Abstract
Objective
To determine bacteriocin producers and the prevalence of structural enterocin genes and to detect the spectrum of activity against foodborne pathogens, from isolates of Enterococcus faecium and Enterococcus faecalis that were isolated from food and the environment.
Results
The entA, entB, entP, ent1071 and entX genes, which encode enterocins were the most frequently observed. Enterocins were thermostable, proteinaceous, and resistant to catalase. None of the isolates produced hemolysin, and inhibition resulting from bacteriophage lysis was excluded. The bactericidal effect of enterocins against L. innocua 12612 was determined by optical density and colony forming units. For the activity spectrum, elimination of mainly Listeria strains, Bacillus sp. and clinical enterococci, was observed. Imaging with scanning electron microscopy after treatment with enterocin Efm22 showed irregular rod-shaped cells and loss of cellular integrity.
Conclusions
The isolates evaluated in this study are candidates for the production of enterocins that will be used as food biopreservatives, because they have high anti-listerial activity even after 24 h of experimentation, and used in the pharmaceutical area because they inhibit clinical microorganisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Food safety is an important global issue because of increasing foodborne diseases and changes in food consumption habits. Therefore, the need to avoid economic losses resulting from microbial-induced spoilage and the preservation of foods by natural methods may be a satisfactory approach to solve many of the current food-related issues (Kaur and Garg 2013).
The development of biopreservation technologies with lactic acid bacteria (LAB) and/or their metabolites represents an additional hurdle in the protection of food against microbial contamination because these bacteria produce several antimicrobial substances, including organic acids, hydrogen peroxide and bacteriocins (Perin et al. 2013). Many bacteriocin-producing LAB strains have been isolated from milk, plants and fermented dairy, vegetable and meats products, many of which have been identified and characterized (Zhang et al. 2018). Studies have demonstrated that bacteriocin from LAB has considerable inhibitory activity against pathogenic and spoilage microorganisms in food such as L. monocytogenes (Zommiti et al. 2018).
Contrary to other bacteriocins, enterocins (produced by Enterococcus sp.) have attracted technological and scientific interest because they exhibit antimicrobial activity against important foodborne pathogens, included Gram positive and negative bacteria, making this peptide of great interest to industry (Schittler et al. 2019). In addition, the search for new enterocin-producing isolates against specific targets or greater action spectra has been constantly highlighted.
Enterocins are found within class I, IIa, IIc and III bacteriocins (Masias et al. 2017), and the cytoplasmic membrane is their primary target. Similar to most bacteriocins, they form pores and thereby deplete transmembrane potential and/or a pH gradient, resulting in cell death. Enterocins show high activity, particularly against Listeria species at low concentrations (Schittler et al. 2019). However, low levels of bacteriocins secreted from natural strains do not meet the requirements of industrial-scale production and have limited applications.
The Listeria genus can be found as a contaminant in meat, milk and other food processing facilities, and the most important species is L. monocytogenes, which causes septicemia, meningitis, encephalitis or death/stillbirth of neonates, especially in high-risk groups in humans (including immunocompromised persons and the elderly) (Khademi and Sahebkar 2019).
Relevant information that must be investigated includes possible bacteriocins that the strains are able to produce, which can be assessed by the identification of specific genes that are related to known bacteriocins and thermal and protease resistance, followed by the inhibitory spectrum (Perin et al. 2013). These data can justify further studies with purified bacteriocins to investigate the diversity of characteristics that allow their use in the food industry as biopreservatives.
In this study, we focused on the isolation and analysis of inhibitory effects of enterocins that are produced by Enterococcus spp. isolates that were obtained from food and environment where common foodborne pathogens strains originate. The inhibitory effect of cell-free culture supernatants (CFSs) was tested against several foodborne microorganisms.
