Abstract
Castrilanthemum debeauxii is a critically endangered annual plant narrowly distributed in mountains of SE Iberia. It occurs in spiny scrublands on limestone soils dominated by Erinacea anthyllis at medium altitudes. Phylogenetic results indicate that the monotypic Castrilanthemum constitutes a paleoendemic lineage that diverged from its closest extant relatives in the Miocene. To achieve a better understanding of its recent evolution and conservation status, we performed an analysis of genetic variation and structure of four subpopulations using plastid haplotypes and AFLPs. We also investigated additional characteristics (ecological requirements, reproductive biology, pre-dispersal seed predation) that are relevant to evaluate the conservation status of the species. Castrilanthemum was found to be a facultative selfer with no major germination limitations. We detected moderate levels of genetic diversity based on AFLPs and plastid DNA sequences. The low genetic structure found for both nuclear and plastid markers, together with species distribution modeling results, pointed to high gene flow, which might counteract processes of genetic erosion caused by the low number of individuals. We found significant levels of pre-dispersal seed predation by the burrowing larvae of a tephritid fly, which may have contributed to the decline of the populations. Based on our results, we suggest directions for the management of Castrilanthemum. In particular, we recommend seeding and collection of seeds for banks, and discuss the possibilities of population reinforcement and reintroduction in localities where the species is currently extinct.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Narrow endemic species have been considered to be especially sensitive to extinction. These species combine low numbers of individuals and populations distributed in small areas (Thompson 2005; López-Pujol et al. 2013). In addition, it has historically been assumed that narrow endemics have low levels of genetic diversity. However, recent studies demonstrate that there is a wide range of possible genetic and ecological scenarios for the so-called narrow endemics: from high to extremely low genetic variation, with or without strong population structuring, with narrow ecological requirements or less ecologically restricted than expected (see Fernández-Mazuecos et al. 2014; Jiménez-Mejías et al. 2015; Fernández-Mazuecos et al. 2016). Such range of unpredictable possibilities makes the decision-making process for the conservation of each narrow endemic unique, and needed of a careful multidisciplinary evaluation (see Gentili et al. 2015).
Annual species are particularly sensitive to demographic disturbances resulting in small population sizes (e.g. water limitation, high temperatures). In addition, populations of annual species are more prone to bottlenecks, genetic drift, and inbreeding depression (Carlsen et al. 2002). The short generation time and lack of overlapping generations accelerate processes of genetic impoverishment, whereas species with long-lived individuals are more capable to buffer threats and maintain levels of genetic diversity in the face of disturbances (Austerlitz et al. 2000; De Castro et al. 2013; Jiménez-Mejías et al. 2015; Silva et al. 2015). High gene flow between populations (Ellstrand and Elam 1993) and soil seed banks (McCue and Holtsford 1998; Levine et al. 2008) are additional strategies that may buffer the negative genetic effects of inter-annual demographic oscillations.
Phylogenetically singular and paleoendemic species are particularly relevant for conservation (Vargas 2010). In recent years, there have been efforts to focus attention on these singular taxa, thus prioritizing the allocation of conservation resources to the most evolutionarily relevant among threatened species. In this respect, recent research on endemic, monotypic plant genera of the Iberian Peninsula and the Balearic Islands that are considered critically endangered, has greatly contributed to the understanding of the evolutionary and conservation value of singular species. Avellara fistulosa (Asteraceae; Fernández-Mazuecos et al. 2016), Castrilanthemum debeauxii (Asteraceae; Tomasello et al. 2015), Gyrocaryum oppositifolium (Boraginaceae; Otero et al. under review) and Naufraga balearica (Apiaceae; Fernández-Mazuecos et al. 2014) are confirmed to be the only extant members of depauperated lineages that diverged from their closest living relatives in the Miocene or earlier.
Castrilanthemum is a monotypic genus comprising C. debeauxii, an annual chamomile-like plant (Asteraceae, tribe Anthemideae). It is a well-characterized case of a paleoendemic, impoverished lineage, which is sister to the clade containing the more diverse genus Leucanthemopsis (Tomasello et al. 2015). It fits a certain definition of “living fossil”, considering its Miocene (>10 Ma) divergence from its sister clade and potential phenotypic stasis, which predate the onset of the Mediterranean climate. In this study we evaluate the genetic characteristics, environmental requirements, reproductive biology, and pre-dispersal seed predation of Castrilanthemum to obtain a detailed assessment of the underlying reasons behind the low number of populations and to lay the foundations of future conservation actions.
Materials and methods
Study species
Castrilanthemum is an annual plant endemic to the northern part of the Baetic Mountain System, located in the southeastern Iberian Peninsula (Fig. 1a, c). In all known populations, it inhabits open scrublands on rocky limestone soils at south-facing slopes, dominated by the spinose shrub Erinacea anthyllis and scattered individuals of Pinus nigra subsp. salzmannii (Blanca et al. 2004). On these slopes with skeletal soils, the E. anthyllis scrubland seems to be the climax community.
a Specimen of Castrilanthemum debeauxii in Sierra de Guillimona (P. Vargas). b Statistical parsimony network of plastid DNA haplotypes. Lines represent single nucleotide substitutions. The rectangle (haplotype A) indicates the haplotype suggested as ancestral by the TCS analysis. Oval and rectangle sizes are proportional to the number of sequences per haplotype. c Maximum entropy distribution model of Castrilanthemum based on seven climatic and one lithological variable. The ‘minimum training presence’ threshold was applied to the logistic output of Maxent in order to obtain presence (red)/absence data. Extant localities used in the model are indicated as red dots. Historical localities with dates are also shown, including the two localities of unknown precise location recorded in the early 20th century (represented as dotted lines). The location of subpopulations sampled for population genetic analyses is indicated (Ca, Cañalonguilla; As, Aserrador). The continuous black lines represent province limits. d Distribution of plastid DNA haplotypes in the four sampled subpopulations (Google Earth imagery, Landsat/Copernicus): Cañalonguilla (left), Aserrador-NW, Aserrador-C and Aserrador-SE (right). Haplotypes are represented as colours (as in b), and pie charts represent haplotype frequencies of sampled subpopulations. (Color figure online)
In 2004, the only known surviving population of Castrilanthemum was in the Sierra de Guillimona mountains, near Puebla de Don Fadrique (Blanca et al. 2004). A later survey in 2013 revealed the existence of three additional populations in neighboring areas (del Río and Gutiérrez 2013), probably forming a single metapopulation. However, previously reported populations from Sierra de Castril and Sierra de la Cabrilla (Fig. 1c) were not found despite exhaustive search by forest rangers and our own prospections, and thus we consider these populations as probably extinct. Overall, the extensive search effort performed in the last few years indicates a reduction of the former distribution area of the species.
