Abstract
Sharks developed life history traits that make them susceptible to overfishing. This is, in turn, a risk for extinction, and several species are affected. The high price of shark fins in the international trade has triggered the widespread capture of sharks at unsustainable levels, prompting illegal and unethical practices, such as finning. To address these concerns, the present study aimed to identify species composition using molecular techniques based on DNA barcoding and DNA polymorphism on samples taken from illegal shark fin seizures conducted by the Federal Environmental Agency of Brazil. A species-specific DNA-based identification from three finning seizures in Brazil found at least 20 species from 747 shark fins, some of which were identified as endangered and protected under Brazilian legislation, while others were representative of restricted catches, according to Appendix II of CITES. In the seizure from Belém, 338 fins were identified as belonging to at least 19 different species, while in the seizure from Natal 211, fins belonging to at least 8 different species were identified. Furthermore, 198 fins from Cananéia were identified through PCR-Multiplex as belonging to Isurus oxyrinchus. These results raise concerns about the environmental and socioeconomic effects of finning on developing countries. Furthermore, this study represents the first finning evaluation from Brazil in the Southwest Atlantic, highlighting the importance of developing policies aimed toward restricting and regulating the shark trade and detecting IUU fisheries and illegal trade of endangered species, mainly in developing countries, where fisheries management, surveillance, and species-specific fisheries catch data are often sporatic.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sharks usually play a major role in marine environments, and large species act mostly as food-web top-predators (Heithaus et al. 2012; Ferretti et al. 2010). Elasmobranchs are naturally more vulnerable to extinction than bony fishes (Myers and Worm 2005) because some species are “k-strategists” characterized by slow growth, late sexual maturity, high longevity, and low fertility, all characteristics which make them particularly susceptible to anthropogenic pressures, such as overfishing (García et al. 2008; Musick et al. 2000). Once overfished, shark populations require long recovery periods (Davidson et al. 2016; Stevens et al. 2000). However, notwithstanding their ecological importance, sharks are among the most threatened vertebrates, mainly because of intense fishing pressure (Bradley and Gaines 2014; Dulvy et al. 2014; Hoffmann et al. 2010).
In the past half century, shark fisheries have expanded rapidly worldwide. At first, they were captured by different fishing gear, such as driftnets, gillnets, purse-seine, and trawling nets, and their products were consumed locally. However, longline fisheries that target tuna (Thunnus spp.) and swordfish (Xiphias gladius) are mostly responsible for shark captures and bycatches (Bonfil 1994; Molina and Cooke 2012; Oliver et al. 2015; Stevens et al. 2000), and now they also supply the demand for shark fins in the international shark fin trade (CITES 2018). According to the Food and Agriculture Organization (FAO) of the United Nations (UN), shark fisheries reached a peak of nearly 880,000 tons in 2003, and, ever since, catches have decreased by 20% (Davidson et al. 2016; Dulvy et al. 2014). Despite this, the reported catches could be highly underestimated, mainly because these statistics have uncertainties with respect to small-scale artisanal catch data (Zeller et al. 2015), and they do not account for by-catches (Zeller and Pauly 2005), illegal catches and discards (Pauly and Zeller 2016), raising concerns about the true magnitude of ecological impact suffered by sharks and rays.
In the South Atlantic, artisanal, industrial and sport fisheries with different types of fishing gear capture several shark species, but land in different fishing ports along the Brazilian coast. For instance, Brazil reported a peak capture of sharks, rays, and chimaeras of 31,259 tons (t) in 1982, but a decrease of 44% in reported captures in 2017 (FAO 2019). As a consequence, different shark species are severely overexploited and now face extinction (e.g., Barreto et al. 2016; Lessa et al. 2016). Even though alarming, the lack of proper policymaking in Brazil’s fisheries management is unlikely to improve as a result of insufficient funding. Moreover, the failure to institute science-based regulations just favors legal breaches and increases the risk of extinction (Fiedler et al. 2017). In fact, sharks are caught on a large scale worldwide, and based on an analysis of average shark weight performed in 2000, mortality of 100 million sharks was estimated and almost 97 million in 2010 with a possible range of values between 63 and 273 million sharks per year (Worm et al. 2013). In their study, Clarke et al. (2006a) stated that the biomass of the global fin trade was four times higher than previously reported by the FAO and largely underestimated. Shark fin supplies come from different sources, but mainly from longline fisheries, by-catches on international waters, and exclusive economic zones of developed and developing countries (Barreto et al. 2016; Jaiteh et al. 2016, 2017; Schiller et al. 2015).
According to Dulvy et al. (2014), a worldwide decline in shark populations is likely associated with the international fin trade. The high demand for shark fin soup in the Asian market triggered widespread shark catches for fins and encouraged finning, an illicit act of removing fins from sharks and discarding the animals back into the ocean, sometimes still alive. This unethical and illegal act has led to unsustainable fishing pressure on these k-strategist species, and many are currently threatened with extinction (Clarke et al. 2006b; de Mitcheson et al. 2018; Dulvy et al. 2014, 2017). The highest incomes from the shark fin trade originate mainly from illegal, unreported, and unregulated (IUU) fisheries, which lack species-specific identification (Dulvy et al. 2014; Stevens et al. 2000) and are often omitted in fisheries catch reports (Pauly and Zeller 2016). As a result of fisheries mismanagement in Brazil, little is known about the shark fin trade. In general, however, we do know that shark meat is exported, but also consumed locally, while fins are mostly traded to Asia and mostly obtained through IUU fisheries. Furthermore, since 1998, Brazilian authorities have dismantled several local finning operations, and it was documented that Brazil is, indeed, a major supplier for the international fin trade (Barreto et al. 2017). Consequently, protected shark species are captured and traded without any regulation or documentation (Fields et al. 2018; Steinke et al. 2017). Nevertheless, field identification of processed fisheries products is, in most cases, unreliable. Dried shark fins are typically missing key diagnostic characteristics, making morphological species identification challenging, or even impossible (Fields et al. 2018), compromising surveillance and law enforcement.
To overcome this obstacle, DNA barcoding (Hebert et al. 2003) is a taxonomic method that employs standardized genetic markers in an organism’s DNA to identify it as belonging to a particular species. This makes it possible to identify unknown DNA samples to a registered species based on comparison to a reference library. Consequently, over the last few years, several studies have aimed to create a Chondrichthyes DNA database and tools for their identification in order to evaluate their biodiversity (Mendonça et al. 2009, 2010; Ribeiro et al. 2012; Ward et al. 2005, 2008, 2009; Wong et al. 2009; Cerutti-Pereyra et al. 2012). Moreover, this technique has been applied for shark fin species identification worldwide (Fields et al. 2015, 2018; Holmes et al. 2009; Liu et al. 2013; Sembiring et al. 2015; Shivji et al. 2002; Steinke et al. 2017). However, a growing number of studies have been dedicated to the application of these genetic approaches for law enforcement (Chapman et al. 2009; Barbuto et al. 2010; De-Franco et al. 2012; Domingues et al. 2013; Steinke et al. 2017; Bunholi et al. 2018; Feitosa et al. 2018; Ferrette et al. 2019). Therefore, the present study aims to apply DNA-based techniques, such as DNA barcoding and PCR-multiplex, to identify the species composition of shark fins from three finning seizures conducted by the Federal Environmental Agency in Brazil totaling more than 13 tons of seized fins. These results highlight the importance of conservation policies that aim to restrict and regulate the shark fin trade, as well as increase knowledge about finning and IUU fisheries in the southwestern Atlantic Ocean.