Materials and methods
Bacterial strains, storage conditions and inoculum
Five Enterococcus faecium (Efm20, Efm22, Efm24, Efm25 and Ent22) and two E. faecalis (Efs27, Efs18) were used. Strains Efm20, Efm22, Efm24, Efm25 and Efs27 were obtained from distinct food (soft cheese) samples over a period of 1 year from 2011 to 2012 as described elsewhere (Furlaneto-Maia et al. 2014). E. faecium Ent22 and E. faecalis Efs18 were obtained from environmental samples (water). These strains were identified using molecular approaches, as previously described (Furlaneto-Maia et al. 2014). Strains demonstrated in vitro antagonistic activity against the indicator strain of Listeria innocua (Ogaki et al. 2016). Other strains were foodborne pathogens and food spoilage (Table 3). Strains belong to the Laboratory of Basic and Applied Microbiology (LAMBA) of the Federal University of Technology—Paraná (Londrina, PR, Brazil) and were maintained at − 80 °C. Before use, frozen stock was inoculated into 10 mL in De Man, Rogosa and Sharpe (MRS) (Enterococcus strains) and brain heart infusion (BHI) broth (Neogen Culture Media, USA) and incubated at 37 °C for 24 h.
Genotyping of genes encoding enterocins
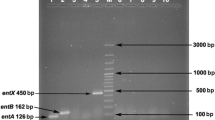
The strains selected were submitted to molecular identification of enterocin-producing genes, which included enterocin A (entA), enterocin B (entB), enterocin P (entP), enterocin L50A/B (entL50A/B), enterocin 1071 (ent1071), enterocin Q (entQ), mundticin KS (entKS), enterocin X (entX), enterocin 31 (ent31) and enterocin AS48 (entAS48), and they were amplified using PCR primers (Table 1). The PCR reactions were performed as previously described in a thermocycler (Esco Technologies, USA). Gene amplification was conducted with an initial denaturation at 95 °C for 5 min, followed by 30 cycles at 95 °C for 30 s, matching the oligonucleotide (Thermo Fisher Scientific, USA) (Table 1) for 30 s, 72 °C for 30 s, and a final extension at 72 °C for 5 min. Amplicons were analyzed on a 1.0% agarose gel (Merck, Germany).
Production and partial purification of enterocins from cell free supernatant
Cell-free supernatants (CFS) of the isolates were previously selected and assayed as described by Tomé et al. (2009), with modifications. The strains were cultured in MRS medium (Neogen Culture Media, USA) overnight, adjusted to 1.0 × 108 CFU mL−1 in MRS (pH 6.2) and maintained at 180 rpm for 18 h at 37 °C. The cultures were centrifuged at 5000 rpm for 15 min, the supernatant was adjusted to pH 6.5 with 1 N NaOH. Partial purification of the enterocins was performed as described by Rocha et al (2019), using 40% saturated ammonium sulfate (Merck). Quantitative determination (UA mL−1) of the partially purified enterocins was conducted as previously described.
Antimicrobial activity and determination of the arbitrary units
In an agar well diffusion assay (AWDA), 30 μL of the enterocin were deposited in 5 mm wells on BHI agar containing L. innocua 12612 (1 × 108 CFU/mL). Finally, the plates were incubated for 24 h at 37 °C. An inhibition halo ≥ 2 mm was considered to be a positive result. Each condition was tested in duplicate. The inhibitory activity of enterocin against L. innocua 12612 was quantified and expressed as arbitrary units (AU) per milliliter. For this experiment, enterocin at 1:2 (v/v) dilutions were deposited onto microplates using MRS. Then, 100 μL of each dilution were deposited onto a new microplate with 100 μL of the indicator bacterium (108 CFU/mL) and incubated at 37 °C for 12 h. Bacterial growth was measured using optical density (OD) in a spectrophotometer (Bio Tek, USA) (600 nm) every 3 h. The arbitrary unit per mL (AU/mL) was defined as the reciprocal of the last dilution that showed growth compared to the control (bacteria without CFS) multiplied by 100 at 6 and 12 h. The OD values were evaluated using one-way ANOVA and Tukey's test, and p < 0.05 was considered a statistically significant difference between the antimicrobial activity of an isolate and the control.