Although there are no specific studies on the pollination biology of Castrilanthemum, it is well known that most of the species of Asteraceae display generalist pollination (Waser et al. 1996). Castrilanthemum fruits lack a pappus, and no evidence for a specialized type of diaspora dispersal has been observed. Therefore, most fruits fall in the vicinity of the mother plant (Blanca et al. 2004), although we cannot rule out a contribution of secondary dispersal by rain or wind. Some of the possible threats reported for the species are nitrification caused by livestock and direct destruction of populations by infrastructure construction (Blanca et al. 2004). Competition with Anthemis tuberculata Boiss., another Asteraceae species that is common in the area and has a similar phenotype, is a serious threat because A. tuberculata seems to be favored in habitats disturbed by the presence of goats and sheep (Blanca et al. 2004).
Population genetics
Sampling and DNA extraction
We obtained 79 samples from four subpopulations within the main population of Castrilanthemum in Sierra de Guillimona (Fig. 1c, d; Table 1). These are placed in the center of the distribution range of the species, and they were the only known subpopulations at the time this study was performed in 2012 (Blanca et al. 2004; cf. del Río and Gutiérrez 2013). Subpopulations within the main population do not form a continuous distribution. They are variably-sized patches separated by gaps of different lengths. Potential limitations to fruit dispersal may restrict gene flow between patches, further suggesting the consideration of each patch as a subpopulation. The subpopulations labeled as Aserrador constitute three well-defined patches along a mountain slope, separated by gaps of 120–240 m. The Cañalonguilla subpopulation is a single continuous patch located c. 4.5 km away from the Aserrador subpopulations.
Collected fresh leaf material was dried and stored in silica gel. Sample size from each subpopulation varied from 11 to 29 individuals depending on the observed subpopulation sizes. Individuals separated by several meters were chosen with the intent of representatively covering the genetic diversity of each subpopulation, thus minimizing the possible effect of seed dispersal limitation. No genetic data were found in the literature, which led us to search for neutral markers throughout the plastid and nuclear genomes. Total DNA was extracted from plant tissue using the DNeasy Plant Mini Kit (QIAGEN, Valencia, CA, USA).
Plastid DNA haplotypes
A pilot study using eight individuals was conducted to test sequence variation of eight plastid DNA (ptDNA) regions used in previous phylogenetic and phylogeographic studies (Shaw et al. 2005, 2007; Hollingsworth et al. 2009): rpl32-trnLUAG, trnQ-rps16, psbJ-petA, ycf6-psbM, atpI-atpH, ndhF-rpl32, psbA-trnH and psbD-trnT. The only two regions that yielded nucleotide variation in the pilot study (rpl32-trnLUAG and psbA-trnH) were sequenced for all samples. Sequences were assembled in Geneious Pro v. 5.6.6 (Kearse et al. 2012), and aligned using the MUSCLE algorithm (Edgar 2004) with default parameters. The resulting concatenated matrix was analysed using the statistical parsimony algorithm (Templeton et al. 1992) as implemented in TCS ver. 1.21 (Clement et al. 2000) to infer genealogical relationships among haplotypes. Given the low number of nucleotide substitutions found, gaps were coded as an additional character. Nucleotide and haplotype diversity were calculated according to Nei (1987) using DnaSP 5.10 (Librado & Rozas 2009). An analysis of molecular variance (AMOVA) was computed using ARLEQUIN ver. 3.5 to evaluate genetic differentiation.
AFLPs
The amplified fragment length polymorphism (AFLP) procedure followed Gaudeul et al. (2000) with minor modifications. Sixteen primer combinations were tested in a pilot study using eight individuals. Of these, four combinations (EcoRI ACT (6-FAM)—MseI CCT, EcoRI AGG (VIC)—MseI CTG, EcoRI ACT (FAM)—MseI CAC, EcoRI AGC (VIC)—MseI CAT) were selected as those with the higher numbers of variable and reproducible markers. Selective PCR products were run on a capillary sequencer (ABI Prism 3700 DNA analyzer; Applied Biosystems, Foster City, California, USA) with the internal size standard GeneScan 500 LIZ (Applied Biosystems). Six replicated samples (7.6% of the sampling) were included to estimate reproducibility and to calculate the average proportion of correctly replicated bands (Bonin et al. 2004). Fragments in the range of 100–500 bp were automatically scored using GeneMapper ver. 3.7 and manually revised. Low reproducibility markers were removed from the matrix following Bonin et al. (2004) to reach a final virtual error of 0%. Phenotype diversity was evaluated using the R-script AFLPdat ver. 2010 (Ehrich 2006, updated on 24th June 2010). The ‘frequency-down-weighted marker’ (DW) value was calculated as a rarity measure (Schönswetter and Tribsch 2005). Gene diversity according to Nei (1987) was computed using ARLEQUIN ver. 3.11 (Excoffier and Lischer 2010). A Bayesian Analysis of Population Structure (BAPS version 4.14; Corander et al. 2003) was used to estimate the population structure. We performed an admixture analysis based on a previous mixture clustering analysis. Numbers of iterations and reference individuals were set to default values. We ran 10 replicates from each of the nine simulations from K = 2 to 20. Population structure was further evaluated using a neighbour-joining (NJ) analysis using Nei–Li distances (Nei and Li 1979) as implemented in PAUP* 4.0b10 (Swofford 2002), with branch support assessed by bootstrapping (10,000 replicates). A principal coordinate analysis (PCoA) was implemented in the package GenAlEx v6.5 (Peakall and Smouse 2006). An AMOVA was computed using ARLEQUIN to evaluate genetic differentiation.
