Abstract
One of the threats that the critically endangered European mink (Mustela lutreola) faces throughout its relict range, including the occidental population, is the impact of the American mink (Mustela vison) invasion in its natural habitat. We aimed to explore the differences in microbiota and genetic diversity between European and American mink to test phylosymbiosis theory. We investigated the gut microbiota composition of European and American mink in controlled environments (captive breeding compounds and fur farms respectively) to account for the impact of the environment on gut bacterial composition. We compared them to the gut microbiota of both mink species in the natural environment across habitats. Our exploratory results showed differences between free-ranging and captive individuals, with more extreme changes in American mink compared to European mink. However, feral American mink from a long-established population exhibited gut bacterial composition closer to the free-ranging native species compared to more recently established feral populations. This result could be explained by dietary shifts in the area sampled based on prey availability through different landscape, but also to a lesser extent due to greater genetic differentiation. This exploratory work contributes to the scarce literature currently available on the dynamics between gut microbiota and mammal invasion.
Graphic Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Invasive alien species have been widely recognized as one of the major threats to biodiversity due to anthropogenic changes at both global and local scales (Lockwood et al. 2007). Invasive species can directly impact the habitat and ecology of native species they interact with as they affect native species’ population sizes and habitat ranges (Genovesi et al. 2012; Zalewski et al. 2010). Before a species establishes and expands to become a successful invader, the colonization of new habitats represents a challenge through a variety of new selective pressures encountered that can be highly costly from an adaptative lens (Amsellem et al. 2017). Therefore, host-associated microbes can play a critical role in the invasive success of an exotic species in a new habitat. These microorganisms (bacteria, archea, virus, fungi and protozoa) range from parasites to obligate mutualists (West et al. 2019). This large range of interactions, often coupled with complex historical and introduction events, can result in a wide variety of ecological dynamics such as colonization resistance and the variation in bacterial niches that promote nutrient absorption by the host (McKenney et al. 2018). Such components of the gut microbiota are thus relevant to host survival and therefore invasion success.
Within the last decade, we have begun to understand the underlying processes driving host-associated microbial community dynamics. The external environment of the host has been reported to be one of the main drivers of variation (Koskella et al. 2017; Spor et al. 2011). Housing facilities such as fur farms and captive breeding facilities in zoos provide intense veterinary care, sanitized enclosures, a standardized diet, and reduced social interactions. Hence, captivity has been shown to alter the microbiota of animals compared to their free-ranging counterparts (Clayton et al. 2016; Wasimuddin et al. 2017; van Leeuwen et al. 2020). Many of these studies show similar trends: a decrease in bacterial phylotype richness (or α-diversity) among captive individuals compared to their free-ranging conspecifics, as well as differences in community composition (or β-diversity) between the groups. However, some host species show the opposite pattern (McKenzie et al. 2017; Greene et al. 2019; Frankel et al. 2019), postulating that the gut microbiota respond differently to captivity according to host taxa, mainly through their feeding strategy and gut physiology. Differences observed in gut microbial communities have largely been attributed to altered diets in captivity that can also lead to the extinction of microbial niches and functions in the host’s gut over multiple generations in captivity (Sonnenburg et al. 2016; van Leeuwen et al. 2020). Diet has therefore been reported to have an important influence on the mammalian gut microbiota (Reese and Dunn 2018; Martinez-Mota et al. 2019).
Despite the strong impact of the host environment on its gut microbial community, the genetics and biology of the host should also be considered to fully understand the complex dynamics that occur in these systems (Koskella et al. 2017; Spor et al. 2011). Phylosymbiosis is described as an increase in compositional similarity between bacterial communities colonizing closely related hosts compared with distantly related hosts (Groussin et al. 2017; Lim and Bordenstein 2020). Many investigated mammals have supported this pattern, such as bats, apes and rodents (Brooks et al. 2016; Kohl et al. 2018; Knowles et al. 2019; Ochman et al. 2010; Phillips et al. 2012), as well as other animal taxa (Pollock et al. 2018; Sevellec et al. 2019; van Opstal and Bordenstein 2019); however, other studies have not detected signals of phylosymbiosis in some mammals (Baxter et al. 2015; Greene et al. 2019; Grond et al. 2020). Groussin et al. (2017) also suggested that the tight associations between some host taxa and some of their associated gut microbes might not generalize to the entire gut microbial community, hence the strong environmental effects on gut microbial composition. No study to date has examined phylosymbiosis in the context of invasion ecology in mammals. Within mammals, carnivores have short transit time and digestive tracts, so the gut microbiota are potentially less impacted by diet (Reese and Dunn 2018; Ley et al. 2008). From the current literature, mustelids are known to harbor relatively low diversity and abundance of gut microbes (Compo et al. 2018; Bahl et al. 2017), potentially making them interesting study species to further understand the link between host environment and associated microbes.
An example of a successful invader is the American mink (Mustela vison) in Europe, which was introduced from North America for fur farming in the early twentieth century. Following accidental escapes, as well as intentional releases, this species became established in 28 European countries (Bonesi and Palazon 2007; Reid et al. 2016). The American mink is also present as an invasive species in parts of South America and Asia (Mora et al. 2018; Shimatani et al. 2010).
The generalist and opportunistic aspects of American mink’s diet strongly impacted populations of 47 reported native species, reducing abundance in prey species of seabirds (Nordström et al. 2003), voles (Banks et al. 2005), and crustaceans (Fischer et al. 2009), six of them being included in the IUCN Red List categories near threatened, vulnerable, endangered and critically endangered (Genovesi et al. 2012). One of the impacted species is the critically endangered European mink (Mustela lutreola), with evidence of competition for resources (Melero et al. 2008; Sidorovich et al. 2010; Podra et al. 2013). Both species have similar ecological niches, being carnivorous mammals in riparian ecosystems and predating on both aquatic and terrestrial prey. The presence of the American mink in the native species habitat was shown to reduce the diet breath of European mink so that it becomes more specialized, while the American mink’s diet became more generalist (Sidorovich et al. 2010).
In France, the American mink was introduced in the 1920s in the Eastern side of the country; in the 1950s, many farms moved to the western side where access to fish by-products for mink feeding was readily available (Léger and Ruette 2005). A long-term monitoring study from 2000 to 2015 recorded evidence of the expansion of the American mink over the Atlantic coast in France with multiple established populations, including: (1) the historical region of Brittany, Normandy and Pays de la Loire, (2) the western region of the Pyrenees up to northern Aquitaine, and (3) the Eastern region of the Pyrenees (Léger et al. 2018; Fig. S1). In contrast, the western distribution of the European mink is reduced to seven departments of southwestern France (Maizeret et al. 2002) and to northern Spain, mainly in Navarra (Põdra and Gomez 2018). Moreover, French populations are probably highly fragmented, especially in departments where the invasive species is abundant. The low density of individuals in these regions and low genetic diversity of the Western population perhaps due to a bottleneck event (Cabria et al. 2015; Michaux et al. 2005) encouraged the creation of a captive breeding program in Spain at the Fundación para la Investigación en Etología y Biodiversidad (FIEB), with individuals originating from free-ranging populations in Spain.
