013) Compensatory evolution and the origins of innovations. Genetics 193:1209–1220. https://doi.org/10.1534/genetics.112.148627
Rands CM, Meader S, Ponting CP, Lunter G (2014) 8.2% of the human genome is constrained: variation in rates of turnover across functional element classes in the human lineage. PLOS Genet 10:e1004525. https://doi.org/10.1371/journal.pgen.1004525
Redon R, Ishikawa S, Fitch KR et al (2006) Global variation in copy number in the human genome. Nature 444:444–454. https://doi.org/10.1038/nature05329
Ren L, Huang W, Cannon EKS et al (2018) A mechanism for genome size reduction following genomic rearrangements. Front Genet. https://doi.org/10.3389/fgene.2018.00454
Rogozin IB, Wolf YI, Sorokin AV et al (2003) Remarkable interkingdom conservation of intron positions and massive, lineage-specific intron loss and gain in eukaryotic evolution. Curr Biol 13:1512–1517
Ségurel L, Wyman MJ, Przeworski M (2014) Determinants of mutation rate variation in the human germline. Annu Rev Genomics Hum Genet 15:47–70. https://doi.org/10.1146/annurev-genom-031714-125740
Shav-Tal Y, Darzacq X, Shenoy SM et al (2004) Dynamics of single mRNPs in nuclei of living cells. Science 304:1797–1800. https://doi.org/10.1126/science.1099754
Struhl K (2007) Transcriptional noise and the fidelity of initiation by RNA polymerase II. Nat Struct Mol Biol 14:103–105. https://doi.org/10.1038/nsmb0207-103
Sung W, Ackerman MS, Dillon MM et al (2016) Evolution of the insertion-deletion mutation rate across the tree of life. G3 Genes Genomes Genet 6:2583–2591. https://doi.org/10.1534/g3.116.030890
Szabo Q, Bantignies F, Cavalli G (2019) Principles of genome folding into topologically associating domains. Sci Adv 5:eaaw668. https://doi.org/10.1126/sciadv.aaw1668
Tawfik DS (2020) Enzyme promiscuity and evolution in light of cellular metabolism. FEBS J 287:1260–1261. https://doi.org/10.1111/febs.15296
Treeck BV, Protter DSW, Matheny T et al (2018) RNA self-assembly contributes to stress granule formation and defining the stress granule transcriptome. PNAS 115:2734–2739. https://doi.org/10.1073/pnas.1800038115
Villar D, Flicek P, Odom DT (2014) Evolution of transcription factor binding in metazoans—mechanisms and functional implications. Nat Rev Genet 15:221–233. https://doi.org/10.1038/nrg3481
Vinogradov AE (2004) Evolution of genome size: multilevel selection, mutation bias or dynamical chaos? Curr Opin Genet Dev 14:620–626. https://doi.org/10.1016/j.gde.2004.09.007
Wang J, Cheng H (2020) Out or decay: fate determination of nuclear RNAs. Essays Biochem. https://doi.org/10.1042/EBC20200005
Ward LD, Kellis M (2012) Evidence of abundant purifying selection in humans for recently acquired regulatory functions. Science 337:1675–1678. https://doi.org/10.1126/science.1225057
Warnecke T, Hurst LD (2011) Error prevention and mitigation as forces in the evolution of genes and genomes. Nat Rev Genet 12:875–881. https://doi.org/10.1038/nrg3092
Waters CK (2007) Causes that make a difference. J Philos 104:551–579
Wimsatt WC (1972) Teleology and the logical structure of function statements. Stud Hist Philos Sci Part A 3:1–80. https://doi.org/10.1016/0039-3681(72)90014-3
Woodward J (2010) Causation in biology: stability, specificity, and the choice of levels of explanation. Biol Philos 25:287–318. https://doi.org/10.1007/s10539-010-9200-z
Zhang F, Saha S, Shabalina SA, Kashina A (2010) Differential arginylation of actin isoforms is regulated by coding sequence-dependent degradation. Science 329:1534–1537. https://doi.org/10.1126/science.1191701
Zinder JC, Lima CD (2017) Targeting RNA for processing or destruction by the eukaryotic RNA exosome and its cofactors. Genes Dev 31:88–100. https://doi.org/10.1101/gad.294769.116
Acknowledgements
We would like to thank Stefan Linquist and Ford Doolittle for organizing and inviting us to their workshop, Evolutionary Roles of Transposable Elements: The Science and the Philosophy, held in Halifax, Nova Scotia in October of 2018. We would also like to thank the other participants at that workshop for their valuable contributions. We are grateful to Adam Streed for producing our graphics. Finally, we additionally thank Stefan and Ford for their efforts as editors and reviewers of this special issue.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Havstad, J.C., Palazzo, A.F. Not functional yet a difference maker: junk DNA as a case study. Biol Philos 37, 29 (2022). https://doi.org/10.1007/s10539-022-09854-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10539-022-09854-1