Species distribution modeling
Species distribution modeling was performed to evaluate the potential distribution of Castrilanthemum under present ecological conditions. We employed the maximum entropy algorithm, as implemented in Maxent v3.3 (Phillips et al. 2006), because of its good performance with small numbers of presence-only data (Elith et al. 2006; Pearson et al. 2007). A set of 19 bioclimatic variables, at a resolution of 30 arc-seconds, was retrieved from the WorldClim website (www.worldclim.org; Hijmans et al. 2005) for a study area including extant and historical localities (37.7–38.2ºN and 2.3–3ºW). Seven variables that are uncorrelated in the Iberian Peninsula (Fernández-Mazuecos and Vargas 2013) were selected: bio3 (isothermality), bio4 (temperature seasonality), bio5 (maximum temperature of warmest month), bio6 (minimum temperature of coldest month), bio13 (precipitation of wettest month), bio14 (precipitation of driest month) and bio15 (precipitation seasonality). Given the apparent preference of Castrilanthemum for calcareous substrates (del Río and Gutiérrez 2013), we included an additional lithological variable at a resolution of 6 arc-seconds (M. Fernández-Mazuecos and B.J. Glover, unpublished), built from the Lithostratigraphic Map of Spain (IGME 2006). This layer included five lithological categories for the study area: ultramafic, coarse-grained detritic, fine-grained detritic, calcareous and dolomitic. The eight variables were used as predictors to calibrate the Maxent model. In the occurrence dataset, we included 24 precise coordinates in three mountain ranges (Sierra Guillimona, Sierra de los Cuartos, Sierra de la Hoya del Espino), which comprehensively represent the geographic range where the plant has been observed in recent times (del Río and Gutiérrez 2013). Historical localities in two other mountain ranges (Sierra de Castril, Sierra de la Cabrilla) were not included due to the lack of precise coordinates (reviewed by Vogt and Oberprieler 1996). Occurrence data were randomly split into training data (80%), used for model building, and test data (remaining 20%), used to evaluate model accuracy. Ten subsample replicates were performed, and fitness of the resulting model was assessed with the area under the receiver operating characteristic (ROC) curve. A jackknife analysis was run to evaluate variable contributions to the models, and response curves were inferred to assess how predictions were affected by variable values. To convert continuous suitability values to discrete presence/absence (1/0) values, we chose the ‘minimum training presence’ threshold, which provides a conservative estimate of suitable areas (Pearson et al. 2007). The extent of the potential range was calculated in ArcMap 9.3 (ESRI, Redlands, CA, USA). For comparison, the extent of occurrence and area of occupancy (based on 1 × 1 km cells) of the species were calculated from extant localities using GeoCAT (Bachman et al. 2011). Given the relative uncertainty of models based on small numbers of localities, projections to past and future conditions were not attempted.
Reproductive biology
Seed germination
In May–June 2011, ripe achenes were collected in the field from decaying plants, thus ensuring that the individuals had completed their reproductive cycle. A total of 464 achenes from 30 plants were collected, including 383 achenes from the Aserrador-C subpopulation and 81 achenes from the Aserrador-SE subpopulation. Fruits from the two subpopulations were kept separate. Immediately after collection, achenes were subjected to a stratification period of two months at 5 °C. They were then placed in seedbeds with sterile substrate (70% commercial potting soil, 30% sand) in a germination chamber with 16 h of light per day and a constant temperature of 18 °C. Seedbeds were checked for fungal infestation and germination was recorded every 7 days for 8 weeks. Germination rate was expressed as a percentage of the total number of fruits. Given that no fungal infestation was observed, no fruits were removed from the seedbeds until the end of the experiment.
Mating system
Plants grown from seedlings generated in the previous experiment and maintained in a greenhouse were used to determine the mating system of the species by controlled hand pollination experiments. The experiment was performed on a total of 33 plants. The treatments included: spontaneous self-pollination (13 capitula, one per individual), manual self-pollination (10 capitula, one per individual), and manual cross-pollination (10 capitula, one per individual). For both self-pollination experiments the capitulum buds were protected with tea bags, which prevented insect pollination after anthesis. In the manual self-pollination treatment, pollen from the same individual (usually from the same capitulum, otherwise from any other capitulum available in the same plant) was added to the stigmas using a brush. In the manual cross-pollination treatment, a mixture of pollen from different donor individuals was added to the stigmas of the treated plant. As the anthesis in Castrilanthemum is protogynous and occurs in a centripetal fashion within the capitulum, the manual treatments were first applied when the outermost florets exposed their stigmas, and again when the central florets did. Ripe capitula were collected, and the numbers of successfully developed achenes counted. Differences between the three treatments were tested in a one-way ANOVA. Data were ln-transformed to meet normality. As the variances of the data were not homogeneous, comparisons between groups were assessed using the post hoc test of Games-Howell. Statistical analyses were performed with the SPSS software (IBM SPSS Statistics, version 22.0. Armonk, NY: IBM Corp.).
Fruits produced from each treatment were germinated as explained above. For the spontaneous self-pollination treatment, we tested 56 achenes, which was the total number of ripe achenes produced by all capitula under this treatment. For the other two treatments, we tested 100 achenes of each.
Inter-annual seed set variation and predispersal seed predation
To evaluate inter-annual differences in seed set, as well as seed damage caused by insects, we studied individuals from the subpopulations Aserrador-NW and Cañalonguilla. Ripe capitula from decaying plants were collected in the field in May–June 2012 and 2013, immediately before fruit dispersal. The total number of capitula studied per subpopulation was 99 and 111 for Aserrador-NW in 2012 and 2013, respectively, and 100 and 120 for Cañalonguilla in 2012 and 2013, respectively. Plants were randomly selected along a transect dividing each subpopulation. Every collected capitulum belonged to a different individual. Capitula were dissected under a binocular microscope with the assistance of tweezers. Achenes and flower remnants were classified into three categories: ripe achenes, immature achenes, and predated achenes. We tested differences in total flower production and percentages of ripe and predated achenes between subpopulations and years. Each dataset obtained each year in each subpopulation was considered as an independent sample. Data met normality in most cases, whereas in others the observed values did not strongly deviate from the expected ones (i.e., P = 0.053, after the Shapiro–Wilk test). According to this and the large sample size, we performed one-way ANOVA to test differences between groups. As the variances of the data were not homogeneous, the comparisons between groups were conducted using the post hoc test of Games-Howell as implemented in SPSS.