The purpose of this study was to understand the relationships between the gut microbiota of related invasive and native host species sharing similar ecological niches. We were interested in: (i) if the environment (free-ranging or captive) had a stronger influence than species or population identity for gut microbial diversity and composition, (ii) if differences in gut microbial communities could be explained by host belonging to a specific population, and (iii) if genetic relationships between host populations were reflected in terms of gut microbial compositional similarity. To study these questions, we examined gut bacterial species (or phylotype) richness, gut microbiota structure, and composition differences between American and European mink in captive settings (fur farm and captive breeding program), and in the natural environment across three different habitats in western France and Spain (Brittany region, the Nive basin in the Pyrénées-Atlantiques departement, and Navarra). To test for a phylosymbiosis signal, we also investigated the genetic diversity and structure of those mink populations using previously collected data through neutral microsatellite markers analysis.
Methods
Sample collection and study sites
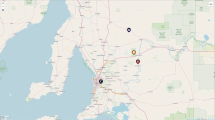
Fecal samples and rectal contents were collected from live or dead animals from five different populations. For free-ranging populations, six European mink were sampled in the Navarra region (Spain), twelve American mink were sampled in the Nive watershed (Southwest, Pyrénées-Atlantiques, France) and sixteen American mink from Brittany (Tomé island and close mainland; Fig. 1). To investigate habitat variation from each free-ranging sampled populations, a map was created using QGIS 3.16.6-Hanover with GPS coordinates for each sample. Layers documenting landscape use, were simplified to agricultural, built, natural and water surfaces from datasets originating from IDENA (Spatial Data Infrastructure of Navarre) and data.gouv.fr from Open Street Map (Alexandre Lexman). For captive populations, ten European mink were sampled in captive settings at the Fundación para la Investigación en Etología y Biodiversidad breeding center (FIEB) and fourteen American mink from a fur farm in the Pyrénées-Atlantiques department (Southwest of France). All samples were collected using sterile tweezers and placed in sterilized microcentrifuge tubes filled with 96% ethanol. Those tubes were stored in a − 20 °C freezer until further processing (Asangba et al. 2019).
DNA extraction and sequencing
Gene amplicon sequencing was used to study the bacterial communities. DNA extractions from the fecal samples collected were conducted using the QiaAmp Mini Kit with Inhibitex (Qiagen) following the manufacturer's instructions. Two blank extractions were made to control for contamination during the extraction process. A mock community sample (HM-783D, BEI resources) containing genomic DNA from 20 bacterial strains at concentrations ranging from 0.6 to 1400 pg/μl was also added in each library to confirm the reliability of our method. After DNA extraction, the targeted gene for taxonomic affiliation (16S rRNA gene—515F & 806R) was amplified through polymerase chain reactions (PCRs). The library preparation and sequencing were performed by Novogene UK. Using their designated library protocol, 2 × 250 bp paired‐end sequencing was completed using broad bacterial primers of the region V4 of the 16S rRNA gene using an Illumina NovaSeq platform in 100k reads/samples depth (Illumina Biotechnology Co.).
Bioinformatics
The quality controls of the paired‐end sequence reads were performed through the software FastQC (Andrews 2010). Sequence reads demultiplexing, denoising and amplicon sequence variants (ASVs) picking steps were done with the QIIME2 tool (Bolyen et al. 2019; v. 2020.8), using the DADA2 pipeline (Callahan et al. 2016, 2017). ASVs—or also referred to as bacterial phylotypes—were then screened to the 97% 16S rRNA gene full‐length reference sequences from the Silva RDP v.138.1 database (Pruesse et al. 2007) for taxonomical association using the VSEARCH classifier implemented in QIIME2 (Bokulich et al. 2018). Sequence alignment and phylogeny building were also conducted in QIIME2.
Analysis of the two blank extractions showed the presence of bacterial sequences that probably derived from contamination during laboratory sample handling. However, the diversity of this control was dissimilar from those of all mink samples (Bray–Curtis dissimilarity > 80.6%). For subsequent analysis of sequences associated with mink samples, sequences found in blank extraction samples were trimmed from the whole dataset. The cumulative sum scaling (CSS) method was used to normalize the data using the metagenomeSeq package (Paulson et al. 2013) in R (R version 3.5.2, R Core Team 2008). It can decrease the fold difference in sampling depth and avoid the rarefying of counts (Weiss et al. 2017).
Statistical analysis for comparison of α‐diversity of gut bacteria between groups
After CSS normalization, mink groups were divided as followed: European mink in captivity (EM Breeding Center; n = 7), American mink in captivity (AM Farm; n = 14), free-ranging European mink (EM Spain; n = 6); and free-ranging American mink distinct populations in Brittany and Nive (AM Brittany; n = 15 and AM Nive; n = 10; Fig. 1). All statistical analyses were conducted in R (R version 3.5.2, R Core Team 2008) using the phyloseq (McMurdie and Holmes 2013) and microbiome packages (Lahti et al. 2017) for manipulation of data. Chao1, Shannon’s diversity index and Faith’s PD in each sample were used as metrics to measure the α‐diversity of gut bacteria between samples. Linear mixed models fit by REML were conducted with according to either mink populations or mink species and environment (captive of wild) and each model included the variable host sex, sample type (feces or rectal content) and animal status (live or dead at time of sampling) as predictors, and microbial richness indexes as response variable. Normality of the residuals were tested with Shapiro–Wilk tests and visual representations.
Statistical analysis for comparison of β‐diversity of gut bacteria between groups and differential abundance
Unweighted and weighted UniFrac distance matrices between samples (Lozupone et al. 2010) were used to investigate differences in gut microbial communities between population, host sex, host environment, and host species with all bacterial taxa. A PERMANOVA model Adonis from the vegan package was constructed with 9999 permutations with reported F, R2, and p values to determine whether there were significant differences between the mink populations, host species, sex, status of the animal and sample type as main effects (Oksanen et al. 2019) after testing the homogeneity of groups variances using the betadisper function from the same package. Pairwise PERMANOVAs were then conducted to investigate variations between groups with 9999 permutations. A principal component analysis (PCoA) using Unifrac distance measures between samples was conducted to visualize the potential similarities between groups. Finally, a UPGMA dendrogram was constructed using the qiime diversity beta-rarefaction function in QIIME 2 by mink populations with weighted Unifrac distances with 20 iterations with mean ceiling at 10,000 sequences rarefaction.
