Abstract
Non-native Phragmites australis decreases biodiversity and produces dense stands in North America. We surveyed the endophyte communities in the stems, leaves and roots of collections of P. australis obtained from two sites with a low and high salt concentration to determine differences in endophyte composition and assess differences in functional roles of microbes in plants from both sites. We found differences in the abundance, richness and diversity of endophytes between the low saline collections (18 species distributed in phyla Ascomycota, Basidiomycota and Stramenopiles (Oomycota); from orders Dothideales, Pleosporales, Hypocreales, Eurotiales, Cantharellales and Pythiales; Shannon H = 2.639; Fisher alpha = 7.335) and high saline collections (15 species from phylum Ascomycota; belonging to orders Pleosporales, Hypocreales, Diaporthales, Xylariales and Dothideales; Shannon H = 2.289; Fisher alpha = 4.181). Peyronellaea glomerata, Phoma macrostoma and Alternaria tenuissima were species obtained from both sites. The high salt endophyte community showed higher resistance to zinc, mercury and salt stress compared to fungal species from the low salt site. These endophytes also showed a greater propensity for growth promotion of rice seedlings (a model species) under salt stress. The results of this study are consistent with the ‘habitat-adapted symbiosis hypothesis’ that holds that endophytic microbes may help plants adapt to extreme habitats. The capacity of P. australis to establish symbiotic relationships with diverse endophytic microbes that enhance its tolerance to abiotic stresses could be a factor that contributes to its invasiveness in saline environments. Targeting the symbiotic associates of P. australis could lead to more sustainable control of non-native P. australis.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Endophytic fungi are ubiquitous in plants where they associate with multiple plant tissues (Bacon and White 2000; Currie et al. 2014). These microorganisms are frequently examined as potential sources of bioactive metabolites (Zhao et al. 2011), enzymes of industrial interest (Grawe et al. 2015); they also play roles in biological control of plant pathogens (Compant et al. 2010), plant growth promotion (You et al. 2012) and bioremediation (Li et al. 2012a, b). Endophytes play important ecological roles for environmental adaptation of their hosts including increasing resistance against pathogens and herbivores (Clay and Schardl 2002), assisting in mineral nutrition of the plant (Kipfer et al. 2011), mitigating the effects of water stress (Atala et al. 2012), and affecting plant responses to other biotic and abiotic stresses (Clay and Schardl 2002; Rodriguez et al. 2008; Cheplick and Faeth 2009; Soares et al. 2015). Symbiotic relationships can have a direct or indirect impact on the structure, function and composition of plant communities, the expansion of host niches (Rudgers et al. 2015), and food webs and ecosystem processes (Clay and Holah 1999; Aschehoug et al. 2014) including invasion processes. Mutualisms often are an important mechanism used in invasion processes (Callaway et al. 2011), so the competitive ability of invasive species can be increased with mutualistic associations established with the autochthonous microbiota (Reinhart and Callaway 2006; Jordan et al. 2008; Andonian and Hierro 2011) or endophytic microorganisms (Aschehoug et al. 2012, 2014). Rodriguez et al. (2008) and Redman et al. (2002) proposed that symbiosis with endophytic microbes may help plants adapt to particular environmental stresses; they termed this hypothesis ‘habitat-adapted symbiosis’.
Common reed (Phragmites australis subsp. australis (Cav.) Trin. Ex Steud.) is a perennial grass native to Eurasian wetlands (Holm et al. 1977; Meyerson and Cronin 2013). In North America, this non-native macrophyte is highly productive and often outcompetes native plants to create large expanses with very low plant and wildlife biodiversity (Silliman and Bertness 2004; Meyerson et al. 2009). Unlike the native genotype (Phragmites australis subsp. americanus), the non-native Phragmite australis subsp. australis (P. australis) is considered an invasive species and very aggressive (Gessner et al. 1996; Saltonstall 2002; Windham and Meyerson 2003; Lambertini et al. 2012; Meyerson and Cronin 2013). Success in invasion has been related to the ecological capability of P. australis to tolerate and grow in a range of soil salinity and fertility levels (Haslam 1972; Lissner et al. 1999a, b; Meyerson et al. 2000), its wide dispersion efficiency (Kettenring and Mock 2012; Meyerson et al. 2012), high genetic diversity (Saltonstall 2002; Lambertini et al. 2006; Fer and Hroudova 2009), and microbial symbiosis (Kowalski et al. 2015). It also has been shown to tolerate heavy metals and other stresses (Quan et al. 2007; Bonanno and Giudice 2010). The ability of P. australis to cope with salt stress plays a crucial role in colonization of coastal habitats (Achenbach and Brix 2014), but it is unclear how much the plant’s microbiome influences the resistance to salt stress.
The microbial endophyte communities of P. australis have been examined in several previous studies (Wirsel et al. 2001; Li et al. 2010; Angelini et al. 2012; Fischer and Rodriguez 2013; Kim et al. 2014; Hipol and Cuevas 2014; Sim et al. 2015; Sauvêtre and Schröder 2015). The differentiation of the composition and functions of the microbiota associated with P. australis in the invasion process are still unknown. We are not aware of any previous studies that compare the structure and diversity of the endophytic fungal community from P. australis between high salt and low salt sites and examines the differentiation of these communities by functional traits.
Our hypothesis is that the capacity of P. australis to tolerate high salt soils is partially the result of endophytic fungi that enhance host tolerance to salt stress. We use culture dependent methods to assess the community structure and diversity of endophytic fungi associated with P. australis (haplotype M from one site in New Jersey) in high salt and low salt soils. In developing this work, we attempted to answer the following questions: Do endophytic fungal communities in grasses at the two sites differ in structure and composition? Do these communities themselves differ in the degree of resistance to environmental stress? Do these communities differ in the capacity to mitigate stress in the plant?
Materials and methods
Collection sites
P. australis plants were collected from two sites along the shore in Sandy Hook, New Jersey. Five plants were collected in each site. In the inland population in site 1(40.45°N; −73.99°W) approximately 500 m from the shoreline, the sparse population margins in dry sand at the edge of a dense stand in fresh standing water was sampled; and at site 2 (40.47°N; −74.00°W) a dense population was located on the shore approximately 4 meters from the water at high tide. Whole plants (1.5–2.0 m height) with undamaged leaves, roots and rhizomes were collected, brought to the laboratory in polythene bags, and processed the same day. Plants were confirmed as the invasive haplotype M morphologically by Dr. Bernd Blossey in the Department of Natural Resources at Cornell University. For analysis of soil sodium content at both sites, soil was sampled at a depth of 10 cm with approximately six samples arbitrarily taken from around the base of plants. Sand was sampled at various places and levels and combined at each site. Samples were sent to the Rutgers University Soil Test Laboratory and analyzed for exchangeable cation extraction which would specifically provide Na+, Ca++, Mg++, and K+ values (typically as milli-equivalents/100 g). The lime requirement index and soil pH also were tested (McLean 1982).
