Abstract
Gene therapy is based on the principle of the genetic manipulation of DNA or RNA for treating and preventing human diseases. The clustered regularly interspaced short palindromic repeats/CRISPR associated nuclease9 (CRISPR/Cas9) system, derived from the acquired immune system in bacteria and archaea, has provided a new tool for accurate manipulation of genomic sequence to attain a therapeutic result. The advantage of CRISPR which made it an easy and flexible tool for diverse genome editing purposes is that a single protein (Cas9) complex with 2 short RNA sequences, function as a site-specific endonuclease. Recently, application of CRISPR/Cas9 system has become popular for therapeutic aims such as gene therapy. In this article, we review the fundamental mechanisms of CRISPR-Cas9 function and summarize preclinical CRISPR-mediated gene therapy reports on a wide variety of disorders.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Studies have shown that gene therapy is both much more challenging and technically complex than the simple idea of gene replacement. Many of these challenges are based on the ability to accurately control the introduction of genetic material into target cells. Techniques for adding exogenous genes have had remarkable improvements so far, however, there are still a few challenges including unforeseeable effects on gene expression and unpremeditated effects on adjacent genes, limitations in the delivery size of the transgene and finally the addition of exogenous genes cannot always directly address dominant mutations or eliminate undesirable genetic material such as viral genomes or receptors. Targeted gene editing with programmable nucleases, such as zinc finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs), and clustered regularly interspaced short palindromic repeats/CRISPR-associated nuclease 9 (CRISPR-Cas9) has emerged to make accurate, targeted corrections to genome sequences for resolving these fundamental limitations of conventional methods for gene addition. CRISPR-Cas9 is derived from the type II acquired immune system in bacteria and archaea and presents several advantages upon its counterparts and displays therapeutic potentials (Maeder and Gersbach 2016; Teimourian and Abdollahzadeh 2015). Unlike ZFNs and TALENs, CRISPR/Cas9 for targeting different regions of the genome do not require the engineering of novel proteins also, able to do efficient multiplex genome editing and has high efficiency of introducing DSBs. In this article, we review the fundamental mechanisms of CRISPR-Cas9 in genome editing and summarize preclinical CRISPR-mediated therapy reports on a wide variety of diseases and disorders (Table 1).
Mechanism of CRISPR-Cas9 in genome editing
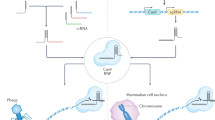
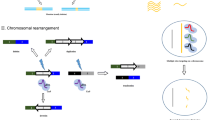
Generally, CRISPR/Cas9 works in a similar manner with other programmable nucleases, through RNA-directed endonuclease cleavage of the invading genomic sequence. Short sequences of invading DNA elements derived from phages or plasmids are incorporated into the host genome to form a CRISPR locus flanked by a repeat sequence. Upon subsequent infection, this locus is transcribed and processed into crRNA that harbors a short area of homology (≈ 20 bp) and guides the complex to a special genetic locus. crRNA in cooperation with tracrRNA forms an RNA structure to recruit Cas9 to identify and cleave target DNA after Watson–Crick base pairing among crRNA and the target DNA. Generally four strategies utilize for double-strand break repair including gene knockout/disruption and gene deletion by NHEJ manner or gene correction and gene knock-in/insertion by HDR manner (Maeder and Gersbach 2016). The CRISPR system, as a powerful tool for targeted manipulation of the genome, has a very high potential for basic and clinical research. Recently, the use of this system has become popular for therapeutic purposes such as gene therapy.
Viral infection
CRISPR-Cas9 system as an adaptive immune defense system has shown therapeutic potential in manipulating and disabling viral genome to obstruct virus infection. There are two strategies to tackle latent HIV infection by CRISPR/Cas9 technology. In the first approach, host genes that needed for HIV-1 T cell infection is targeted. The most advanced genome editing-based antiviral therapy to date is the ablation of CCR5 gene by ex vivo adjustment of T cells and CD34+ hematopoietic stem and progenitor cells (HSPCs). In the second approach viral DNA is targeted for mutating vital sites or cleaving proviral DNA from infected cells. CRISPR/Cas9 strategies for targeting viral genome have also been applied to several other viral pathogens, including human papillomavirus, Hepatitis B virus, Herpesviruses, JC Virus and hepatitis C virus (Soppe and Lebbink 2017).