The differential abundance analysis was conducted on the raw ASVs count, using the DESeq2 package (Love et al. 2014), with a negative binomial Wald test to test significance between each group. Only phylotypes with a significance level (α) below 0.001 after false discovery rate (FDR) corrections were considered using the Benjamin–Hochberg method. All phylotypes were tested in contrast, meaning that differential abundance was done pairwise between each mink population. ASVs below the significance level and with a negative log2 fold change had thus their abundance significantly lower in the first group tested, and a positive log2 fold change indicated that the phylotype was significantly higher in the first group compared to the other group. We conducted this analysis to test differential abundance first, between captive and free-ranging populations within mink species and second between free-ranging populations between and within mink species.
Microsatellite markers genotyping, and analysis
A total of 94 hair and tissue samples were extracted from a larger dataset of samples from European and American mink over a ten-year period between 2000 and 2019 (unpublished data). All samples derived from the same population that the fecal samples originated from, but many from different individuals. Eighteen free-ranging American mink were sampled in Brittany (Côtes d’Armor), thirty American mink were sampled in the Pyrenees Atlantiques (Southwest region of France), as well as thirty individuals from the same fur farm in the southwest of France. Finally, ten captive European mink were sampled in captive settings at the FIEB breeding center and six free-ranging European mink were sampled in Navarra (Spain).
Genomic DNA was isolated using the DNeasy Blood and Tissue Kit (QIAGEN) from tissue and hair samples. Negative controls were also used. Multilocus genotypes were obtained by PCR amplification of 10 autosomal microsatellites (Fleming et al. 1999; Cabria et al. 2007). The forward primer of each locus was 5′-end labeled with a fluorescent dye. The following multiplex sets were designed: mix 1 (MLUT 25, MLUT 27, Mvis 099) and mix 2 (MLUT 04, MER009, Mvis075, Mvis072, MER41, MER022). PCR and genotyping steps were carried out following Pigneur et al. (2019). Length variation determination (alleles and genotypes) was performed using Genemapper 4.0 (Applied Biosystems). To construct consensus multilocus genotypes, an allele was only accepted if observed at least twice. We thus accepted heterozygous genotypes that were observed twice. A homozygote was accepted after three positive PCRs gave the same single allele.
The genetic structure of the mink populations was inferred using Bayesian clustering analysis with Structure 2.3 software (Pritchard et al. 2000). We ran 10 iterations for each K value from 1 to 10 using the admixture model. A total of 106 MCMC repetitions were performed after a burn-in period of 20%. The results of the 10 iterations for each K value were summarized and averaged using the Clumpp method (Jakobsson and Rosenberg 2007). The optimal number of clusters was investigated using the ΔK method (Evanno et al. 2005). For subsequent analyses, individuals were sorted according to their geographic origin (sorted into 5 main populations: Brittany, Nive basin, Navarra, Farm and Breeding Center, Fig. 1). Mean allelic richness by locus (Ar), the expected (He) and observed (Ho) heterozygosity were calculated for each defined group using diveRsity (Keenan et al. 2013). An Euclidian distance matrix was constructed using GenAlEx 6.5 (Peakall and Smouse 2006), and a PERMANOVA model Adonis was conducted in a similar manner to β‐diversity gut bacterial analysis with species and population. A UPGMA dendrogram was also constructed based with average linkage based on Fst values between mink populations.
Results
Microsatellite markers analysis
Overall, the three American mink populations had greater total allele count (A), percentage of heterozygous locus (%H), allelic richness (Ar), observed heterozygosity (Ho) and expected heterozygosity (He) than the European mink populations (Table 1). This suggests greater heterozygosity and genetic diversity in neutral markers for the American mink, and we observed even higher for American mink in the fur farm compared to feral conspecifics. Bayesian clustering assignment recovered three distinct genetic clusters within our populations (Fig. 2; Table S1); the European mink individuals form one cluster (K2), American mink from the farm and the Nive basin another (K1), and individuals from Brittany overlap on 2 clusters (K1 and K3; Figs. 2, 3A). Only three American mink had admixture patterns between the two American mink clusters and belong to the fur farm population. Genetic distances between individuals’ analysis through a PERMANOVA model indicated significantly greater distance between mink species than within, as well as according to mink population (Adonis: F = 7.6547; R2 = 0.07206; p = 0.0009; F = 3.1927; R2 = 0.09016; p = 0.0089, respectively). Finally, a dendrogram based on Fst distances between populations revealed that the mink population sampled had lower distance within species than between species (Fig. 3A).
Comparison of α‐diversity in gut bacterial
Samples of a mock community containing known concentrations of genomic DNA from 20 bacterial strains were sequenced. Nineteen of the twenty different strains originally included in the sample were detected. Therefore, our protocol allowed bacterial DNA detection and identification to the genus level if its concentration in the DNA extract was at least 2.8 pg/μl, and provided that the sequence was included in the reference database. Following the raw data processing, we obtained 1,947,964 sequences belonging to 3036 distinct bacterial phylotypes (ASVs) after removal of negative control sequences, for 57 samples.
Gut bacterial phylotypes richness did not significantly vary according to host species, sex, animal status or sample type when considering three richness measures. However, both captive mink species exhibit lower bacterial diversity through lower Shannon indexes compared to conspecific free-ranging mink (Table 2; Fig. 4B; F = 2.718; p value < 0.05). The Shannon index also significantly varied according to mink populations (Table 2; F = 2.505; p value < 0.05) and captivity seemed to have a strong negative impact on gut bacterial richness for both host species, especially compared to the American mink population from Brittany (Fig. 4A, B).
Boxplots representing Shannon Index variation of the gut microbiota depending on A host’s environmental group, ** represents the p value meeting the standard cutoff p < 0.01 and ***p < 0.001 from by Dunn test of Kruskal–Wallis multiple comparisons with Benjamini and Hochberg correction, and depending on B host’s environment for both mink species
Comparison of β‐diversity of gut bacteria between groups
As expected in mustelid gastrointestinal tracts, all samples were dominated by the Firmicutes and Proteobacteria phyla, mostly belonging to the Clostridiaceae and Peptostreptococcaceae families (Fig. 5; Compo et al. 2018; Bahl et al. 2017). The gut bacterial community composition of male and female mink for both species considered in the study (Adonis: F = 0.314; R2 = 0.0058; p = 0.725) were not significantly different and explained around 0.5% of the variation, neither did host status (Adonis: F = 1.189; R2 = 0.0208; p = 0.451) or sample type (Adonis: F = 1.987; R2 = 0.0348; p = 0.109). However, 15% of gut bacterial composition variation was explained by mink belonging to the different populations in both weighted and unweighted Unifrac distances (Adonis: F = 2.411; R2 = 0.1581; p = 0.0115; and F = 2.3067; R2 = 0.1503; p = 0.0001, respectively, Fig. 6). The variation seemed to be mainly explained by free-ranging or captive conditions (Figure S2). We did observe significantly greater distances between feral American mink in Brittany and other American mink groups, but no differences were detected between both captive and free-ranging European mink and American mink in Brittany (Figs. 6 & S2). A wide interindividual variation in gut bacterial composition was also observed in free-ranging European mink (Figure S2). Overall, feral American mink in Brittany and free-ranging European mink had lower β‐diversity between them than any other mink populations (Fig. 3B).