Isolation and identification of microorganisms
In the laboratory, apical meristems, fully expanded apical leaves and roots (approximately 0.5–1-mm-thick) were sampled from each plant for endophyte analysis. The plant material was superficially disinfected with 70 % ethanol for 1 min, 2.5 % sodium hypochlorite for 5 min (meristematic tissues and leaves) or 7 min (roots), and then rinsed 5 times with autoclaved distilled water. The edges of the leaf fragments were removed and fragments of tissues (2 × 2 mm) were placed on Petri plates containing 10 % trypticase soy agar (10 % TSA), yeast extract-sucrose agar (1 % yeast extract + 1 % sucrose; 1 % YES) and potato dextrose agar plus three antibiotics (ampicillin + tetracycline + streptomycin—50 µg ml−1 for each antibiotic; PDA + 3). We used three media to maximize the likelihood that endophytes in samples would be cultured and detected. Meristem and root fragments (approximately 2-mm-long) were analyzed on the same three media types. The plates were incubated at room temperature and growth was assessed as it occurred. We analyzed 504 fragments (252 from site 1 and 252 from site 2) of each tissue on the media (7 plates/12 fragments each plate). Representative isolates were stored at −80 °C in the Department of Plant Biology and Pathology, Rutgers University, New Brunswick, New Jersey, USA.
The endophytic fungi growing from the samples were grouped into morphotypes following the method used by Lacap et al. (2003). To confirm the grouping, microscopic traits were observed via slides obtained from microculturing. DNA was extracted from a representative of each morphological group with QIAGEN miniprep kits according to the manufacturer’s recommendations. The validation of morphotypes was confirmed by fingerprinting analysis (data not shown) using molecular markers inter-retrotransposon amplified polymorphism (IRAP) and inter-simple sequence repeat ISSR with primers CLIRAP1 (5′-CGT CGA GAA ACA GCT CAC GA-3′) CLIRAP 4 (5′-CTT CCA TTG GCA AGG TGC-3′) (Santos et al. 2012) and BH1 (5′-GTG GTG GTG GTG GTG-3′), respectively. The primers ITS5 (TCC GTA GGT GAA CCT GCG G) and ITS4 (TCC TCC GCT TAT TGA TAT GC) were used for amplification of the ITS region (White et al. 1990). The PCR products were purified with QIAquick PCR Purification Kit and sequenced by the Sanger method (Genewiz, South Plainfield, New Jersey). The sequences were aligned and fixed in MEGA 6 (Tamura et al. 2013). The ITS sequences were compared with sequences deposited in GenBank database using BLASTn (http://www.ncbi.nlm.nih.gov). Identifications were based on Blastn and morphological analysis. ITS sequences with ≥97 % identity (Morris et al. 2008) and morphological similar characteristics were grouped in same species. Colonization frequency (CF%) was determined as the percentage of the endophyte-infected fragments of the total observed fragments in each site. Diversity was measured as Shannon–Wiener and Fisher’s alpha index to characterize the species diversity (Fisher et al. 1943; Spellerberg and Fedor 2003) and species evenness was estimated with Pielou’s evenness index (Pielou 1966) in Past 3.x (Hammer et al. 2001).
Metal and salt resistance of isolates
Resistance and susceptibility to metal toxicity and salt stress were examined for each isolate using the PDA medium amended with HgCl2 (1 mM), ZnSO4·7H2O (10 mM) (Aleem et al. 2003) and NaCl (200 mM). In parallel, cultures without metal and salt were performed as the control treatments. For each factor, the endophytic strains were incubated at 28 °C for 10 days. The radial growth was evaluated from two perpendicular measurements (in mm) that passed through the center of the inoculated portion every other day. The tolerance index (TI), an indication of the organism response to stress was calculated from the growth rate of the strain exposed to metals or salt divided by the growth rate in the control plate. TI > 1 and TI = 1 indicated a resistance strain and TI < 1 indicated a sensible strain. A TI = 0 result indicated that the strain was killed by stress. Three replicates were used for each fungal species and control. The TI data were subsequently analyzed using the multivariate ordination technique principal component analysis (PCA) to assess whether different endophytic communities differ by tolerance stress. PCA scores were further analyzed by ANOVA using R version 3.0.2 (R Core Team 2013) to determine the significance of separations observed in PCA plots to examine similarities between TI and fungi species site.
Effect of endophytic community on rice under salt stress
The ability of the strains to assist the host under salt stress was evaluated in rice seedlings (Oryza sativa L.) because it was not possible to remove native microbes from P. australis seedlings (see Fischer and Rodriguez 2013); furthermore, P. australis and rice have similar responses against pollutants (Chu et al. 2006) and anoxic conditions (Armstrong and Armstrong 2001). To reduce microbial populations on rice seedlings, seeds were immersed in distilled water and maintained at 60 °C for 30 min. The seeds were superficially disinfected with sodium hypochlorite 2.5 % for 15 min and then rinsed 5 times with autoclaved distilled water. Then, the cultivable endophyte-free (E−) seeds were germinated in 10 % TSA for 5 days. To create the experimentally inoculated plants, seedlings (E−) were placed in Petri dishes containing fungal endophytes cultured on potato dextrose agar and allowed to remain in contact with fungal hyphae for 12 h to ensure adequate inoculation. Then, seedlings were transplanted into magenta boxes containing autoclaved vermiculite:soil mix (3:1) moistened with Murashige and Skoogs salt solution (Sigma-Aldrich) plus 0 or 200 mM NaCl. Fafard® Growing Mix 2 was autoclaved 2× for 1 h each autoclaving (1 atm at 121 °C). Magenta boxes were incubated at ambient laboratory temperature in a 12-h alternating light/dark cycle. Eight seedlings were used for each treatment. Plants in vitro were collected after 20 days and rinsed in tap water. The lengths of shoots and roots were measured. Growth promotion efficiency (GPE) was estimated to elucidate the relative effect of tested strains on plants according to Almoneafy et al. (2014). The experiment was set up as a completely randomized design. Data of plant growth and others parameters were analyzed using R version 3.0.2 (R Core Team 2013). Logarithm and natural logarithm transformed data were used for the ANOVA when non-homogeneity of variances was observed.
Results
Isolation and identification of microorganisms
Soil from site 1 exhibited lower values of sodium concentration 25.0 mg/kg soil (low sodium concentration—LS) compared with the soil collected from site 2, which was 175.2 mg/kg soil (high sodium concentration—HS) (Table 1). The two sites were representative of distinct environments in salinity. Populations of P. australis were very uniform in these environments, as well as other soil chemical properties. Therefore, the choice of the two sites is based on the homogeneity of the two collection sites to assess our hypothesis about the effect of salinity on the community of endophytic fungi.
There was no fungal growth from fragments of leaves and meristems over a period of 45 days. We observed abundant growth of fungi from fragments of roots. These strains were isolated and separated into 33 morphotypes from 225 strains of endophytes isolated from 252 fragments of the roots from site 1 (LS) (18 morphotypes) and 252 fragments of the roots from site 2 (HS) (15 morphotypes).
The colonization frequency (CF) of roots differed significantly (t test: t 4.2716, p 0.00011) according to collection site. Regardless of the medium used, roots collected in LS resulted in CF 15.10 ± 10.67 % colonization, while roots in HS reached 32.45 ± 6.56 % colonization. Considering each culture medium and collection site, the CF in PDA did not differ between the sites, unlike of CF in 10 % TSA and 1 % YES. The CF obtained in different media for HS did not differ between culture media. But the CF in the roots of LS varied with the culture medium used (ANOVA: F 9.269, p 0.002). The CF in PDA resulted in 27.38 ± 10.64 % of the fragments with endophytes (Tukey’s test, p < 0.05) compared with 10 % TSA (9.52 ± 9.37 %) and 1 % YES (8.33 ± 4.45 %).