Effect of heat and enzymes on enterocin activity
The thermostability of partially concentrated enterocins was evaluated by the treatment at 80 °C for 10 min and 100 °C for 20 min. To determine the sensitivity of the antimicrobial components against proteolytic enzymes, the selected enterocin were treated with α-chymotrypsin (50 mg/mL), protease (50 mg/mL) and trypsin (20 mg/mL) enzymes (Merck) in addition to the catalase enzyme (Merck) at a final concentration of 1 mg/mL. Then, the antagonistic activity of the treated enterocin was evaluated using the AWDA assay against L. innocua 12612. Each condition was tested in duplicate.
Mode of inhibition
The antimicrobial effect of partially purified enterocins was evaluated using L. innocua 12612, as described by Rocha et al. (2019) with modifications. The indicator bacteria was adjusted to 1.5 × 108 CFU/mL in MRS medium at an OD of 600 nm at 37 °C. Culture medium (10 mL) was placed into tubes, and 0.5 mL of partially purified enterocin was added, except for the control, which only contained ultrapure water (MilliQ). The OD and cell count (CFU/mL) were measured at 0, 1, 2, and 4 h of incubation time. The cell counts were determined on BHI agar medium.
Spectrum of antimicrobial activity
Enterocins selected by the previous experiments against L. innocua 12612 were used to evaluate their activity spectrum against 24 indicator bacteria (Table 2). For this experiment, the antagonistic activity of enterocins was evaluated using a diffusion technique in agar as previously described. The antimicrobial spectrum was determined quantitatively against L. innocua 12612 and qualitatively against other bacteria.
Hemolytic and bacteriophage activity
The hemolytic activity of enterococci isolates was analyzed as described by Eaton and Gasson (2001) in BHI supplemented with 5% sheep blood (Newprov, Brazil). The bacteriophage activity was analyzed as described by Ogaki et al. (2016). Each condition was tested in duplicate.
Scanning electron microscopy (SEM)
The L. innocua 12612 and B. subtilis cells grown for 18 h in the presence of enterocin were washed twice with PBS and then fixed with 2.5% glutaraldehyde (Electron Microscopy Sciences) in 0.1 M sodium cacodylate buffer for 18 h at 4 °C. The samples were carefully washed with 0.1 M sodium cacodylate buffer, and post-fixation was performed for 1 h at 25 °C with 1% osmium tetroxide in 0.1 M sodium cacodylate buffer. Samples were gently dehydrated in graded ethanol (50% to 100% ethanol), critical-point-dried in CO2 (BALTEC DCP 030 Critical Point Dryer), coated with gold (BALTEC SDC 050 Sputter Coater) and viewed in a FEI Quanta 200 Scanning Electron Microscope.
Results and discussion
The presence of ten common structural enterocin genes, singly or in varying combinations, in the genome of Enterococcus spp. was tested. All seven enterococci, E. faecium 20 (Efm20), E. faecium 22 (Efm22), E. faecium 24 (Efm24), E. faecium 25 (Efm25), E. faecium 22 (Ent22), E. faecalis 27 (Efs27) and E. faecalis 18 (Efs18) harbored at least one enterocin-encoding gene (Table 2), of which 71% harbored entA, entB, entP and entX genes, concomitantly. Strains comprising single or multiple-enterocin-encoding genes may possess unique combinations of beneficial and desirable biotechnological properties, particularly antimicrobial activity.
The structural enterocin genes entA, entB, entP and entX were the most frequently observed. E. faecium isolates that had the most complex enterocin gene profile (entA/entB/entP/entX) were capable of causing weak inactivation of Listeria strains, corroborating the data from Rocha et al. (2019). Vandera et al. (2018) have observed the genomes of E. faecalis and E. faecium, which indicated the presence of entA, entB and entP genes. The genes entA and entB are used by the same carrier responsible for externalizing enterocin, and they are controlled by the same regulatory system and commonly found together (Hassan et al. 2012). Differently, genes encoding for enterocin L50A/B, Q, K, 31 and AS48 were not found in any of the enterococci evaluated.