Differential bacterial abundance analysis
The assessment of the differential abundance of bacterial phylotypes using a negative binomial Wald test was conducted on the core microbiota of 391 phylotypes. From those, 141 phylotypes from nine phyla varied significantly among the mink populations with 82% belonging to Firmicutes and Proteobacteria. When comparing captive and free-ranging populations within mink species, feral American mink had phylotypes differentially abundant to captive conspecifics, from 100 to 65 ASVs, most of them decreasing (Table S2). Feral American mink had a ratio of 1.77 and 2.6, expressing more decreases than increases in taxa abundance in the natural environment compared to captive conspecifics. This decrease in taxon abundance between free-ranging and captive populations is higher in American than European mink (0.7). A large portion of those phylotypes belonged to the Bacteroida (families Flavobacteriaceae, Muribacculaceae and Chitinobacteraceae) and Clostridia (genera Rhomboustia mostly) classes (Table S2). However, when comparing free-ranging populations of both species, we observed more taxa abundance variation between the two populations of free-ranging American mink (64 taxa differentially abundant), than variation between American and European mink (53 taxa for the Nive basin and 42 taxa for Brittany). Feral American mink in Brittany had more phylotypes abundances in common with free-ranging native European mink than its conspecifics from the Nive basin (Table S2). Most of the abundance variation was attributed to reduction in ASVs belonging to the Firmicutes phylum (Lactobacillus, Clostridium genera and Peptostreptococcaceae family) and Gammaproteobacteria class between the two American mink populations.
Discussion
On the influence of human impacts on the mink gut microbiota
This study is the first to examine how the gut bacteria of riparian carnivores vary between related species with similar ecological niches in the context of farming, invasion, and conservation. Our results did not find any support for phylosymbiosis, as genetic relationships were not reflected in the composition of the gut microbiota (Fig. 3). There was also a reduction in the richness of the bacterial community in captivity that surpassed host species differences. A similar pattern was further observed in β‐diversity measures. This trend has already been observed in other host taxa with a carnivorous diet (Canidae, McKenzie et al. 2017). It is currently well established that animals living in captivity experience a range of changes that can influence their gut bacteria, from diet change, veterinary care, specific and uniform environmental substrates, as well as reduced contact with conspecifics and other species. While most of the current literature compared free-ranging animals to individuals kept in zoos (Clayton et al. 2016; Borbón-García et al. 2017; Wasimuddin et al. 2017), the same trend is expected between feral and farmed mink.
We also observed differentially abundant taxa in free-ranging mink compared to captive conspecifics. In addition, bacterial loss was stronger in the invasive American mink than the native European species when comparing free-ranging populations to captive conspecifics. In this regard, feral American mink would have experienced less recolonization from gut bacteria in natural habitats than the European mink, when compared to their captive conspecifics. By nestedness and turnover of bacterial communities, feral American mink would have left a subset of captive gut microbes during the invasion process, potentially leading to pathogen loss. However, many successful invasions have occurred without any pathogen loss and further investigation on targeted bacteria would be needed (Amsellem et al. 2017).
There are three potential ways that can explain a stronger pattern of differentiation in gut bacteria communities between feral to captive settings in the American mink compared to the European mink. First, the two species have very different conditions in captivity. Farmed American mink are held in individual and open-air elevated cages with minimal substrate and enrichment, while European mink are held in an enclosure with access to a pond, natural substrates and enrichment (branches, vegetation, mud). Moreover, the diet of the American mink is composed of processed fish and chicken, whereas the diet of the European mink consists of whole fish, chicken and mice. Those differences in captive conditions could explain the significant difference in the bacterial communities between wild and captive American mink, compared to the European mink.
Second, when considering free-ranging European mink in their natural habitat, they could be more likely to select specific gut bacterial taxa because of their shared coevolutionary history with the environmental microbes in western France (Bankers et al. 2021). On the other hand, the invasive American mink may lack host-microbes coevolutionary history with native bacteria and could thus be less likely to retain newly acquired microbes when becoming feral. It is worth noting that the estimated divergence time between the two mink species is 8.28 million years ago (Hedges et al. 2006), and further research with other native mustelids such as the European polecat (Mustela putorius), that diverged more recently from the European mink could give more insight into gut bacteria colonization from wild to captive settings.
The third explanation relies on the evolutionary history of the American mink itself. The domestication process of this species started in the 1860s in Canada (Morris et al. 2020), as humans selected animals with dense, soft, and shiny fur, as well as increased fertility to maximize their revenue. Docility, also termed confidence towards humans, was another behavioural trait that many European breeders favoured to ease daily handling and improved welfare (Thirstrup et al. 2019). Thus, genetic and phenotypic differences have already been observed between free-ranging and farmed American mink, including smaller brain size, longer transit time and increased nitrogen metabolism in farmed animals (Morris et al. 2020; Bowman et al. 2017; Gugolek et al. 2013; Kruska 1996). This explanation seems consistent with the high genetic diversity in mink from the fur farm observed in this study compared to free-ranging American mink populations. There is increasing evidence of the important interactions between the gut microbiota and the gut-brain axis in many species, including farm animals (Collins et al. 2012; Kraimi et al. 2019). It would be likely that artificial selection might have impacted the overall gut microbiota composition of the American mink through morphological and physiological variation, and thus changes in the gut-brain-axis, compared to the European mink that has not experienced domestication. The effects of domestication on gut bacteria have already been observed in other mammals (Prabhu et al. 2020). However, to confirm either of both explanations on those exploratory results, further investigation with larger sampling size should be conducted.
No phylosymbiosis signal observed in mink
In general, our results did not support the phylosymbiosis hypothesis, and it was observed that the host environment had a strong influence on the mink gut microbiota. First, neither gut bacterial α- or β‐diversity varied according to host species. Second, the feral American mink groups were more distinct from one another than with the free-ranging European mink, despite belonging to the same species. Furthermore, feral American mink in the Nive basin had less similarly abundant bacterial taxa in common with free-ranging European mink than feral American mink in Brittany. The absence of a phylosymbiosis signal is consistent with the fact that despite not being the most diverse population genetically, the invasive American mink from Brittany are the most genetically differentiated from the other American mink populations, being composed of at least three different genetic clusters. Three genetic pools have already been documented in this long-established population due to accidental releases over multiple introduction events, fostering diversity but also genetic drift (Bifolchi et al. 2010).
