Abstract
The use of microbial phosphotriesterases in the degradation of organophosphorus compounds employed as pesticides, plasticizers and petroleum additives is a sustainable alternative for bioremediation of water and soils, decontamination of particular foods and as poisoning antidote. Whole cells of six wild type microorganisms—Streptomyces phaeochromogenes, Streptomyces setonii, Nocardia corynebacterioides, Nocardia asteroides and two Arthrobacter oxydans—selected in our lab as phosphotriesterase sources, were further tested as biocatalysts in the hydrolysis of paraoxon, methyl paraoxon, methyl parathion, coroxon, coumaphos, dichlorvos and chlorpyrifos, highlighting 98% conversion of chlorpyrifos into its hydrolysis products using whole cells of S. phaeochromogenes at pH 8 and 40 °C. Immobilized whole cells and enzyme extracts were also assessed, observing as a general trend, that there is no significant variation in hydrolytic activity between them. These results suggest that according to the circumstances, immobilized whole cells (avoiding cellular disruption and centrifugation) or enzyme extracts (which can be handled more easily) could be used.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Organophosphorus compounds (OPs) have been synthetized as chemical warfare agents during World War II and then began to be used as pesticides, plasticizers and petroleum additives (Singh 2009). Their excessive use in agriculture and homes produces environmental pollution as well as high toxicity in mammals (Kim et al. 2014). OPs prevent the hydrolysis of the neurotransmitter acetylcholine to choline by acting as irreversible inhibitors of acetylcholinesterase. Consequently, acetylcholine is accumulated in the body causing an overstimulation of nerve impulses, convulsions, respiratory failures and eventually death (Theriot and Grunden 2011). Therefore, the identification of microorganisms capable of degrading OPs is relevant for applications such as environmental monitoring and remediation. Furthermore, purified microbial OP hydrolases might be used as antidote against OPs poisoning by using liposomes and other biocompatible polymer-based carriers to deliver these enzymes (Iyer and Iken 2015; Liu et al. 2015). For this reason, a variety of organophosphate degrading bacteria has been isolated and studied such as Brevundimonas diminuta, Flavobacterium sp., Alteromonas sp., Bacillus stearothemophilus, Nocardia sp., Escherichia coli, Arthrobacter, Corynebacterium glutamicum and Burkholderia sp. (Manco et al. 2018).
Within the group of OP hydrolases, phosphotriesterases (PTEs, E.C. 3.1.8.1) constitute a group of enzymes that catalyze the stereoselective hydrolysis of a large number of phosphotriesters (Elias et al. 2008). They exert their activity by cleavage of the P–O, P–N or P–S bond, differing in their substrate specificity and molecular mass (Sogorb et al. 2004). As a consequence, a large number of OP structures such as paraoxon, chlorpyrifos, parathion, methyl parathion, coumaphos, monocrotophos and fenamiphos can be hydrolyzed by these enzymes, being it the first and main step in this detoxification process. Then, OPs hydrolysis products can be used as carbon and/or phosphorus source by a wide variety of soil and water bacteria (Iyer et al. 2013; Kumar et al. 2018).
A cost-effective alternative to the large-scale application of isolated organophosphorus hydrolases, consists in the use of microorganisms as either growing or resting whole cells (Singh 2009; de Carvalho 2017). In particular for microbial bioremediation purposes different parameters play important role, but among them pH and temperature are specially relevant (Niti et al. 2013). Singh et al. have reported effects of soil pH and temperature on chlorpyrifos and fenamiphos degradation observing an increase in activity at higher pHs and in the temperature range from 15 to 35 °C (Singh et al. 2003a, b).
In our laboratory we have previously evaluated and optimized the hydrolysis conditions (pH and temperature) of methyl paraoxon employing as biocatalysts six new sources of bacterial PTEs: Streptomyces phaeochromogenes, Streptomyces setonii, Nocardia corynebacterioides, Nocardia asteroides and two Arthrobacter oxydans (Santillan et al. 2016). In the present work these microorganisms were assessed in the hydrolysis of different OPs such as paraoxon, methyl parathion, coroxon, coumaphos, dichlorvos and chlorpyrifos (Fig. 1). PTE activity was studied employing whole cells and enzyme extracts, free as well as immobilized in calcium alginate beads.