We obtained 225 isolates comprising 30 species in 23 genera from both HS and LS plant populations (Table 2). Analyses of ITS rDNA sequences revealed the correspondence of the morphotypes with distinct species of filamentous fungi. Roots from plants at the HS and LS sites were colonized by 14 and 16 genera, respectively. The sequences have been deposited in GenBank under accession numbers: KT827254–KT827285.
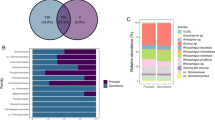
The endophyte community from plants at the LS site showed higher species richness (Table 2): 18 species of fungi, distributed into five classes (Dothideomycetes, Sordariomycetes, Agaricomycetes, Eurotiomycetes and Oomycetes) and six orders (Dothideales, Pleosporales, Hypocreales, Eurotiales, Cantharellales and Pythiales). Sordariomycetes, Pleosporales and Hypocreales, were represented by the greatest numbers of species. The endophyte community in roots from the HS sites showed lower richness, consisting exclusively of species of Ascomycota (Table 2), almost equally distributed in the Dothideomycetes and Sordariomycetes classes, and included orders Pleosporales, Hypocreales, Diaporthales, Xylariales and Dothideales; orders Pleosporales and Hypocreales were the most frequently isolated.
The species Peyronellaea glomerata, Phoma macrostoma and Alternaria tenuissima were encountered in both low and high saline sites (Table 2). The species richness difference between the plants at the two sites is reflected in the indices of diversity. The indices were higher for plants at the low saline site (Shannon H = 2.64; Fisher alpha = 7.33) compared to those at the high saline site (Shannon H = 2.29; Fisher alpha = 4.18). The dominant species in the low saline site were Peyronellaea glomerata-FN3 (relative abundance 19.23 %) and Fusarium sporotrichioides-FN23 (relative abundance 19.23 %); and Fusarium oxysporum-FI28 (relative abundance 28.57 %) and Peyronellaea glomerata-FI47 (relative abundance 20.40 %) were more frequently isolated in plants from the high saline site (Table 2).
Stress tolerance
The addition of mercury (HgCl2 1 mM) improved growth (TI ≥ 1) of Phomopsis sp.-FI26, Phomopsis mali-FI15, Fusarium oxysporum-FI28 and Paraphoma radicina-FN8 (Table 2). The other species were sensitive to mercury resulting in TI values <1. The resistance profiles on ZnSO4.7H2O (10 mM) were differentiated by isolation source. Approximately 61 and 26.5 % of the endophytic microbes from the low and high saline sites, respectively, did not grow in the medium containing Zn2+ (IT = 0) (Table 2). The addition of zinc to the culture medium improved the Phomopsis sp.-FI26 growth (TI > 1). The other strains showed TI < 1 values indicating sensitivity to ZnSO4·7H2O (10 mM). The TI values varied according to the host source, ranging from 0.05 to 0.33 (species from low saline site) and 0.07–0.94 (species from high saline site), respectively.
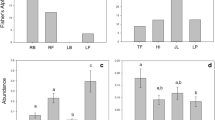
The profiles of salt tolerance in endophytes differed based on whether plants grew in a low or high salt soil. The addition of 300 mM NaCl favored growth (TI > 1) of approximately 87 % of fungi obtained from the high saline site. Only Fusarium avenaceum-FI25 and Pseudoseptoria obscura-FI48 were sensitive (TI < 1) to salt stress. In contrast, only Alternaria sp.-FN20, Fusarium sporotrichioides-FN23 and Ilyonectria radicícola-FN28, isolated from the low saline site, were resistant (TI > 1) to NaCl 300 mM. The other species (approximately 83 %) were sensitive (TI < 1) to NaCl. The PCA analysis indicated that the set of effects (TI values: Hg2+, Zn2+ and NaCl) could differentiate fungal community from the site 1 = LS and site 2 = HS (Fig. 1). The PC1 and PC2 axes explain 82.94 % of the variations found. The two endophytic communities were different in the response to Hg2+, Zn2+ and NaCl in accordance to the mean test for the values of PC1 (t = −3.5756, df = 31, p value = 0.001169) and PC2 (t = 3.4341, df = 31, p value = 0.001709).
The ability to protect rice seedlings from salt stress was clearly different between the two fungal communities (Tables 2, 3). A. tenuissima-FI31, Arthrinium arundinis-FI6, Paraphaeosphaeria michotii-FI21, Phomopsis sp.-FI26, Purpureocillium lilacinum-FI41, Septoriella hubertusii-FI4 Phomopsis mali-FI15 and Bipolaris buchloes-FI10 all increased stress tolerance with TI-rice >1.9 (Table 2). Pythium dissotocum-FN24 was the only species isolated from LS site that showed TI-rice >1.9 (Table 2). Nine species from the high saline site induced increased length (root + shoot) of the rice seedlings GPE values ranging from 72.0 to 131.8 % (Table 3). Only two isolates from the LS site resulted in GPE 62.0 and 119.1 % (Table 3). Other species of endophytes resulted in negative effects on the rice seedlings, drastically reducing total seedling lengths and consequently resulting in negative values of GPE (Table 3). In total, seven and four endophytic species from low and high saline sites, respectively, had pathogenic (i.e., reducing growth of seedlings over the un-inoculated controls) effects in rice under salt stress. The other species (nine and two from low and high saline sites, respectively) did not result in significantly different effects compared to the control (no inoculation seedlings) (Table 3).
Discussion
Our results show that variation in soil salt content between the two sites studied may influence the endophytic community in P. australis root systems. Understanding whether and how endophytic microbes may be affecting competitive ability of plants is important basic information and may lead to new management strategies of invasive populations (Kowalski et al. 2015). Knowledge of the symbiotic microbial communities that enhance invasiveness contributes to our understanding of the roles that microbes play in enabling invasive plants to outcompete native species.
Distribution and diversity of endophytes in P. australis plants
We could not isolate endophytes from young leaves and shoot meristematic tissues even after 45 days of incubation. This may be explained in that we used very young leaf and meristem tissues. Many fungal endophytes, especially the class 2 and 3 endophytes, colonize plant tissues as they age and may not be present in younger tissues (Rodriguez et al. 2009). Although other authors have reported fungal isolation from leaves of P. australis, the ages of leaves were not always clear (Wirsel et al. 2001; Neubert et al. 2006; Angelini et al. 2012; Fischer and Rodriguez 2013; Sim et al. 2015). Venkatachalam et al. (2015) also did not detect endophytes in leaves of Cymodocea serrulata and Thalassia sp. when analyzing 100 leaf fragments from 10 different grasses species.