Similar to formation of a distinct population through genetic drift within farms, an analogous concept termed ecological drift might have occurred in gut microbes between mink populations, in relation with the ecology of the host (Kohl 2020). These shifts in bacterial composition between free-ranging mink species could be explained by variation in prey availability due to habitat differences between the areas sampled. Studies in other parts of Europe showed that the American mink has a plastic diet (Maran et al. 1998; Zalewski and Bartoszewicz 2012; Chibowski et al. 2019). When found in agricultural landscapes, the mink tend to feed on ground-dwelling small mammals, such as Microtus sp. that are highly abundant in rural habitats (Krawczyk et al. 2013). Considering the variation in landscapes in our study (Fig. 1), the Côtes d’Armor area in Brittany is more subject to anthropogenic activities compared to the Nive watershed in the Southwest. The latter is mainly composed of forests (48%) and meadows (30%; MNHN 2015), while the Côtes d’Armor landscape was dominated in 2015 by agricultural areas (56%), then forests (21%) and very few meadows (9%; DRAAF Bretagne 2021). A study conducted in northeastern Spain observed that the free-ranging American mink mostly predated on crayfish and this might be reflective of the mink diet in the Nive watershed (Melero et al. 2008). This difference in diet related to landscape variation between the two American mink populations could thus be reflected in the different composition of the gut microbial communities (Reese and Dunn 2018; San Juan et al. 2020).
Regarding the free-ranging European mink habitat, the land uses of Navarra in 2015 was primarily agricultural areas (34.8%) and forests (28.2%), followed by meadows (15.7%; Vicente et al. 2005). The greatest proportion of agricultural lands in both Navarra and Brittany could thus indicate similar prey availability compared to the Nive watershed. Palazon et al. (2004) observed that the European mink diet in Navarra and La Rioja was predominantly composed of small mammals and fish, thus supporting the hypothesis that gut microbial composition of both mink species according to prey availability based on land occupation. A recent survey of the European mink diet in Navarra supports this hypothesis, mammals such as Arvicola sp., Microtus sp. and Apodemus sp. representing 36% of the diet from 30 mink sampled, and squamates like Natrix sp. 25% of the diet, while fishes and crustaceans respectively represented 6% and 5% of their diet (Podra, unpublished data 2022). Further work on their diet and gut bacteria, as well as prey surveys in all mink territory could validate this hypothesis.
In conclusion, this study provides insight into the relationship between the gut bacteria of invasive and native carnivorous mammal hosts, with no observable signals of phylosymbiosis due to the strong influence of the environment and diet of the host on its associated microbes. Studying gut microbiota differences between mink farms in multiple countries, as well as individuals in their native habitat could also give more insight into the effects of domestication on microbe-host relationships. The American mink could also play a role in disease transmission among native species, as they can carry the Aleutian Disease Virus (ADV), the Canine Distemper Virus (CDV) as well as many eukaryote parasites that can be transmitted to other mustelids, feral cats, and even humans (Leimann et al. 2015; Martínez-Rondán et al. 2017; Torres et al. 2008). While the methods in our study did not allow to address deleterious microbes in the gut of the American mink as a reservoir that impacts European mink, investigation on functional diversity within those microbes could shed more light on this question. Studying the invasive success of a carrier species like the American mink becomes even more critical, especially because mink, feral and/or in farms, interact with many other species including humans.
Data availability
Supporting information has been made available online. Final DNA sequences, ASV table, taxonomy table and mapping file have been uploaded: Dryad link TBD.
References
Amsellem L, Brouat C, Duron O, Porter SS, Vilcinskas A, Facon B (2017) Importance of microorganisms to macroorganisms invasions: is the essential invisible to the eye? In: Advances in ecological research, vol 57, 1 edn. https://doi.org/10.1016/bs.aecr.2016.10.005
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc
Asangba AE, Halajian A, Lamb A, Wright PC, Leigh SR, Stumpf RM (2019) Variations in the microbiome due to storage preservatives are not large enough to obscure variations due to factors such as host population, host species, body site, and captivity. Am J Primatol. https://doi.org/10.1002/ajp.23045
Bahl MI, Hammer AS, Clausen T, Jakobsen A, Skov S, Andresen L (2017) The gastrointestinal tract of farmed mink (Neovison vison) maintains a diverse mucosa-associated microbiota following a 3-day fasting period. MicrobiologyOpen 6(3):1–8. https://doi.org/10.1002/mbo3.434
Bankers L, Dahan D, Neiman M, Adrian-Tucci C, Frost C, Hurst GDD, King KC (2021) Invasive freshwater snails form novel microbial relationships. Evol Appl 14(3):770–780. https://doi.org/10.1111/eva.13158
Banks PB, Nordstrom M, Ahola M, Korpimaki E (2005) Variable impacts of alien mink predation on birds, mammals and amphibians of the Finnish archipelago: a long-term experimental study. In: IX International Mammalogical Congress. Sapporo, Japan
Baxter NT, Wan JJ, Schubert AM, Jenior ML, Myers P, Schloss PD (2015) Intra- and interindividual variations mask interspecies variation in the microbiota of sympatric Peromyscus populations. Appl Environ Microbiol 81(1):396–404. https://doi.org/10.1128/AEM.02303-14
Bifolchi A, Picard D, Lemaire C, Cormier JP, Pagano A (2010) Evidence of admixture between differentiated genetic pools at a regional scale in an invasive carnivore. Conserv Genet 11(1):1–9. https://doi.org/10.1007/s10592-008-9780-1
Bokulich NA, Kaehler BD, Rideout JR, Dillon M, Bolyen E, Knight R, Gregory Caporaso J (2018) Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6(1):1–17. https://doi.org/10.1186/s40168-018-0470-z