Materials and methods
Chemicals
Paraoxon, methyl parathion, coumaphos, dichlorvos and chlorpyrifos were purchased from Sigma-Aldrich (St. Louis, MO, USA). Methyl paraoxon and coroxon were synthesized by oxidative desulfuration from methyl parathion and coumaphos, respectively (Bielawski and Casida 1988).
Analytical and HPLC grade solvents were from Biopack, Sintorgan or J.T. Baker. Culture media reagents were obtained from Merck or Biopack.
Microorganisms
All microorganisms were cultured at optimized conditions according to CECT. B. diminuta CIP 7129 (Bd), A. oxydans ATCC 14358 (C55) and A. oxydans ATCC 14359 (C64) were cultured in nutrient agar medium II (1 g meat extract, 5 g peptone, 2 g yeast extract, 5 g NaCl, 1 L H2O). Streptomyces phaeochromogenes CCRC 10811 (C13) and S. setonii ATCC 39116 were grown in Streptomyces medium (4 g glucose, 10 g malt extract, 4 g yeast extract, 2 g CaCO3, 1 L H2O). N. asteroides ATCC 19296 (C49) was cultured in Bennett´s medium (4 g glucose, 10 g malt extract, 4 g yeast extract, 1 L H2O), whereas N. corynebacterioides ATCC 14898 (C39) in Nutrient agar I with 1% maltose (5 g meat extract, 10 g peptone, 10 g NaCl, 10 g maltose, 1 L H2O). All cultures were conducted for 72 h.
Biocatalysts
Whole cells
Bacteria strains were grown as previously mentioned (in “Microorganisms” section) and optical density was measured at 600 nm. An aliquot containing the required amount of cells was collected by centrifugation at 8600 × g for 5 min. The resulting cell pellets were directly employed as biocatalysts.
Enzyme extract
The membrane fraction from each bacteria was obtained and studied as biocatalyst. Cellular pellet containing 1.2 × 1010 cells (obtained as in “Whole cells” section) was resuspended in 10 mL of distilled water and was disrupted by ultrasonication using the Vibra-cell disruptor, VCX130 (Sonics, USA), at 50% amplitude for 5 cycles at 4 °C. After that, this suspension was centrifuged at 8600 × g for 15 min and the resulting pellet containing the membrane fraction was further tested as biocatalyst since PTEs are usually associated with cellular membrane (Mulbry and Karns 1989).
Immobilization
1.2 × 1010 Cells or enzyme extracts obtained from the same number of cells were resuspended in a 2% (w/v) sodium alginate solution. These suspensions were then added dropwise to a well-stirred, 100 mM CaCl2 solution. The obtained calcium-alginate gel beads were kept in the same CaCl2 solution with gentle stirring for 1 h to allow them to harden and finally were washed with physiological solution (NaClaq 0.9%).
Activity assays
The pellets containing 2 × 109 whole cells, obtained as mentioned in the “Microorganisms” and “Whole cells” sections, were added to the reaction mix consisting of 2 mM OPs in 1 mL of Tris–HCl buffer 50 mM pH 8. Enzymatic hydrolysis was assayed in standard conditions (pH 8 and 30 °C) and optimized conditions for each biocatalyst: S. phaeochromogenes at pH 8–40 °C; S. setonii, N. asteroides and A. oxydans C64 at pH 8–50 °C; B. diminuta, N. corynebacterioides and A. oxydans C55 at pH 8–60 °C (Santillan et al. 2016) using paraoxon, methyl parathion, coumaphos, coroxon, dichlorvos and chlorpyrifos as substrates. The same reaction mix without cells was used as chemical control, whereas this reaction mix but using B. diminuta cells as biocatalyst was used as positive control. The determination of the initial reaction rates (V0) was carried out by analyzing 50 µL samples taken at 0, 15, 30, 45, 60, 90, 120 and 150 min.