Materials and methods
Sampling
Seized shark fins were collected from three different surveillance operations conducted by the Brazilian Institute of Environment and Renewable Natural Resources (IBAMA), a federal agency under the Brazilian Ministry of Environment. In the first seizure, in the city of Belém in Pará state, northern Brazil, 8 tons of dried fins were seized from a finning recidivist Brazilian company exporting to China, and a total of 400 dried fins were randomly sampled. In the second seizure, more than 5 tons of dried fins from two foreign fishing companies, one Spanish and the other Chinese, both accused of finning in the city of Natal, Rio Grande do Norte state, northeastern Brazil, were randomly sampled, totaling 300 fins. During the third seizure, 198 fresh fin tissues were sampled from a Brazilian pelagic longline fishing company during landings in the city of Cananéia, São Paulo, southeastern Brazil, and this company was also accused of finning (Fig. 1). All sampled tissue fragments were stored in 100% ethyl alcohol at − 20 °C and were supplied by IBAMA.
DNA-based species identification
Genomic DNA from fins was extracted using the NucleoSpin® Tissue kit (MACHEREY–NAGEL GmbH and Co. KG). Samples from the first two seizures were identified by amplifying cytochrome c oxidase subunit I (COI), a mitochondrial DNA (mtDNA) gene. Polymerase chain reaction (PCR) was carried out according to the parameters and specifications of Platinum®Taq DNA Polymerase (Invitrogen™) using two primers (Forward: FishF1: 5′-TCA ACC AAC CAC AAA GAC ATG GGC AC-3′ and Reverse: FishR1: 5′-TAG AGT TCT GGG TGG CCA AAG AAT CA-3′) (Ward et al. 2005). For sequencing reaction, PCR products were submitted to an enzymatic purification protocol using the ExoSAP-IT™ PCR Clean-Up kit (Applied Biosystems™). The sequencing protocols followed the procedures of the BigDye™ Terminator v3.1 Sequencing Kit (Applied Biosystems™), and sequences were generated on the ABI 3130 Genetic Analyzer (Applied Biosystems™). In addition, shark fin seizures from Cananéia came exclusively from pelagic longline fisheries. Accordingly, a PCR-multiplex developed by Shivji et al. (2002) for species-specific pelagic shark identification was used. This system was applied to differentiate DNA sequences from among different pelagic shark species using the nuclear ribosomal internal transcribed spacer (ITS) 2 locus. Such method combines six pelagic shark primers (P. glauca, Isurus oxyrinchus, I. paucus, Lamna nasus, Carcharhinus falciformis, and C. obscurus) with two universal primers in an eight-primer single tube reaction. This PCR-multiplex was performed by changing the annealing temperature to 65 °C to provide a diagnostic-sized amplicon for each shark species. The reaction effectiveness was analyzed on a 1.5% agarose gel under UV light.
Samples from Natal and Belém seizures were identified using two methods: (i) a sequence similarity-based identification and (ii) a tree-based identification (van Velzen et al. 2012). First, a sequence similarity comparison was undertaken to match the closest sequence on the public library databases, such as the Barcode of Life Data (BOLD) system, version 4 (http://v4.boldsystems.org/) (Ratnasingham and Hebert 2007) and GenBank® (https://www.ncbi.nlm.nih.gov/genbank/) (Benson et al. 2017), using similarity scores (e.g., BLAST, https://blast.ncbi.nlm.nih.gov/Blast.cgi) (Altschul et al. 1990) and assigning the species label of the closest reference to the query. Second, a tree-based strategy used a classic phylogenetic method (Ronquist and Huelsenbeck 2003) to assign unidentified barcodes to species based on their clusters in the tree, attempting to estimate the phylogeny between the reference barcodes and the query sequence. The query was assigned to the species within which it was clustered.
Following this, we performed the two phylogenetic methods with the generated sequences coupled with the highest similarity scores for species sequences downloaded from Barcode Index Numbers (BINs) (Ratnasingham and Hebert 2013) in BOLD v4, as described in Online Resource 1. The sequence of Collared Carpetshark Parascyllium_collare (FMVIC281-08) was used as outgroup. All sequences were aligned using MUSCLE (multiple sequence comparison by log-expectation) algorithms (Edgar 2004) in Geneious R11.1.5 (Kearse et al. 2012).
The COI matrix was tested for nucleotide substitution saturation (Xia et al. 2003) with DAMBE6 (Xia 2017) and showed no significant saturation. The best-fit nucleotide substitution model, HKY+I+G (pinv = 0.6, α = 1.65), was evaluated by the Bayesian Information Criterion (BIC) and Decision Theory (DT) performance-based selection with jModelTest2 on XSEDE (Darriba et al. 2012; Towns et al. 2014) through CIPRES Science Gateway v3.3 (Miller et al. 2010). A Maximum-Likelihood (ML) phylogenetic tree was constructed by PhyML v 3.3 (Guindon and Gascuel 2003; Guindon et al. 2010) implemented in Geneious R11.1.5. A BioNJ (Gascuel 1997) tree was used as the starting tree to define the strategy of the tree topology search as the best of Nearest Neighbor Interchanges (NNI) and Subtree Pruning and Regrafting (SPR) with 1000 non-parametric bootstrap replicates. The PhyML tree was set as the initial tree of Bayesian inference performed in BEAST v1.10.4 (Suchard et al. 2018), adopting an unregulated relaxed lognormal clock (Drummond et al. 2006) and the Yule speciation process (Gernhard 2008; Yule 1925) as priors. The posterior probability of parameters was estimated using Markov chain Monte Carlo (MCMC) with 500 million generations and 10% of burn-in. The model convergence, effective sample size (ESS) and 95% of highest posterior density (HPD) intervals were all calculated by TRACER, v1.7.1 (Rambaut et al. 2018). TreeAnnotator, v1.10.4, summarized the information of a sample of trees produced by BEAST onto a single Highest Log Clade Credibility Tree. FigTree, v1.4.4 (Rambaut 2012) was used to view the phylogenies.
Results
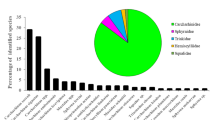
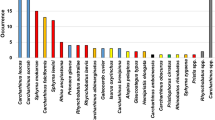
A total of 747 shark fins were sequenced and identified at the lowest taxonomic level, using 651 base pairs (bp) of the COI gene and PCR-Multiplex. Two orders were identified (Carcharhiniformes and Lamniformes), four families (Carcharhinidae, Sphyrnidae, Alopiidae, and Lamnidae) and at least 20 shark species (Table 1). The seizure from Belém was the most sampled and biodiverse, with a total of 338 shark fins identified, and at least 19 coastal and pelagic species were found. The seizure from Natal was the second in sampling size (n = 211), accounting for at least seven pelagic species (Table 1; Online Resources 2 and 3).
Prionace glauca, Isurus oxyrinchus, and Carcharhinus porosus were the most abundant identified species, representing 33.73%, 27.71%, and 13.12%, respectively (Table 1). Miscellaneous shark species from Carcharhinidae, Sphyrnidae, Alopiidae, and Lamnidae accounted for 25.43% of the total sampling (Table 1). At least nine identified species are listed in the threatened categories of the IUCN Red List, accounting for almost 40% of all individuals (Fig. 2a; Table 1), whereas 20.61% of the total is also on the Brazilian Red List (Fig. 2b; Table 1). In addition, six identified species (i.e., C. falciformis, Sphyrna lewini, S. mokarran, Isurus oxyrinchus, I. paucus and Alopias superciliosus) are currently listed in Appendix II of CITES.