Cancer therapy
CRISPR-Cas9 technology for treating cancer has so far been used in three areas: cancer immunotherapy, manipulation of cancer genome and epigenome and elimination or inactivation of carcinogenic viral infections.
To confer higher affinity for tumor antigen, T cells have been genetically engineered to express altered αβ T-cell receptors (TCRs). Genome editing by CRISPR-Cas9 system for production of the chimeric antigen receptor (CAR) has been broadly recognized as one of the largest progress in personalized cancer immunotherapy. In CAR T cell therapy, autologous T cells are gathered, genetically engineered to attack cancer antigens ex vivo and then transferred back into the patient. Improving of CAR T cell function could be done by CRISPR-Cas9 system via interrupting the genes that coding T cell inhibitory receptors or signaling molecules (Zych et al. 2018).
Oncogenic virus infections are associated with tumorigenesis, for example, HBV and hepatitis C virus in liver cancer, human papillomavirus (HPV) in cervical cancer and Epstein-Barr virus (EBV) in nasopharyngeal carcinoma. CRISPR-mediated knock out of miRNAs can be used in cancer therapy, since miRNAs are involved in the regulation of cellular processes in both normal and pathological events (Chang et al. 2016). Suppression of the transcriptional activities of DNA elements can be done by binding of catalytically inactivated cas9 (dCas9) to them, a method called CRISPRi. Moreover, dCas9 that does not cleave DNA can fuse to histone modifiers and proteins involved in altering DNA methylation and under the steerage of sgRNAs, dCas9 and the modifiers are directed to the goal sites and perform epigenetic regulations (Fineran and Dy 2014).
Cardiovascular disease
Studies have been shown individuals with naturally occurring loss-of-function mutation in proprotein convertase subtilisin/kexin type 9 (PCSK9) have lower levels of LDLC thus disruption of the PCSK9 gene by CRISPR/Cas9 system is a hoping therapeutic target for the prevention of cardiovascular disease (Wang et al. 2016).
Selectively disrupting disease-causing mutation in WPW syndrome by use of SNP-derived PAM (a pathogenic mutation that leads to the occurrence of a novel PAM) promises an efficient approach in the treatment of WPW syndrome and other dominant inherited diseases (Xie et al. 2016).
Recently, CRISPR/Cas9 system has been used to correct a pathogenic mutation in the MYBPC3 gene in human embryos that causes hypertrophic cardiomyopathy (HCM) (Ma et al. 2017).
Eye disease
Cataracts were the first eye disorder that edited by CRISPR/Cas9 system for therapeutic aim. Wu et al. have corrected a dominant mutation in the Crygc gene via modification of mutant alleles in a CRISPR/Cas9-mediated HDR manner.
Leber Congenital Amaurosis type 10 (LCA10) is the most prevalent type of LCA result from mutations in the CEP290 gene. CRISPR/Cas9-mediated NHEJ approach has been used for deletion of an intronic region in the CEP290 gene comprising a mutation that disrupts the gene coding sequence by generating an aberrant splice site.
Age-related macular degeneration (AMD), Meesmann’s epithelial corneal dystrophy (MECD), and Retinitis Pigmentosa (RP) are the other retinal diseases targeted by CRISPR/Cas9 system (Peddle and MacLaren 2017).
Hematologic disease
Two strategies have been used to treat sickle cell disease (SCD) using CRISPR/Cas9 system: correction of the sickle cell mutation and induction of fetal hemoglobin (HbF) expression. In gene correction approach haematopoietic stem cells derived from patients and corrected Ex vivo by CRISPR/Cas9 system, then transplanted to the patient (Dever et al. 2016). BCL11A is a transcriptional regulator that acts as a strong HbF silencer and suppresses the expression of γ-globin, thus suppression of BCL11A can be used to treat both SCD and β-thalassemia (Canver et al. 2015).
By the use of CRISPR/Cas9 system, β-thalassemia-causing mutations in patient-derived induced pluripotent stem cells (iPSCs) can be corrected and then transplanted the corrected iPSCs to the patient (Liu et al. 2017).