Biocatalysed organophosphorus compounds degradation
The reactions were performed in 6 mL reaction mixtures containing a solution 0.15 mM OPs in 50 mM Tris–HCl pH 8 buffer, and the free or immobilized biocatalysts from S. phaeochromogenes, S. setonii, N. asteroides, A. oxydans C55 and A. oxydans C64 (depending on the OP under study) using standard and optimized conditions (as detailed in “Activity assays” section. Santillan et al. 2016) for 21 days. Moreover, a chemical control was assayed in the same conditions but without biocatalyst.
Samples analysis methods
Methyl paraoxon, paraoxon and methyl parathion
The production of p-nitrophenol (PNF) was quantified by measuring absorbance at 405 nm in 96 wells plate using a Cytation 5 (BioTek Instruments, Inc., USA). The calibration curve was constructed using standard solutions of PNF (0, 0.01, 0.025, 0.05, 0.075 and 0.1 mM).
Coroxon and coumaphos
Chlorferon (CF), the fluorescent product of coroxon and coumaphos enzymatic hydrolysis (λem = 355 nm, λex = 460 nm), was detected employing a Cytation 5 (BioTek Instruments, Inc., USA). Chlorferon quantification was carried out employing the corresponding calibration curve (0, 0.01, 0.025, 0.05, 0.075 and 0.1 mM).
Dichlorvos and chlorpyrifos
Hydrolysis of dichlorvos and chlorpyrifos was determined by substrate consumption using a HPLC (Gilson, USA) with UV detection, and a C18 Column (Grace Apollo 5 µm, 150 × 4,6 mm, HiChrom, England). Dichlorvos analytical conditions were: mobile phase acetonitrile:water 48:52 for 16 min, flow 0.9 mL min−1, 280 nm; while for chlorpyrifos analytical conditions were acetonitrile:1% aqueous phosphoric acid 80:20 as mobile phase for 16 min, flow 0.9 mL min−1, 235 nm. The quantification was performed employing a calibration curve with dichlorvos and chlorpyrifos standards (0.01 to 0.1 mM).
Statistical analysis
Statistical significance was evaluated using Prism 5 statistical software (Graph Pad, Inc., USA). All experiments were performed in triplicate. Results were expressed as mean values ± SD. For multiple comparisons between experimental groups two-way ANOVA [followed by mean 95% confidence interval (CI) comparison test] were performed. Significant levels were defined as p < 0.05.
Results and discussion
PTE activity against different organophosphorus compounds by microbial whole cells
Previously in our laboratory, six wild-type bacteria were selected and studied as whole cell biocatalysts for the hydrolysis of methyl paraoxon, chosen as model substrate, and using B. diminuta as the reference biocatalyst. The hydrolytic activity was determined spectrophotometrically under standard conditions (30 °C, pH 8) and the initial rates of hydrolysis (V0), measured during the first 2.5 h of reaction, were used to compare activity. Subsequently, different pH and temperature conditions were evaluated, selecting an optimized condition for each biocatalyst. S. setonii was the microorganism that exhibited the greatest degrading capacity in the treatment of the model substrate, displaying 33 times more activity than B. diminuta and a 30-fold increase in activity under optimized conditions with respect to standard conditions (Fig. 2a) (Santillan et al. 2016).