Alluvial diagrams, built by RAWGraphs (Mauri et al. 2017), represent correlations between the number of identified species and their conservation status according to the IUCN Red List of Threatened Species™ on the left and the Chico Mendes Institute for Biodiversity Conservation (ICMBio) from Brazil on the right. a Belém seizure (n = 400). b Natal seizure (n = 300). CR critically endangered, EN endangered, VU vulnerable, NT near threatened, LC least concern, DD data deficient
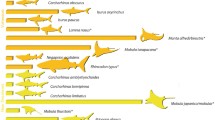
All sequences matched BOLD and GenBank® reference libraries with high similarity (> 99%). Most identified species clustered into high-support clades with DNA Barcode BIN sequences in the phylogenetic tree, evidencing success in genetic identification, except C. obscurus and C. galapagensis that could not be differentiated by only a single-locus approach and thus clustered within the same clade (Fig. 3). All sequences from the Belém (accession nos. MH911009–MH911346) and Natal (accession nos. MH719774–MH719984) seizures were deposited in GenBank®. In the third evaluated finning seizure in Cananéia, using the PCR-Multiplex approach, all shark fins (n = 198) were identified as belonging to only a single species, I. oxyrinchus (Fig. 2; Online Resource 4), according to the amplicon size pattern determined by gel electrophoresis (Fig. 4).
Bayesian phylogenetic inference tree of cytochrome c oxidase subunit I (COI) using 132 haplotypes with 652 base pairs of sequences from BOLD, GenBank®, and from this study. Node values indicate the posterior probability of trees using an evolutionary model based on prior probabilities, producing the plausible phylogenetic tree for the given data. PA and RN codes comprise the sequenced individuals from this study
Discussion
This evaluation is the first genetic identification of shark finning seizures from Brazil in the Southwest Atlantic. Two of the biggest finning seizures in Brazil were sampled, resulting in multimillion-dollar penalties against the law-breaking companies (Barreto et al. 2017). In Belém, at least 19 shark species were identified, a quantity of pelagic species equal to that of coastal species, a statistic indicating that fins came from different fishing modalities and gear. On the other hand, in Natal and Cananéia, mainly pelagic species were identified, indicating that they were caught mostly by a longline fishing fleet on the high seas.
Our results are similar to other evaluations from finning, such as those of Steinke et al. (2017), who also identified 20 shark species from dried fins and gill plates obtained in Canada, China, and Sri Lanka. Despite high species diversity, according to Fields et al. (2018), the contemporary shark fin trade is dominated by only 8 species or complexes, likely comprising more than 1%: P. glauca, C. falciformis, S. lewini and S. zygaena, Carcharhinus spp., C. brevipina, C. leucas, and Isurus spp., as evidenced also by our results. The most abundant identified shark species were P. glauca, I. oxyrinchus, and C. porosus, accounting altogether for almost 75% of all individuals (Table 1). Although, P. glauca is not considered overfished in the Atlantic (ICCAT 2015), Indian (IOTC 2017), or Pacific (ISC 2017) Oceans, it is the most captured shark worldwide by high seas fisheries (Coelho et al. 2018; Oliver et al. 2015) and also the species supplying the international shark fin trade (Clarke et al. 2006a, b; Fields et al. 2018). Isurus oxyrinchus was the second most frequently identified species. This species is also the second most frequently found in the Hong Kong fin trade (Fields et al. 2018), and it is currently overfished in the North Atlantic. The International Commission for the Conservation of Atlantic Tuna (ICCAT) recommends a catch-quota reduction, while in the South Atlantic, the stocks could be overfished, but owing to assessment uncertainties, ICCAT recommends that catch-quota should not be raised (de Bruyn 2017). The species is currently globally assessed as “Endangered” (EN) (Rigby et al. 2019) since its weighted global population trend estimated a high probability of 50–79% reduction over three generation lengths (72–75 years) and slow stock rebuilding, even with prohibited capture (Sims et al. 2018).
Moreover, ecological risk assessment based on biological productivity and susceptibility, as recently adopted by the ICCAT for data-poor fisheries, has demonstrated that P. glauca and I. oxyrinchus are among the most vulnerable shark stocks from longline fisheries in the Atlantic Ocean (Cortés et al. 2015). Most fishing of pelagic sharks in the South Atlantic Ocean occurs on the high seas, mostly as by-catch by longline fisheries targeting tunas (Scombridae) and billfishes (Xiphiidae and Istiophoridae), and they are retained on board for their fins (Bonfil 1994). Over the last 40 years, few data were available for catch rates, relative abundances, and distributions of the captured species. Most information was from small-scale longline fleets operating off Brazil and Uruguay where sharks historically have been landed and sold in local markets (Hazin et al. 2008). Atlantic high seas longline fisheries are responsible for about 25% of reported global shark catches (Clarke 2008). Fishing has been historically high in the northern and southern Atlantic, but it intensified after the 1990s (Amorim et al. 1998; Camhi et al. 2008). Simultaneous exploitation in the South Atlantic by multinational fleets may have also resulted in high levels of underreporting, which, when coupled with species migratory patterns, has interfered with data collection. Since these fleets changed their target strategies over time owing to market demands, technological advancements, and declines in the abundance of target species (Hazin et al. 2008).
Carcharhinus porosus was the third most commonly identified species. This shark species is a known bycatch of artisanal gillnet fisheries in Brazil’s northern region, the center of abundance for this species (Lessa et al. 1999). The increasing fishing pressure, evidence of declines and the lack of management combine to make this species “Critically Endangered” (CR) in Brazil (Lessa et al. 2006). In fact, almost 40% of shark fins belong to species threatened by extinction, and at least a fifth is also protected in Brazil (Table 1; Fig. 2a, b) by Ordinance No. 445 of the Ministry of Environment (MMA), December 17, 2014, which prohibits their capture, transportation, trade, and storage.
Carcharhinus falciformis, Sphyrna lewini, S. mokarran, and Alopias superciliosus, as well as Isurus oxyrinchus and I. paucus with recent declines from 60–96% worldwide (CITES-CoP18 2019), are also listed in Appendix II of CITES. This includes species not necessarily threatened, but for which trade must be regulated in order to allow their sustainability. Such situation has arisen as a result of inadequate management measures, poor law enforcement and lack of control over the level of international trade, causing declines of over 70% across their range and driving them to extinction in some areas. In other finning evaluations, a high number of CITES-listed and endangered species were also identified (de Mitcheson et al. 2018; Cardeñosa et al. 2018, 2019).
This fact demonstrates the harm that illegal practices, such as finning, causes to the marine environment and shark conservation and management efforts. Even so, the biomass of most large pelagic predators has shown a vigorous decline, and the stock of shark, billfish and tuna cannot support ongoing fishing. Thus, the recovery of these populations, especially for large sharks, toward a sustainable fishery could only be accomplished with a significant reduction in fishing in the South Atlantic (Bornatowski et al. 2018). Otherwise, the removal of large pelagic predators could trigger cascading top-down effects (Pauly et al. 2001; Arancibia and Neira 2005; Sibert et al. 2006; Baeta et al. 2009; Ferretti et al. 2010; Freire and Pauly 2010).