Enterocins are commonly classified as class II bacteriocins and are characterized as small, non-lantibiotics with a strong anti-listerial effect, which are desired characteristics for their use in foods (Masias et al. 2017); genes entA and entP are related to anti-listerial activity with the presence of class IIa bacteriocins (Avcı and Özden 2017). Hassan et al. (2012) observed the presence of the ent1071 and entL50A/B genes, which differs from the results in this study. The presence of four enterocin genes in several strains (Table 2) indicates their potential to produce various enterocins.
In this study enterocins were characterized as proteinaceous in nature, with the inhibition of hydrogen peroxide activity through the use of catalase, because of the lost of antibacterial activity after treatment with α-chymotrypsin, protease and trypsin. In addition, the studied enterocins were thermally stable at both evaluated temperatures since they had the same halo measurements relative to the untreated control.
None of the strains showed hemolytic activity or bacteriophage activity. These characteristics were observed by other authors (Tomé et al. 2009; Ben Braïek et al. 2018). Studies to evaluate the strain safety, especially the inability to cause hemolysis using sheep blood, have been conducted successfully for Enterococcus spp. (Zhang et al. 2016; Vandera et al. 2020). Hemolysin production may increase the risk of enterococcal infections (Schittler et al. 2019).
The activity spectrum of enterocins that were characterized and selected against L. innocua 12612 was evaluated against 23 other indicator bacteria (Table 3). The enterocins showed anti-listerial activity against L. innocua, L. monocytogenes, L. ivanovii, E. faecalis 29112 and Bacillus strains. The anti-listerial activity of CFSs from E. faecalis and E. faecium has been observed in reports by several authors, including Hassan et al. (2012), Jaouani et al. (2014) and Vandera et al. (2018).
These enterocins were observed to more efficiently inhibit Listeria and Enterococcus sp. isolates; B. subtilis was also sensitive to the action of enterocins Efm20, Efm22 and Efm25, with inhibitory activity against 62% of foodborne strains.
Although L. monocytogenes is well known for its pathogenicity, L. ivanovii has been well characterized by infection in ruminants, and it is considered a sporadic occurrence, an infection caused by L. ivanovii in immunocompromised humans has been observed (Khademi and Sahebkar 2019). In addition, no inhibitory activity of enterocins against Gram-negative bacteria was observed, which corroborated with the observations of Tomé et al. (2009) and Ben Braïek et al. (2018), and is likely due to the presence of the outer membrane that hinders the entry of enterocins. Notably, there was no inhibition of bacterial that produced enterocin, showing that there is immunity in the producing cells (data not shown). Immunity of bacteriocin-producing cells is conferred by a peptide expressed along with bacteriocin (Benz and Meinhart 2014; Ogaki et al. 2016).
For determination of the arbitrary unit activity (AU), partially purified enterocins produced by all seven enterococci strains were used. As shown in Fig. 1a, high AU values were observed, indicating that the enterocins identified in this study had high bactericidal activity. There was an increase in the AU values from 6 to 12 h (400 to 6400 AU/mL, respectively) for most of enterocins that were tested.
These data are a good indication of the action of these enterocins over time because their main attribute as food biopreservatives is to remain active as long as possible for the safety of food products. In addition, their susceptibility to proteases demonstrates that they are easily digested by enzymes in the gastrointestinal tract without affecting the normal microbiota.
The effect of enterocin activity as bactericidal substances was evaluated against L. innocua 12612 (Fig. 2). Our data showed that bacteriocins of enterococci were effective at decreasing L. innocua cell viability, in the 0–4 h period, except for the enterocin that was obtained from Efs18 h, and this effect was observed after 2 h. The bactericidal characteristic was confirmed because there was a decrease in OD and viability measured by CFU. L. innocua was previously deemed to be a suitable biological indicator for L. monocytogenes (Rocha et al. 2019) and the strain showed a similar sensitivity to the bacteriocin as eight other L. monocytogenes isolates that were tested. It was used as a pathogen surrogate throughout the study.
