Abstract
Invasive Indo-Pacific red lionfish (Pterois volitans) have become well-established residents within reef communities across the western Atlantic Ocean where they pose substantial threats to native fish communities and reef ecosystems. Species-specific identification of prey is necessary to elucidate predator–prey interactions, but can be challenging with traditional visual identification methods given prey are often highly digested, thus not identifiable visually. To supplement visual diet analysis of lionfish (n = 934) sampled in the northern Gulf of Mexico, we applied DNA barcoding to identify otherwise unidentifiable fish prey (n = 696) via amplification of the cytochrome c oxidase subunit I (COI) of the mitochondrial genome. Barcoding nearly doubled the number of identifiable fish prey, thereby greatly enhancing our ability to describe lionfish diet. Thirty-three fish prey species were identified via barcoding, twenty-four of which were not previously detected by traditional methods. Some exploited reef fishes were newly reported (e.g., red snapper, Lutjanus campechanus) or found to constitute higher proportions of lionfish diet than previously reported (e.g., vermilion snapper, Rhomboplites aurorubens). Barcoding added a significant amount of new dietary information, and we observed the highest prey diversity reported to date for invasive lionfish. Potential cannibalism on juveniles also was identified via DNA barcoding, with the highest incidence corresponding to high lionfish densities, thus suggesting density-dependent prey demand may have driven this response. Overall, DNA barcoding greatly enhanced our ability to describe invasive lionfish diet in this study, suggesting that even studies with relatively large diet sample sizes could benefit from barcoding analysis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Indo-Pacific red lionfish, Pterois volitans (Linnaeus, 1758) (hereafter lionfish) are piscivorous scorpionfish that have exhibited an unprecedented invasion in the western Atlantic Ocean over the past 30 years (Whitfield et al. 2002; Schofield 2010). While invasions by marine predators are atypical, lionfish have exhibited an invasion so extensive and rapid in this region that they are considered to be the most successful marine fish invader to date (Morris and Akins 2009; Green et al. 2011). Lionfish (Family: Scorpaenidae) were first introduced into the waters off southeastern Florida in the late 1980s (Schofield 2010), then spread throughout the US South Atlantic Bight (SAB) in the 1990s, and the Caribbean Sea in the 2000s (Whitfield et al. 2002; Schofield 2010). The Gulf of Mexico (GOM) is the most recently invaded basin, where lionfish were first reported in 2009 off the northern Yucatan Peninsula, Mexico (Aguilar-Perera and Tuz-Sulub 2010), in the Florida Keys, and along the west Florida shelf (Schofield 2010). By late 2010, lionfish had been observed in eastern, northern and western regions of the GOM (Schofield 2010; Fogg et al. 2013; Dahl and Patterson 2014; Nuttall et al. 2014). To date, lionfish have established populations in diverse habitats covering over 7 million km2 across the US southeast Atlantic coast, Caribbean Sea, and portions of the GOM (Schofield 2010; Côté et al. 2013a; Schofield et al. 2014). Throughout their invaded range, lionfish pose a threat to fisheries resources, native fish communities, reef ecosystems, and human health (Morris and Akins 2009, Morris and Whitfield 2009).
Invasive lionfish have been described as generalist mesopredators that consume a broad range of fish and invertebrate prey (Morris and Akins 2009; Muñoz et al. 2011; Dahl and Patterson 2014), including the juvenile stages of ecologically and economically important species (Lesser and Slattery 2011). Their distinction as highly efficient predators has been attributed to unique predatory behaviors and prey naïveté (Fishelson 1997; Hornstra et al. 2004; Albins and Lyons 2012; Cure et al. 2012). These factors, along with fast growth, early maturity, high fecundity, and a lack of native predators have resulted in higher lionfish densities and larger body sizes than are observed in their native range (Darling et al. 2011; Hackerott et al. 2013; Dahl and Patterson 2014; Valdivia et al. 2014).
Invasions by predators are expected to have the most damaging impacts on native ecosystems given that predator–prey dynamics can greatly influence community assemblages in both terrestrial and marine systems (Paine 1966; Hixon and Carr 1997; Grosholz et al. 2000; McDonald et al. 2001; Caut et al. 2008). It is essential to understand how invasive predators and native prey interact in a trophic context to effect change in the structure and function of invaded marine communities (Rilov 2009). Visual diet (i.e., gut content) analysis is conventionally used to examine such trophic relationships (Hyslop 1980), but digestion often results in substantial (>40%) portions of the stomach contents being unidentifiable (Morris and Akins 2009; Dahl and Patterson 2014). Unidentifiable portions of diet represent lost information given that visually identifiable portions may not fully characterize diet. Thus, traditional visual diet analyses may provide incomplete data on trophic relationships between invasive and native species. Furthermore, unidentifiable diet portions are often excluded when describing taxonomic contributions to diet, which can heavily bias results if differential digestion rates cause some species or life stages to become visually unidentifiable more quickly than others (Hyslop 1980; Collis et al. 2002). More complete diet information could also be useful in understanding the transfer and bioaccumulation of natural toxins, such as those associated with ciguatera fish poisoning, in ciguatera-endemic regions of the Caribbean (Robertson et al. 2014).
In the case of invasive lionfish, as much as 70% of prey items observed in their stomachs cannot be visually identified to species with a dissecting microscope (Valdez-Moreno et al. 2012; Côté et al. 2013b; Dahl and Patterson 2014). However, molecular tools can be employed to identify otherwise unidentifiable prey items. For example, mitochondrial DNA (mtDNA) barcoding offers an approach to genetically identify prey based on a target fragment (~650 base pair region) of the cytochrome c oxidase subunit I (COI) gene (Hebert et al. 2003; Ivanova et al. 2007; Valdez-Moreno et al. 2012; Côté et al. 2013b). COI sequences are species-specific, highly conserved, and have high reliability and resolution for species identification in fish, even in partially digested or archived samples (Carreon-Martinez et al. 2011). Valdez-Moreno et al. (2012) employed barcoding to examine invasive lionfish diet in the Mexican Caribbean, while Côté et al. (2013b) employed barcoding to examine lionfish diet diversity (i.e., richness) in The Bahamas. In both of those studies, barcoding greatly increased the power of the diet analysis, thus the ability of researchers to evaluate the trophic ecology and impacts of lionfish in those systems.
We previously reported results from visual analysis of lionfish (n = 934) stomach contents sampled in the northern GOM (nGOM), a region with lionfish densities among the highest in the western Atlantic (Dahl and Patterson 2014; Dahl et al. 2016). Despite the relatively large sample size of that study, the potential exists that diet estimates were incomplete or biased given 43% by mass of observed prey could not be identified visually due to digestion (Dahl and Patterson 2014). In the current study, we applied DNA barcoding to identify previously unidentifiable fish prey items reported by Dahl and Patterson (2014). Specific objectives of this work were to (1) evaluate the effectiveness of DNA barcoding to identify visually unidentifiable lionfish prey, and (2) determine whether lionfish diet estimates are enhanced when DNA-barcoded prey items are included in diet analysis.