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Caporaso JG (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37(8):852–857. https://doi.org/10.1038/s41587-019-0209-9
Bonesi L, Palazon S (2007) The American mink in Europe: status, impacts, and control. Biol Conserv 134(4):470–483. https://doi.org/10.1016/j.biocon.2006.09.006
Borbón-García A, Reyes A, Vives-Flórez M, Caballero S (2017) Captivity shapes the gut microbiota of Andean bears: insights into health surveillance. Front Microbiol 8:1–12. https://doi.org/10.3389/fmicb.2017.01316
Bowman J, Beauclerc K, Farid AH, Fenton H, Klütsch CFC, Schulte-Hostedde AI (2017) Hybridization of domestic mink with wild American mink (Neovison vison) in eastern Canada. Can J Zool 53(9):1689–1699. https://doi.org/10.1017/CBO9781107415324.004
DRAAF Bretagne (2021) Fiches territoriales des départements bretons. Site officiel du service régional du ministère en charge de l’agriculture. https://draaf.bretagne.agriculture.gouv.fr/Environnement Accessed 10 June 2021
Brooks AW, Kohl KD, Brucker RM, van Opstal EJ, Bordenstein SR (2016) Phylosymbiosis: relationships and functional effects of microbial communities across host evolutionary history. PLoS Biol 14(11):1–29. https://doi.org/10.1371/journal.pbio.2000225
Cabria MT, González EG, Gómez-Moliner BJ, Zardoya R (2007) Microsatellite markers for the endangered European mink (Mustela lutreola) and closely related mustelids. Mol Ecol Notes 7(6):1185–1188. https://doi.org/10.1111/j.1471-8286.2007.01825.x
Cabria MT, Gonzalez EG, Gomez-Moliner BJ, Michaux JR, Skumatov D, Kranz A, Zardoya R (2015) Patterns of genetic variation in the endangered European mink (Mustela lutreola L., 1761). BMC Evolut Biol 15(1):141. https://doi.org/10.1186/s12862-015-0427-9
Callahan BJ, Mcmurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13(7):581–588. https://doi.org/10.1038/nmeth.3869
Callahan BJ, Mcmurdie PJ, Holmes SP (2017) Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J 11(12):2639–2643. https://doi.org/10.1038/ismej.2017.119
Chibowski P, Zalewski A, Suska-Malawska M, Brzeziński M (2019) Study on geographical differences in American mink diets reveals variations in isotopic composition of potential mink prey. Mammal Res 64(3):343–351. https://doi.org/10.1007/s13364-019-00419-4
Clayton JB, Vangay P, Huang H, Ward T, Hillmann BM, Al-Ghalith GA, Knights D (2016) Captivity humanizes the primate microbiome. Proc Natl Acad Sci 113(37):10376–10381. https://doi.org/10.1073/pnas.1521835113
Collins SM, Surette M, Bercik P (2012) The interplay between the intestinal microbiota and the brain. Nat Rev Microbiol 10(11):735–742. https://doi.org/10.1038/nrmicro2876
Compo NR, Gomez DE, Tapscott B, Weese JS, Id VT (2018) Fecal bacterial microbiota of Canadian commercial mink (Neovison vison): yearly, life stage, and seasonal comparisons. PLoS ONE 13(11):1–18. https://doi.org/10.5683/SP/OEIP77
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620
Fischer D, Pavluvcik P, Sedlacek F, Salek M (2009) Predation of the alien American mink, Mustela vison on native crayfish in middle-sized streams in central and western Bohemia. Folia Zool 58(1):45–56
Fleming MA, Ostrander EA, Cook JA (1999) Microsatellite markers for American mink (Mustela vison) and ermine (Mustela erminea). Mol Ecol 8(8):1351–1352. https://doi.org/10.1046/j.1365-294X.1999.00701.x
Frankel JS, Mallott EK, Hopper LM, Ross SR, Amato KR (2019) The effect of captivity on the primate gut microbiome varies with host dietary niche. Am J Primatol 81(12):1–9. https://doi.org/10.1002/ajp.23061
Genovesi P, Carnevali L, Alonzi A, Scalera R (2012) Alien mammals in Europe: updated numbers and trends, and assessment of the effects on biodiversity. Integr Zool 7(3):247–253. https://doi.org/10.1111/j.1749-4877.2012.00309.x
Greene LK, Clayton JB, Rothman RS, Semel BP, Semel MA, Gillespie TR, Drea CM (2019) Local habitat, not phylogenetic relatedness, predicts gut microbiota better within folivorous than frugivorous lemur lineages. Biol Lett 15(6):5–11. https://doi.org/10.1098/rsbl.2019.0028
Grond K, Bell KC, Demboski JR, Santos M, Sullivan JM, Hird SM (2020) No evidence for phylosymbiosis in western chipmunk species. FEMS Microbiol Ecol 96(1):1–10. https://doi.org/10.1093/femsec/fiz182
Groussin M, Mazel F, Sanders JG, Smillie CS, Lavergne S, Thuiller W, Alm EJ (2017) Unraveling the processes shaping mammalian gut microbiomes over evolutionary time. Nat Commun 8:14319. https://doi.org/10.1038/ncomms14319
Gugołek A, Zalewski D, Strychalski J, Konstantynowicz M (2013) Food transit time, nutrient digestibility and nitrogen retention in farmed and feral American mink (Neovison vison)—a comparative analysis. J Anim Physiol Anim Nutr 97(6):1030–1035. https://doi.org/10.1111/jpn.12006
Hedges SB, Dudley J, Kumar S (2006) TimeTree: a public knowledged-base of divergence times among organisms. Bioinformatics 22:2971–2972
Jakobsson M, Rosenberg NA (2007) Clumpp: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Keenan K, McGinnity P, Cross TF, Crozier WW, Prodöhl PA (2013) diveRsity: an R package for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol Evol 4:782–788
Knowles SCL, Eccles RM, Baltrūnaitė L (2019) Species identity dominates over environment in shaping the microbiota of small mammals. Ecol Lett 22(5):826–837. https://doi.org/10.1111/ele.13240
Kohl KD (2020) Ecological and evolutionary mechanisms underlying patterns of phylosymbiosis in host-associated microbial communities. Philos Trans R Soc B Biol Sci 375(1798):20190251. https://doi.org/10.1098/rstb.2019.0251
Kohl KD, Dearing MD, Bordenstein SR (2018) Microbial communities exhibit host species distinguishability and phylosymbiosis along the length of the gastrointestinal tract. Mol Ecol 27:1874–1883. https://doi.org/10.1111/mec.14460
Koskella B, Hall LJ, Metcalf CJE (2017) The microbiome beyond the horizon of ecological and evolutionary theory. Nat Ecol Evolut. https://doi.org/10.1038/s41559-017-0340-2
Kraimi N, Dawkins M, Gebhardt-Henrich SG, Velge P, Rychlik I, Volf J, Leterrier C (2019) Influence of the microbiota-gut-brain axis on behavior and welfare in farm animals: a review. Physiol Behav 210:11265. https://doi.org/10.1016/j.physbeh.2019.112658
Krawczyk AJ, Bogdziewicz M, Czyz MJ (2013) Diet of the American mink Neovison vison in an agricultural landscape in western Poland. Folia Zool 62(4):304–310. https://doi.org/10.25225/fozo.v62.i4.a8.2013