Apart from the identified threatened species, most other species have been classified as non-threatened, such as “Near Threatened” (NT), “Least Concern” (LC) or “Data Deficient” (DD) (Table 1; Fig. 2). According to the IUCN, more than 80% of all Chondrichthyans are in the non-threatened categories, mostly as a result of uncertainty about the true level of extinction risk, as well as the failure to understand the real patterns of threats and risks (Butchart and Bird 2010), all of which causes truly threatened data-deficient species to be neglected by conservation programs (Bland et al. 2015). Determining the true conservation status of these species is essential to developing an accurate picture of global biodiversity and enabling the protection of threatened species. In Brazil, according to Ordinance MMA No. 43, January 31, 2014, species assessed as non-threatened should be prioritized for research and conservation measures. The results of this evaluation demonstrate that several shark species in the non-threatened categories face finning pressure, putting at risk of extinction data-deficient species that could already be threatened.
Miscellaneous carcharhinid shark species, accounting for almost 18% of the total seized, were identified as C. acronotus, C. brachyurus, C. limbatus, C. perezi, C. porosus, C. signatus, C. galapagensis and/or C. obscurus, G. cuvier, R. lalandii, and R. porosus. Many of these species have been previously identified in the international fin trade (Clarke et al. 2006a, b; Steinke et al. 2017; Feitosa et al. 2018; Fields et al. 2018). Among these species, C. galapagensis and C. obscurus cannot be distinguished by only a single-locus approach since they show a pervasive mitochondrial admixture whereby they could only be distinguished by Single Nucleotide Polymorphisms (SNPs) (Corrigan et al. 2017). Thus, with the COI gene, both species were clustered in the same BIN in BOLD and within the same clade in the phylogenetic reconstruction.
Among the Carcharhinidae family, C. falciformis is a common species in purse-seine and pelagic longline fisheries. Currently, their population size has plummeted in the Atlantic Ocean (Barreto et al. 2016; Cortés et al. 2010) by their highly valuable fins. Consequently, in the Atlantic Ocean, the capture and commercialization of this species are prohibited (ICCAT 2011). Globally, this shark species has been recently listed in Appendix II of CITES, and in spite of all these facts, C. falciformis is one of the most traded species in Hong Kong (Clarke et al. 2006a, b; Fields et al. 2018). Obviously, we evidenced here that this and other species listed in CITES continue to be illegally captured, landed and traded in Brazil (Almerón-Souza et al. 2018; Barreto et al. 2016).
Other endangered shark species, some also listed in Appendix II of CITES, such as hammerhead sharks (e.g., S. lewini, S. mokarran, S. tiburo, and S. tudes) and the bigeye thresher shark, Alopias superciliosus, were also identified in lower frequencies. According to the CITES Trade Database (CITES 2018), Hong Kong is the main CITES-listed species importer. Fins reportedly traded in 2015 for the three large hammerhead sharks, S. lewini (40.8%) by weight, S. zygaena, (35%), and S. mokarran (16.8%), account for the majority of fins reportedly traded (Cardeñosa et al. 2018). Regarding A. superciliosus, the major threat to thresher sharks is fishery-related mortality, as they are taken mainly for their highly valuable fins. Besides our identification, the species was previously detected in the Hong Kong international fin trade (Clarke et al. 2006a, b; Cardenõsa et al. 2018; Fields et al. 2018), and estimations suggest a huge underreporting of these species (Clarke et al. 2006a).
The most recent and significant genetic assessment of international fin trade of Hong Kong was conducted by Fields et al. (2018) who identified at least 76 different species from 4800 sampled fins. Most traded fins corresponded to the orders Carcharhiniformes and Lamniformes, as also revealed by this study and by other evaluations (Clarke et al. 2006a, b; Steinke et al. 2017). In their results, Fields et al. (2018) reported that almost half of the species were from the family Carcharhinidae, and one-third was categorized as critically endangered. Moreover, in our results, most fins belonged to the Carcharhinidae and Lamnidae families, accounting for 64.8% and 29.8%, respectively. However, almost 40% of the identified species in our evaluation were assessed as EN or VU. The rising consumption of threatened species associated with the high demand for luxury wildlife products, such as the shark fin soup, is putting a heavy pressure on several species increasingly unable to sustain the current levels of exploitation and trade. Lack of proper management and science-based decision making, as well as poor law enforcement, influential lobbying interests from fishing companies, and weak, or no, regulation in the product’s source countries and/or demand centers all raise the risk of extinction of exploited species (de Mitcheson et al. 2018).
Hong Kong has been traditionally a major consumer of shark fins, as well as the principal international hub for about 50% of globally traded dried shark fins, much of which is re-exported to mainland China (Clarke et al. 2006a, b; de Mitcheson et al. 2018; Shea and To 2017). Imports of dried shark fins into Hong Kong grew six-fold between 1960 and 2002, declining to one-third of 1960 levels by 2016 to almost 6000 tons (de Mitcheson et al. 2018). Hong Kong government data show a drop in shark fin re-exports to China of nearly 80% in the last decade. The overall consumption outside mainland China has not declined significantly, nor has commercial fishing pressure on sharks (Clarke 2004; Clarke et al. 2007; Steinke et al. 2017; Fields et al. 2018; Cardeñosa et al. 2018). Furthermore, trade in Hong Kong appears to be declining, possibly from consumer and retailer educational campaigns around shark fin consumption, and evidence of this also comes from other trade hubs, such as Macau and Vietnam (Fabinyi et al. 2012).
On the other hand, Brazil is one of the many worldwide fin exporters to Hong Kong, and since 2004, more than 83 tons of fins were seized in Brazil and companies summarily charged with illegal finning (Barreto et al. 2017). Of the 20 identified shark species studied, the outcome of our genetic evaluation shows that most are threatened with extinction. Some are even CITES-listed as internationally trade-regulated, and a fifth is protected. These results represent the first DNA-based assessment of finning seizures from Brazil and demonstrate that IUU fisheries have an impact on endangered shark species when coupled with illegal practices like finning, raising, in turn, concerns about the lack of fishery surveillance in developing countries which continue to supply the international shark fin trade. In 2016, Brazil was ranked as the 10th country for reported shark landings, accounting for almost 18,700 tons, while it is the 11th shark fin exporter (FAO 2018).
Brazil has its own set of regulations for shark landings and transportation on board, namely requiring that all fins be attached to the shark during landings (MPA/MMA No. 14, November 26, 2012). Despite this, no nationally standardized fisheries data collection system has been available since 2007 in Brazil, and total catches may be almost two-fold those reported to FAO (Freire et al. 2015). The absence of fisheries species-specific catch data hinders efforts to monitor the capture of threatened and protected species, to implement fisheries good practices, and to maintain sustainable stocks (Di Dario et al. 2015). According to Barreto et al. (2017), only a few companies are authorized to export shark fins. However, no reliable information specifies which species or quantities are being traded. In addition, at least fifteen raids to disrupt illegal fin trading in Brazil were carried out between 2004 and 2013. Therefore, owing to the absence of a reliable standardized fisheries species-specific collection data system and the widespread presence of IUU fisheries, we can conclude that Brazil is currently one of the major hotspots for the international shark fin black market.
The Brazilian fisheries management system is obsolete and cannot promote sustainable fisheries or prevent overexploitation in the absence of shared strategic management in the southwestern Atlantic (Griffith 2008; Symes 2006). Furthermore, fisheries legislation is complex, conservation efforts are weak, as are surveillance and law enforcement (Cochrane and Doulman 2005; Pinheiro et al. 2015; Fiedler et al. 2017). Therefore, an efficient fisheries management policy should not be based on models that do not account for the complex effects of the interaction among the marine ecosystem, capture quotas, and the exploited species. Instead, they should be based on scientific data for mid- and long-term sustainable management. This will require an investment in scientific research, new technologies and the application of viable public policies, conservation plans and fisheries management (Walters 2007; Fromentin et al. 2014; Fiedler et al. 2017) to the exclusion of fisheries lobbying interests.