Comparison of initial reaction rates (V0) obtained for organophosphorus compounds degradation at standard (30 °C, pH 8) and optimized conditions (S. phaeochromogenes (C13): pH 8 and 40 °C; S. setonii (C35), N. asteroides (C49) and A. oxydans (C64): pH 8 and 50 °C; B. diminuta (Bd), N. corynebacterioides (C39) and A. oxydans (C55): pH 8 and 60 °C) by microbial whole cells. The results are expressed as mean ± SD, two-way ANOVA followed by Bonferroni post hoc test. Statistical analyses were performed comparing the reaction conditions with each other for each strain (***P < 0.001, **P < 0.01, *P < 0.05). PTE activities among biocatalysts for each substrate were compared (###P < 0.001)
In the present work, the microorganisms previously studied were used as biocatalysts for the degradation of different OPs: paraoxon, methyl parathion, coroxon, coumaphos, dichlorvos and chlorpyrifos. In order to select an optimal biocatalyst for the biodegradation of each substrate, degradation data over time were determined at both standard and optimized conditions (Fig. S1). Figure 2 shows the corresponding V0 values for these compounds. In general, when methyl parathion was assessed as substrate, a marked decrease in hydrolysis was observed compared to that obtained for the corresponding oxygenated analogue (methyl paraoxon). The only structural difference between these substrates is the replacement of the phosphoryl oxygen of the methyl paraoxon by sulfur in methyl parathion. The lower electronegativity of the sulfur causes a decrease in the electrophilic nature of the phosphor atom and consequently a decrease in its reactivity towards hydrolysis (Hong and Raushel 1996). However, it is worth highlighting that A. oxydans C64 strain at optimized conditions catalyzed hydrolysis of methyl parathion more efficiently (2-fold higher Vo) than hydrolysis of methyl paraoxon. Furthermore, this biocatalyst was 10 times more active in methyl parathion hydrolysis than the control strain B. diminuta under the same conditions (Fig. 2b).
When paraoxon was employed as substrate (Fig. 2c) most of the microorganisms showed a decrease in the activity with respect to that obtained for methyl paraoxon, except for N. asteroides that exhibited a similar reaction rate under optimized conditions. Moreover, N. asteroides in those reaction conditions showed the largest activity, exceeding 148 times the V0 value obtained under standard conditions. These variations could be correlated with the report of Dornaski (1989) who studied the relationship between the structure of the substrate and the activity of PTE from B. diminuta. The substitution of ethoxy for methoxy groups in the substrate produced an increase in Km without no significant change in Vmax. However, when these groups were changed to the propyl and butyl groups, the Vmax and km values decreased substantially, suggesting that the active site that binds alkyl portion of the substrate would be hydrophobic (Donarski et al. 1989).
When structural diverse OPs such as coroxon and coumaphos were assessed, N. asteroides was the strain that most efficiently catalyzed the degradation of coroxon, while S. phaeochromogenes was the best biocatalyst for the hydrolysis of coumaphos (Fig. 2d, e). This last biocatalyst was 3 times more active under optimized conditions and also twice as active as the control strain. When comparing the hydrolysis of coroxon and coumaphos, as expected and in agreement with the report of Horne (2002) who evaluated the hydrolytic activity of Pseudomonas monteilli C11 for these two OPs, the oxon was faster degraded (Horne et al. 2002). In addition, these bulkier OPs were hydrolysed less efficiently than methyl paraoxon, paraoxon and methyl parathion.
For dichlorvos, most of the assayed biocatalysts showed higher hydrolytic activities than for the other oxon derivatives. This result could be attributed to the fact that, structurally, this OP is the smallest tested substrate. Furthermore, the leaving group of dichlorvos (dichlorovinyl alcohol) tautomerizes to dichloroacetaldehyde and, being highly volatile, it does not accumulate in the reaction medium, shifting the equilibrium and thus favoring hydrolysis. In particular, A. oxydans C55 exhibited the highest hydrolysis rates, up to 100 and 2.5 times higher than B. diminuta under standard and optimized reaction conditions, respectively (Fig. 2f).
Chlorpyrifos is a one of the most widely used OPs because of its low cost and easy access. When its degradation was analysed, four out of the six wild type microorganisms studied in this work: S. phaeochromogenes, A. oxydans C55, S. setonii and N. corynebacterioides showed high activity, being 3.5, 3, 2.5 and 2 times more active than B. diminuta at standard conditions (Fig. 2g). These results are particularly interesting considering that chlorpyrifos is a thiophosphotriester derivative and, as previously mentioned, these compounds are less susceptible to hydrolysis. It is remarkable the fact that for this substrate, in contrast to most of the obtained results, higher hydrolysis rates were observed at standard conditions rather than at optimized ones. This shows that although the selection of general reaction conditions may be usually useful, in this particular case, optimization for each substrate would be more appropriate.