We found that roots of P. australis plants from both sites were colonized abundantly by endophytic microbes. Despite the lower CF in the roots from the LS site, endophytic microbial diversity (Shannon diversity) was higher than that supported by the HS site. Biotic and abiotic factors influence the CF, richness and diversity of endophytes (Arnold 2007; Arnold and Lutzoni 2007). The Shannon’s diversity values 2.64 (roots from LS site) and 2.28 (roots from HS site) were between 1.5 and 3.5; therefore, the Shannon’s index for each P. australis population in the present study was similar to those reported in another study on endophytes from Hevea brasiliensis (Gazis and Chaverri 2010).
We recovered a high diversity of endophytes from the P. australis root samples, with isolates from roots belonging to species of the phylum Ascomycota, Basidiomycota and Oomycota (Stramenopiles), including 23 genera and 30 species (Table 2). These results clearly indicate that a high diversity of endophytic microbes is associated with roots of P. australis. The endophytic community differentiated between high and low saline sites in high-level taxa as observed in other macrophytes (Sandberg et al. 2014). In general, endophytic fungi of plant root systems tend to be highly diverse (Vandenkoornhuyse et al. 2002). The dominant species we encountered in plants of the HS site were Fusarium oxysporum and Peyronellaea glomerata, and in plants of the LS site were Fusarium sporotrichioides and P. glomerata. F. oxysporum has been reported as an endophytic fungus in nearly 100 plant species (Kuldau and Yates 2000). F. sporotrichioides (Szécsi et al. 2013) and P. glomerata (Zhang et al. 2012) are endophytes found in a lower diversity of hosts.
The endophytic species isolated in our study were not previously detected in Phragmites roots (Wirsel et al. 2001; Angelini et al. 2012; Kim et al. 2014; Hipol and Cuevas 2014; Sim et al. 2015). Fischer and Rodriguez (2013) found Arthrinium arundinis, F. oxysporum and F. sporotrichioides in shoots of non-native P. australis collected in Michigan. Paraphaeosphaeria michotii was found in leaf sheaths of P. autralis along a salinity gradient in Belgium (van Ryckegem and Verbeken 2005).
Species of genera Arthrinium, Diaporthe, Fusarium, Phoma and Trichoderma have been previously isolated from P. australis roots (Wirsel et al. 2001; Angelini et al. 2012; Sim et al. 2015). Species of the genera Phomopsis, Alternaria, Paecilomyces and Xylaria have been isolated as endophytes from many other grass plants (Kleczewski et al. 2012; Márquez et al. 2012). We suggest that composition of the endophytic communities in P. australis may differ due to various factors, including plant genotype, tissue type, soil composition, geographic and seasonal factors (Collado et al. 1999; Higgins et al. 2007; Suryanarayanan et al. 2011; U’Ren et al. 2012; Gehring et al. 2014).
Invasive species may thrive due to escape from soil-borne phytopathogens when they are introduced into new habitat (Keane and Crawley 2002; Parker and Gilbert 2007; Reinhart et al. 2010). Some species in our study are known causative agents of plant diseases, including F. oxysporum (Guo et al. 2014), A. tenuissima (Blodgett and Swart 2002) and F. sporotrichioides (Mielniczuk et al. 2004). Many grasses are rich sources of endophytic Fusarium (Szécsi et al. 2013). The detection of known plant pathogens is relatively common in endophytic fungal communities (Moricca et al. 2012; Sun et al. 2012; Wearn et al. 2012; Pawlowska et al. 2014) reinforcing the idea that the symbiosis between plant and endophyte oscillates between parasitism and mutualism (Schulz and Boyle 2006). The root colonization of pathogenic strains varies depending on the plant species and the forma specialis as in F. oxysporum colonization (Alabouvette et al. 2009). Since none of the P. australis plants in our study showed symptoms of disease, it seems possible that pathogenicity by these and other root endophytes may be suppressed in the symbiotic relationship.
Root endophytes and plant stress tolerance
Salt-affected soils are characterized by excess levels of soluble salts (salinity) and/or Na+ in the solution phase as well as in a cation exchange complex (sodicity) (see Qadir et al. 2007). Plants are exposed to water stress, ion toxicity and disturbances in mineral nutrition when they are growing under hypersaline conditions (Sabzalian and Mirlohi 2010). Although our study was limited, the differences between the endophyte communities obtained from the two sites are proposed to be caused, at least in part, by the site conditions that played a role in influencing which microbes survived and associated with host roots. In this respect, it is notable that soils (predominantly sand) associated with the plants at the inland site contained significantly less salt than we measured in sand associated with the plants at the other site (Na+ = 0.11 meq/100 g at LS site (site 1) vs. Na+ = 0.76 meq/100 g at HS site (site 2).
Endophytes often increase the competitive ability of plants, especially under stress (e.g., Saikkonen et al. 2013). Osmotic and specific ionic stresses from salinity can cause stunted growth and reduced plant yield (Todaka et al. 2015). Excess salinity in the soil can compromise the availability of water and nutrients for the plants by affecting osmotic potential of the soil solution. High sodium levels also cause degradation of the soil structure, dispersion clay, and toxicity in plants, and may even prevent the germination of seeds and inhibit root development (Smith et al. 2009).
Many of the endophytes from the LS site were not found to be resistant to salt, while many endophytes found in plants from the HS site were found to be resistant to salt. This, in tandem with results of experiments showing that the endophytes from plants in the HS environment promote the growth of plants (rice) under salt stress, lend support to the ‘habitat-adapted symbiosis hypothesis’ proposed by Rodriguez et al. (2008). According to this hypothesis, plants living under stressful conditions may enhance their resistance to stress by associating with endophytes that increase their tolerance to stress. Our use of rice to screen P. australis endophytes for their capacity to enhance salt tolerance is supported by studies that have demonstrated that endophytic fungi that provide increased stress tolerance in one host also may provide stress tolerance in other more distantly related hosts (Waller et al. 2005; Redman et al. 2011). In addition, rice and P. australis are both grasses, and several studies have shown that they possess similar physiological characteristics (see Armstrong and Armstrong 2001, 2005; Soukup et al. 2002; Serghat et al. 2005; Chu et al. 2006).
Most saline resistant plants have biochemical and physiological mechanisms to reduce this abiotic stress (Munns and Tester 2008). The community of endophytic fungi obtained from plants at the HS site (>50 % of species resulted in TI-rice >1.9) may contribute more effectively to enhanced salt tolerance than the community from roots of P. australis collected at the LS site (~5 % of species). It seems reasonable that some plants may naturally associate with endophytic microbiota to enhance their competiveness or invasiveness (Nuñez et al. 2009; Molina-Montenegro et al. 2015; Van der Putten et al. 2007; Kowalski et al. 2015). Nevertheless, due to the limitations of our experiments expanded studies examining a greater number of sampling sites and assessment of endophyte effects on changes in stress tolerance of seedlings of P. australis rather than rice would be desirable to confirm our experimental results.