To conclude, the results demonstrate the efficiency of DNA-based identification methodologies and evidence the disastrous impact of finning on marine ecosystems and developing countries. Endangered and protected species are illegally traded and demand new tools for proper surveillance, highlighting the importance of public–private partnerships to encourage the implementation of DNA-based technology for effective law enforcement and monitoring of wildlife trade, as well as the conservation of protected and endangered species and the detection of commercial fraud.
References
Almerón-Souza F, Sperb C, Castilho CL, Figueiredo PI, Gonçalves LT, Machado R, Oliveira LR, Valiati VH, Fagundes NJ (2018) Molecular identification of shark meat from local markets in southern brazil based on DNA barcoding: evidence for mislabeling and trade of endangered species. Front Genet 9:138. https://doi.org/10.3389/fgene201800138
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Amorim AF, Arfelli CA, Fagundes L (1998) Pelagic elasmobranchs caught by longliners off southern Brazil during 1974–97: an overview. Mar Freshw Res 49(7):621–632. https://doi.org/10.1071/MF97111
Arancibia H, Neira S (2005) Long-term changes in the mean trophic level of Central Chile fishery landings. Sci Mar 69(2):295–300. https://doi.org/10.3989/scimar.2005.69n2295
Baeta F, Costa MJ, Cabral H (2009) Changes in the trophic level of Portuguese landings and fish market price variation in the last decades. Fish Res 97(3):216–222. https://doi.org/10.1016/j.fishres.2009.02.006
Barbuto M, Galimberti A, Ferri E, Labra M, Malandra R, Galli P, Casiraghi M (2010) DNA barcoding reveals fraudulent substitutions in shark seafood products: the Italian case of “palombo” (Mustelus spp.). Food Res Int 43(1):376–381. https://doi.org/10.1016/j.foodres.2009.10.009
Barreto R, Ferretti F, Flemming JM, Amorim A, Andrade H, Worm B, Lessa R (2016) Trends in the exploitation of South Atlantic shark populations. Conserv Biol 30:792–804. https://doi.org/10.1111/cobi12663
Barreto RR, Bornatowski H, Motta FS, Santander-Neto J, Vianna GMS, Lessa R (2017) Rethinking use and trade of pelagic sharks from Brazil. Mar Policy 85:114–122. https://doi.org/10.1016/j.marpol.2017.08.016
Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW (2017) GenBank. Nucleic Acids Res 46:D41–D47. https://doi.org/10.1093/nar/gkx1094
Bland LM, Collen BEN, Orme CDL, Bielby JON (2015) Predicting the conservation status of data-deficient species. Conserv Biol 29:250–259. https://doi.org/10.1111/cobi12372
Bonfil R (1994) Overview of world elasmobranch fisheries. Food and Agriculture Organization of the United Nations, Rome
Bornatowski H, Angelini R, Coll M, Barreto RR, Amorim AF (2018) Ecological role and historical trends of large pelagic predators in a subtropical marine ecosystem of the South Atlantic. Rev Fish Biol Fish 28(1):241–259. https://doi.org/10.1007/s11160-017-9492-z
Bradley D, Gaines SD (2014) Extinction risk: counting the cost of overfishing on sharks and rays. eLife 3:e02199. https://doi.org/10.7554/eLife02199
Bunholi IV, Ferrette BLS, De Biasi JB, Magalhães CO, Rotundo MM, Oliveira C, Foresti F, Mendonça FF (2018) The fishing and illegal trade of the angelshark: dNA barcoding against misleading identifications. Fish Res 206:193–197. https://doi.org/10.1016/jfishres201805018
Butchart SH, Bird JP (2010) Data deficient birds on the IUCN Red List: what don’t we know and why does it matter? Biol Conserv 143:239–247. https://doi.org/10.1016/jbiocon200910008
Camhi MD, Fordham SV, Fowler SL (2008) Domestic and international management for pelagic sharks. Sharks of the open ocean. Blackwell, Oxford, pp 418–444
Cardeñosa D, Fields AT, Babcock EA, Zhang H, Feldheim K, Shea SK, Fischer GA, Chapman DD (2018) CITES-listed sharks remain among the top species in the contemporary fin trade. Conserv Lett 11:e12457. https://doi.org/10.1111/conl12457
Cardeñosa D, Shea KH, Zhang H, Feldheim K, Fischer GA, Chapman DD (2019) Small fins, large trade: a snapshot of the species composition of low-value shark fins in the Hong Kong markets. Anim Conserv. https://doi.org/10.1111/acv.12529
Cerutti-Pereyra F, Meekan MG, Wei NWV, O’Shea O, Bradshaw CJ, Austin CM (2012) Identification of rays through DNA barcoding: an application for ecologists. PLoS ONE 7(6):e36479. https://doi.org/10.1371/journal.pone.0036479
Chapman DD, Pinhal D, Shivji MS (2009) Tracking the fin trade: genetic stock identification in western Atlantic scalloped hammerhead sharks Sphyrna lewini. Endanger Species Res 9:221–228. https://doi.org/10.3354/esr00241
CITES (2018) CITES trade statistics derived from the CITES Trade Database. UNEP World Conservation Monitoring Centre, Cambridge. https://trade.cites.org/. Accessed 14 Dec 2018
CITES-CoP18 (2019) Eighteenth meeting of the conference of the parties of the convention on international trade in endangered species of Wild Fauna and Flora. https://citessharks.org/. Accessed 30 Aug 2019
Clarke S (2004) Shark product trade in Hong Kong and mainland China and implementation of the CITES shark listings. TRAFFIC East Asia, Hong Kong, China.