CRISPR/Cas9-mediated HDR is one of the strategies has been used to correct mutation in patients with haemophilia B and group C Fanconi Anemia (FA-C). In another approach, the insertion of an exon 2–8 cDNA into intron 1 of F9 has been used with regard to the subject that the majority of identifying F9 mutations located in the coding sequences of exons 2–8 (Ohmori et al. 2017). Near half of all severe haemophilia A are caused by two large (140 or 600 kb) chromosomal inversions in coagulation Factor VIII (F8) and correction of these inversions in patient-derived iPS cells have been made via transfer of an RNP complex of Cas9 and a pair of sgRNAs for targeting two different positions to induce reinversion of the mutations (Park et al. 2015).
Immunological disorders
Mutations in the JAK3 gene result in an autosomal recessive form of T-B+ SCID (Severe combined immunodeficiency). Recently, Chang et al. by use of CRISPR/Cas9-mediated HDR approach corrected point mutation in exon 14 of the JAK3 gene in patient-derived iPSC (Chang et al. 2015).
Patient-derived iPSC from X-linked chronic granulomatous disease (X-CGD) patients containing point mutations in the CYBB gene repaired by CRISPR/Cas9-mediated HDR approach and then transplant to the patients (De Ravin et al. 2017).
Knockout of miR-155, a key proinflammatory regulator in rheumatoid arthritis (RA), by CRISPR/CAS9 system showed inhibition of proinflammatory cytokine production in macrophage cell line and offered a hoping therapeutic strategy in RA treatment (Jing et al. 2015).
Inborn errors of metabolism (IEM)
Mutation in fumarylacetoacetate hydrolase (FAH) that catalyzes the ultimate step of tyrosine catabolism results in Hereditary Tyrosinemia type I (HT-I). By CRISPR/Cas9-mediated HDR homozygous G → A point mutation of the final nucleotide of exon 8 in FAH has been corrected (Yin et al. 2016). In a new gene therapy strategy, HPD gene deleted in liver via CRISPR/Cas9 system to reprogram the metabolic pathways showed higher therapeutic effect in the treatment of HT-I than directly correcting the pathogenic mutation in FAH (Pankowicz et al. 2016).
Ornithine transcarbamylase (OTC) deficiency, a disease resulting from an X-linked deficiency of the OTC enzyme, leading to hyperammonemia. Yang et al. corrected a G → A point mutation of the OTC gene by co-delivery of two AAV vectors, one carrying a sgRNA and the donor DNA and the other carrying Cas9 nuclease (Yang et al. 2016). Arginase deficiency corrected by addition of arginase 1 cDNA into exon 1 of hypoxanthine–guanine phosphoribosyltransferase (HPRT) locus in iPSCs (Lee et al. 2016).
Muscular dystrophy
The strategies used to treat Duchenne muscular dystrophy (DMD) include: correcting of point mutations, knock-in of the full missing exon, and restoring DMD’s reading frame by deletion of the mutated exons. CRISPR/Cas9-mediated HDR approach has been used for correction of point mutation (Zhu et al. 2017) and the addition of missing exon 44 (Li et al. 2015). Deletion of one or more exons from the mutated transcript can restore the reading frame and generate a truncated but yet functional protein. This approach has been used to remove exon 23, exons 20–23, exons 50–54 and exons 45–55 by using multiple gRNAs that targeted regions surrounding desired exon(s) to generate more than one DSB simultaneously and removing large segments (El Refaey et al. 2017).
Limb-girdle muscular dystrophies types 2B (LGMD2B), types 2D (LGMD2D) and Myotonic dystrophy type 1 (DM1) are other muscular dystrophy that targeted by CRISPR/Cas9 (Turan et al. 2016; van Agtmaal et al. 2017).
Neurological disorders
So far, two strategies for gene therapy of progressive neurodegenerative diseases have been used, including deletion of pathogenic mutations and targeted gene correction. CRISPR/Cas9 system can be used for targeted deletion of the CAG repeats by sgRNA/Cas9 collections that flanked this domain and produced double strand breaks which subsequently induced non-homologous end joining (NHEJ) process (Monteys et al. 2017). Deletion of the open reading frame of HTT can also decrease mutant huntingtin mass (Talan 2015). Friedreich Ataxia and Amyotrophic lateral sclerosis (ALS) are other disease that targeted by CRISPR/Cas9 (Ouellet et al. 2017; Wang et al. 2017).