Methods
Study location and specimen collection
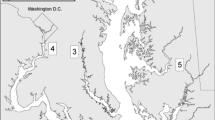
Lionfish sampling was performed seasonally from April 2013 through March 2014 on nGOM natural and artificial reefs ranging in depth from 24 to 35 m (Fig. 1). Individuals were captured by divers via spearing immediately posterior to the spinal column and were placed in a saltwater ice slurry upon surfacing. Speared lionfish were ranked by size and randomly sampled to retain every nth fish so that approximately 100 lionfish per habitat type per season were sampled for diet analyses. Each lionfish was weighed (nearest 0.1 g) and measured (nearest mm total length, TL). All applicable institutional and/or national guidelines for the care and use of animals were followed during the course of this study.
Visual gut content analysis
Red lionfish (Pterois volitans) stomachs were extracted from fish after inspection of esophagus and gills for regurgitated prey. Stomach contents of each sampled lionfish were fixed in 100% (200 proof) ethanol in plastic bags. Preserved prey items were visually identified to the lowest taxonomic level possible, counted, and dried for at least 48 h at 70 °C to obtain dry mass (Hoese and Moore 1998; Kells and Carpenter 2011). Visual identification was performed either with the naked eye or under an Olympus SZX12 dissecting microscope. Identifiable prey taxa were grouped into seven categories: shrimps, crabs, other benthic invertebrates, pelagic invertebrates, reef fishes, non-reef benthic fishes, and pelagic fishes, while unidentifiable prey items were grouped into fish and invertebrates. To assess dietary contribution of each visually identified prey category, percent mass (%M) was computed by dividing the dry mass of each prey taxa by the total dry mass of all prey from each individual, including visually unidentified prey. Diet was assessed by habitat type (natural versus artificial reefs) and lionfish size class (small: <200 mm TL; medium: 200–250 mm TL; large: >250 mm TL).
DNA barcoding preparation and analysis
Any prey item that was identified as a fish but could not be identified to a taxonomic level lower than family was considered unidentifiable and retained for DNA barcoding analysis. These prey items were categorized into two digestion stages, whole fish (i.e., most of carcass was present) or fish remains, and weighed wet to the nearest 0.01 g. Samples were processed for barcoding by first removing any external layer of tissue that had been in contact with the lionfish sample’s stomach wall and/or fluids with sterile scalpels and forceps, and then a small (approximately 1 mm3, 15–25 mg) plug of muscle tissue was excised from each unidentifiable prey item. Tissue samples were blotted dry and placed in a 1.5 ml sterile microcentrifuge tube. To prevent cross-contamination between tissue extractions, tools were rinsed with 70% ethanol and then flame sterilized. Tissue plugs were frozen at −80 °C until DNA extraction.
DNA extractions were performed following the optimized standardized protocols of Handy et al. (2011) with DNeasy Blood & Tissue Kits (Qiagen, CA). Tissue was placed in a solution of lysis buffer and proteinase K and homogenized with sterile disposable polypropylene pestles (Thermo Fisher Scientific, MA), and then digested for 1–2 h at 56 °C until lysed. DNA extractions were quantified on a Nanodrop 1000 spectrophotometer (Thermo Fisher Scientific, MA) for DNA concentration and purity (i.e., ratio of absorbance at 260 and 280 nm) and then visualized on 1% agarose gels to verify these estimates. Samples with high DNA yield (>100 ng/ul) were diluted 10X with ultrapure water.
Amplification of the 650 bp barcode region of the COI gene was performed with universal M13 tailed fish primer cocktails C_FishF1t1and C_FishR1t1 developed by Ivanova et al. (2007) for barcoding (Table S1). The 20 µl polymerase chain reaction (PCR) mixes included 10 µl of 10% trehalose, 4.4 µl ultrapure water, 2 µl 10X PCR buffer, 1 µl 50 mM MgCl2, 0.2 µl each 10 µM primer cocktail, and 0.1 µl each of 10 mM dNTPs and Platinum® Taq polymerase (Thermo Fisher Scientific, MA) and 1.6 µl DNA template. DNA template concentration ranged from approximately 1–40 ng/µl. Thermal conditions for COI PCR amplification included initial denaturation at 95 °C for 2 min, followed by 35 cycles of 94 °C for 40 s, 53.5 °C for 40 s, and 72 °C for 1 min, with a final extension at 72 °C for 5 min and a hold at 4 °C. Positive and negative controls were used for each 96-well test-plate of amplification reactions to test for PCR reaction quality and contamination in reagents. Template DNA was replaced with ultrapure water in negative controls, and template DNA extracted from Pterois volitans muscle tissue was utilized for positive controls.
PCR products were visualized on 1% agarose gels and positive reactions were identified by a clear band aligned to the 650 bp fragment of the TrackIt™ 1 Kb Plus DNA ladder (Thermo Fisher Scientific, MA). All positive PCR products, along with positive and negative controls, were sent to Beckman Coulter Genomics (Danvers, MA) where they underwent PCR cleanup and bidirectional sequencing using M13F and M13R primers (Online Resource 1).
Barcoding sequence analysis
The COI barcode sequences of unidentified prey items were analyzed to assess length and quality using Geneious (version 8.1.6; Kearse et al. 2012). Forward and reverse sequences were trimmed to remove ambiguous and/or low quality bases and remnant primer from amplification or sequencing reactions and then aligned using ClustalW to produce consensus sequences. Sequences were then visualized and edited in Geneious to resolve less reliable base calls. Final contigs were ranked by length as long (>500 bp), medium (300–500 bp) and short (<300 bp). Long and medium ranked sequences were submitted to nucleotide BLAST searches through the NCBI GenBank database to classify unidentified prey item sequences to the closest match in the COI database. BLAST searches returned the closest ten sequences in the reference databases based on values of sequence similarity (i.e., % identity), E-value (i.e., number of expected hits by chance) and grade (i.e., percentage combining % identity, E-value and query coverage). The species-level identification function of the Barcode of Life Database was used to resolve any potential database gaps in GenBank (BOLD, www.boldsystems.org; Ratnasingham and Hebert 2007). Collection and sequence data are available from GenBank and Gulf of Mexico Research Initiative Information and Data Cooperative (http://www.ncbi.nlm.nih.gov/; Dahl and Patterson 2017).
We adopted three criteria to establish accurate fish identification from BLAST results. A species match was identified for long sequences with >99% grade. A genus level match was identified for long sequences with a 95–99% grade. Finally, a family level match was identified by either long sequences with <95% grade or medium sequences with >95% grade. Sequences with genus and family level identifications from BLAST were also submitted into the BOLD ID function which assigned species level identification based on <1% sequence divergence and genus level identification based on <3% divergence with a reference sequence. Classification to genus and/or family level was necessary in some cases due to incomplete taxonomic coverage of GOM fishes in the reference databases. Sequences of <300 bp were not included in further analyses. These criteria for sequence inclusion were considered to be conservative given recent studies have utilized COI sequences as short as 100 bp if they had a high (90%) probability of being accurately assigned to species level (Meusnier et al. 2008; Valentini et al. 2009).