Clarke S (2008) Use of shark fin trade data to estimate historic total shark removals in the Atlantic Ocean. Aquat Living Resour 21(4):373–381. https://doi.org/10.1051/alr:2008060
Clarke SC, Magnussen JE, Abercrombie DL, McAllister MK, Shivji MS (2006a) Identification of shark species composition and proportion in the Hong Kong shark fin market based on molecular genetics and trade records. Conserv Biol 20:201–211. https://doi.org/10.1111/j1523-1739200500247x
Clarke SC, McAllister MK, Milner-Gulland EJ, Kirkwood GP, Michielsens CG, Agnew DJ, Pikitch EK, Nakano H, Shivji MS (2006b) Global estimates of shark catches using trade records from commercial markets. Ecol Lett 9:1115–1126. https://doi.org/10.1111/j1461-0248200600968x
Clarke S, Milner-Gulland EJ, Bjørndal T (2007) Social, economic, and regulatory drivers of the shark fin trade. Mar Res Econ 22(3):305–327
Cochrane KL, Doulman DJ (2005) The rising tide of fisheries instruments and the struggle to keep afloat. Philos Trans R Soc B 360(1453):77–94. https://doi.org/10.1098/rstb.2004.1568
Coelho R, Mejuto J, Domingo A, Yokawa K, Liu KM, Cortés E, Romanov EV, Silva C, Hazin FHV, Arocha F, Mwilima AM (2018) Distribution patterns and population structure of the blue shark (Prionace glauca) in the Atlantic and Indian Oceans. Fish Fish 19:90–106. https://doi.org/10.1111/faf12238
Corrigan S, Delser PM, Eddy C, Duffy C, Yang L, Li C, Bazinet AL, Mona S, Naylor GJ (2017) Historical introgression drives pervasive mitochondrial admixture between two species of pelagic sharks. Mol Phylogenet Evol 110:122–126. https://doi.org/10.1016/jympev201703011
Cortés E, Arocha F, Beerkircher L, Carvalho F, Domingo A, Heupel M, Holtzhausen H, Santos MN, Ribera M, Simpfendorfer C (2010) Ecological risk assessment of pelagic sharks caught in Atlantic pelagic longline fisheries. Aquat Living Resour 23:25–34. https://doi.org/10.1051/alr/2009044
Cortés E, Domingo A, Miller P, Forselledo R, Mas F, Arocha F, Campana S, Coelho R, Da Silva C, Hazin FHV, Holtzhausen H (2015) Expanded ecological risk assessment of pelagic sharks caught in Atlantic pelagic longline fisheries. Collect Vol Sci Pap ICCAT 71:2637–2688
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772. https://doi.org/10.1038/nmeth2109
Davidson LN, Krawchuk MA, Dulvy NK (2016) Why have global shark and ray landings declined: improved management or overfishing? Fish Fish 17:438–458. https://doi.org/10.1111/faf12119
de Bruyn P (2017) Report of the 2017 ICCAT Shortfin mako assessment meeting. International Commission for the Conservation of Atlantic Tunas, Madrid
de Mitcheson YS, Andersson AA, Hofford A, Law CS, Hau LC, Pauly D (2018) Out of control means off the menu: the case for ceasing consumption of luxury products from highly vulnerable species when international trade cannot be adequately controlled; shark fin as a case study. Mar Policy 98:115–120
De-Franco B, Mendonça FF, Oliveira C, Foresti F (2012) Illegal trade of the guitarfish Rhinobatos horkelii on the coasts of central and southern Brazil: genetic identification to aid conservation. Aquat Conserv 22:272–276. https://doi.org/10.1002/aqc2229
Di Dario F, Alves CB, Boos H, Frédou FL, Lessa RP, Mincarone MM, Pinheiro MA, Polaz CN, Reis RE, Rocha LA, Santana FM (2015) A better way forward for Brazil’s fisheries. Science 347(6226):1079. https://doi.org/10.1126/science.347.6226.1079-a
Domingues RR, Amorim AF, Hilsdorf AWS (2013) Genetic identification of Carcharhinus sharks from the southwest Atlantic ocean (Chondrichthyes: Carcharhiniformes). J Appl Ichthyol 29:738–742. https://doi.org/10.1111/jai12154
Drummond AJ, Ho SY, Phillips MJ, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLoS Biol 4:e88. https://doi.org/10.1371/journalpbio0040088
Dulvy NK, Fowler SL, Musick JA, Cavanagh RD, Kyne PM, Harrison LR, Carlson JK, Davidson LN, Fordham SV, Francis MP, Pollock CM (2014) Extinction risk and conservation of the world’s sharks and rays. eLife 3:e00590. https://doi.org/10.7554/eLife00590
Dulvy NK, Simpfendorfer CA, Davidson LN, Fordham SV, Bräutigam A, Sant G, Welch DJ (2017) Challenges and priorities in shark and ray conservation. Curr Biol 27:R565–R572. https://doi.org/10.1016/jcub201704038
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Fabinyi M, Pido M, Harani B, Caceres J, Uyami-Bitara A, De las Alas A, Buenconsejo J, Ponce de Leon EM (2012) Luxury seafood consumption in China and the intensification of coastal livelihoods in Southeast Asia: the live reef fish for food trade in Balabac, Philippines. Asia Pac Viewp 53:118–132. https://doi.org/10.1111/j1467-8373201201483x
FAO (2018) FishStatJ—software for fishery statistical time series. FAO Fisheries and Aquaculture Department, Rome. http://www.fao.org/fishery/. Accessed 16 Sept 2018
FAO (2019) Fisheries and Aquaculture Department. Capture fisheries. FAO Fisheries and Aquaculture Department, Rome. http://www.fao.org/fishery/. Assessed 12 Aug 2019
Feitosa LM, Martins APB, Giarrizzo T, Macedo W, Monteiro IL, Gemaque R, Nunes JLS, Gomes F, Schneider H, Sampaio I, Souza R (2018) DNA-based identification reveals illegal trade of threatened shark species in a global elasmobranch conservation hotspot. Sci Rep 8:3347. https://doi.org/10.1038/s41598-018-21683-5
Ferrette BLS, Domingues RR, Rotundo MM, Miranda MP, Bunholi IV, De Biasi JB, Oliveira C, Foresti F, Mendonça FF (2019) DNA barcode reveals the bycatch of endangered batoids species in the southwest atlantic: implications for sustainable fisheries management and conservation Efforts. Genes 10(4):304. https://doi.org/10.3390/genes10040304
Ferretti F, Worm B, Britten GL, Heithaus MR, Lotze HK (2010) Patterns and ecosystem consequences of shark declines in the ocean. Ecol Lett 13:1055–1071. https://doi.org/10.1111/j1461-0248201001489x
Fiedler FN, Port D, Giffoni BB, Sales G, Fisch F (2017) Pelagic longline fisheries in southeastern/south Brazil. Who cares about the law? Mar Policy 77:56–64. https://doi.org/10.1016/j.marpol.2016.12.011
Fields AT, Abercrombie DL, Eng R, Feldheim K, Chapman DD (2015) A novel mini-DNA barcoding assay to identify processed fins from internationally protected shark species. PLoS ONE 10(2):e0114844. https://doi.org/10.1371/journal.pone.0114844
Fields AT, Fischer GA, Shea SK, Zhang H, Abercrombie DL, Feldheim KA, Babcock EA, Chapman DD (2018) Species composition of the international shark fin trade assessed through a retail-market survey in Hong Kong. Conserv Biol 32:376–389. https://doi.org/10.1111/cobi13043
Freire KMF, Pauly D (2010) Fishing down Brazilian marine food webs, with emphasis on the east Brazil large marine ecosystem. Fish Res 105(1):57–62. https://doi.org/10.1016/j.fishres.2010.02.008