The results obtained in this work allowed the selection of novel bacterial species capable of degrading efficiently each of the OPs tested (Table 1). Although, the use of Streptomyces chattanoogensis and Streptomyces olivochromogenes has been reported in chlorpyrifos degradation in minimal media at similar rates to those obtained in this work for S. phaeochromogenes, the fact that this the last biocatalyst does not require nutrients to carry out the hydrolysis of the OPs, confers it an advantage over biodegradation using growing cells.
Organophosphorus compounds biodegradation using different biocatalyst forms
Enzyme extract vs. whole cells
With a view to developing a biocatalyst suitable to a bioreactor, the bacteria that exhibited the highest PTE activity for each OP were further studied. Since PTE often is a cell membrane protein, OPs must diffuse through the outer membrane to interact with the enzyme, what represents a disadvantage with respect to extracellular enzymes. In order to overcome this limitation, the expression of the protein on bacteria surfaces has been developed (Ghanem and Raushel 2005; Kapoor and Rajagopal 2011; Yuan et al. 2011). As an alternative, we propose in this work the use of enzyme extracts generated by sonication and centrifugation of the selected bacteria. This procedure produces pellets containing the PTE associated to the membrane debris. This enzyme extract exposes the enzyme to the reaction medium, and in addition, would help to preserve the natural environment of the protein, keeping its folding and activity. OPs biodegradation was carried out for 21 days under standard and optimized conditions since after this period of time no significant differences in the hydrolysis profile were observed (Fig. S2). Solutions composed of 0.15 mM OPs were used considering that in general this is the concentration of the commercial formulations used in agriculture as pesticides (Álvarez et al. 2013).
In most cases, enzyme extracts showed higher PTE activities than the corresponding whole cells. S. setonii enzyme extract produced 21% more degradation of methyl paraoxon than whole cells under standard conditions, whereas under optimized conditions no significant differences were observed (Fig. 3a). Methyl parathion was completely hydrolyzed under optimized conditions by A. oxydans C64 enzyme extract, while only 15% degradation was observed with whole cells (Fig. 3b). Additionally, the hydrolytic efficiency reached by these microorganisms at both standard and optimized conditions could be exploited by combining them to decontaminate environments treated with methyl parathion, since methyl paraoxon is always present in these environments, as a product of photolytic oxidation of methyl parathion (Jaga and Dharmani 2006).
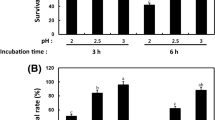
Percentage of organophosphorus compounds biodegradation employing free or immobilized whole cells and enzyme extracts at standard (30 °C, pH 8) and optimized conditions (S. setonii, N. asteroides and A. oxydans C64: pH 8 and 50 °C; S. phaeochromogenes: pH 8 and 40 °C and A. oxydans C55: pH 8 and 60 °C) at 21 days of reaction. The reaction mixture without biocatalyst was used as chemical degradation control. The results are expressed as mean ± SD, two-way ANOVA followed by Bonferroni post hoc test: ***P < 0.001, **P < 0.01 and *P < 0.05 significances when comparing the reaction condition for each kind of biocatalyst, ###P < 0.001 and ##P < 0.01 significance when whole cells and enzymatic extracts were compared. +++P < 0.001 and ++P < 0.01 significance when free and immobilized biocatalysts were compared
When the hydrolysis of paraoxon was studied using the enzyme extract of N. asteroides as biocatalyst, 80% hydrolysis was reached, representing an increase of 15% compared to the conversion obtained with whole cells under optimized conditions (Fig. 3c). Furthermore, degradation reached using N. asteroides whole cells at standard conditions are similar or even higher (1.5 mg L−1 d−1) than those reported employing Stenotrophomonas maltophilia, Aeromonas hydrophilia and Exiguobacterium indicum, which degraded 1.4 mg L−1 d−1 and 0.46 mg L−1 d−1 of paraoxon at pH 7, 30 °C (Iyer and Iken 2013).