To improve sequence quality or verify the accuracy of the extraction and barcoding process, replicates were performed by extracting new tissue plugs and repeating the entire DNA barcoding process. This was performed for all samples that had poor sequence quality from the original sequencing and also had sufficient tissue mass remaining. Replicates were also randomly chosen among a subset of samples that had high initial quality, as well as among those that returned species of interest (e.g., new or exploited species) as the closest COI match.
Incorporating barcode information into diet
To incorporate new species identified by barcoding into existing %M estimates of lionfish diet, wet masses of unidentified prey items were converted to simulated original dry masses using the digestion stage (i.e., whole fish or fish remains) and a wet-dry mass conversion factor from visually identified fish prey. Prey identified by barcoding were then sorted into the original prey categories for further analysis (Dahl and Patterson 2017). The proportion of visually unidentifiable fish prey that was not able to be resolved through barcoding remained in the unidentifiable fish category.
Sample-based prey taxa accumulation curves were computed in Primer (v6; Clarke and Gorley 2006) to assess diet richness in nGOM lionfish as a function of sample size, and whether estimates of prey diversity changed when DNA barcoded prey were added. The order in which samples were successively incorporated was randomly resampled by permutation (999 permutations) to satisfy the assumption of random sample order. Given the number of individuals (n = 934) was equal between methods, the shapes of the curves (i.e., rate of species accumulation, asymptote reached) and 95% confidence intervals (i.e., overlapping or not) could be compared between methods to assess whether DNA barcoding added significant information to diet estimates (Colwell et al. 2004). Curves were computed for lowest taxonomic level of unique prey taxa as well as higher order identification (i.e., fish family and invertebrate infraorder) to examine trends in diet diversity.
Results
Lionfish (n = 934) examined in this study ranged in size from 67 to 377 mm TL and were collected from natural and artificial reefs over the course of one year (mean n = 117 fish per treatment combination). With traditional visual diet identification, 1485 (64.8% by number; 57% by mass) prey items were identified in lionfish stomach samples, while 807 (35.2% by number; 43% by mass) were not visually identifiable. Unidentified fish prey items (n = 696) were processed for barcoding as described above, but unidentified invertebrate prey items (n = 73 items) were not barcoded.
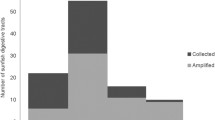
DNA extracts from the 696 visually unidentified fish prey items yielded 575 (82.6%) readable sequences that could be identified to species, genus or family levels. Mean length (±SD) of barcode sequences identified to species was 627 ± 26 bp (Online Resource 2). Mean length of barcode sequences identified to genus and family level were 622 ± 24 and 455 ± 115 bp, respectively. Among readable COI sequences, 444 (77.2%) were positively identified to species level (63.8% of total visually unidentified fish prey items). The remaining 131 sequences were identified to either genus (n = 73) or family level (n = 58) based our classification criteria. With the addition of barcoding the number of identifiable fish prey nearly doubled (932 vs. 499 prey items), resulting in 78.0% of total fish prey being identified. Whole fish prey had a slightly higher probability (85%) of being identified using barcoding than more highly digested fish remains (79%). As many as five unique species were identified by DNA barcoding from a single lionfish stomach. For all test plates, positive controls produced a COI barcode with a correct species match. Negative controls did not produce any COI barcodes, indicating satisfactory quality control across all test plates.
Dahl and Patterson (2014) previously reported 11 species, 3 genera, 16 families and 8 orders or infraorders among the visually-identified invertebrate prey, along with 18 fish prey species from 15 families (Table 1). Prey diversity increased greatly when DNA barcoding was incorporated into prey identification, including 24 species, 19 genera, and 9 families not previously identified with visual methods (Table 1). Of these, 30 are novel prey taxa unreported from other lionfish diet studies (Table 1). Notable species found in lionfish diet for the first time via DNA barcoding were red snapper (Lutjanus campechanus), wenchman (Pristipomoides aquilonaris), roughtongue bass (Pronotogrammus martinicensis) and red porgy (Pagrus pagrus), which are all ecologically or economically important reef fishes in the region. Potential cannibalism also was revealed by DNA barcoding, with red lionfish being the most frequently identified prey taxon (n = 100 of 696 or 14.4%) via barcoding. Among red lionfish sequences in the GenBank database, our sequences matched voucher specimens from the western and southern Caribbean, and Brazil (Accession Numbers: KJ739816, KM488633, and KP641132, respectively) most closely (>99.7% similarity). Prey items barcoded as red lionfish occurred in lionfish of all size classes, but was found to be most prevalent in medium to large fish size categories.
A subset of prey samples (n = 150) was selected for duplicate DNA barcoding of new dissections of prey tissue. Samples were chosen to represent a range of species and tissue quality levels, and where further species verification was deemed essential (e.g., red lionfish, exploited species) (Online Resource 3). Duplication of barcoding for the 39 initially short sequences (<300 bp) and 72 initially poor sequences (<100 bp) resulted in 67 sequences >300 bp in length that were sufficient for inclusion in further analyses (Online Resource 4). Out of 39 duplicates run from initially short sequences, 30 resulted in similar or longer length sequences than initial runs (Online Resource 3). For these samples, all but one returned identical matches from reference databases, and this sample was excluded from further analyses (Online Resource 3, 4). Species identity was confirmed for duplicate analysis of the prey samples that originally were barcoded red snapper, as was true for 15 other species of interest (Online Resource 3, 4).
Species identification was confirmed for the 16 duplicates of samples that originally were barcoded as Pterois volitans (Online Resource 3,4). It should be noted that while cannibalism was not documented visually among the samples collected in this study, Dahl and Patterson (2014) visually identified 28 partially digested individual prey as family: Scorpaenidae. Furthermore, subsequent stomach content analysis of lionfish sampled in the nGOM has produced clearly identifiable juvenile P. volitans in the stomachs of larger conspecifics; however, such observations have been infrequent (Fig. 5).
Differences in diet among size classes, habitats and seasons were apparent from visually identified prey, with piscivory increasing with fish size. Diet breadth was greatest on artificial reefs due in part to consumption of non-reef associated fishes, and invertebrates contributing increasingly to diet in winter (Fig. 2a). Dahl and Patterson (2014) originally characterized nGOM lionfish diet based on visually identified prey only, thus excluding unidentified prey. When visually unidentified prey are included in estimates of diet percentages by prey category, changes in mean %M were apparent, especially with respect to contributions of invertebrate prey relative to fish prey (Fig. 2b). Percent mass contribution of invertebrates to lionfish diet considering only visually-identified prey was 6.8% on natural reefs and 17.0% on artificial reefs. With the addition of barcoded fish prey these contributions declined to 4.5 and 8.0%, respectively (Fig. 2c). Prey categorized as unidentified fishes (Fig. 2b) was reduced from >40% of diet by mass to ~7% after DNA barcoding (Fig. 2c; Table 1). The importance of fishes to the diet of lionfish was more pronounced in lionfish collected from natural reef habitat; however, the trend of increased foraging on invertebrates in winter was observed in both habitats (Fig. 2c; Table 1). Prey identified as reef-associated fish was greatest in spring and summer seasons on artificial reefs, due to large numbers of vermilion snapper (~16% diet by mass) identified via barcoding (Fig. 2c; Table 1). An ontogenetic shift toward piscivory with increasing lionfish size remained apparent with the addition of barcoded prey, where juvenile lionfish consume more invertebrates by mass across all seasons (Fig. 2c).