Although several enterocins differ in their molecular structures, the mode of bactericidal action is similar. Contrary to Jaouani et al. (2014), in this study, there was a correlation between the presence of enterocin structural genes as well as the values of arbitrary units and the activity spectrum of bacteriocinogenic strains.
The enterocins that were evaluated in this study showed characteristics that were of interest as well as potential activity against clinical isolates and species that are known as contaminants and pathogens in food, which make them potential food biopreservatives. This can help to increase the shelf life of food as well as the food supply and human safety.
Proper evaluation of the potential application of these isolates and their antimicrobial peptides in food processing should be based on their activity and mode of action under conditions that reproduce those used in food products, as performed by Pingitore et al. (2012) and Mogoşanu et al. (2017). However, Maia et al. (2019) showed that the conditions of food products that did not affect the antagonistic action of enterocins Efm20 and Efm22.
SEM analyses revealed that exposure of L. innocua and B. subtilis to bacteriocin Efm22 resulted in a coarse, collapsed surface of the cell with surface protuberances that may indicate cellular leakages (Fig. 3b and d). According to Masias et al. (2017) class-IIa bacteriocins bind to bacterial cell envelope associated mannose phosphotransferase system, leading to pore formation. Our results suggest that bacteriocins produced by enterococci represent potential character for promising future applications to control pathogenic Listeria species. The SEM images showed that non-treated Listeria and Bacillus cells had typical rod shapes with smooth surfaces (Fig. 3a and c), while enterocin-treated cells displayed alterations resembling cell membrane damage and the presence of extracellular material (Fig. 3b and d).
The results obtained provide new insights on antibacterial activity produced by E. faecium and E. faecalis strains against food borne and spoilage bacteria, concomitantly.
Conclusions
The antimicrobial capacity of enterococci has been well studied because of the search for alternative forms of antimicrobials and food biopreservatives in the pharmaceutical and food industries, respectively. The antimicrobial activity that was observed in the supernatants from enterococcal cultures against bacteria that are important in the contamination and pathogenicity of food and against bacteria of clinical origin makes these isolates promising candidates in an alimentary and/or pharmaceutical context.
References
Avci M, Özden TB (2017) Safety evaluation of enterocin producer Enterococcus sp. strains isolated from traditional turkish cheeses. Pol J Microbiol 6:223–233
Ben Braïek O, Cremonesi P, Morandi S, Smaoui S, Hani K, Ghrairi T (2018) Safety characterisation and inhibition of fungi and bacteria by a novel multiple enterocin-producing Enterococcus lactis 4CP3 strain. Microb Pathogenesis 118:32–38. https://doi.org/10.1016/j.micpath.2018.03.005
Benz J, Meinhart A (2014) Antibacterial effector/immunity systems: it's just the tip of the iceberg. Curr Opin Microbiol 17:1–10. https://doi.org/10.1016/j.mib.2013.11.002
Citti L, Rovero P, Colombo MG, Mariani L, Poliseno L, Rainaldi G (2002) Efficacy of an amphipathic oligopeptide to shuttle and release a cis-acting DNA decoy into human cells. Biotechniques 32:172–177. https://doi.org/10.3389/fmicb.2015.00782
De Vuyst L, Moreno MF, Revets H (2003) Screening for enterocins and detection of hemolysin and vancomycin resistance in enterococci of different origins. Int J Food Microbiol 84:299–318. https://doi.org/10.1016/s0168-1605(02)00425-7