Respiratory disorders
The ΔF508 mutation is the first mutation identified in CFTR and present in 90% of cystic fibrosis (CF) patients and two studies have been done for the correction of ΔF508 mutation in iPCS via CRISPR/Cas9-mediated HDR approach (Firth et al. 2015). Alpha-1 Antitrypsin Deficiency (AATD) was another respiratory disease that edited by CRISPR/Cas9 system via both HDR based precise gene correction and NHEJ-mediated gene disruption (Smith et al. 2015).
Skin disease
Dystrophic Epidermolysis bullosa, a blistering skin disorder, caused by mutations in the COL7A1 gene and can be inherited in an autosomal recessive (RDEB) or dominant manner (DDEB). CRISPR/Cas9-mediated HDR approach can correct the causative mutation in patient-derived RDEB fibroblasts (Hainzl et al. 2017). Dominant negative mutations in the COL7A1 gene leading to DDEB and reading frame disruption of the mutant allele by CRISPR/Cas9-mediated NHEJ, while wild-type allele remains intact, has been applied for correction of these dominant negative disorder (Shinkuma, Guo, and Christiano 2016).
Commercializing the CRISPR technology
CRISPR/Cas9, as an incredibly powerful technology, will play a key role in how we diagnose and treat disease, thus a number of companies have invested in CRISPR technology. There are three big players in CRISPR-based gene-editing biotech companies including Editas Medicine (editas medicine. 2018), Intellia Therapeutics (Intellia Therapeutics. 2018), and CRISPR Therapeutics (CRISPR Therapeutics .2018). For example, CRISPR Therapeutics approaches classify in four main areas: (i) ex vivo approaches involving gene editing of hematopoietic cells for hemoglobinopathies, (ii) ex vivo approaches in immuno-oncology for cancer, (iii) in vivo approaches targeting the liver (haemophilia) and (iv) additional in vivo approaches targeting other organ systems, such as muscle (DMD) and lung (CF).So, if CRISPR is able to target all monogenic diseases diagnosed every year, this market worth billions dollar globally, also by addition of cancers, this market becomes more valuable.
Challenges and outlooks
CRISPR/Cas9 system showing new era in correcting gene defects to treating a numerous variety of human diseases by gene therapy, but several technical challenges still need to be solved regarding accuracy and efficiency issues. Various approaches have been explored to minimize Cas9-mediated off-target events that are mentioned in referenced papers (Chira et al. 2017; Ren and Zhao 2017). Several approaches for increasing HDR rate have been described, including inhibition of the NHEJ pathway, use of longer homologous donor DNA, cell cycle synchronization techniques for control the transfer time of Cas9 protein and sgRNA complexes (because HDR takes place mainly in S and G2 phases) and targeting non-target DNA strand that was free of protein interaction and located in the PAM – distal (Lin et al. 2014; Gibson and Yang 2017). Also, Homology-Independent Targeted Integration (HITI) approach may solve low HDR efficiency (Nygaard et al. 2016). After solving these current challenges and improvement of the delivery methods, the CRISPR-Cas9 system can apply for clinical applications in patients. Recently this system has been approved by an advisory committee at the US National Institute of Health as the first clinical trial to attack cancer cells (Reardon 2016).