Stacked bar plots of mean percent red lionfish (P. volitans) diet by mass for prey categories observed across habitat types, seasons, and size class for: a visually identified prey items, excluding unidentifiable prey; b visually identified prey items, including unidentifiable prey; and c prey items identified visually and with DNA barcoding. Size categories: S ≤ 200 mm total length (TL), M = 200–250 mm TL, and L ≥ 250 mm TL and habitat types: natural = natural reefs, artificial = artificial reefs
Prey exhibited a wide range in estimated dry mass, from 0.01 to 11.20 g, with a mean (±SD) of 0.72 ± 1.1 g (Fig. 3). Mean (±SD) mass (0.71 ± 1.46 g) of fish prey that remained unidentified following barcoding was similar to the mean of the overall distribution. Masses of prey identified to family were larger on average (mean ± SD = 1.0 ± 1.59 g) than of either species- or genus-level (mean ± SD = 0.68 ± 0.84 and 0.64 ± 0.57 g, respectively) identification. High prevalence of relatively large jawfishes (family: Opistognathidae) and seabasses (family: Serranidae) that could be identified only to family with barcoding explains this trend. The size of fish prey that were barcoded as lionfish (i.e., potential cannibalism) was much smaller than the majority of the samples that underwent barcoding. Half of the prey samples identified by barcoding as P. volitans had a mass <0.25 g and >90% were under 1 g. Vermilion snapper were similarly small in size, with nearly 80% of samples identified via barcoding being under 1 g in mass.
The number of unique prey taxa contributing to visually-identified lionfish diet accumulated rapidly with increasing numbers of lionfish sampled (Fig. 4a). At a sample size of 100 individuals, already thirty prey taxa were observed, which continued to increase through all 934 individuals, although the curve begins to reach an asymptote after approximately 800 samples. When taxa identified with DNA barcoding were included, that curve did not reach an asymptote (Fig. 4a). At 100 individuals sampled with DNA barcoding, the number of prey taxa increased to forty-seven unique taxa (Fig. 4a). Overall, this represents an addition of 36 unique prey taxa via DNA barcoding. Prey richness is significantly different with the addition of barcoded fish prey given that 95% CIs do not overlap the curve for visually-identified prey alone. When curves were fit only to fish family and invertebrate infraorder, prey richness was significantly higher when DNA barcoding results were included (Fig. 4b).
Species accumulation curves of red lionfish (P. volitans) prey taxa identified during this study. a Cumulative number of unique prey identified to species or lowest taxonomic level of identification, and b fish families and invertebrate infraorders, as a function of number of lionfish sampled and analyzed with visual gut content analysis (solid line) or visual identification plus DNA barcoding (dashed line). Every 5th 95% confidence interval is plotted to allow comparison of curves between visual identification and DNA barcoding methods
Discussion
DNA barcoding greatly enhanced the ability to describe lionfish diet, and together with visual identification this study reveals the richest diversity of prey reported to date for this invasive species. At the family level alone, these diet data nearly double the highest estimates of prey diversity reported for lionfish (Morris and Akins 2009; Valdez-Moreno et al. 2012; Côté et al. 2013b). Including visually identified prey, we recorded a total of 41 fish species, 37 genera, and 24 families among lionfish prey. DNA barcoding revealed lionfish diet in the nGOM to be far broader than previously reported from traditional visual identification results where we were able to identify 24 species, 19 genera and nine families of fishes that were previously undetected by visual gut content analysis (Dahl and Patterson 2014).
Our estimates of prey diversity remain high even when compared to other studies in which DNA barcoding was utilized to describe lionfish diet (Valdez-Moreno et al. 2012; Côté et al. 2013b). Valdez-Moreno et al. (2012) reported fish prey from 14 families, 22 genera and 34 species, while Côté et al. (2013b) reported fish prey from 16 families, 27 genera and 37 species. While species richness of prey was comparable among all studies, the diversity of prey constituting lionfish diet at the level of family and genus were far greater in the current study. Our study was conducted outside of the Caribbean, thus prey communities lionfish encountered were inherently different than where lionfish diet was previously explored with DNA barcoding. Côté et al. (2013b) estimated the occurrence of 90 potential prey fish species that were small enough (<13 cm) for lionfish to ingest in the region of The Bahamas in which they conducted their barcoding diet study. In our study region, Patterson et al. (2014) reported 91 reef fishes present on nGOM natural and artificial reefs, but <70 of these species had some life stage present on reefs which would meet Côté et al.’s (2013b) 13-cm threshold. Therefore, the higher diversity of lionfish prey observed in the current study occurred despite lower diversity of potential reef fish prey.
Potential cannibalism
DNA barcoding not only increased our ability to characterize lionfish diet, but also revealed potential cannibalism in the nGOM, a relatively recently invaded region. While we can definitely say that lionfish mitochondrial DNA was associated with tissue samples isolated from these prey samples in lionfish stomachs, unequivocal evidence of lionfish cannibalism does not exist just because some prey samples were barcoded as lionfish. There are three possible explanations for lionfish identifications via DNA barcoding: (1) prey tissue may have been contaminated with DNA from the consumer lionfish during initial dissections; (2) prey DNA was degraded due to digestion to the point that the only viable DNA present and amplified was that from the consumer lionfish; or, (3) lionfish DNA was present in prey tissue because the prey was a cannibalized lionfish.
A rigorous sample processing protocol was put in place to guard against contaminating prey tissue samples with DNA from consumer lionfish, but contamination cannot be completely discounted as the source of lionfish DNA in barcoded samples. Great effort was expended to ensure sterile techniques, perform positive and negative barcoding controls, and extract tissue from prey items after removing surface layers of tissue. Despite these techniques, it remains a possibility that DNA from the stomach lumen of consumers could have been shed onto prey fragments and not removed prior to PCR amplification. However, the mass of potentially contaminating consumer lionfish stomach tissue, cells, or DNA would have been orders of magnitude lower than the mass, hence DNA, of prey tissue sampled for barcoding. Therefore, it seems unlikely that contamination alone could have resulted in the degree to which lionfish DNA was present in barcoded prey samples, especially given the techniques utilized in this study.
Amplification of a species’ DNA via PCR is not only a function of the abundance of that species’ DNA versus other DNA present in a given sample, but also the quality of the DNA present (Gonzalez et al. 2012). While the mass of any consumer lionfish DNA present in prey tissue dissected for DNA amplification was likely orders of magnitude lower than the mass of prey DNA, it is possible that highly digested prey items had DNA of such poor quality that most of the DNA capable of being amplified was that of the consumer lionfish. Clearly, the likelihood of this phenomenon occurring would increase with more advanced states of digestion. In future barcoding studies, qualitative scoring of the degree of prey digestion would be one way to test whether the incidence of apparent cannibalism increases with increasing stage of prey digestion.