Du Toit M, Franz CMAP, Dicks LMT, Holzapfel WH (2000) Preliminary characterization of bacteriocins produced by Enterococcus faecium and Enterococcus faecalis isolated from pig faeces. J Appl Microbiol 88:482–494. https://doi.org/10.1046/j.1365-2672.2000.00986.x
Eaton TJ, Gasson MJ (2001) Molecular screening of Enterococcus virulence determinants and potential for genetic exchange between food and medical isolates. Appl Environ Microbiol 67:1628–1635. https://doi.org/10.1128/AEM.67.4.1628-1635.2001
Edalatian MR, Najafi MBH, Mortazavi SA, Alegría Á, Delgado S, Bassami MR, Mayo B (2012) Production of bacteriocins by Enterococcus spp. isolated from traditional, Iranian, raw milk cheeses, and detection of their encoding genes. Eur Food Res Technol 234:789–796. https://doi.org/10.1007/s13594-011-0045-2
Furlaneto-Maia L, Rocha KR, Henrique FC, Giazzi A, Furlaneto MC (2014) Antimicrobial resistance in Enterococcus sp isolated from soft cheese in Southern Brazil. Adv Microbiol 4:175–181. https://doi.org/10.4236/aim.2014.43023
Hassan M, Diep DB, Javadzadeh Y, Dastmalchi S, Nes IF, Sharifi Y, Saber Y, Safar F, Lotfipour F (2012) Prevalence of bacteriocin activities and bacteriocin-encoding genes in enterococcal clinical isolates in Iran. Can J Microbiol 58:359–368. https://doi.org/10.1139/W11-136
Jaouani I, Abbassi MS, Alessandria V, Bouraoui J, Ben SR, Kilani H, Mansouri R, Messadi L, Cocolin L (2014) High inhibition of Paenibacillus larvae and Listeria monocytogenes by Enterococcus isolated from different sources in Tunisia and identification of their bacteriocin genes. Lett Appl Microbiol 59:17–25. https://doi.org/10.1111/lam.12239
Kaur B, Garg N (2013) Characteristics of bacteriocin BA28 produced by Pediococcus acidi lactici BA28. Mintage J Pharm Med Sci 2:17–20
Khademi F, Sahebkar A (2019) A systematic review and meta-analysis on the prevalence of antibiotic-resistant Listeria species in food, animal and human specimens in Iran. J Food Sci Technol 56:5167–5183. https://doi.org/10.1007/s13197-019-04040-w
Maia LF, Costa LC, Rocha KR, Schueller J, Tosoni NF, Furlaneto MC (2019) Influence of optimised commercial medium on bacteriocin production by Enterococcus faecium. Acta Sci Technol 41:e42324. https://doi.org/10.4025/actascitechnol.v41i1.42324
Martin M, Gutierrez J, Criado R, Herranz C, Cintas LM, Hernandez PE (2006) Genes encoding bacteriocins and their expression and potential virulence factors of enterococci isolated from wood pigeons (Columba palumbus). J Food Prot 69:520–531. https://doi.org/10.4315/0362-028x-69.3.520
Masias E, Dupuy FG, da Silva Sanches PR, Farizano JV, Cilli E, Bellomio A, Minahk C (2017) Impairment of the class IIa bacteriocin receptor function and membrane structural changes are associated to enterocin CRL35 high resistance in Listeria monocytogenes. BBA 1861:1770–1776. https://doi.org/10.1016/j.bbagen.2017.03.014
Mogoşanu GD, Grumezescu AM, Bejenaru C, Bejenaru LE (2017) Natural products used for food preservation, food preservation. Elsevier, Amsterdam, pp 365–411. eBook ISBN: 9780128043745
Ogaki MB, Rocha KR, Terra MR, Furlaneto MC, Furlaneto-Maia L (2016) Screening of the enterocin-encoding genes and antimicrobial activity in Enterococcus species. J Microbiol Biotechnol 26:1026–1034. https://doi.org/10.4014/jmb.1509.09020
Özdemir GB, Oryaşın E, Bıyık HH, Özteber M, Bozdoğan B (2011) Phenotypic and genotypic characterization of bacteriocins in enterococcal isolates of different sources. Indian J Microbiol 51:182–187. https://doi.org/10.1007/s12088-011-0143-0
Perin LM, Miranda RO, Camargo AC, Colombo M, Carvalho AF, Nero LA (2013) Antimicrobial activity of the Nisin Z producer Lactococcus lactis subsp. lactis Lc08 against Listeria monocytogenes in skim milk. Arq Bras Med Vet Zootec 65:1554–1560. https://doi.org/10.1590/S0102-09352013000500037