Results
Population genetics
Plastid DNA haplotypes
The complete aligned length of the concatenated psbA-trnH and rpl32-trnLUAG regions was 1148 bp. Two variable sites plus one gap were found that resulted in four haplotypes (A–D). Haplotype B was the most widespread (three subpopulations), followed by haplotype A (two subpopulations). Haplotypes C and D were detected in a single subpopulation each (Fig. 1d; Table 1). Haplotype diversity across all subpopulations averaged 0.629, and nucleotide diversity 0.00066. The most diverse subpopulation was Aserrador-NW (0.517, 0.00046) and the least diverse Aserrador-C (null values for both statistics) (Table 1). A simple network was produced by the statistical parsimony analysis (Fig. 1b). No loops or missing haplotypes were inferred. A central position in the network was inferred for haplotype A. The other three haplotypes differed from it by a single mutational step. AMOVA analysis (Table 2) showed that most observed variation was explained by differences among subpopulations (60.25%) but a considerable amount of variation was also found within subpopulations (39.73%).
AFLPs
AFLP data resulted in a matrix of 87 fragments with 100% reproducibility according to the replicated individuals. Gene diversity across all subpopulations averaged 0.279 ± 0.143. The most diverse subpopulation was Aserrador-SE (0.343 ± 0.141) and the least diverse Cañalonguilla (0.221 ± 0.136) (Table 1). The two subpopulations (Cañalonguilla and Aserrador-NW) with lower values of genetic diversity displayed DW values three times higher than the other two subpopulations. AMOVA analysis (Table 2) showed that most observed variation was explained by differences among individuals within subpopulations (91.54%) and only 8.46% was due to differences among subpopulations. Neither the NJ tree nor the PCoA (Fig. 2a, b) revealed a strong population structure, although the subpopulation Aserrador-SE was somewhat differentiated from the others according to the plot of axes 1 and 3 of the PCoA (Fig. 2a). BAPS found K = 2 as the optimal number of groups (I and II; Fig. 2c). Group I was widespread, whereas group II was confined to subpopulation Aserrador-SE (Table 1). All the individuals assigned to group II displayed plastid haplotype C, although haplotype C was also found in a few individuals assigned to group I. Both groups presented certain degree of admixture (Fig. 2c).
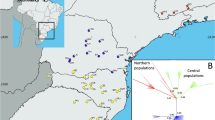
Genetic structure of four subpopulations of Castrilanthemum based on AFLP data. a Principal coordinate analysis (PCoA); values for the first two axes (above) and the first and third axes (below) are plotted; percentage of variation explained by each axis is shown in brackets. b Neighbour-joining tree. c Genetic clustering of individuals based on a Bayesian admixture analysis in BAPS; each vertical bar represents a single individual, with colours representing the genetic contributions of the two genetic clusters (roman numerals) detected in the mixture analysis. Subpopulations are denoted by the same symbols across panels (see legend in a). (Color figure online)
Species distribution modeling
The Maxent model (with mean area under the receiver-operating characteristic curve for testing data 0.982) inferred a potential distribution of Castrilanthemum spanning the three mountain ranges where the species currently occurs (Sierra de Guillimona, Sierra de los Cuartos, Sierra de la Hoya del Espino) together with relatively smaller areas in adjacent mountains (Fig. 1c). These included small and fragmented suitable areas in two mountain ranges (Sierra de Castril, Sierra de la Cabrilla) where the species was observed in the early 20th century (Vogt and Oberprieler 1996). We estimated a total potential range of 361 km2, a relatively large area that contrasts with an extent of occurrence of 26 km2 and an area of occupancy of 14 km2 estimated from extant populations. According to the jackknife analysis, the three most important variables for the model were the maximum temperature of warmest month, precipitation seasonality and precipitation of driest month, and the response analysis confirmed a clear preference of the species for calcareous substrates (Supporting Information S1).
Reproductive biology
Seed germination
The observed germination success rate was moderate to high. Germination percentages were 70.5% for the subpopulation Aserrador-C, and 53.1% for the subpopulation Aserrador-SE, resulting in an overall success rate of 67.5%. It took up to five weeks for all the viable seeds to germinate (Fig. 3a). A germination peak was observed in the first week (50.2%) followed by a decrease in the number of germinated seeds in the following three weeks (10.8, 4.1, and 0.2%, respectively), and a final weak second peak in the fifth week (2.2%).
Mating system
The pollination treatments significantly affected the seed set (Table 3; Fig. 3b). Whereas spontaneous and manual self-pollination treatments did not show significant differences (average seed set per capitulum of 2.1 and 12.3% respectively), the manual cross-pollination treatment differed significantly from the other two, with a much higher seed set per capitulum (47.3%). Germination rate of the fruits produced in all treatments was high (94.6% for spontaneous self-pollination, 97% for manual self-pollination, and 74% for manual cross-pollination).
Inter-annual seed set variation and pre-dispersal seed predation
Three species of insects were recovered from the analyzed capitula (Fig. 4): (1) Burrowing larvae of a tephritid fly (Diptera) predated on 59.5–81.8% of all the analyzed capitula; we only found one larva per capitulum. (2) A cecidomyid fly (Diptera), likely assignable to the genus Ozirhincus (Dorchin et al. 2015), induced galls in the achenes; it was observed only in four capitula, and several pupae were detected per capitulum. (3) Braconid wasps (Hymenoptera), presumably more than one species and apparently parasitizing the tephritid larvae, were identified three times. It was unclear whether the mortality of the tephritid larvae was partly due to the parasitoid, or simply the result of the plant wilting.
Insects recovered from Castrilanthemum capitula. a–c Tephritid fly: adult (a), burrowing larvae (b, the one on the left is shown in a burrow between Castrilanthemum achenes) and empty puparium (c). d, e Cecidomyid fly: adult (d) and pupa (e). f Braconid wasp. In all images, the black bar represents 1 mm
Differences in seed set were significant between years but not between subpopulations within the same year (Tables 4, 5; Fig. 3c–e). Production of fruits per capitulum was lower in 2012 than in 2013, and the same pattern was found for the percentage of ripe achenes detected. Conversely, predation percentages were higher in 2012 than in 2013.