The final possibility for why lionfish mitochondrial DNA was so pervasive among barcoded prey samples is that at least some of those samples were in fact cannibalized lionfish. The likelihood of that being the case might be somewhat diminished by the lack of direct observation of cannibalized lionfish for samples examined here. However, we have observed evidence of lionfish cannibalism in subsequent sampling of lionfish stomach contents (see Fig. 5), and several small (<5 cm) prey items were reported as Family:Scorpaenidae when lionfish prey items were originally identified visually (Dahl and Patterson 2014). One potential reason for the lack of visual evidence for cannibalism in the current study could be the size of lionfish being consumed by larger conspecifics. Prey samples that were barcoded as lionfish were smaller than the majority of prey samples identified with barcoding. This would suggest that any cannibalism that did occur resulted primarily from larger adults cannibalizing small, recently settled juveniles, which is similar to the pattern reported by Valdez-Moreno et al. (2012). Conversely, these results could also be explained by tissue of small individuals of other species being digested so quickly that consumer lionfish DNA amplified rather than degraded DNA of non-lionfish prey.
Examples of prey fish visually identified as P. volitans from lionfish stomach contents. a A red lionfish prey item (30 mm in length) visually identified from the stomach contents of a red lionfish measuring 127 mm TL, collected from an artificial reef south of Pensacola, FL, and b a red lionfish prey item (37 mm in length) visually identified from the stomach contents of a red lionfish measuring 218 mm TL, collected from an artificial reef south of Mexico Beach, FL
Very little is known about lionfish larval and early juvenile stages, particularly the size at which lionfish settle onto reefs following their planktonic larval stage (Ahrenholz and Morris 2011). The late post-flexion larval stage for the species is reported to be between 9.5 and 11 mm standard length (SL) (Imamura and Yabe 1996), and a newly settled (i.e., not fully pigmented) juvenile collected in the nGOM was 15 mm SL (Byron et al. 2014). This is similar to the most often encountered size range for prey samples that barcoded as P. volitans in this study. The next largest component of diet revealed by DNA barcoding was vermilion snapper, and these prey items were also most likely newly settled recruits given their relatively small size. Early juvenile prey fishes lose most or all identifiable characters rapidly (<60 min) after ingestion by other fishes (Schooley et al. 2008; Legler et al. 2010). Therefore, predation on these early life stages is very difficult to detect visually and may be missed completely without molecular techniques.
While highly useful in identifying species, DNA barcoding alone cannot conclusively detect cannibalism within a species. Since predator specimens were not subjected to DNA barcoding, we cannot directly compare lionfish prey sequence haplotypes to those of their consumer. However, even if this had been done, the COI haplotype diversity in nGOM lionfish populations is likely too low to detect differences between predator and prey (Hamner et al. 2007; Betancur-R et al. 2011; Toledo-Hernández et al. 2014). As stated above, one of the possible explanations for lionfish mitochondrial DNA associated with prey tissue samples is contamination by DNA from consumers, which then amplified during PCR. Molecular blocking can be used to prevent amplification of consumer (or otherwise erroneous) DNA (De Barba et al. 2014), but was not used here.
Prey samples from 89 adult lionfish barcoded as P. volitans, which occurred across small, medium, and large size classes from both natural and artificial reefs. The frequency of this potential cannibalism increased from spring to winter and occurred most frequently in larger adult lionfish. The highest incidence of P. volitans prey barcodes were observed in winter when lionfish were at their highest densities and their largest mean size (Dahl and Patterson 2014). If cannibalism was in fact occurring, these results would suggest size-structured and density-dependent cannibalism in nGOM lionfish. Intraspecific competition for prey resources has most likely increased with the rapid population growth observed in invasive lionfish in the nGOM, with all size classes competing for the same pool of resources and space on reefs. On nGOM reefs, fish diversity declines during winter (Dance et al. 2011), which corresponds to a higher proportion of invertebrates in lionfish diets (Dahl and Patterson 2014). Therefore, the higher incidence of potential cannibalism during winter may have reflected lower availability of preferred prey (Rudolf 2008). However, any inferences drawn about potential cannibalism at this stage cannot be substantiated given uncertainties about the likelihood of contamination of prey samples with consumer lionfish DNA. Genotyping of consumer lionfish and prey samples that barcoded as lionfish could provide an unequivocal test of whether cannibalism is occurring in the nGOM, and could also be employed to quantify the extent to which it occurs (Koskinen and Piironen 2000; Guichoux et al. 2011).
Trends in diet with unidentified fish resolved
Including prey taxa that were identified via DNA barcoding among lionfish stomach samples clearly expands our knowledge of invasive lionfish diet in the nGOM. However, re-examining visually-identified prey taxa previously reported highlighted an issue with how the data were originally presented. In Dahl and Patterson (2014), percent diet composition by mass was only reported for taxa that had been visually identified. However, unidentified fish prey constituted a larger percentage of the diet than did unidentified invertebrate prey (by mass, 43% to <1%). Therefore, by only considering visually identifiable prey when computing diet composition, the importance of fish, in general, to lionfish diet was understated. While lionfish clearly are generalist mesopredators, they are more piscivorous in the nGOM than was previously reported (see Fig. 2).
With the inclusion of DNA barcoding results, general conclusions reported earlier about lionfish diet still hold, but some differences are apparent. Again, the overall breadth of lionfish diet in the nGOM is substantially higher than previously reported. Reef-associated fish prey contributed >10% more (by mass) to lionfish diet on artificial reefs, which is mostly attributable to juvenile snappers (e.g., vermilion snapper, red snapper, wenchman) and potential cannibalism on conspecific juveniles. Reef-associated fish prey on natural reefs declined nearly 10% with the addition of non reef-associated fishes such as flounders (family: Bothidae), jawfishes (family: Opistognathidae), and lizardfishes (family: Synodontidae). Contribution of pelagic fishes to the diet remained similar for lionfish captured on natural reefs, but declined by nearly 5% on artificial reefs. These differences taken together do not affect the overall trends that lionfish on natural reefs predominantly consume reef-associated fishes, and that lionfish on artificial reefs have broader diets with higher contributions from non reef-associated and pelagic fishes.
Several exploited fish species were identified in invasive lionfish diet via DNA barcoding, which may be of particular interest to resource managers. Vermilion snapper (Rhomboplites aurorubens) was the second most commonly barcoded prey species with 98 prey items from 61 adult lionfish, and constituted nearly 16% of lionfish diet by mass on artificial reefs across the study. There were 24 lionfish that had multiple juvenile vermilion snapper identified with barcoding, and in 7 cases more than three vermilion snapper juveniles were found in the stomach of an individual lionfish. Red snapper and red porgy are other notable fishery species found in lionfish diet for the first time via DNA barcoding. Predation on juvenile red snapper by lionfish was not expected given red snapper typically do not recruit to reef habitat until their second year of life when they are >250 mm TL (Patterson et al. 2005; Workman et al. 2002). However, precocious recruitment to adult habitat is sometimes observed (Bailey et al. 2001).