Pingitore EV, Todorov SD, Sesma F, Franco BDGM (2012) Application of bacteriocinogenic Enterococcus mundtii CRL35 and Enterococcus faecium ST88Ch in the control of Listeria monocytogenes in fresh Minas cheese. Food Microbiol 32:38–47. https://doi.org/10.1016/j.fm.2012.04.005
Rehaiem A, Martínez B, Manai M, Rodríguez A (2012) Technological performance of the enterocin A producer Enterococcus faecium MMRA as a protective adjunct culture to enhance hygienic and sensory attributes of traditional fermented milk ‘Rayeb’. Food Bioprocess Technol 5:2140–2150. https://doi.org/10.1007/s11947-010-0501-7
Rocha KR, Perini HF, Souza CM, Schueler J, Tosoni NF, Furlaneto MC, Furlaneto-Maia L (2019) Inhibitory effect of bacteriocins from enterococci on developing and preformed biofilms of Listeria monocytogenes, Listeria ivanovii and Listeria innocua. World J Microb Biot 35:96–106. https://doi.org/10.1007/s11274-019-2675-0
Schittler L, Perin LM, de Lima MJ, Lando V, Todorov SD, Nero LA, da Silva WP (2019) Isolation of Enterococcus faecium, characterization of its antimicrobial metabolites and viability in probiotic Minas Frescal cheese. J Food Sci Technol 56:5128–5137. https://doi.org/10.1007/s13197-019-03985-2
Tomé E, Todorov SD, Gibbs PA, Teixeira PC (2009) Partial characterization of nine bacteriocins produced by lactic acid bacteria isolated from cold-smoked salmon with activity against Listeria monocytogenes. Food Biotechnol 23:50–73. https://doi.org/10.1080/08905430802671956
Vandera E, Tsirka G, Kakouri A, Koukkou AI, Samelis J (2018) Approaches for enhancing in situ detection of enterocin genes in thermized milk, and selective isolation of enterocin-producing Enterococcus faecium from Baird-Parker agar. Int J Food Microbiol 281:23–31. https://doi.org/10.1016/j.ijfoodmicro.2018.05.020
Vandera E, Parapouli M, Kakouri A, Koukkou AI, Hatziloukas E, Samelis J (2020) Structural enterocin gene profiles and mode of antilisterial activity in synthetic liquid media and skim milk of autochthonous Enterococcus spp. isolates from artisan Greek Graviera and Galotyri cheeses. Food Microbiol 86:103335.
Zhang F, Jiang M, Wan C, Chen X, Chen X, Tao X, Shah NP, Wei H (2016) Screening probiotic strains for safety: evaluation of virulence and antimicrobial susceptibility of enterococci from healthy Chinese infants. Dairy Sci 99:4282–4290. https://doi.org/10.3168/jds.2015-10690
Zhang J, Yang Y, Yang H, Bu Y, Yi H, Zhang L, Han X, Ai L (2018) Purification and partial characterization of bacteriocin Lac-B23, a novel bacteriocin production by Lactobacillus plantarum J23, isolated from Chinese traditional fermented milk. Front Microbiol 9:2165. https://doi.org/10.3389/fmicb.2018.02165
Zommiti M, Cambronel M, Maillot O, Barreau M, Sebei K, Feuilloley M, Ferchichi M, Connil N (2018) Evaluation of probiotic properties and safety of Enterococcus faecium isolated from artisanal tunisian meat “Dried Ossban”. Front Microbiol 6:1685. https://doi.org/10.3389/fmicb.2018.01685
Acknowledgements
This work was supported by Fundação Araucária/ Governo do Paraná – Brazil, PROPPG/ UTFPR. This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brasil (CAPES)—Finance Code 001.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest. The founding sponsors had no role in the design of the study, nor in the data collection, analyses, or interpretation of data, the writing of the manuscript, nor the decision to publish the results.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Furlaneto-Maia, L., Ramalho, R., Rocha, K.R. et al. Antimicrobial activity of enterocins against Listeria sp. and other food spoilage bacteria. Biotechnol Lett 42, 797–806 (2020). https://doi.org/10.1007/s10529-020-02810-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-02810-7