Following the same trend, coroxon degradation studies showed the best results using the enzyme extract from N. asteroides under optimized conditions (82%) (Fig. 3d). In contrast, when the degradation of 0.15 mM coumaphos was studied, no production of chlorferon was observed (data not shown), probably due to its low solubility in comparison to other OPs. However, similar to the methyl parathion–methyl paraoxon pair, coumaphos can be oxidized by photolysis, resulting in coroxon, so the use of N. asteroides could be an alternative for the remediation of both contaminants.
On the other hand, the treatment of dichlorvos with A. oxydans C55 whole cells under optimized conditions produced 70% hydrolysis, which represents 17% more degradation respect to the enzyme extract (Fig. 3e). According to this result, S. phaeochromogenes whole cells were the best biocatalyst for chlorpyrifos biodegradation (98%) at optimized conditions (pH 8, 40 °C) (Fig. 3f). This result is better than that reported by Vidya Lakshmi et al., who employed microbial consortia that allowed 46–72% degradation of 50 mg L−1 chlorpyrifos as the sole carbon source in aqueous medium at 37 °C after 20 days. (Vidya Lakshmi et al. 2008).
Free vs. immobilized biocatalysts
With prospects to develop a bioreactor for the treatment of water contaminated with organophosphorus pesticides, both whole cells and enzyme extracts were immobilized by entrapment in calcium alginate beads. In most cases, a decrease in the hydrolytic activity for the immobilized biocatalysts was observed under both standard and optimized conditions (Fig. 3), which is an usual consequence of entrapment immobilization techniques. This lower activity was related to possible mass transfer issues since the substrates should pass through the calcium alginate matrix to access the biocatalyst. Despite this disadvantage, the immobilization allows greater stability, easier separation of the biocatalyst from the reaction media and, as a direct consequence, more efficient reuse (Polakovič et al. 2017).
On the other hand, when both free and immobilized catalysts were compared, no relevant variations were observed in the degradation profile. The exception was the immobilized S. phaeochromogenes whole cells in the hydrolysis of chlorpyrifos under optimized conditions, which degraded 75% of the substrate initial amount, being these results similar to those recently obtained by our research group with Sphingomonas sp. (Santillan et al. 2020). This and the report of Garcia-Mancha et al., regarding the use of an anaerobic expanded granular sludge bed reactor (EGSB) the reached higher degradation percentages (96%) at 55 °C that under mesophilic conditions (35 °C), suggest that immobilized S. phaeochromogenes whole cells could be applied in a packed bed bioreactor for the treatment of wastewater (García-Mancha et al. 2017). Furthermore, when the immobilized enzyme extract from S. phaeochromogenes was used as biocatalyst, it hydrolyzed 20% of the starting chlorpyrifos, whereas the free biocatalyst only showed 4% degradation, suggesting that the immobilization process could enhance enzyme extract stability.
The versatility of the studied biocatalysts offers the possibility of choosing, depending on the circumstances (environmental conditions or type of system: water or soil), whole cells (avoiding cellular disruption and centrifugation) or enzyme extracts (which can be easily handled), free or immobilized.
Conclusions
In this work, with a view to the bioremediation of water contaminated with OPs, we have studied six wild type microorganisms in the degradation of different OPs. Due to the wide use as insecticides of dichlorvos and chlorpyrifos, the results achieved using whole cells of A. oxydans and S. phaeochromogenes, which under optimized conditions reached 70 and 98% degradation respectively, are noteworthy. We have also studied the corresponding enzyme extracts, as well as the immobilized form of all the assayed biocatalysts. The obtained results showed that both, the immobilized whole cells and enzyme extracts, are potentially useful biocatalysts to be applied in water decontamination. In addition, the studied biocatalysts showed differential activities at different pHs and temperatures, which also make them attractive for the bioremediation of soils and for the decontamination of particular foods (Yu et al. 2006; Akram and Mushtaq 2017).