Potential applications of P. australis endophytes
Redman et al. (2011) argued that endophytes may be useful in mitigating impacts of climate change and expanding agricultural production into stressful environments. Redman et al. (2011) also demonstrated that endophytic fungi isolated from Leymus mollis and Dichanthelium lanuginosum promoted plant growth in rice seedlings under salt stress similar to what we observed for the P. australis root endophytes. Fungal endophytes isolated from hosts on saline soils may have potential application to increase tolerance of agriculturally and horticulturally important plants to salt or arid environments (Waller et al. 2005; Khan et al. 2011; Yin et al. 2014; Azad and Kaminskyj 2015). P. australis has also been used widely to treat industrial wastewater containing heavy metals in wetland systems (Jean and De 1997; Vymazal and Kropfelova 2005). The success of P. australis in invasion of diverse environments has been related to phenotypic, genetic and reproductive plasticity (Haslam 1972; Lissner et al. 1999a, b; Saltonstall 2002; Lambertini et al. 2006; Fer and Hroudova 2009; Kettenring and Mock 2012; Meyerson et al. 2012; Cronin et al. 2015). However, it is possible that the capacity of P. australis to grow in heavy metal contaminated sites could stem at least in part from endophytic microbes. In this respect it has been shown that P. australis is tolerant to toxic concentrations of Zn (Weis and Weis 2004) and Hg (Stoltz and Greger 2002). Our analysis revealed the presence of several heavy metal tolerant taxa of fungi. These heavy metal-tolerant fungi could contribute to the capacity of non-native P. australis to grow in heavy metal contaminated sites. Future experiments with our isolated strains are needed to evaluate the effect of P. australis endophytes on plant resistance to heavy metals (HgCl2 and ZnSO4·7H2O). If such endophytes are confirmed, they could be used to produce cultivars of plants for bioremediation of contaminated sites (Stoltz and Greger 2002; Idris et al. 2004; Weis and Weis 2004; Li et al. 2012a, b).
Conclusions
This study shows that endophytic microbes of P. australis are diverse, may be distinctive at low and high taxonomic levels, and are influenced by ecosystem conditions. Endophytic fungal communities from P. australis differed depending on soil saline conditions. Experiments using rice revealed that some of the endophytes found at the high salinity site increased tolerance of rice seedlings to elevated levels of salinity. This result is consistent with the hypothesis of ‘habitat-adapted symbiosis’ (Rodriguez et al. 2008), where endophytes are hypothesized to help plant hosts adapt to specific, often harsh, habitats. Our results suggest that endophytes play a role in increasing the capacity of P. australis to grow in high salinity soils, probably contributing to invasion in saline environments.
References
Achenbach L, Brix H (2014) Can differences in salinity tolerance explain the distribution of four genetically distinct lineages of Phragmites australis in the Mississippi River Delta? Hydrobiologia 737:5–23
Alabouvette C, Olivain C, Migheli Q, Steinberg C (2009) Micobiological control of soil-borne phytopathogenic fungi with special emphasis on wilt-inducing Fusarium oxysporum. New Phytol 184:529–544
Aleem A, Isar J, Malik A (2003) Impact of long-term application of industrial wastewater on the emergence of resistance traits in Azotobacter chroococcum isolated from rhizospheric soil. Bioresour Technol 86:7–13. doi:10.1016/S0960-8524(02)00134-7
Almoneafy AA, Kakar KU, Nawaz Z, Li B, Saand MA, Chun-lan Y, Xie GL (2014) Tomato plant growth promotion and antibacterial related-mechanisms of four rhizobacterial Bacillus strains against Ralstonia solanacearum. Symbiosis 63:59–70. doi:10.1007/s13199-014-0288-9
Andonian K, Hierro JL (2011) Species interactions contribute to the success of a global plant invader. Biol Invasions 13:2957–2965. doi:10.1007/s10530-011-9978-x
Angelini P, Rubini A, Gigante D, Reale L, Pagiotti R, Venanzoni R (2012) The endophytic fungal communities associated with the leaves and roots of the common reed (Phragmites australis) in Lake Trasimeno (Perugia, Italy) in declining and healthy stands. Fungal Ecol 5:683–693. doi:10.1016/j.funeco.2012.03.001
Armstrong J, Armstrong W (2001) Rice and Phragmites: effects of organic acids on growth, root permeability, and radial oxygen loss to the rhizosphere. Am J Bot 88:1359–1370
Armstrong J, Armstrong W (2005) Rice: sulfide-induced barriers to root radial oxygen loss, Fe2+ and water uptake, and lateral root emergence. Ann Bot 96:625–638
Arnold AE (2007) Understanding the diversity of foliar endophytic fungi: progress, challenges, and frontiers. Fungal Biol Rev 21:51–66. doi:10.1016/j.fbr.2007.05.003
Arnold AE, Lutzoni F (2007) Diversity and host range of foliar fungal endophytes: are tropical leaves biodiversity hotspots? Ecology 88:541–549. doi:10.1890/05-1459
Aschehoug ET, Metlen KL, Callaway RM, Newcombe G (2012) Fungal endophytes directly increase the competitive effects of an invasive forb. Ecology 93:3–8
Aschehoug ET, Callaway RM, Newcombe G, Tharayil N, Chen S (2014) Fungal endophyte increases the allelopathic effects of an invasive forb. Oecologia 175:285–291
Atala C, Capponi EM, Pereira G, Navarrete E, Oses R, Montenegro MM (2012) Impact of mycorrhizae and irrigation in the survival of seedlings of Pinus radiata D. Don subject to drought. Gayana Bot 69:296–304
Azad K, Kaminskyj S (2015) A fungal endophyte strategy for mitigating the effect of salt and drought stress on plant growth. Symbiosis. doi:10.1007/s13199-015-0370-y
Bacon CW, White JF (2000) Microbial Endophytes. Marcel Dekker Inc., New York
Blodgett JT, Swart WJ (2002) Infection, colonization, and disease of Amaranthus hybridus leaves by the Alternaria tenuissima group. Plant Dis 86:1199–1205. doi:10.1094/PDIS.2002.86.11.1199
Bonanno G, Giudice RL (2010) Heavy metal bioaccumulation by the organs of Phragmites australis (common reed) and their potential use as contamination indicators. Ecol Indic 10:639–645. doi:10.1016/j.ecolind.2009.11.002
Callaway RM, Bedmar EJ, Reinhart KO, Silvan CG, Klironomos J (2011) Effects of soil biota from different ranges on Robinia invasion: acquiring mutualists and escaping pathogens. Ecology 92:1027–1035
Cheplick GP, Faeth SH (2009) Ecology and evolution of the grass-endophyte symbiosis. Oxford University Press, Oxford