DNA barcoding efficacy
DNA barcoding was highly effective in identifying lionfish fish prey that were not identifiable visually. Our optimized DNA barcoding methodology yielded >80% efficiency for producing reads with high enough quality for prey taxon identification. Combining DNA barcoding with traditional visual identification methods resulted in identification of >90% of all lionfish prey by mass. Furthermore, the addition of fish prey identified with barcoding resulted in approximately 250% more fish species identified in diets relative to visual diet analysis. Digestion level may have contributed to DNA barcoding success, where lightly digested whole prey accounted for approximately 60% of molecular identifications and more highly digested prey only accounted for approximately 40%. Validation studies on the reliability of COI as a taxonomic tool are numerous across a diversity of taxa (Hebert et al. 2003; Clare et al. 2007; Dawnay et al. 2007) including fishes (Ward et al. 2005) and reproducibility of COI barcodes based on novel DNA extractions and amplification from prey samples was high in this study.
Higher-level taxonomic identifications (i.e., genus and family level) from barcoding were necessitated in part due to poor sequence quality but also incomplete taxonomic coverage for COI in reference databases. For example, we visually identified purple reeffish (Chromis scotti) in the stomachs of lionfish (Dahl and Patterson 2014), but it was not possible to obtain a species level match with DNA barcoding in the GenBank or BOLD databases. Sequences from 24 prey specimens most closely matched Chromis scotti, however, a verified barcode does not currently exist for the species given that fewer than three voucher sequences have been submitted to BOLD (Ratnasingham and Hebert 2007). Thus, these individuals were classified as Chromis sp. It is highly likely that these prey were actually C. scotti given high sequence quality and complete taxonomic coverage of remaining Chromis spp. that occur in the nGOM. Similar issues with reference database gaps occurred for genera within the families Apogonidae, Gobiidae, Labridae, Opistognathidae, Serranidae, and Triglidae. For example, the presence of a cryptic species of cardinalfish (Astrapogon sp.) in the nGOM is inferred by the lack of a species match from taxonomically complete databases for this genus, which was also reported by Valdez-Moreno et al. (2012). Ultimately, this study highlights that gaps exist in COI databases with respect to nGOM reef fishes. Global barcoding initiatives (e.g. FISH-BOL, CBOL) that aim to obtain COI records of all fishes may result in more nGOM fishes being barcoded in the near future (Ward et al. 2009).
DNA barcoding is often used to study diet in fishes with sample size limitations as a result of rare occurrence or logistical challenges in sampling. However, this method can be applied more broadly if enough information is gained by its use. Our study demonstrates that even for a diet study with a relatively large sample size, for which visual diet analysis might be expected to have detected most prey taxa, DNA barcoding added a significant amount of new dietary information. This was clearly evident from prey accumulation curves comparing lionfish prey richness between visual identification and barcoding methodologies. Using visual identification, 30 prey taxa were observed at a sample size of 100, which is comparable to the richness that Côté et al. (2013b) observed at a similar sample size using DNA barcoding in The Bahamas. In contrast, we found 47 unique taxa at a sample size of 100 individuals combining visual identification with DNA barcoding. Despite the high proportion of prey identified with DNA barcoding, an asymptote in prey richness was not reached in this study. The slope of the curve for visually identified prey noticeably flattened after sampling 400 individuals, but no obvious asymptote was reached. This is a notable finding given the large sample size (nearly 1000 fish) and prior studies indicating diet saturation at ~700 individuals (Morris and Akins 2009).
Lionfish prey diversity appears to be higher in the nGOM region than in other regions, which was not evident from visual diet analysis alone. The broad diet of invasive lionfish diet in the nGOM supports the inference that lionfish are generalists that will opportunistically prey on small demersal or benthic species or early life stages, suites of which differ between regions of the invaded range (Morris and Akins 2009; Green et al. 2011; Muñoz et al. 2011; Green and Côté 2014). The diversity of prey identified within lionfish stomachs in the nGOM indicates they are a substantial threat to numerous taxa in this region. As lionfish population size, mean body size, and biomass continue to increase on both natural and artificial reefs these interactions could only be expected to intensify (Dahl et al. 2016; Dahl et al. unpublished data).
Our ability to determine with high resolution exactly what lionfish are eating on nGOM reefs will allow for more accurate assessment of direct lionfish impacts on native reef fish communities of the nGOM and shed light on potential indirect impacts to other species. Of particular importance is the detection of several regionally exploited reef fish species as lionfish prey, which has important implications for fishery managers. The diversity of non-reef and reef fishes alike in lionfish stomach samples may portend population declines due to increasing lionfish predation. Furthermore, lionfish predation also may indirectly impact reef fishes that also depend on those species for food. Lionfish feeding ecology data at regional scales are important inputs for ecosystem modeling efforts (e.g., Chagaris et al. 2015; Chagaris et al. in press). Such efforts are critical to understanding current and predicted future direct and indirect impacts of lionfish in the nGOM ecosystem, as well as for simulating potential management strategies to mitigate these impacts.
Change history
03 July 2018
In the original publication, the Acknowledgements section has been published incorrectly. The corrected Acknowledgements is given in this correction:
References
Aguilar-Perera A, Tuz-Sulub A (2010) Non-native, invasive red lionfish (Pterois volitans [Linnaeus, 1758]: Scorpaenidae), is first recorded in the southern Gulf of Mexico, off the northern Yucatan Peninsula, Mexico. Aquat Invasions 5:S9–S12
Ahrenholz DW, Morris JA Jr (2011) Larval duration of the lionfish, Pterois volitans along the Bahamian Archipelago. Environ Biol Fish 88(4):305–309
Albins MA, Lyons PJ (2012) Invasive red lionfish Pterois volitans blow directed jets of water at prey fish. Mar Ecol Prog Ser 448:1–5