References
Akram S, Mushtaq M (2017) Techniques to detect and detoxify organophosphorus pesticides from fruit juices. Elsevier, Inc., Amsterdam
Álvarez M, Du Mortier C, Sokolic T, Cirelli AF (2013) Studies on the persistence of a commercial formulation of chlorpyrifos on an agricultural soil from provincia de Buenos Aires, República Argentina. Water Air Soil Pollut. https://doi.org/10.1007/s11270-013-1571-8
Bielawski J, Casida JE (1988) Phosphorylating intermediates in the peracid oxidation of phosphorothionates, phosphorothiolates, and phosphorodithiolates. J Agric Food Chem 36:610–615. https://doi.org/10.1021/jf00081a052
de Carvalho CCCR (2017) Whole cell biocatalysts: essential workers from Nature to the industry. Microb Biotechnol 10:250–263. https://doi.org/10.1111/1751-7915.12363
Donarski WJ, Dumas DP, Heitmeyer DP et al (1989) Structure–activity relationships in the hydrolysis of substrates by the phosphotriesterase from Pseudomonas diminuta. Biochemistry 28:4650–4655. https://doi.org/10.1021/bi00437a021
Elias M, Dupuy J, Merone L et al (2008) Structural basis for natural lactonase and promiscuous phosphotriesterase activities. J Mol Biol 379:1017–1028. https://doi.org/10.1016/j.jmb.2008.04.022
García-Mancha N, Monsalvo VM, Puyol D et al (2017) Enhanced anaerobic degradability of highly polluted pesticides-bearing wastewater under thermophilic conditions. J Hazard Mater. https://doi.org/10.1016/j.jhazmat.2017.06.032
Ghanem E, Raushel FM (2005) Detoxification of organophosphate nerve agents by bacterial phosphotriesterase. Toxicol Appl Pharmacol 207:459–470. https://doi.org/10.1016/j.taap.2005.02.025
Hong SB, Raushel FM (1996) Metal–substrate interactions facilitate the catalytic activity of the bacterial phosphotriesterase. Biochemistry 35:10904–10912. https://doi.org/10.1021/bi960663m
Horne I, Harcourt RL, Sutherland TD et al (2002) Isolation of a Pseudomonas monteilli strain with a novel phosphotriesterase. FEMS Microbiol Lett 206:51–55. https://doi.org/10.1016/S0378-1097(01)00518-3
Iyer R, Iken B (2015) Protein engineering of representative hydrolytic enzymes for remediation of organophosphates. Biochem Eng J 94:134–144. https://doi.org/10.1016/j.bej.2014.11.010
Iyer R, Iken B (2013) Identification of water-borne bacterial isolates for potential remediation of organophosphate contamination. Adv Biol Chem 03:146–152. https://doi.org/10.4236/abc.2013.31018
Iyer R, Iken B, Damania A (2013) A comparison of organophosphate degradation genes and bioremediation applications. Environ Microbiol Rep 5:787–798. https://doi.org/10.1111/1758-2229.12095
Jaga K, Dharmani C (2006) Methyl parathion: an organophosphate insecticide not quite forgotten. Rev Environ Health 21:57–67. https://doi.org/10.1515/REVEH.2006.21.1.57
Kapoor M, Rajagopal R (2011) Enzymatic bioremediation of organophosphorus insecticides by recombinant organophosphorus hydrolase. Int Biodeterior Biodegrad 65:896–901. https://doi.org/10.1016/j.ibiod.2010.12.017
Kim CS, Seo JH, Kang DG, Cha HJ (2014) Engineered whole-cell biocatalyst-based detoxification and detection of neurotoxic organophosphate compounds. Biotechnol Adv 32:652–662. https://doi.org/10.1016/j.biotechadv.2014.04.010
Kumar S, Kaushik G, Dar MA et al (2018) Microbial degradation of organophosphate pesticides: a review. Pedosphere 28:190–208. https://doi.org/10.1016/S1002-0160(18)60017-7
Liu Y, Li J, Lu Y (2015) Enzyme therapeutics for systemic detoxification. Adv Drug Deliv Rev 90:24–39. https://doi.org/10.1016/j.addr.2015.05.005
Manco G, Porzio E, Suzumoto Y (2018) Enzymatic detoxification: a sustainable means of degrading toxic organophosphate pesticides and chemical warfare nerve agents. J Chem Technol Biotechnol 93:2064–2082. https://doi.org/10.1002/jctb.5603