Chu WK, Wong MH, Zhang J (2006) Accumulation, distribution and transformation of DDT and PCBs by Phragmites australis and Oryza sativa L.: I. Whole plant study. Environ Geochem Health 28:159–168. doi:10.1007/s10653-005-9027-8
Clay K, Holah J (1999) Fungal endophyte symbiosis and plant diversity in successional fields. Science 285:1742–1745
Clay K, Schardl C (2002) Evolutionary origins and ecological consequences of endophyte symbiosis with grasses. Am Nat 160:99–127
Collado J, Platas G, González I, Peláez F (1999) Geographical and seasonal influences on the distribution of fungal endophytes in Quercus ilex. New Phytol 144:525–532
Compant S, Clément C, Sessitsch A (2010) Plant growth-promoting bacteria in the rhizo- and endosphere of plants: their role, colonization, mechanisms involved and prospects for utilization. Soil Biol Biochem 42:669–678
Cronin JT, Bhattarai GP, Allen WJ, Meyerson LA (2015) Biogeography of a plant invasion: plant–herbivore interactions. Ecology 96:1115–1127
Currie AF, Wearn J, Hodgson S, Wendt H, Broughton SJ, Jin L (2014) Foliar fungal endophytes in herbaceous plants: a marriage of convenience. In: Verma VC, Gange AC (eds) Advances in Endophytic Research. Springer, New Delhi, pp 61–81
Fer T, Hroudova Z (2009) Genetic diversity and dispersal of Phragmites australis in a small river system. Aquat Bot 90:165–171
Fischer MS, Rodriguez RJ (2013) Fungal endophytes of invasive Phagramites australis populations vary in species composition and fungicide susceptibility. Symbiosis 61:55–62. doi:10.1007/s13199-013-0261-z
Fisher RA, Corbet AS, Williams CB (1943) The relation between the number of species and the number of individuals in a random sample of an animal population. J Anim Ecol 12:42–58. doi:10.2307/1411
Gazis R, Chaverri P (2010) Diversity of fungal endophytes in leaves and stems of wild rubber trees (Hevea brasiliensis) in Peru. Fungal Ecol 3:240–254. doi:10.1016/j.funeco.2009.12.001
Gehring CA, Ji B, Fong S, Whitham TG (2014) Hybridization in Populus alters the species composition and interactions of root-colonizing fungi: consequences for host plant performance. Botany 92:287–293. doi:10.1139/cjb-2013-0174
Gessner MO, Schieferstein B, Müller U, Barkmann S, Lenfers UA (1996) A partial budget of primary organic carbon flows in the littoral zone of a hardwater lake. Aquat Bot 55:93–105
Grawe GF, de Oliveira TR, de Andrade Narciso E, Moccelini SK, Terezo AJ, Soares MA, Castilho M (2015) Electrochemical biosensor for carbofuran pesticide based on esterases from Eupenicillium shearii FREI-39 endophytic fungus. Biosens Bioelectron 63:407–413. doi:10.1016/j.bios.2014.07.069
Guo L, Han L, Yang L et al (2014) Genome and transcriptome analysis of the fungal pathogen Fusarium oxysporum f. sp. cubense causing banana vascular wilt disease. PLoS ONE. doi:10.1371/journal.pone.0095543
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:1–9
Haslam SM (1972) Biological flora of the British Isles: Phragmites communis Trin. J Ecol 60:585–610
Higgins KL, Arnold AE, Miadlikowska J, Sarvate SD, Lutzoni F (2007) Phylogenetic relationships, host affinity, and geographic structure of boreal and arctic endophytes from three major plant lineages. Mol Phylogenet Evol 42:543–555
Hipol RM, Cuevas VC (2014) Copper tolerance and copper accumulation of culturable endophytic yeasts of Phragmites australis cav. (trin) ex steud. from the mine tailings pond in Mankayan, Benguet, Philippines. Asian J Appl Sci 2:636–643
Holm LG, Plucknett DL, Pancho JV, Herberger JP (1977) The World’s Worst Weeds: distribution and biology. The University Press of Hawaii, Honolulu
Idris R, Trifonova R, Puschenreiter M, Welzel WW, Seissitsch A (2004) Bacterial communities associated with flowering plants of the Ni hyperaccumulator Thlaspi goesingense. Appl Environ Microbiol 70:2667–2677
Jean L, De M (1997) Constructed wetlands for sludge dewatering. Water Sci Technol 35:279–285
Jordan NR, Larson DL, Huerd SC (2008) Soil modification by invasive plants: effects on native and invasive species of mixed-grass prairies. Biol Invasions 10:177–190
Keane RM, Crawley MJ (2002) Exotic plant invasions and the enemy release hypothesis. Trends Ecol Evol 17:164–170. doi:10.1016/S0169-5347(02)02499-0
Kettenring KM, Mock KE (2012) Genetic diversity, reproduction mode, and dispersal differ between cryptic invader, Phragmites australis, and its native conspecific. Biol Invasions 14:2489–2504. doi:10.1007/s10530-012-0246-5
Khan AL, Hamayun M, Kim YH, Kang SM, Lee IJ (2011) Ameliorative symbiosis of endophyte (Penicillium funiculosum LHL06) under salt stress elevated plant growth of Glycine max L. Plant Physiol Biochem 49:852–861. doi:10.1016/j.plaphy.2011.03.005
Kim H, You YH, Yoon H, Seo Y, Kim YE, Choo YS, Lee IJ, Shin JH, Kim JG (2014) Culturable fungal endophytes isolated from the roots of coastal plants inhabiting Korean east coast. Mycobiology 42:100–108. doi:10.5941/MYCO.2014.42.2.100
Kipfer T, Moser B, Egli S, Wohlgemuth T, Ghazoul J (2011) Ectomycorrhiza succession patterns in Pinus sylvestris forests after stand-replacing fire in the Central Alps. Oecologia 167:219–228
Kleczewski MM, Bauer JT, Bever JD, Clay K, Reynolds HL (2012) A survey of endophytic fungi of switchgrass (Panicum virgatum) in the Midwest, and their putative roles in plant growth. Fungal Ecol 5:521–529. doi:10.1016/j.funeco.2011.12.006
Kowalski KP, Bacon C, Bickford W, Braun H, Clay K, Leduc-Lapierre M, Lillard E, McCormick MK, Nelson E, Torres M, White J, Wilcox DA (2015) Advancing the science of microbial symbiosis to support invasive species management: a case study on Phragmites in the Great Lakes. Front Microbiol 6:1–14. doi:10.3389/fmicb.2015.00095
Kuldau GA, Yates IE (2000) Evidence for Fusarium endophytes in cultivated and wild plants. In: Bacon CW, White JF Jr (eds) Microbial endophytes. Marcel Dekker, New York, pp 85–117
Lacap DC, Hyde KD, Liew ECY (2003) An evaluation of the fungal ‘morphotype’ concept based on ribosomal DNA sequences. Fungal Divers 12:53–66
Lambertini C, Gustafsson MH, Frydenberg J, Lissner J, Speranza M, Brix H (2006) A phylogeographic study of the cosmopolitan genus Phragmites (Poaceae) based on AFLPs. Plant Syst Evol 258:161–182
Lambertini C, Mendelssohn IA, Gustafsson MH, Olesen B, Riis T, Sorrell BK, Brix H (2012) Tracing the origin of Gulf Coast Phragmites (Poaceae): a story of long-distance dispersal and hybridization. Am J Bot 99:538–551. doi:10.3732/ajb.1100396
Li YH, Zhu JN, Zhai ZH, Zhang Q (2010) Endophytic bacterial diversity in roots of Phragmites australis in constructed Beijing Cuihu wetland (China). FEMS Microbiol Lett 309:84–93. doi:10.1111/j.1574-6968.2010.02015.x