Freire KMF, Aragão JAN, Araújo ARR, Ávila-da-Silva AO, Bispo MCS, Canziani GV, Carneiro MH, Carneiro MH, Gonçalves FDS, Keunecke KA, Mendonça JT, Divovich E (2015) Reconstruction of catch statistics for Brazilian marine waters (1950-2010). In: Freire KMF, Pauly D (eds) Fisheries catch reconstructions for Brazil’s mainland and oceanic islands. Fisheries centre research reports, vol 23. Fisheries Centre. University of British Columbia, Vancouver, pp 3–30
Fromentin JM, Bonhommeau S, Arrizabalaga H, Kell LT (2014) The spectre of uncertainty in management of exploited fish stocks: the illustrative case of Atlantic bluefin tuna. Mar Policy 47:8–14. https://doi.org/10.1016/j.marpol.2014.01.018
García VB, Lucifora LO, Myers RA (2008) The importance of habitat and life history to extinction risk in sharks, skates, rays and chimaeras. Proc R Soc Lond B 275:83–89. https://doi.org/10.1098/rspb20071295
Gascuel O (1997) BIONJ: an improved version of the NJ algorithm based on a simple model of sequence data. Mol Biol Evol 14:685–695. https://doi.org/10.1093/oxfordjournalsmolbeva025808
Gernhard T (2008) The conditioned reconstructed process. J Theor Biol 253:769–778. https://doi.org/10.1016/jjtbi200804005
Griffith DR (2008) The ecological implications of individual fishing quotas and harvest cooperatives. Front Ecol Environ 6(4):191–198. https://doi.org/10.1890/050060
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704. https://doi.org/10.1080/10635150390235520
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. https://doi.org/10.1093/sysbio/syq010
Hazin FH, Broadhurst MK, Amorim AF, Arfelli CA, Domingo A (2008) Catches of pelagic sharks by subsurface longline fisheries in the South Atlantic Ocean during the last century: a review of available data with an emphasis on Uruguay and Brazil. Sharks Open Ocean Biol Fish Conserv. https://doi.org/10.1002/9781444302516
Hebert PD, Cywinska A, Ball SL, Dewaard JR (2003) Biological identifications through DNA barcodes. Proc R Soc Lond B 270(1512):313–321. https://doi.org/10.1098/rspb.2002.2218
Heithaus MR, Wirsing AJ, Dill LM (2012) The ecological importance of intact top-predator populations: a synthesis of 15 years of research in a seagrass ecosystem. Mar Freshw Res 63:1039–1050. https://doi.org/10.1071/MF12024
Hoffmann M, Hilton-Taylor C, Angulo A, Böhm M, Brooks TM, Butchart SH, Carpenter KE, Carpenter KE, Chanson J, Collen B, Cox NA, Darwall WR (2010) The impact of conservation on the status of the world’s vertebrates. Science 330:1503–1509. https://doi.org/10.1126/science1194442
Holmes BH, Steinke D, Ward RD (2009) Identification of shark and ray fins using DNA barcoding. Fish Res 95(2–3):280–288. https://doi.org/10.1016/j.fishres.2008.09.036
ICCAT (2011) Recommendation by ICCAT on the conservation of silky sharks caught in association with ICCAT fisheries, 11-08 BYC. International Commission for the Conservation of Atlantic Tunas, Spain. https://www.iccat.int/Documents/Recs/compendiopdf-e/2011-08-e.pdf. Accessed 11 Dec 2018
ICCAT (2015) Report of the 2015 ICCAT blue shark stock assessment session ICCAT Collect Vol Sci Pap 72:866–1019. https://www.iccat.int/Documents/SCRS/DetRep/BSH_ASS_ENG.pdf. Accessed 9 Dec 2018
IOTC (2017) Status summary for species of tuna and tuna-like species under the IOTC mandate, as well as other species impacted by the IOTC fisheries Stock Status Dashboard. http://www.iotc.org/science/status-summary-species-tuna-and-tuna-species-under-iotc-mandate-well-other-species-impacted-iotc. Accessed 12 Dec 2018
ISC (2017) Stock assessment and future projections of blue shark in the north pacific ocean. WCPFC-SC13-2017/SA-WP-10 WCPFC-SC, Rarotonga, Cook Islands, 9–17. https://www.wcpfc.int/node/19204. Accessed 10 Dec 2018
Jaiteh VF, Hordyk AR, Braccini M, Warren C, Loneragan NR, Thurstan R (2016) Shark finning in eastern Indonesia: assessing the sustainability of a data-poor fishery. ICES J Mar Sci 74:242–253. https://doi.org/10.1093/icesjms/fsw170
Jaiteh VF, Loneragan NR, Warren C (2017) The end of shark finning? Impacts of declining catches and fin demand on coastal community livelihoods. Mar Policy 82:224–233. https://doi.org/10.1016/jmarpol201703027
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Lessa R, Santana F, Menni R, Almeida Z (1999) Population structure and reproductive biology of the smalltail shark (Carcharhinus porosus) off Maranhão (Brazil). Mar Freshw Res 50:383–388. https://doi.org/10.1071/MF98127
Lessa R, Quijano SM, Santana FM, Monzini J (2006) Rhizoprionodon porosus. The IUCN Red List of Threatened Species 2006:eT61407A12473033 https://doi.org/10.2305/iucnuk2006rltst61407a12473033en. Accessed 14 Sept 2018
Lessa R, Batista VS, Santana FM (2016) Close to extinction? The collapse of the endemic daggernose shark (Isogomphodon oxyrhynchus) off Brazil. Glob Ecol Conserv 7:70–81. https://doi.org/10.1016/j.gecco.2016.04.003
Liu SYV, Chan CLC, Lin O, Hu CS, Chen CA (2013) DNA barcoding of shark meats identify species composition and CITES-listed species from the markets in Taiwan. PLoS ONE 8(11):e79373. https://doi.org/10.1371/journal.pone.0079373
Mauri M, Elli T, Caviglia G, Uboldi G, Azzi M (2017) RAWGraphs: a visualisation platform to create open outputs. In: Proceedings of the 12th biannual conference on Italian SIGCHI chapter (p. 28). ACM. https://doi.org/10.1145/3125571.3125585
Mendonça FF, Hashimoto DT, Porto-Foresti F, Oliveira C, Gadig OBF, Foresti F (2009) Identification of the shark species Rhizoprionodon lalandii and R. porosus (Elasmobranchii, Carcharhinidae) by multiplex PCR and PCR-RFLP techniques. Mol Ecol Resour 9:771–773. https://doi.org/10.1111/j1755-0998200902524x
Mendonça FF, Hashimoto DT, De-Franco B, Porto-Foresti F, Gadig OBF, Oliveira C, Foresti F (2010) Genetic identification of lamniform and carcharhiniform sharks using multiplex-PCR. Conserv Genet Resour 2:31–35. https://doi.org/10.1007/s12686-009-9131-7
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES science gateway for inference of large phylogenetic trees. In: Gateway computing environments workshop (GCE), pp 1–8. https://doi.org/10.1109/gce20105676129
Molina JM, Cooke SJ (2012) Trends in shark bycatch research: current status and research needs. Rev Fish Biol Fish 22:719–737. https://doi.org/10.1007/s11160-012-9269-3
Musick JA, Burgess G, Cailliet G, Camhi M, Fordham S (2000) Management of sharks and their relatives (Elasmobranchii). Fisheries 25:9–13. https://doi.org/10.1577/1548-8446(2000)025%3C0009:MOSATR%3E20CO;2
Myers RA, Worm B (2005) Extinction, survival or recovery of large predatory fishes. Philos Trans R Soc B 360(1453):13–20. https://doi.org/10.1098/rspb.2007.1295
Oliver S, Braccini M, Newman SJ, Harvey ES (2015) Global patterns in the bycatch of sharks and rays. Mar Policy 54:86–97. https://doi.org/10.1016/jmarpol201412017
Pauly D, Zeller D (2016) Catch reconstructions reveal that global marine fisheries catches are higher than reported and declining. Nat Commun 7:n10244. https://doi.org/10.1038/ncomms10244