Mulbry WW, Karns JS (1989) Purification and characterization of three parathion hydrolases from gram-negative bacterial strains. Appl Environ Microbiol 55:289–293
Niti C, Sunita S, Kamlesh K, Rakesh K (2013) Bioremediation: an emerging technology for remediation of pesticides. Res J Chem Environ 17:18
Polakovič M, Švitel J, Bučko M et al (2017) Progress in biocatalysis with immobilized viable whole cells: systems development, reaction engineering and applications. Biotechnol Lett 39:667–683. https://doi.org/10.1007/s10529-017-2300-y
Santillan JY, Dettorre LA, Lewkowicz ES, Iribarren AM (2016) New and highly active microbial phosphotriesterase sources. FEMS Microbiol Lett. https://doi.org/10.1093/femsle/fnw276
Santillan JY, Rojas NL, Ghiringhelli PD et al (2020) Organophosphorus compounds biodegradation by novel bacterial isolates and their potential application in bioremediation of contaminated water. Bioresour Technol 317:124003. https://doi.org/10.1016/j.biortech.2020.124003
Singh BK (2009) Organophosphorus-degrading bacteria: ecology and industrial applications. Nat Rev Microbiol 7:156–164. https://doi.org/10.1038/nrmicro2050
Singh BK, Walker A, Morgan JAW, Wright DJ (2003) Effects of soil pH on the biodegradation of chlorpyrifos and isolation of a chlorpyrifos-degrading bacterium. Appl Environ Microbiol 69:5198–5206. https://doi.org/10.1128/AEM.69.9.5198-5206.2003
Singh BK, Walker A, Morgan JAW, Wright DJ (2003) Role of soil pH in the development of enhanced biodegradation of fenamiphos. Appl Environ Microbiol 69:7035–7043. https://doi.org/10.1128/AEM.69.12.7035-7043.2003
Sogorb MA, Vilanova E, Carrera V (2004) Future applications of phosphotriesterases in the prophylaxis and treatment of organophosphorus insecticide and nerve agent poisonings. Toxicol Lett 151:219–233. https://doi.org/10.1016/j.toxlet.2004.01.022
Theriot CM, Grunden AM (2011) Hydrolysis of organophosphorus compounds by microbial enzymes. Appl Microbiol Biotechnol 89:35–43. https://doi.org/10.1007/s00253-010-2807-9
Vidya Lakshmi C, Kumar M, Khanna S (2008) Biotransformation of chlorpyrifos and bioremediation of contaminated soil. Int Biodeterior Biodegrad 62:204–209. https://doi.org/10.1016/j.ibiod.2007.12.005
Yu YL, Fang H, Wang X et al (2006) Characterization of a fungal strain capable of degrading chlorpyrifos and its use in detoxification of the insecticide on vegetables. Biodegradation 17:487–494. https://doi.org/10.1007/s10532-005-9020-z
Yuan Y, Yang C, Song C et al (2011) Anchorage of GFP fusion on the cell surface of Pseudomonas putida. Biodegradation 22:51–61. https://doi.org/10.1007/s10532-010-9375-7
Donarski WJ, Dumas DP, Heitmeyer DP et al. (1989) Structure–activity relationships in the hydrolysis of substrates by the phosphotriesterase from Pseudomonas diminuta. Biochemistry 28:4650–4655. https://doi.org/10.1021/bi00437a021
Horne I, Harcourt RL, Sutherland TD et al. (2002) Isolation of a Pseudomonas monteilli strain with a novel phosphotriesterase. FEMS Microbiol Lett 206:51–55. https://doi.org/10.1016/S0378-1097(01)
Acknowledgements
This study was supported by a Grant of OPCW (L/ICA/ICB/190041/14), UNQ and ANPCyT (PICT 2011–2007). ESL and AMI are Members of the Scientific Researcher Career of CONICET. JYS and AM are CONICET Fellows.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Santillan, J.Y., Muzlera, A., Molina, M. et al. Microbial degradation of organophosphorus pesticides using whole cells and enzyme extracts. Biodegradation 31, 423–433 (2020). https://doi.org/10.1007/s10532-020-09918-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-020-09918-7