References
Canver MC, Smith EC, Sher F, Pinello L, Sanjana NE, Shalem O, Chen DD, Schupp PG, Vinjamur DS, Garcia SP, Luc S, Kurita R, Nakamura Y, Fujiwara Y, Maeda T, Yuan GC, Zhang F, Orkin SH, Bauer DE (2015) BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature 527:192–197
Chang CW, Lai YS, Westin E, Khodadadi-Jamayran A, Pawlik KM, Lamb LS Jr, Goldman FD, Townes TM (2015) Modeling human severe combined immunodeficiency and correction by CRISPR/Cas9-enhanced gene targeting. Cell Rep 12:1668–1677
Chang H, Yi B, Ma R, Zhang X, Zhao H, Xi Y (2016) CRISPR/cas9, a novel genomic tool to knock down microRNA in vitro and in vivo. Sci Rep 6:22312
Chira S, Gulei D, Hajitou A, Zimta AA, Cordelier P, Berindan-Neagoe I (2017) CRISPR/Cas9: transcending the reality of genome editing. Mol Ther Nucl Acids 7:211–222
De Ravin SS, Li L, Wu X, Choi U, Allen C, Koontz S, Lee J, Theobald-Whiting N, Chu J, Garofalo M, Sweeney C, Kardava L, Moir S, Viley A, Natarajan P, Su L, Kuhns D, Zarember KA, Peshwa MV, Malech HL (2017) CRISPR-Cas9 gene repair of hematopoietic stem cells from patients with X-linked chronic granulomatous disease. Sci Transl Med 9:eaah3480
Dever DP, Bak RO, Reinisch A, Camarena J, Washington G, Nicolas CE, Pavel-Dinu M, Saxena N, Wilkens AB, Mantri S, Uchida N, Hendel A, Narla A, Majeti R, Weinberg KI, Porteus MH (2016) CRISPR/Cas9 beta-globin gene targeting in human haematopoietic stem cells. Nature 539:384–389
El Refaey M, Xu L, Gao Y, Canan BD, Adesanya TMA, Warner SC, Akagi K, Symer DE, Mohler PJ, Ma J, Janssen PML, Han R (2017) In vivo genome editing restores dystrophin expression and cardiac function in Dystrophic mice. Circ Res 121:923–929
Fineran PC, Dy RL (2014) Gene regulation by engineered CRISPR-Cas systems. Curr Opin Microbiol 18:83–89
Firth AL, Menon T, Parker GS, Qualls SJ, Lewis BM, Ke E, Dargitz CT, Wright R, Khanna A, Gage FH, Verma IM (2015) Functional gene correction for cystic fibrosis in lung epithelial cells generated from patient iPSCs. Cell Rep 12:1385–1390
Gibson GJ, Yang M (2017) What rheumatologists need to know about CRISPR/Cas9. Nat Rev Rheumatol 13:205–216
Hainzl S, Peking P, Kocher T, Murauer EM, Larcher F, Del Rio M, Duarte B, Steiner M, Klausegger A, Bauer JW, Reichelt J, Koller U (2017) COL7A1 editing via CRISPR/Cas9 in recessive dystrophic Epidermolysis bullosa. Mol Ther 25:2573–2584
Jing W, Zhang X, Sun W, Hou X, Yao Z, Zhu Y (2015) CRISPR/CAS9-mediated genome editing of miRNA-155 inhibits proinflammatory cytokine production by RAW264.7 cells. Biomed Res Int 2015:326042
Lee PC, Truong B, Vega-Crespo A, Gilmore WB, Hermann K, Angarita SA, Tang JK, Chang KM, Wininger AE, Lam AK, Schoenberg BE, Cederbaum SD, Pyle AD, Byrne JA, Lipshutz GS (2016) Restoring ureagenesis in hepatocytes by CRISPR/Cas9-mediated genomic addition to arginase-deficient induced pluripotent stem cells. Mol Ther Nucl Acids 5:e394
Li HL, Fujimoto N, Sasakawa N, Shirai S, Ohkame T, Sakuma T, Tanaka M, Amano N, Watanabe A, Sakurai H, Yamamoto T (2015) Precise correction of the Dystrophin gene in duchenne muscular dystrophy patient induced pluripotent stem cells by TALEN and CRISPR-Cas9. Stem Cell Rep 4:143–154
Lin S, Staahl BT, Alla RK, Doudna JA (2014) Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. Elife 3:e04766
Liu Yali, Yang Yi, Kang Xiangjin, Lin Bin, Qian Yu, Song Bing, Gao Ge, Chen Yaoyong, Sun Xiaofang, Li Xiaoping, Lei Bu, Fan Yong (2017) One-step biallelic and scarless correction of a β-thalassemia mutation in patient-specific iPSCs without drug selection. Mol Ther Nucl Acids 6:57–67