Discussion
High genetic diversity and gene flow
Relatively high levels of genetic diversity were found for Castrilanthemum debeauxii despite its critically endangered status. This is a new case where narrow endemicity and endangered status of a species are not related to low levels of genetic diversity (see Fernández-Mazuecos et al. 2014; Jiménez-Mejías et al. 2015; Forrest et al. 2017).
As an annual plant in the Mediterranean floristic region, Castrilanthemum is subject to demographic fluctuations due to inter-annual climatic variation (del Río and Gutiérrez 2013). Indeed, our study has detected significant differences in seed set between two years (Tables 4, 5). The underlying reason might be related to differences in precipitation between 2012 and 2013 (average precipitation 183.2 and 468.1 mm respectively; data from Puebla de Don Fadrique meteorological station, Consejería de Agricultura, Pesca y Desarrollo Rural 2016), and thus differences in levels of predation. Nevertheless, our data do not demonstrate such causality. In annual plants with small population sizes, dry years have been shown to result in fewer plants and decreased flowering synchrony, with a high potential for bottlenecks, rapid diversity loss, spatial differentiation and inbreeding depression (Austerlistz et al. 2000; Carlsen et al. 2002). Remarkably, none of these is clearly shown by our results. Despite its reduced area of occurrence (observed and potential; Fig. 1c), Castrilanthemum does not display signs of genetic erosion. The haplotype diversity and number of plastid haplotypes (0.629, four haplotypes; Table 1; Fig. 1b, d) reached high levels considering such a small occupation area. Likewise, the level of genetic diversity, as estimated by AFLPs, was indeed high (0.279) when compared with other rare species of the family Asteraceae: Avellara fistulosa (0.18; Fernández-Mazuecos et al. 2016); Lasthenia conjugens (0.37; Ramp Neale et al. 2008); Olearia gardneri (0.17; Barnaud and Houliston 2010); Sonchus sect. Pustulati spp. (0.17–0.09; Silva et al. 2015). It is noteworthy that among these species only L. conjugens and Castrilanthemum (the two with the highest genetic diversity) are pappusless species, while in the others the fruits bear pappus and are primarily dispersed by wind. Also, Castrilanthemum displays low differentiation among subpopulations and overall weak geographic structure (Figs. 1d, 2; Tables 1, 2) with a characteristic pattern of more structure for maternally inherited ptDNA haplotypes than for predominatly nuclear AFLP markers (see Trapnell and Hamrick 2004). Nevertheless, incipient differentiation was detected for one of the subpopulations (Aserrador-SE) by both AFLPs (Fig. 2a, c) and plastid haplotypes (Fig. 1d).
Lower investment in pollen transfer and seed production has been proposed as a key feature of narrow endemic species (Lavergne et al. 2004). Castrilanthemum does not display strong signs of pollen limitation. Differences of the order of <50% in the production of seeds are observed between the pollen-saturated specimens of our outcrossing experiment and specimens in the wild, as well as between these and the specimens of our self-pollination treatments. In the absence of vegetative persistence as a mechanism to buffer diversity loss (Austerlitz et al. 2000; De Castro et al. 2013; Jiménez-Mejías et al. 2015; Silva et al. 2015; Amat et al. 2013), our results point to gene flow between subpopulations via both pollen and seeds as the mechanism counteracting the deleterious effects of inter-annual oscillations on the genetic diversity of Castrilanthemum. This gene flow may be facilitated by the generalist pollination and by habitat suitability in the mountain ranges where the species occurs (Fig. 1c). In another paleoendemism with a similarly small distribution range (Naufraga balearica), a strong spatial structure and lack of genetic admixture between isolated populations have been associated with limited pollen dispersal by ants and lack of seed dispersal mechanisms, together with strict microecological requirements (Cursach and Rita 2012; Fernández-Mazuecos et al. 2014). In contrast, the fragmented distribution of Castrilanthemum, with the four sampled subpopulations in close proximity to each other, appears to have enabled relatively pervasive movement of alleles and haplotypes between subpopulations (Frey et al. 2012). While the species is relatively abundant within the center of its potential distribution area, where common rocky soils are found, peripheral isolated populations of the species seem to have disappeared (e.g. Sierra de Castril; Vogt and Oberprieler 1996; Fig. 1c). In these extinct populations, the effects of bottlenecks and genetic drift, coupled with the additional pressure of herbivory, may have been more severe than in the core area.
The breeding system of Castrilanthemum may also have an effect on its population genetics. Although the production of seeds by selfing cannot be underestimated, the values of seed set observed in the field were more in agreement with those obtained by the outcrossing rather than selfing experiments (Tables 3, 4). Differences in spatial structure and genetic diversity observed in other rare plants have been attributed to differences in breeding system, with more structuring and less diversity in selfers in comparison to outcrossers (Bacchetta et al. 2011; Coppi et al. 2008; Ramp Neale et al. 2008; Crema et al. 2009; Gómez-Gómez et al. 2012; but see Wolfe et al. 2014).
The role of a potential soil seed bank remains to be evaluated (Levine et al. 2008). It could play a part in buffering the effect of population fluctuations, including potential bottlenecks generated by seed predators (Fig. 3e). However, a permanent seed bank would not explain the genetic admixture or the low spatial structuring. On the contrary, an important contribution of the seed bank has been proposed for annual taxa with stronger geographical structure (e.g. Chloropyron palmatum, Ayres et al. 2015).
Damaging effect of predating insects
Predispersal consumption of flowers and seeds may have an important negative effect on population dynamics (Bertness et al. 1987). Two insects were found to feed on Castrilanthemum fruits: a tephritid fly and a cecidomyid fly. The role of the cecidomyid fly in seed number reduction appears to be rather minor because the life cycle of each larva is confined to a gall within a single ovary. However, the tephritid fly may significantly affect the population dynamics of Castrilanthemum. Fruit damage by larvae of this species was commonly observed, and sometimes involved the destruction of entire capitula.
Changes in phenological patterns and seed production limitation are strategies against seed consumers known from perennial plants (Bertness et al. 1987). However, such strategies cannot be afforded by annual plants, whose population survival highly depends on the synchrony of individuals and final production of seeds. An effect of resource availability on predation is revealed by our data. The higher production of fruits in 2013 compared with 2012 (Fig. 3c–e) coincided with less pre-dispersal predation (Fig. 3e). On the other hand, no significant differences between subpopulations within the same year were found (Tables 4, 5), which suggests that pre-dispersal predation operates at the whole population level.