Kruska D (1996) The effect of domestication on brain size and composition in the mink (Mustela vison). J Zool 239(4):645–661. https://doi.org/10.1111/j.1469-7998.1996.tb05468.x
Lahti L et al (Bioconductor, 2017) Tools for microbiome analysis in R. Microbiome package version. http://microbiome.github.io/microbiome
Léger F, Ruette S (2005) Le Vison d’Amérique, une espèce qui se développe en France: résultat d’une enquête nationale réalisée en 1999. Faune Sauvage 266:29–36
Léger F, Steinmetz J, Laoué E, Maillard JF, Ruette S (2018) L’expansion du vison d’Amérique en France Période 2000–2015. Faune Sauvage 138:23–31
Leimann A, Knuuttila A, Maran T, Vapalahti O, Saarma U (2015) Molecular epidemiology of Aleutian mink disease virus (AMDV) in Estonia, and a global phylogeny of AMDV. Virus Res 199:56–61. https://doi.org/10.1016/j.virusres.2015.01.011
Ley RE, Hamady M, Lozupone C, Turnbaugh PJ, Ramey RR, Bircher JS, Gordon JI (2008) Evolution of mammals and their gut microbes. Science 320:1647–1651. https://doi.org/10.1126/science.1155725
Lim SJ, Bordenstein SR (2020) An introduction to phylosymbiosis. Proc R Soc B Biol Sci 287(1922):20192900. https://doi.org/10.1098/rspb.2019.2900
Lockwood JL, Hoopes MF, Marchetti MP (2007) Invasion ecology, 1st edn. Wiley-Blackwell, New York
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Lozupone C, Lladser ME, Knights D, Stombaugh J, Knight R (2010) UniFrac: an effective distance metric for microbial community comparison. ISME J 5(2):169–172. https://doi.org/10.1038/ismej.2010.133
Maizeret C, Migot P, Rosoux R, Chusseau JP, Gatelier T, Maurin H, Fournier-Chambrillon C (2002) The distribution of the European mink (Mustela lutreola) in France: towards a short term extinction? Mammalia 66:525–532
Maran T, Kruuk H, Macdonald DW, Polma M (1998) Diet of two species of mink in Estonia: displacement of Mustela lutreola by M. vison. Commun Mammal Soc 76(245):218–222
Martinez-Mota R, Kohl KD, Orr TJ, Dearing DM (2019) Natural diets promote retention of the native gut microbiota in captive rodents. ISME J. https://doi.org/10.1038/s41396-019-0497-6
Martínez-Rondán FJ, Ruiz de Ybáñez MR, Tizzani P, López-Beceiro AM, Fidalgo LE, Martínez-Carrasco C (2017) The American mink (Neovison vison) is a competent host for native European parasites. Vet Parasitol 247:93–99. https://doi.org/10.1016/j.vetpar.2017.10.004
McKenney EA, Koelle K, Dunn RR, Yoder AD (2018) The ecosystem services of animal microbiomes. Mol Ecol 27:2164–2172. https://doi.org/10.1111/mec.14532
McKenzie VJ, Song SJ, Delsuc F, Prest TL, Oliverio AM, Korpita TM, Knight R (2017) The effects of captivity on the mammalian gut microbiome. Integr Comp Biol. https://doi.org/10.1093/icb/icx090
McMurdie PJ, Holmes S (2013) phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE. https://doi.org/10.1371/journal.pone.0061217
Melero Y, Palazon S, Bonesi L, Gosalbez J (2008) Feeding habits of three sympatric mammals in NE Spain: the American mink, the spotted genet, and the Eurasian otter. Acta Theriol 53(3):263–273. https://doi.org/10.1007/bf03193123
Michaux JR, Hardy OJ, Justy F, Fournier P, Kranz A, Cabria M, Libois R (2005) Conservation genetics and population history of the threatened European mink Mustela lutreola, with an emphasis on the west European population. Mol Ecol 14(8):2373–2388. https://doi.org/10.1111/j.1365-294X.2005.02597.x
Mora M, Medina-Vogel G, Sepúlveda MA, Noll D, Álvarez-Varas R, Vianna JA (2018) Genetic structure of introduced American mink (Neovison vison) in Patagonia: Colonisation insights and implications for control and management strategies. Wildl Res 45(4):344–356. https://doi.org/10.1071/WR18026
Morris KY, Bowman J, Schulte-Hostedde A, Wilson PJ (2020) Functional genetic diversity of domestic and wild American mink (Neovison vison). Evolut Appl. https://doi.org/10.1111/eva.13061
Muséum national d’Histoire naturelle (ed) (2015) Inventaire National du Patrimoine Naturel. https://inpn.mnhn.fr. Accessed 10 June 2021
Nordström M, Hogmander J, Laine J, Nummelin J, Laanetu N, Korpimaki E (2003) Effects of feral mink removal on seabirds, waders and passerines on small islands of the Baltic Sea. Biol Conserv 109:359–368
Ochman H, Worobey M, Kuo CH, Ndjango JBN, Peeters M, Hahn BH, Hugenholtz P (2010) Evolutionary relationships of wild hominids recapitulated by gut microbial communities. PLoS Biol 8(11):3–10. https://doi.org/10.1371/journal.pbio.1000546
Oksanen J, Blanchet GF, Friendly M, Kindt R, Legendre P, McGlinn D, Wagner H (2019) Vegan: community ecology package. R package version 2.5-4. https://CRAN.R-project.org/package=vegan
Palazón S, Ruiz-Olmo J, Gosàlbez J (2004) Diet of European mink (Mustela lutreola) in Northern Spain. Mammalia 68(2–3):159–165. https://doi.org/10.1515/mamm.2004.016
Paulson JN, Colin Stine O, Bravo HC, Pop M (2013) Differential abundance analysis for microbial marker-gene surveys. Nat Methods 10(12):1200–1202. https://doi.org/10.1038/nmeth.2658
Peakall R, Smouse PE (2006) GENALEX 6: Genetic analysis in. Excel population genetic software for teaching and research. Mol Ecol Notes 6(1):288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Phillips CD, Phelan G, Dowd SE, McDonough MM, Ferguson AW, Delton Hanson J, Baker RJ (2012) Microbiome analysis among bats describes influences of host phylogeny, life history, physiology and geography. Mo Ecol 21(11):2617–2627. https://doi.org/10.1111/j.1365-294X.2012.05568.x
Pigneur LM, Caublot G, Fournier-Chambrillon C, Fournier P, Giralda-Carrera G, Grémillet X, Le Roux B, Marc D, Simonnet F, Smitz N, Sourp E, Steinmetz J, Urra-Maya F, Michaux JR (2019) Current genetic admixture between relictual populations might enhance the recovery of an elusive carnivore. Conserv Genet 20:1133–1148. https://doi.org/10.1007/s10592-019-01199-9
Põdra M, Gómez A (2018) Rapid expansion of the American mink poses a serious threat to the European mink in Spain. Mammalia 82(6):580–588. https://doi.org/10.1515/mammalia-2017-0013
Põdra M, Maran T, Sidorovich VE, Johnson PJ, Macdonald DW (2013) Restoration programmes and the development of a natural diet: a case study of captive-bred European mink. Eur J Wildl Res 59(1):93–104. https://doi.org/10.1007/s10344-012-0653-z