Li HY, Li DW, He CM, Zhou ZP, Mei T, Xu HM (2012a) Diversity and heavy metal tolerance of endophytic fungi from six dominant plant species in a Pb-Zn mine wasteland in China. Fungal Ecol 5:309–315. doi:10.1016/j.funeco.2011.06.002
Li HY, Wei DQ, Shen M, Zhou ZP (2012b) Endophytes and their role in phytoremediation. Fungal Divers 54:11–18. doi:10.1007/s13225-012-0165-x
Lissner J, Schierup HH, Comin FA, Astorga V (1999a) Effect of climate on the salt tolerance of two Phragmites australis populations. I. Growth, inorganic solutes, nitrogen relations and osmoregulation. Aquat Bot 64:317–333
Lissner J, Schierup HH, Comin FA, Astorga V (1999b) Effect of climate on the salt tolerance of two Phragmites australis populations. II. Diurnal CO2 exchange and transpiration. Aquat Bot 64:335–350
Márquez SS, Bills GF, Herrero N, Zabalgogeazcoa I (2012) Non-systemic fungal endophytes of grasses. Fungal Ecol 5:289–297. doi:10.1016/j.funeco.2010.12.001
McLean EO (1982) Soil pH and lime requirement. In: Page AL, Miller LH, Keeney DR (eds) Chemical and microbiological properties. Methods of soil analysis. Part 2, 2nd edn. American Society of Agronomy, Madison, WI
Meyerson LA, Cronin JT (2013) Evidence for multiple introductions of Phragmites australis to North America: detection of a new non-native haplotype. Biol Invasions 15:2605–2608. doi:10.1007/s10530-013-0491-2
Meyerson LA, Saltonstall K, Windham L, Kiviat E, Findlay S (2000) A comparison of Phragmites australis in freshwater and brackish marsh environments in North America. Wetlands Ecol Manag. doi:10.1023/A:1008432200133
Meyerson L, Saltonstall K, Chambers R (2009) Phragmites australis in eastern North America: a historical and ecological perspective. In: Silliman BR, Gorsholz T, Bertness M (eds) Human impacts in salt marshes: a global perspective. California Press, Oakland
Meyerson LA, Lambertini C, McCormick MK, Whigham DF (2012) Hybridization of common reed in North America? The answer is blowing in the wind. AoB Plants. doi:10.1093/aobpla/pls022
Mielniczuk E, Kiecana I, Perkowski J (2004) Susceptibility of oat genotypes to Fusarium crookwelense Burgess, Nelson and Toussoun infection and mycotoxin accumulation in kernels. Biologia 59:809–816
Molina-Montenegro MA, Oses R, Torres-Díaz C, Atala C, Núñez MA, Armas C (2015) Fungal endophytes associated with roots of nurse cushion species have positive effects on native and invasive beneficiary plants in an alpine ecosystem. Perspect Plant Ecol 17:218–226
Moricca S, Ginetti B, Ragazzi A (2012) Species- and organ-specificity in endophytes colonizing healthy and declining Mediterranean oaks. Phytopathol Mediterr 51:587–598
Morris MH, Smith ME, Rizzo DM, Rejmánek M (2008) Bledsoe CS (2008) Contrasting ectomycorrhizal fungal communities on the roots of co-occurring oaks (Quercus spp.) in a California woodland. New Phytol 178:167–176. doi:10.1111/j.1469-8137.2007.02348.x
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681. doi:10.1146/annurev.arplant.59.032607.092911
Neubert K, Mendgen K, Brinkman H, Wirsel SGR (2006) Only few fungal species dominate highly diverse mycofloras associated with the common reed. Appl Environ Microbiol 72:1118–1128
Nuñez MA, Horton TR, Simberloff D (2009) Lack of belowground mutualisms hinders Pinaceae invasions. Ecology 90:2352–2359
Parker IM, Gilbert GS (2007) When there is no escape: the effects of natural enemies on native, invasive, and noninvasive plants. Ecology 88:1210–1224
Pawłowska J, Wilk M, Śliwińska-Wyrzychowska A, Mętrak M, Wrzosek M (2014) The diversity of endophytic fungi in the above-ground tissue of two Lycopodium species in Poland. Symbiosis 63:87–97. doi:10.1007/s13199-014-0291-1
Pielou EC (1966) The measurement of diversity in different types of biological collections. J Theor Biol 13:131–144. doi:10.1016/0022-5193(66)90013-0
Qadir M, Oster JD, Schubert S, Noble AD, Sahrawat KL (2007) Phytoremediation of sodic and saline-sodic soils. Adv Agron 96:197–247
Quan WM, Han JD, Shen AL, Ping XY, Qian PL, Li CJ, Shi LY, Chen YQ (2007) Uptake and distribution of N, P and heavy metals in three dominant salt marsh macrophytes from Yangtze River estuary, China. Mar Environ Res 64:21–37
R Core Team (2013) A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/
Redman RS, Sheehan KB, Stout RG, Rodriguez RJ, Henson JM (2002) Thermotolerance generated by plant/fungal symbiosis. Science 298:1581
Redman RS, Kim YO, Woodward CJDA, Greer C, Espino L, Doty SL, Rodriguez RJ (2011) Increased fitness of rice plants to abiotic stress via habitat adapted symbiosis: a strategy for mitigating impacts of climate change. PLoS ONE 6(7):e14823. doi:10.1371/journal.pone.0014823
Reinhart KO, Callaway RM (2006) Soil biota and invasive plants. New Phytol 170:445–457
Reinhart KO, Van der Putten WH, Tytgat T, Clay K (2010) Virulence of soil-borne pathogens and invasion by Prunus serotina. New Phytol 186:484–495
Rodriguez RJ, Henson J, Van Volkenburgh E, Hoy M, Wright L, Beckwith F, Kim YO, Redman RS (2008) Stress tolerance in plants via habitat-adapted symbiosis. ISME J 2:404–416
Rodriguez RJ, White JF Jr, Arnold AE, Redman RS (2009) Fungal endophytes: diversity and functional roles. New Phytol 182:314–330. doi:10.1111/j.1469-8137.2009.02773.x/pdf
Rudgers JA, Dereske LB, Crawford KM, Emery SM (2015) Fungal symbiont effects on dune plant diversity depend on precipitation. J Ecol 103:219–230. doi:10.1111/1365-2745.12338
Sabzalian MR, Mirlohi A (2010) Neotyphodium endophytes trigger salt resistance in tall and meadow fescues. J Plant Nutr Soil Sci 173:952–957
Saikkonen K, Ruokolainen K, Huitu O, Gundel PE, Piltti T, Hamilton CE, Helander M (2013) Fungal endophytes help prevent weed invasions. Agric Ecosyst Environ 165:1–5. doi:10.1016/j.agee.2012.12.002
Saltonstall K (2002) Cryptic invasion by a non-native genotype of the common reed, Phragmites australis, into North America. Proc Natl Acad Sci USA 99:2445–2449
Sandberg D, Battista L, Arnold AE (2014) Fungal endophytes of aquatic macrophytes: diverse host-generalists characterized by tissue preferences and geographic structure. Microb Ecol 67:735–747
Santos LV, de Queiroz MV, Santana MF, Soares MA, de Barros EG, de Araújo EF, Langin T (2012) Development of new molecular markers for the Colletotrichum genus using RetroCl1 sequences. World J Microbiol Biotechnol 28:1087–1095. doi:10.1007/s11274-011-0909-x
Sauvêtre A, Schröder P (2015) Uptake of carbamazepine by rhizomes and endophytic bacteria of Phragmites australis. Front Plant Sci 6:1–11. doi:10.3389/fpls.2015.00083