Mutualistic relationships between parasitic flies and plants have been previously reported, with flies facilitating pollination through their visit to flowers for oviposition (see Jaeger el al. 2000). However, this does not seem the case for Castrilanthemum, given the strong deleterious effect of the fly (about 50% of achenes predated some years; Table 4), as well as the generalist pollination in Asteraceae (Waser et al. 1996). However, field observations of the pollinators of Castrilanthemum are needed to confirm this hypothesis.
Conservation measures
Given the well-distributed diversity and lack of geographic structure, the management of this species seems straightforward from a genetic-based point of view. Collection of seeds for seed banks, evaluation of their viability and other measures of ex situ conservation should representatively cover the entire area of the species, especially focusing on the most genetically diverse ones (Aserrador-SE and Aserrador-C according to AFLPs, Aserrador-NW according to haplotype diversity). The certain degree of genetic distinctness of the Aserrador-SE and Aserrador-NW subpopulations (according to BAPS grouping and DW values respectively; Table 1) should also be taken into account. A more extensive study would be required to assess the genetic features of populations not included in our population genetic analysis.
Reintroduction in historical localities is advisable when a limited number of surviving populations, inherently vulnerable to extinction, remain (Maunder 1992). The distribution modeling analysis revealed small and fragmented areas as suitable for Castrilanthemum in historical localities where it seems to be currently extinct (Sierra de Castril, Sierra de la Cabrilla; Fig. 1c). Reintroduction in these localities may be problematic, given the apparent small size of the suitable spots and their isolation from the main distribution area, which might prevent gene flow and condemn reintroduced populations to genetic impoverishment and eventually extinction. However, the extent of potential distribution in these areas may have been underestimated, given that historical localities were not included in our modeling analysis. Reintroduction should not be ruled out, but the causes behind the extinction of historical populations and the suitability of target habitats should be assessed beforehand.
In areas of the extant distribution with low population densities, controlled seeding might have a positive effect (Ramp et al. 2006), both in terms of increasing genetic variability and decreasing pre-dispersal seed predation. In addition, to buffer the effects of global warming, assisted upslope migration could be performed to help the species access suitable areas in the immediate vicinity of the main distribution (Ferrarini et al. 2016).
References
Amat ME, Silvertown J, Vargas P (2013) Strong spatial genetic structure reduces reproductive success in the critically endangered plant genus Pseudomisopates. J Heredity. doi:10.1093/jhered/est042
Austerlitz F, Mariette S, Machon N, Gouyon P-H, Godelle B (2000) Effects of colonization processes on genetic diversity: differences between annual plants and tree species. Genetics 154:1309–1321
Ayres DR, Fleishman E, Launer A, Lee AK, Zippin D (2015) Genetic structure and demography of Chloropyron palmatum, an endangered annual plant. Madroño 62:139–149. doi:10.3120/madr-62-03-139-149.1
Bacchetta G, Fenu G, Mattana E, Zecca G, Grassi F, Casazza G, Minuto L (2011) Genetic variability of the narrow endemic Rhamnus persicifolia Moris (Rhamnaceae) and its implications for conservation. Biochem Syst Ecol 39:477–484. doi:10.1016/j.bse.2011.06.018
Bachman S, Moat J, Hill AW, de la Torre J, Scott B (2011) Supporting Red List threat assessments with GeoCAT: geospatial conservation assessment tool. Zookeys 150:117–126. doi:10.3897/zookeys.150.2109
Barnaud A, Houliston GJ (2010) Population genetics of the threatened tree daisy Olearia gardneri (Asteraceae), conservation of a critically endangered species. Conserv. Genet 11:1515–1522. doi:10.1007/s10592-009-9981-2
Bertness MD, Wise C, Ellison AM (1987) Consumer pressure and seed set in a salt marsh perennial plant community. Oecologia 71:190–200
Blanca G, Gutiérrez L, Luque P, Benavente A, Morales C, Garrido A (2004) Castrilanthemum debeauxii (Degen, Hervier & E.Rev.) Vogt & oberprieler. In: Bañares A, Blanca G, Güemes J, Moreno JC, Ortiz S (eds) Atlas y Libro Rojo de la flora vascular amenazada de España. Taxones prioritarios. Dirección General de Conservación de la Naturaleza, Madrid, pp 164–165
Bonin A, Bellemain E, Bronken Eidesen P, Pompanon F, Brochmann C, Taberlet P (2004) How to track and assess genotyping errors in population genetics studies. Mol Ecol 13:3261–3273. doi:10.1111/j.1365-294X.2004.02346.x
Carlsen TM, Espeland EK, Pavlik BM (2002) Reproductive ecology and the persistence of an endangered plant. Biodivers & Conservation 11:1247–1268. doi:10.1023/A:1016066618824
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659. doi:10.1046/j.1365-294x.2000.01020.x
Consejería de Agricultura, Pesca y Desarrollo Rural (2016, continuously updated) Estaciones agroclimáticas. https://www.juntadeandalucia.es/agriculturaypesca/ifapa/ria/servlet/FrontController?action=Init. Accessed 11 December 2016
Coppi A, Mengoni A, Selvi F (2008) AFLP fingerprinting of Anchusa (Boraginaceae) in the Corso-Sardinian system: genetic diversity, population differentiation and conservation priorities in an insular endemic group threatened with extinction. Biol Conserv 141:2000–2011. doi:10.1016/j.biocon.2008.05.015
Corander J, Waldmann P, Sillanpää MJ (2003) Bayesian analysis of genetic differentiation between populations. Genetics 163:367–374