Pollock FJ, McMinds R, Smith S, Bourne DG, Willis BL, Medina M, Zaneveld JR (2018) Coral-associated bacteria demonstrate phylosymbiosis and cophylogeny. Nat Commun 9(1):1–13. https://doi.org/10.1038/s41467-018-07275-x
Prabhu VR, Kamalakkannan R, Arjun MS, Nagarajan M (2020) Consequences of domestication on gut microbiome: a comparative study between wild gaur and domestic mithun. Front Microbiol 11:1–12. https://doi.org/10.3389/fmicb.2020.00133
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig WG, Peplies J, Glöckner FO (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35:7188–7196. https://doi.org/10.1093/nar/gkm864
R Development Core Team (2008) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Reese AT, Dunn RR (2018) Drivers of microbiome biodiversity: a review of general rules, feces, and ignorance. Mbio 9:e01294-18. https://doi.org/10.1128/mBio.01294-18
Reid F, Schiaffini M, Schipper J (2016) Neovison vison. The IUCN Red List of Threatened Species 2016: e.T41661A45214988. https://doi.org/10.2305/IUCN.UK.2016-1.RLTS.T41661A45214988.en
San Juan PA, Hendershot JN, Daily GC, Fukami T (2020) Land-use change has host-specific influences on avian gut microbiomes. ISME J 14(1):318–321. https://doi.org/10.1038/s41396-019-0535-4
Sevellec M, Laporte M, Bernatchez A, Derome N, Bernatchez L (2019) Evidence for host effect on the intestinal microbiota of whitefish (Coregonus sp.) species pairs and their hybrids. Ecol Evol 9(20):11762–11774. https://doi.org/10.1002/ece3.5676
Shimatani Y, Fukue Y, Kishimoto R, Masuda R (2010) Genetic variation and population structure of the feral American mink (Neovison vison) in Nagano, Japan, revealed by microsatellite analysis. Mammal Study 35(1):1–7. https://doi.org/10.3106/041.035.0101
Sidorovich VE, Polozov AG, Zalewski A (2010) Food niche variation of European and American mink during the American mink invasion in north-eastern Belarus. Biol Invasions 12(7):2207–2217. https://doi.org/10.1007/s10530-009-9631-0
Sonnenburg ED, Smits SA, Tikhonov M, Higginbottom SK, Wingreen NS, Sonnenburg JL (2016) Diet-induced extinctions in the gut microbiota compound over generations. Nature 529(7585):212–215. https://doi.org/10.1038/nature16504
Spor A, Koren O, Ley R (2011) Unravelling the effects of the environment and host genotype on the gut microbiome. Nat Rev Microbiol 9(4):279–290. https://doi.org/10.1038/nrmicro2540
Thirstrup JP, Villumsen TM, Malmkvist J, Lund MS (2019) Selection for temperament has no negative consequences on important production traits in farmed mink. J Anim Sci 97(5):1987–1995
Torres J, Miquel J, Fournier P, Liberge M, Fons R, Feliu C (2008) Helminth communities of the autochthonous mustelids Mustela lutreola and M. putorius and the introduced Mustela vison in south-western France. J Helminthol 82:349–355. https://doi.org/10.1017/S0022149X08046920
van Leeuwen P, Mykytczuk N, Mastromonaco GF, Schulte-Hostedde AI (2020) Effects of captivity, diet, and relocation on the gut bacterial communities of white-footed mice. Ecol Evol 10(11):4677–4690. https://doi.org/10.1002/ece3.6221
Van Opstal EJ, Bordenstein SR (2019) Phylosymbiosis impacts adaptive traits in Nasonia wasps. Mbio 10(4):1–11. https://doi.org/10.1128/mBio.00887-19
Vicente AM, Donézar M, Del Barrio F (2005) Mapa de cultivos y aprovevhamientos de Navarra. Departamento de Agricultura, Ganadería y Alimentación del Gobierno de Navarra
Wasimuddin, Menke S, Melzheimer J, Thalwitzer S, Heinrich S, Wachter B, Sommer S (2017) Gut microbiomes of free-ranging and captive Namibian cheetahs: diversity, putative functions and occurrence of potential pathogens. Mol Ecol 26(20):5515–5527. https://doi.org/10.1111/mec.14278
Weiss S, Xu ZZ, Peddada S, Amir A, Bittinger K, Gonzalez A, Hyde ER (2017) Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome 5(27):1–18. https://doi.org/10.1186/s40168-017-0237-y
West AG, Waite DW, Deines P, Bourne DG, Digby A, McKenzie VJ, Taylor MW (2019) The microbiome in threatened species conservation. Biol Conserv 229:85–98. https://doi.org/10.1016/j.biocon.2018.11.016
Zalewski A, Bartoszewicz M (2012) Phenotypic variation of an alien species in a new environment: The body size and diet of American mink over time and at local and continental scales. Biol J Linn Soc 105(3):681–693. https://doi.org/10.1111/j.1095-8312.2011.01811.x
Zalewski A, Michalska-Parda A, Bartoszewicz M, Kozakiewicz M, Brzeziński M (2010) Multiple introductions determine the genetic structure of an invasive species population: American mink Neovison vison in Poland. Biol Conserv 143(6):1355–1363. https://doi.org/10.1016/j.biocon.2010.03.009
Acknowledgements
FIEB (Ciprian Petrescu), INRAE (Olivier Lorvelec, Patricia le-Quilliec), Madis Podra, OFB (Christelle Bellanger, Maylis Fayet), Augustin Granel, Alice Mouton, Jasmine Veitch, Adrien André, EEP program for the European mink, Tiit Maran, the government of Navarra, and the Ministry for the Ecological Transition and the Demographic Challenge (Spain).
Funding
Funding was supported by the NSERC CREATE grant, ReNewZoo and the Conversation Genetics Laboratory of the University of Liège.
Author information
Authors and Affiliations
Contributions
JM, ASH and PVL planned and designed the study. CA, CFC, FU-M, and PC provided samples and contact for other sample access. PVL performed the sample preparation for sequencing. JM provided sequencing services and together with ASH advised on laboratory and sampling procedures. PVL performed bioinformatics, statistical analyses and the interpretation of results with feedbacks provided by ASH and JM. PVL wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Conflict of interest
All authors certify that they have no affiliations with or involvement in any organization or entity with any financial interest or non-financial interest in the subject matter or materials discussed in this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
van Leeuwen, P.M.L., Schulte-Hostedde, A.I., Fournier-Chambrillon, C. et al. A microbial tale of farming, invasion and conservation: on the gut bacteria of European and American mink in Western Europe. Biol Invasions 25, 1693–1709 (2023). https://doi.org/10.1007/s10530-023-03007-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-023-03007-5