Pauly D, Palomares ML, Froese R, Sa-a P, Vakily M, Preikshot D, Wallace S (2001) Fishing down Canadian aquatic food webs. Can J Fish Aquat Sci 58(1):51–62. https://doi.org/10.1139/f00-193
Pinheiro HT, Di Dario F, Gerhardinger LC, Melo MR, Moura RL, Reis RE, Hazin FHV, Zuanon J, Rocha LA (2015) Brazilian aquatic biodiversity in peril. Science 350(6264):1043–1044. https://doi.org/10.1126/science.350.6264.1043-a
Rambaut A (2012) FigTree v143 molecular evolution, phylogenetics and epidemiology. University of Edinburgh, Institute of Evolutionary Biology, Edinburgh. http://tree.bio.ed.ac.uk/software/figtree/. Accessed 20 Nov 2018
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarisation in Bayesian phylogenetics using Tracer 1.7. Syst Biol. https://doi.org/10.1093/sysbio/syy032
Ratnasingham S, Hebert PD (2007) BOLD: the barcode of life data system. Mol Ecol Notes 7:355–364. https://doi.org/10.1111/j1471-8286200701678x
Ratnasingham S, Hebert PD (2013) A DNA-based registry for all animal species: the Barcode Index Number (BIN) system. PLoS ONE 8:e66213. https://doi.org/10.1371/journalpone0066213
Ribeiro ADO, Caires RA, Mariguela TC, Pereira LHG, Hanner R, Oliveira C (2012) DNA barcodes identify marine fishes of São Paulo State, Brazil. Mol Ecol Resour 12(6):1012–1020. https://doi.org/10.1111/1755-0998.12007
Rigby CL, Barreto R, Carlson J, Fernando D, Fordham S, Francis MP, Jabado RW, Liu KM, Marshall A, Pacoureau N, Romanov E, Sherley RB, Winker H (2019) Isurus oxyrinchus. The IUCN Red List of Threatened Species 2019:e.T39341A2903170. https://doi.org/10.2305/IUCN.UK.2019-1.RLTS.T39341A2903170.en
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Schiller L, Alava JJ, Grove J, Reck G, Pauly D (2015) The demise of Darwin’s fishes: evidence of fishing down and illegal shark finning in the Galápagos Islands. Aquat Conserv 25:431–446. https://doi.org/10.1002/aqc2458
Sembiring A, Pertiwi NPD, Mahardini A, Wulandari R, Kurniasih EM, Kuncoro AW, Cahyani ND, Anggoro AW, Ulfa M, Madduppa H, Carpenter KE (2015) DNA barcoding reveals targeted fisheries for endangered sharks in Indonesia. Fish Res 164:130–134. https://doi.org/10.1016/j.fishres.2014.11.003
Shea KH, To AWL (2017) From boat to bowl: patterns and dynamics of shark fin trade in Hong Kong—implications for monitoring and management. Mar Policy 81:330–339
Shivji M, Clarke S, Pank M, Natanson L, Kohler N, Stanhope M (2002) Genetic identification of pelagic shark body parts for conservation and trade monitoring. Conserv Biol 16:1036–1047. https://doi.org/10.1046/j1523-1739200201188x
Sibert J, Hampton J, Kleiber P, Maunder M (2006) Biomass, size, and trophic status of top predators in the Pacific Ocean. Science 314(5806):1773–1776. https://doi.org/10.1126/science.1135347
Sims DW, Mucientes G, Queiroz N (2018) Shortfin mako sharks threatened by inaction. Science 359:1342. https://doi.org/10.1126/scienceaat0315
Steinke D, Bernard AM, Horn RL, Hilton P, Hanner R, Shivji MS (2017) DNA analysis of traded shark fins and mobulid gill plates reveals a high proportion of species of conservation concern. Sci Rep 7:9505. https://doi.org/10.1038/s41598-017-10123-5
Stevens JD, Bonfil R, Dulvy NK, Walker PA (2000) The effects of fishing on sharks, rays, and chimaeras (chondrichthyans), and the implications for marine ecosystems. ICES J Mar Sci 57:476–494. https://doi.org/10.1006/jmsc20000724
Suchard MA, Lemey P, Baele G, Ayres DL, Drummond AJ, Rambaut A (2018) Bayesian phylogenetic and phylodynamic data integration using BEAST 110. Virus Evol 4:vey016. https://doi.org/10.1093/ve/vey016
Symes D (2006) Fisheries governance: a coming of age for fisheries social science? Fish Res 81(2-3):113–117. https://doi.org/10.1023/A:1008946522213
Towns J, Cockerill T, Dahan M, Foster I, Gaither K, Grimshaw A, Hazlewood V, Hazlewood V, Lathrop S, Lifka D, Peterson GD, Roskies R (2014) XSEDE: accelerating scientific discovery. Comput Sci Eng 16:62–74. https://doi.org/10.1109/MCSE201480
van Velzen R, Weitschek E, Felici G, Bakker FT (2012) DNA barcoding of recently diverged species: relative performance of matching methods. PLoS ONE 7:e30490. https://doi.org/10.1371/journalpone0030490
Walters CJ (2007) Is adaptive management helping to solve fisheries problems. Ambio 36(4):304–308. https://doi.org/10.1579/0044-7447(2007)36%5b304:IAMHTS%5d2.0.CO;2
Ward RD, Zemlak TS, Innes BH, Last PR, Hebert PD (2005) DNA barcoding Australia’s fish species. Philos Trans R Soc Lond B 360:1847–1857. https://doi.org/10.1098/rstb20051716
Ward RD, Holmes BH, White WT, Last PR (2008) DNA barcoding Australasian chondrichthyans: results and potential uses in conservation. Mar Freshw Res 59(1):57–71. https://doi.org/10.1071/MF07148
Ward RD, Hanner R, Hebert PD (2009) The campaign to DNA barcode all fishes. J Fish Biol 74(2):329–356. https://doi.org/10.1111/j.1095-8649.2008.02080.x
Wong EHK, Shivji MS, Hanner RH (2009) Identifying sharks with DNA barcodes: assessing the utility of a nucleotide diagnostic approach. Mol Ecol Res 9:243–256
Worm B, Davis B, Kettemer L, Ward-Paige CA, Chapman D, Heithaus MR, Kessel ST, Gruber SH (2013) Global catches, exploitation rates, and rebuilding options for sharks. Mar Policy 40:194–204. https://doi.org/10.1016/jmarpol201212034
Xia X (2017) DAMBE6: new tools for microbial genomics, phylogenetics, and molecular evolution. J Hered 4:431–437. https://doi.org/10.1093/jhered/esx033
Xia X, Xie Z, Salemi M, Chen L, Wang Y (2003) An index of substitution saturation and its application. Mol Phylogenet Evol 26:1–7. https://doi.org/10.1016/S1055-7903(02)00326-3
Yule GU (1925) A mathematical theory of evolution, based on the conclusions of Dr JC Willis, FRS. Philos Trans R Soc Lond B 213:21–87. https://doi.org/10.1098/rstb19250002
Zeller D, Pauly D (2005) Good news, bad news: global fisheries discards are declining, but so are total catches. Fish Fish 6:156–159. https://doi.org/10.1111/j1467-2979200500177x
Zeller D, Harper S, Zylich K, Pauly D (2015) Synthesis of underreported small-scale fisheries catch in Pacific island waters. Coral Reefs 34:25–39. https://doi.org/10.1007/s00338-014-1219-1
Acknowledgements
Funding was provided by the São Paulo Research Foundation (FAPESP) (Grant Numbers 2009/54660-6; 2011/23787-0; 2017/02420-8).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Angus Jackson.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article belongs to the Topical Collection: Biodiversity exploitation and use.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
da Silva Ferrette, B.L., Domingues, R.R., Ussami, L.H.F. et al. DNA-based species identification of shark finning seizures in Southwest Atlantic: implications for wildlife trade surveillance and law enforcement. Biodivers Conserv 28, 4007–4025 (2019). https://doi.org/10.1007/s10531-019-01862-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10531-019-01862-0