Crema S, Cristofolini G, Rossi M, Conte L (2009) High genetic diversity detected in the endemic Primula apennina Widmer (Primulaceae) using ISSR fingerprinting. Pl Syst Evol 280:29–36. doi:10.1007/s00606-009-0167-7
Cursach J, Rita J (2012) Implications of the reproductive biology of the narrow endemic Naufraga balearica (Apiaceae) for its conservation status. Pl Syst Evol 298:581–596. doi:10.1007/s00606-011-0568-2
De Castro O, Sepe F, Di Maio A, Cennamo P, De Luca P, Gianguzzi L, Menale B (2013) Genetic structure in the paleoendemic and endangered Petagnaea gussonei (Spreng.) Rauschert (Saniculoideae, Apiaceae) and implications for its conservation. Pl Syst Evol 299:209–223. doi:10.1007/s00606-012-0716-3
del Río J, Gutiérrez L (2013) Novedades sobre la distribución del macroendemismo bético Castrilanthemum debeauxii (Degen, Hervier & É. Rev.) Vogt & Oberprieler. Anales Biol 35:101–104. doi:10.6018/analesbio.0.35.15
Dorchin N, Astrin JJ, Bodner L, Harris KM (2015) Morphological and molecular revision of the genus Ozirhincus (Diptera: cecidomyiidae)—long-snouted seed-feeding gall midges on Asteraceae. PLoS ONE 10(7):e0130981. doi:10.1371/journal.pone.0130981
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucl Acids Res 32:1792–1797. doi:10.1093/nar/gkh340
Ehrich D (2006) AFLPdat: a collection of R functions for convenient handling of AFLP data. Mol Ecol Notes 6:603–604. doi:10.1111/j.1471-8286.2006.01380.x
Elith J, Graham CH, Anderson RP, Dudík M, Ferrier S, Guisan A, Hijmans RJ, Huettmann F, Leathwick JR, Lehmann A (2006) Novel methods improvepredictionof species’distributions fromoccurrencedata. Ecography 29:129–151. doi:10.1111/j.2006.0906-7590.04596.x
Ellstrand NC, Elam DR (1993) Population genetic consequences of small population size: implications for plant conservation. Ann Rev Ecol Syst 24:217–242
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567. doi:10.1111/j.1755-0998.2010.02847.x
Fernández-Mazuecos M, Vargas P (2013) Congruence between distribution modelling and phylogeographical analyses reveals Quaternary survival of a toadflax species (Linaria elegans) in oceanic climate areas of a mountain ring range. New Phytol 198:1274–1289. doi:10.1111/nph.12220
Fernández-Mazuecos M, Jiménez-Mejías P, Rotllan-Puig X, Vargas P (2014) Narrow endemics to Mediterranean islands: moderate genetic diversity but narrow climatic niche of the ancient, critically endangered Naufraga (Apiaceae). Perspect Pl Ecol Evol Syst 16:190–202. doi:10.1016/j.ppees.2014.05.003
Fernández-Mazuecos M, Jiménez-Mejías P, Martín-Bravo S, Buide ML, Álvarez I, Vargas P (2016) Narrow endemics on coastal plains: miocene divergence of the critically endangered genus Avellara (Compositae). Pl Biol 18:729–738. doi:10.1111/plb.12458
Ferrarini A, Selvaggi A, Abeli T, Alatalo JM, Orsenigo S, Gentili R, Rossi G (2016) Planning for assisted colonization of plants in a warming world. Sci Rep 6:28542. doi:10.1038/srep28542
Forrest A, Escudero M, Heuertz M, Wilson Y, Cano E, Vargas P (2017) Testing the hypothesis of low genetic diversity and population structure in narrow endemic species: the endangered Antirrhinum charidemi (Plantaginaceae). Bot J Linn Soc 183:260–270. doi:10.1093/botlinnean/bow002
Frey DJ, Haag CR, Kozlowski G, Tison J-M, Mráz P (2012) High genetic and morphological diversity despite range contraction in the diploid Hieracium eriophorum (Asteraceae) endemic to the coastal sand dunes of south-west France. Bot J Linn Soc 169:365–377. doi:10.1111/j.1095-8339.2012.01215.x
Gaudeul M, Taberlet P, Till-Bottraud I (2000) Genetic diversity in an endangered alpine plant, Eryngium alpinum L. (Apiaceae), inferred from amplified fragment length polymorphism markers. Mol Ecol 9:1625–1637. doi:10.1046/j.1365-294x.2000.01063.x
Gentili R, Bacchetta G, Fenu G, Cogoni D, Abeli T, Rossi G, Salvatore MC, Baroni C, Citterio S (2015) From cold to warm-stage refugia for boreo-alpine plants in southern European and Mediterranean mountains: the last chance to survive or an opportunity for speciation? Biodiversity 16:1–15. doi:10.1080/14888386.2015.1116407
Gómez-Gómez L, Ahrazem O, Herranaz JM, Ferrandis P (2012) Genetic characterization and variation within and among populations of Anthyllis rupestris Coss., and endangered endemism of southern Spain. Biochem Syst Ecol 45:138–147. doi:10.1016/j.bse.2012.07.015
Hijmans RJ, Cameron SE, Parra JL, Jones PG, Jarvis A (2005) Very high resolution interpolated climate surfaces for global land areas. Int J Climatol 25:1965–1978. doi:10.1002/joc.1276
Hollingsworth PM, Forrest LL, Spouge JL, Hajibabaei M, Ratnasingham S, van der Bank M, Chase MW, Cowan RS, Erickson DL, Fazekas AJ, Graham SW, James KE, Kim K-J, Kress WJ, Schneider H, van AlphenStahl J, Barrett SCH, van den Berg C, Bogarin D, Burgess KS, Cameron KM, Carine M, Chacón J, Clark A, Clarkson JJ, Conrad F, Devey DS, Ford CS, Hedderson TAJ, Hollingsworth ML, Husband BC, Kelly LJ, Kesanakurti PR, Kim JS, Kim Y-D, Lahaye R, Lee H-L, Long DG, Madriñán S, Maurin O, Meusnier I, Newmaster SG, Park C-W, Percy DM, Petersen G, Richardson JE, Salazar GA, Savolainen V, Seberg O, Wilkinson MJ, Yi D-K, Little DP (2009) A DNA barcode for land plants. Proc Natl Acad Sci USA 106:12794–12797. doi:10.1073/pnas.0905845106
IGME (2006) Mapa litoestratigráfico y de permeabilidades de España 1/200.000. Instituto Geológico y Minero de España, Madrid
Jaeger N, Till-Bottraud I, Després L (2000) Evolutionary conflict between Trollius europaeus and its seed-parasite pollinators Chiastocheta flies. Evol Ecol Res 2:885–896