Bailey HK, Cowan JH Jr, Shipp RL (2001) Experimental evaluation of potential effects of habitat size and presence of conspecifics on habitat association by young-of-the-year red snapper. Gulf Mexico Sci 19:119–131
Betancur-R R, Hines A, Acero AP, Orti G, Wilbur AE, Freshwater DW (2011) Reconstructing the lionfish invasion: insights into Greater Caribbean biogeography. J Biogeog 38(7):1281–1293
Byron D, Heck KL Jr, Kennedy MA (2014) Presence of juvenile lionfish in a northern Gulf of Mexico nursery habitat. Gulf Mexico Sci 32:75–77
Carreon-Martinez L, Johnson TB, Ludsin SA, Heath DD (2011) Utilization of stomach content DNA to determine diet diversity in piscivorous fishes. J Fish Biol 78(4):1170–1182
Caut S, Angulo E, Courchamp F (2008) Dietary shift of an invasive predator: rats, seabirds and sea turtles. J Appl Ecol 45:428–437. doi:10.1111/j.1365-2664.2007.01438.x
Chagaris D, Binion S, Bodanoff A, Dahl K, Granneman J, Harris H, Mohan J, Rudd M, Swenarton M, Ahrens, R, Patterson W, Morris J, Allen M An ecosystem-based approach to evaluating impacts and management of invasive lionfish. AFS Fish (in press)
Chagaris D, Binion S, Bodanoff A, Dahl K, Granneman J, Harris H, Mohan J, Rudd M, Swenarton M, Ahrens R, Allen M, Morris J, Patterson W (2015) Modeling lionfish management strategies on the West Florida Shelf: workshop summary and results. University of Florida, Gainesville, p 31
Clare EL, Lim BK, Engstrom MD, Eger JL, Hebert PD (2007) DNA barcoding of Neotropical bats: species identification and discovery within Guyana. Mol Ecol Notes 7(2):184–190
Clarke KR, Gorley RN (2006) PRIMER version 6: user manual/tutorial. PRIMER-E, Plymouth
Collis K, Roby DD, Craig DP, Adamany S, Adkins JY, Lyons DE (2002) Colony size and diet composition of piscivorous waterbirds on the lower Columbia River: implications for losses of juvenile salmonids to avian predation. T Am Fish Soc 131(3):537–550
Colwell RK, Mao CX, Chang J (2004) Interpolating, extrapolating, and comparing incidence-based species accumulation curves. Ecology 85(10):2717–2727
Côté IM, Green SJ, Hixon MA (2013a) Predatory fish invaders: insights from Indo-Pacific lionfish in the western Atlantic and Caribbean. Biol Conserv 164:50–61
Côté IM, Green SJ, Morris JA Jr, Akins JL, Steinke D (2013b) Diet richness of invasive Indo-Pacific lionfish revealed by DNA barcoding. Mar Ecol Prog Ser 472:249–256
Cure K, Benkwitt CE, Kindinger TL, Pickering EA, Pusack TJ, McIlwain JL, Hixon MA (2012) Comparative behavior of red lionfish Pterois volitans on native Pacific versus invaded Atlantic coral reefs. Mar Ecol Prog Ser 467:181–192
Dahl KA, Patterson WF III (2014) Habitat-specific density and diet of rapidly expanding invasive Red Lionfish, Pterois volitans, populations in the northern Gulf of Mexico. PLoS ONE 9(8):e105852
Dahl KA, Patterson WF III (2017) DNA barcoding of invasive lionfish diet in the northern Gulf of Mexico. Dataset distributed by: Gulf of Mexico research initiative information and data cooperative (GRIID-C). University of Florida. Available from: UDI: R4.x267.000:0026
Dahl KA, Patterson WF III, Snyder RA (2016) Experimental assessment of lionfish removals to mitigate reef fish community shifts on northern Gulf of Mexico artificial reefs. Mar Ecol Prog Ser 558:207–221
Dance MA, Patterson WF III, Addis DT (2011) Fish community and trophic structure of reef fishes in the northeastern Gulf of Mexico. B Mar Sci 87:301–324
Darling ES, Green SJ, O’Leary JK, Côté IM (2011) Indo-Pacific lionfish are larger and more abundant on invaded reefs: a comparison of Kenyan and Bahamian lionfish populations. Biol Invasions 13:2045–2051
Dawnay N, Ogden R, McEwing R, Carvalho GR, Thorpe RS (2007) Validation of the barcoding gene COI for use in forensic genetic species identification. Forensic Sci Int 173(1):1–6
De Barba M, Miquel C, Boyer F, Mercier C, Rioux D, Coissac E, Taberlet P (2014) DNA metabarcoding multiplexing and validation of data accuracy for diet assessment: application to omnivorous diet. Mol Ecol Res 14(2):306–323
Fishelson L (1997) Experiments and observations on food consumption, growth and starvation in Dendrochirus brachypterus and Pterois volitans (Pteroinae Scorpaenidae). Environ Biol Fish 50(4):391–403
Fogg AQ, Hoffmayer ER, Driggers WB III, Campbell MD, Pellegrin GJ, Stein W (2013) Distribution and length frequency of invasive lionfish (Pterois sp.) in the northern Gulf of Mexico. Gulf Caribb Res 25:111–115
Gonzalez JM, Portillo MC, Belda-Ferre P, Mira A (2012) Amplification by PCR artificially reduces the proportion of the rare biosphere in microbial communities. PLoS ONE 7(1):e29973. doi:10.1371/journal.pone.0029973
Green SJ, Côté IM (2014) Trait-based diet selection: prey behaviour and morphology predict vulnerability to predation in reef fish communities. J Anim Ecol 83(6):1451–1460
Green SJ, Akins JL, Côté IM (2011) Foraging behaviour and prey consumption in the Indo-Pacific lionfish on Bahamian coral reefs. Mar Ecol Prog Ser 433:159–167
Grosholz ED, Ruiz GM, Dean CA, Shirley KA, Maron JL et al (2000) The impacts of a nonindigenous marine predator in a California bay. Ecology 81:1206–1224
Guichoux E, Lagache L, Wagner S, Chaumeil P, Léger P, Lepais O, Lepoittevin C, Malausa T, Revardel E, Salin F, Petit RJ (2011) Current trends in microsatellite genotyping. Mol Ecol Res 11(4):591–611
Hackerott S, Valdivia A, Green SJ, Côté IM, Cox CE (2013) Native predators do not influence invasion success of Pacific lionfish on Caribbean reefs. PLoS ONE 8(7):e68259
Hamner RM, Freshwater DW, Whitfield PE (2007) Mitochondrial cytochrome b analysis reveals two invasive lionfish species with strong founder effects in the western Atlantic. J Fish Biol 71(sb):214-22
Handy SM, Deeds JR, Ivanova NV, Herbert PDN, Hanner RH, Ormos A, Weigt LA, Moore MM, Yancy HF (2011) A single laboratory validated method for the generation of DNA barcodes for the identification of fish for regulatory compliance. J AOAC Int 94:201–209
Hebert PD, Penton EH, Burns JM, Janzen DH, Hallwachs W (2003) Ten species in one: dNA barcoding reveals cryptic species in the neotropical skipper butterfly Astraptes fulgerator. P Natl Acad Sci USA 101(41):14812–14817
Hixon MA, Carr MH (1997) Synergistic predation, density dependence and population regulation in marine fish. Science 277:946–949
Hoese HD, Moore RH (1998) Fishes of the Gulf of Mexico, 2nd edn. College Station, Texas
Hornstra HM, Herrel A, Montgomery WL (2004) Gas bladder movement in lionfishes: a novel mechanism for control of pitch. J Morphol 260:299–300
Hyslop EJ (1980) Stomach contents analysis—a review of methods and their application. J Fish Biol 17(4):411–429
Imamura H, Yabe M (1996) Larval record of a red firefish, Pterois volitans, from northwestern Australia (Pisces: Scorpaeniformes). Bull Fac Fish-Hokkaido Univ 47:41–46