Schulz B, Boyle C (2006) What are endophytes? In: Schulz B, Boyle CJ, Sieber TN (eds) Microbial root endophytes. Springer, Berlin, pp 1–13
Serghat S, Mradmi K, Touhami AO, Douira A (2005) Rice leaf pathogenic fungi on wheat, oat, Echinochloa phyllopogon and Phragmites australis. Phytopathol Mediterr 44:44–49
Silliman BR, Bertness MD (2004) Shoreline development drives invasion of Phragmites australis and the loss of plant diversity on New England salt marshes. Conserv Biol. doi:10.1111/j.1523-1739.2004.00112.x
Sim CSF, Tan WS, Ting ASY (2015) Endophytes from Phragmites for metal removal: evaluating their metal tolerance, adaptive tolerance behaviour and biosorption efficacy. Desalination Water Treat. doi:10.1080/19443994.2015.1013507
Smith AP, Chen D, Chalk PM (2009) N2 fixation by faba bean (Vicia faba L.) in a gypsum-amended sodic soil. Biol Fertil Soils 45:329–333
Soares MA, Li HY, Bergen M, Silva JM, Kowalski KP, White JF (2015) Functional role of an endophytic Bacillus amyloliquefaciens in enhancing growth and disease protection of invasive English ivy (Hedera helix L.). Plant Soil. doi:10.1007/s11104-015-2638-7
Soukup A, Votrubova O, Cizkova H (2002) Development of anatomical structure of roots of Phragmites australis. New Phytol 153:277–287
Spellerberg IF, Fedor PJ (2003) A tribute to Cluade Shannon (1916–2001) and a plea for more rigorous use of species richness, species diversity, and the “ShannonWiener” Index. Glob Ecol Biogeogr 12:177–179. doi:10.1046/j.1466-822X.2003.00015.x
Stoltz E, Greger M (2002) Accumulation properties of As, Cd, Cu, Pb and Zn by four wetland plant species growing on submerged mine tailings. Environ Exp Bot 47:271–280
Sun X, Ding Q, Hyde KD, Guo LD (2012) Community structure and preference of endophytic fungi of three woody plants in a mixed forest. Fungal Ecol 5:624–632. doi:10.1016/j.funeco.2012.04.001
Suryanarayanan TS, Murali TS, Thirunavukkarasu N, Rajulu MBG, Venkatesan G, Sukumar R (2011) Endophytic fungal communities in woody perennials of three tropical forest types of the Western Ghats, southern India. Biodivers Conserv 20:913–928
Szécsi Á, Magyar D, Tóth S, Szõke C (2013) Poaceae: a rich source of endophytic fusaria. Acta Phytopathol Entomol Hung. doi:10.1556/APhyt.48.2013.1.2
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. doi:10.1093/molbev/mst197
Todaka D, Shinozaki K, Yamaguchi-Shinozaki K (2015) Recent advances in the dissection of drought-stress regulatory networks and strategies for development of drought-tolerant transgenic rice plants. Front Plant Sci 6:84. doi:10.3389/fpls.2015.00084
U’Ren JM, Lutzoni F, Miadlikowska J, Laetsch AD, Arnold AE (2012) Host and geographic structure of endophytic and endolichenic fungi at a continental scale. Am J Bot 99:898–914
Van der Putten WH, Klironomos JN, Wardle DA (2007) Microbial ecology of biological invasions. ISME J 1:28–37
Van Ryckegem G, Verbeken A (2005) Fungal diversity and community structure on Phragmites australis (Poaceae) along a salinity gradient in the Scheldt estuary (Belgium). Nova Hedwigia 80:173–197
Vandenkoornhuyse P, Baldauf SL, Leyval C, Straczek J, Young JPW (2002) Extensive fungal diversity in plant roots. Science 295:2051
Venkatachalam A, Thirunavukkarasu N, Suryanarayanan TS (2015) Distribution and diversity of endophytes in seagrasses. Fungal Ecol 13:60–65
Vymazal J, Krőpfelova L (2005) Growth of Phragmites australis and Phalaris arundinacea in constructed wetlands for wastewater treatment in the Czech Republic. Ecol Eng 25:606–621. doi:10.1016/j.ecoleng.2005.07.005
Waller F, Achatz B, Baltruschat H et al (2005) The endophytic fungus Piriformospora indica reprograms barley to salt-stress tolerance, disease resistance, and higher yield. Proc Natl Acad Sci USA 102:13386–13391. doi:10.1073/pnas.0504423102
Wearn JA, Sutton BC, Morley NJ, Gange AC (2012) Species and organ specificity of fungal endophytes in herbaceous grassland plants. J Ecol 100:1085–1092
Weis JS, Weis P (2004) Metal uptake, transport and release by wetland plants: implications for phytoremediation and restoration. Environ Int 30:685–700
White TJ, Bruns T, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetic. In: Innis MA, Gelfald DH, Sninsky JJ, White TJ (eds) PCR Protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Windham L, Meyerson LA (2003) Effects of common reed (Phragmites australis) expansions on nitrogen dynamics of tidal marshes of the northeastern US. Estuaries 26:452–464
Wirsel SGR, Leibinger W, Ernst M, Mendgen K (2001) Genetic diversity of fungi closely associated with common reed. New Phytol 149:589–598
Yin L, Ren A, Wei M, Wu L, Zhou Y, Li X, Gao Y (2014) Neotyphodium coenophialum-infected tall fescue and its potential application in the phytoremediation of saline soils. Int J Phytorem 16:235–246. doi:10.1080/15226514.2013.773275
You YH, Yoon H, Kang SM, Shin JH, Choo YS, Lee IJ, Lee JM, Kim JG (2012) Fungal diversity and plant growth promotion of endophytic fungi from six halophytes in suncheon bay. J Microbiol Biotechnol 22:1549–1556
Zhang XY, Bao J, Wang GH, He F, Xu XY, Qi SH (2012) Diversity and antimicrobial activity of culturable fungi isolated from six species of the south China sea gorgonians. Microb Ecol 64:617–627. doi:10.1007/s00248-012-0050-x
Zhao J, Shan T, Mou Y, Zhou L (2011) Plant-derived bioactive compounds produced by endophytic fungi. Mini Rev Med Chem 11:159–168
Acknowledgments
The Federal University of Mato Grosso (UFMT), Department of Plant Biology and Pathology of Rutgers University; The Brazilian National Council for Scientific and Technological Development (CNPq) for Post Doctoral Fellowship; International Institute of Science and Technology in Wetlands (INAU). The authors are also grateful to support from the John E. and Christina C. Craighead Foundation, USDA-NIFA Multistate Project W3147 and the New Jersey Agricultural Experiment Station. Any use of trade, product or firm names is for descriptive purposes only and does not imply endorsement by the U.S. Government. This article is Contribution 1957 of the USGS Great Lakes Science Center.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Guest editors: Laura A. Meyerson and Kristin Saltonstall/Phragmites invasion.
Rights and permissions
About this article
Cite this article
Soares, M.A., Li, HY., Kowalski, K.P. et al. Evaluation of the functional roles of fungal endophytes of Phragmites australis from high saline and low saline habitats. Biol Invasions 18, 2689–2702 (2016). https://doi.org/10.1007/s10530-016-1160-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-016-1160-z