Jiménez-Mejías P, Fernández-Mazuecos M, Amat ME, Vargas P (2015) Narrow endemics in European mountains: high genetic diversity within the monospecific genus Pseudomisopates (Plantaginaceae) despite isolation since the late Pleistocene. J Biogeogr 42:1455–1468. doi:10.1111/jbi.12507
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Mentjies P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. doi:10.1093/bioinformatics/bts199
Lavergne S, Thompson JD, Garnier E, Debussche M (2004) The biology and ecology of narrow endemic and widespread plants: a comparative study of trait variation in 20 congeneric pairs. Oikos 107:505–518. doi:10.1111/j.0030-1299.2004.13423.x
Levine JM, McEachern AK, Cowan C (2008) Rainfall effects on rare anual plants. J Ecol 96:795–806. doi:10.1111/j.1365-2745.2008.01375.x
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452. doi:10.1093/bioinformatics/btp187
López-Pujol J, Martinell MC, Massó S, Blanché C, Sáez L (2013) The ‘paradigm of extremes’: extremely low genetic diversity in an extremely narrow endemic species, Coristospermum huteri (Umbelliferae). Plant Syst Evol 299:439–446. doi:10.1007/s00606-012-0732-3
Maunder M (1992) Plant reintroduction: an overview. Biodivers & Conserv 1:51–61. doi:10.1007/BF00700250
McCue KA, Holtsford TP (1998) Seed bank influence on genetic diversity in the rare anual Clarkis springvillensis. Am J Bot 85:30–36
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. doi:10.1111/j.1471-8286.2005.01155.x
Pearson RG, Raxworthy CJ, Nakamura M, Townsend Peterson A (2007) Predicting species distributions from small numbers of occurrence records: a test case using cryptic geckos in Madagascar. J Biogeogr 34:102–117. doi:10.1111/j.1365-2699.2006.01594.x
Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecol Model 190:231–259. doi:10.1016/j.ecolmodel.2005.03.026
Ramp Neale JM, Ranker TA, Collinge SK (2008) Conservation of rare species with island-like distributions: a case study of Lasthenia conjugens (Asteraceae) using population genetic structure and the distribution of rare markers. Pl Spec Biol 23:97–110. doi:10.1111/j.1442-1984.2008.00211.x
Ramp JM, Collinge SK, Ramker TA (2006) Restoration genetics of the vernal pool endemic Lasthenia conjugens (Asteraceae). Conserv Genet 7:631–649. doi:10.1007/s10592-005-9052-2
Schönswetter P, Tribsch A (2005) Vicariance and dispersal in the alpine perennial Bupleurum stellatum L. (Apiaceae). Taxon 54:725–732. doi:10.2307/25065429
Shaw J, Lickey EB, Beck JT, Farmer SB, Liu W, Miller J, Siripun KC, Winder CT, Schilling EE, Small RL (2005) The tortoise and the hare II: relative utility of 21 noncoding chloroplast DNA sequences for phylogenetic analysis. Am J Bot 92:142–166. doi:10.3732/ajb.92.1.142
Shaw J, Lickey EB, Schilling EE, Small RL (2007) Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. Am J Bot 94:275–288. doi:10.3732/ajb.94.3.275
Silva JL, Lim S-Y, Kim S-C, Mejías JA (2015) Phylogeography of cliff-dwelling relicts with a highly narrow and disjunct distribution in the western Mediterranean. Am J Bot 102:1538–1551. doi:10.3732/ajb.1500152
Swofford DL (2002) PAUP*: phylogenetic analysis using parsimony (*and other methods), version 4.0b10. Sinauer, Sunderland, MA
Templeton AR, Crandall KA, Sing CF (1992) A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III Cladogram estimation. Genetics 132:619–633
Thompson JD (2005) Plant evolution in the mediterranean. Oxford University Press, New York
Tomasello S, Álvarez I, Vargas P, Oberprieler C (2015) Is the extremely rare Iberian endemic plant species Castrilanthemum debeauxii (Compositae, Anthemideae) a ‘living fossil’? Evidence from a multi-locus species tree reconstruction. Mol Phylogen Evol 82:118–130. doi:10.1016/j.ympev.2014.09.007
Trapnell DW, Hamrick JL (2004) Partitioning nuclear and chloroplast variation at multiple spatial scales in the neotropical epiphytic orchid, Laelia rubescens. Mol Ecol 13:2655–2666. doi:10.1111/j.1365-294X.2004.02281.x
Vargas P (2010) Estudio de las plantas amenazadas en España. ¿Hay fósiles vivientes aún desconocidos? Lychnos 3:19–23
Vogt R, Oberprieler C (1996) Castrilanthemum Vogt & Oberprieler, a new genus of the Compositeae-Anthemideae. Anales Jard Bot Madrid 54:336–346
Waser NM, Chittka L, Price MV, Williams NM, Ollerton J (1996) Generalization in pollination systems, and why it matters. Ecology 77:1043–1060. doi:10.2307/2265575
Wolfe AD, McMullen-Sibul A, Tepedino VJ, Kubatko L, Necamp T, Fassnacht S (2014) Conservation genetics and breeding system of Penstemon debilis (Plantaginaceae), a rare beardtongue endemic to oil shale talus in western Colorado, USA. J Syst Evol 52:598–611. doi:10.1111/jse.12100
Acknowledgements
The authors wish to thank E. Cano and A. Herrero for technical support; M. Carles-Tolrá for help identifying the insects found in Castrilanthemum capitula; and the local government of Andalusia for collecting permits. This research was funded by Banco Santander, the Spanish National Research Council (CSIC) and Fundación General CSIC (FGCSIC) through a Proyecto Cero on endangered species 2010 (“Do all endangered species hold the same value?: origin and conservation of living fossils of flowering plants endemic to Spain”).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Daniel Sanchez Mata.
P. Jiménez-Mejías and M. Fernández-Mazuecos contributed equally to the preparation of this paper.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiménez-Mejías, P., Fernández-Mazuecos, M., Gutiérrez, L. et al. Narrow endemics in Mediterranean scrublands: high gene flow buffers genetic impoverishment in the annual monospecific Castrilanthemum (Asteraceae). Biodivers Conserv 26, 2607–2626 (2017). https://doi.org/10.1007/s10531-017-1374-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10531-017-1374-0