Ivanova NV, Zemlak TS, Hanner R, Hebert PDN (2007) Universal primer cocktails for fish DNA barcoding. Mol Ecol Notes 7:544–548
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Mentjies P, Drummond A (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28(12):1647–1649
Kells V, Carpenter K (2011) A field guide to coastal fishes: from Maine to Texas. Baltimore, Maryland
Koskinen MT, Piironen J (2000) The one that did not get away: individual assignment using microsatellite data detects a case of fishing competition fraud. Proc R Soc Lond B Biol Sci 267(1453):1699–1704
Legler ND, Johnson TB, Heath DD, Ludsin SA (2010) Water temperature and prey size effects on the rate of digestion of larval and early juvenile fish. T Am Fish Soc 139(3):868–875
Lesser MP, Slattery M (2011) Phase shift to algal dominated communities at mesophotic depths associated with lionfish (Pterois volitans) invasion on a Bahamian coral reef. Biol Invasions 13:1855–1868
McDonald PS, Jensen GC, Armstrong DA (2001) The competitive and predatory impacts of the nonindigenous crab Carcinus maenas (L.) on early benthic phase Dungeness crab Cancer magister Dana. J Exp Mar Biol Ecol 258:39–54. doi:10.1016/S0022-0981(00)00344-0
Meusnier I, Singer GA, Landry JF, Hickey DA, Hebert PD, Hajibabaei M (2008) A universal DNA mini-barcode for biodiversity analysis. BMC Genom 9(1):214
Morris JA Jr, Akins JL (2009) Feeding ecology of invasive lionfish (Pterois volitans) in the Bahamian archipelago. Environ Biol Fish 86(3):389–398
Morris JA Jr, Whitfield PE (2009) Biology, ecology, control and management of the invasive Indo-Pacific lionfish: an integrated assessment. NOAA Technical Memorandum NOS NCCOS 99. p 57
Muñoz RC, Currin CA, Whitfield PE (2011) Diet of invasive lionfish on hard bottom reefs of the Southeast USA: insights from stomach. Mar Ecol Prog Ser 432:181–193
Nuttall MF, Johnston MA, Eckert RJ, Embesi JA, Hickerson EL (2014) Lionfish (Pterois volitans [Linnaeus, 1758] and P. miles [Bennett, 1828]) records within mesophotic depth ranges on natural banks in the Northwestern Gulf of Mexico. BioInvasion Rec 3:111–115
Paine RT (1966) Food web complexity and species diversity. Am Nat 100(910):65–75
Patterson WF III, Wilson CA, Bentley SJ, Cowan JH Jr, Henwood T, Allen YC, Dufrene TA (2005) Delineating juvenile red snapper habitat on the northern Gulf of Mexico continental shelf. Am Fish Soc Symp 41:277–288
Patterson WF III, Tarnecki JH, Addis DT, Barbieri LR (2014) Reef fish community structure at natural versus artificial reefs in the northern Gulf of Mexico. P Gulf Caribb Fish Ins 67:4–8
Ratnasingham S, Hebert P (2007) BOLD: the barcode of life data system. Mol Ecol Notes 7:355–364
Rilov G (2009) Predator–prey interactions of marine invaders. Springer, Berlin Heidelberg, pp 261–285
Robertson A, Garcia AC, Quintana HA, Smith TB II, Bernard F, Reale-Munroe K, Gulli JA, Olsen DA, Hooe-Rollman JI, Jester EL, Klimek BJ (2014) Invasive lionfish (Pterois volitans): a potential human health threat for ciguatera fish poisoning in tropical waters. Mar Drugs 12(1):88–97
Rudolf VH (2008) Impact of cannibalism on predator-prey dynamics: size-structured interactions and apparent mutualism. Ecology 89(6):1650–1660
Schofield PJ (2010) Update on geographic spread of invasive lionfishes (Pterois volitans [Linnaeus, 1758] and P. miles [Bennett, 1828]) in the Western North Atlantic Ocean, Caribbean Sea and Gulf of Mexico. Aquat Invasions 5:S117–S122
Schofield PJ, Akins L, Gregoire-Lucente DR, Pawlitz RJ (2014) Invasive lionfish use a diversity of habitats in Florida: U.S. Geological Survey Fact Sheet 2014–3032, p 2, http://dx.doi.org/10.3133/fs20143032
Schooley JD, Karam AP, Kesner BR, Marsh PC, Pacey CA, Thornbrugh DJ (2008) Detection of larval remains after consumption by fishes. T Am Fish Soc 37(4):1044–1049
Toledo-Hernández C, Vélez-Zuazo X, Ruiz-Diaz CP, Patricio AR, Mège P, Navarro M, Sabat AM, Betancur-R R, Papa R (2014) Population ecology and genetics of the invasive lionfish in Puerto Rico. Aquat Invasions 9:227–237
Valdez-Moreno M, Quintal-Lizama C, Gómez-Lozano R, del Carmen García-Rivas M (2012) Monitoring an alien invasion: DNA barcoding and the identification of lionfish and their prey on coral reefs of the Mexican Caribbean. PLoS ONE 7(6):e36636
Valdivia A, Bruno JF, Cox CE, Hackerott S, Green SJ (2014) Re-examining the relationship between invasive lionfish and native grouper in the Caribbean. PeerJ 2:e348
Valentini A, Pompanon F, Taberlet P (2009) DNA barcoding for ecologists. Trends Ecol Evol 24(2):110–117
Ward RD, Zemlak TS, Innes BH, Last PR, Hebert PD (2005) DNA barcoding Australia’s fish species. Philos T Roy Sos B 360(1462):1847–1857
Ward RD, Hanner R, Hebert PD (2009) The campaign to DNA barcode all fishes, FISH-BOL. J Fish Biol 74(2):329–356
Whitfield PE, Gardner T, Vives SP, Gilligan MR, Courtenay WR Jr, Ray GC, Hare JA (2002) Biological invasion of the Indo-Pacific lionfish Pterois volitans along the Atlantic coast of North America. Mar Ecol Prog Ser 235:289–297
Workman I, Shah A, Foster D, Hataway B (2002) Habitat preferences and site fidelity of juvenile red snapper (Lutjanus campechanus). ICES J Mar Sci 59:S43–S50
Acknowledgements
We thank Dalton Kennedy, Scott Bartel, Clint Retherford, Bryan Clark, Andy Ross, Anna Clark, Jeremy Porter, Michael Day and Kylie Gray for assistance acquiring lionfish samples for this study. We thank Lei Wang, Pearce Cooper, and Natalie Ortell for minor assistance with molecular equipment. We thank Joseph Tarnecki, Brian Klimek, Justin Lewis, Steve Garner, and Michael Norberg for assistance with the processing of whole lionfish. This research was made possible by a Grant from The Gulf of Mexico Research Initiative/C-IMAGE II and Mississippi Alabama SeaGrant (USM-GR03924-R-HCE-04-PD). Data are publicly available through the Gulf of Mexico Research Initiative Information and Data Cooperative (GRIIDC) at https://data.gulfresearchinitiative.org. Sequences are accessible through the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/).
Author contributions
WFP and AR conceived and designed study; AR and ACO and KAD developed methodology; KAD conducted fieldwork; KAD generated sequencing data and molecular analyses under advice of AR and ACO; KAD and WFP analyzed data and performed statistical analyses; KAD wrote the manuscript; all co-authors provided review and editorial advice.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dahl, K.A., Patterson, W.F., Robertson, A. et al. DNA barcoding significantly improves resolution of invasive lionfish diet in the Northern Gulf of Mexico. Biol Invasions 19, 1917–1933 (2017). https://doi.org/10.1007/s10530-017-1407-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-017-1407-3