Ma H, Marti-Gutierrez N, Park SW, Wu J, Lee Y, Suzuki K, Koski A, Ji D, Hayama T, Ahmed R, Darby H, Van Dyken C, Li Y, Kang E, Park AR, Kim D, Kim ST, Gong J, Gu Y, Xu X, Battaglia D, Krieg SA, Lee DM, Wu DH, Wolf DP, Heitner SB, Belmonte JCI, Amato P, Kim JS, Kaul S, Mitalipov S (2017) Correction of a pathogenic gene mutation in human embryos. Nature 548:413–419
Maeder ML, Gersbach CA (2016) Genome-editing Technologies for gene and cell therapy. Mol Ther 24:430–446
Monteys AM, Ebanks SA, Keiser MS, Davidson BL (2017) CRISPR/Cas9 editing of the mutant huntingtin allele in vitro and in vivo. Mol Ther 25:12–23
Nygaard S, Barzel A, Haft A, Major A, Finegold M, Kay MA, Grompe M (2016) A universal system to select gene-modified hepatocytes in vivo. Sci Transl Med 8:342
Ohmori T, Nagao Y, Mizukami H, Sakata A, Muramatsu SI, Ozawa K, Tominaga SI, Hanazono Y, Nishimura S, Nureki O, Sakata Y (2017) CRISPR/Cas9-mediated genome editing via postnatal administration of AAV vector cures haemophilia B mice. Sci Rep 7:4159
Ouellet DL, Cherif K, Rousseau J, Tremblay JP (2017) Deletion of the GAA repeats from the human frataxin gene using the CRISPR-Cas9 system in YG8R-derived cells and mouse models of Friedreich ataxia. Gene Ther 24:265–274
Pankowicz FP, Barzi M, Legras X, Hubert L, Mi T, Tomolonis JA, Ravishankar M, Sun Q, Yang D, Borowiak M, Sumazin P, Elsea S, Bissig-Choisat B, Bissig KD (2016) Reprogramming metabolic pathways in vivo with CRISPR/Cas9 genome editing to treat hereditary tyrosinaemia. Nat Commun 7:12642
Park CY, Kim DH, Son JS, Sung JJ, Lee J, Bae S, Kim JH, Kim DW, Kim JS (2015) Functional correction of large factor VIII gene chromosomal inversions in hemophilia A patient-derived iPSCs using CRISPR-Cas9. Cell Stem Cell 17:213–220
Peddle CF, MacLaren RE (2017) The application of CRISPR/Cas9 for the treatment of retinal diseases. Yale J Biol Med 90:533–541
Reardon S (2016) First CRISPR clinical trial gets green light from US panel. Nat Methods 5:374–375
Ren Jiangtao, Zhao Yangbing (2017) Advancing chimeric antigen receptor T cell therapy with CRISPR/Cas9. Protein Cell 8:634–643
Shinkuma S, Guo Z, Christiano AM (2016) Site-specific genome editing for correction of induced pluripotent stem cells derived from dominant dystrophic epidermolysis bullosa. Proc Natl Acad Sci USA 113:5676–5681
Smith C, Abalde-Atristain L, He C, Brodsky BR, Braunstein EM, Chaudhari P, Jang YY, Cheng L, Ye Z (2015) Efficient and allele-specific genome editing of disease loci in human iPSCs. Mol Ther 23:570–577
Soppe JA, Lebbink RJ (2017) Antiviral goes viral: harnessing CRISPR/Cas9 to combat viruses in humans. Trends Microbiol 25(10):833–850
Talan Jamie (2015) News from the Society for Neuroscience Annual Meeting: gene editing techniques show promise in silencing or inhibiting the mutant Huntington’s disease gene. Neurol Today 15:14–16
Teimourian S, Abdollahzadeh R (2015) Technology developments in biological tools for targeted genome surgery. Biotechnol Lett 37:29–39
Turan S, Farruggio AP, Srifa W, Day JW, Calos MP (2016) Precise correction of disease mutations in induced pluripotent stem cells derived from patients with limb girdle muscular Dystrophy. Mol Ther 24:685–696
van Agtmaal EL, Andre LM, Willemse M, Cumming SA, van Kessel IDG, Wjaa van den Broek G, Gourdon D, Furling V, Mouly DG, Monckton DG, Wansink DG (2017) CRISPR/Cas9-Induced (CTGCAG)n repeat instability in the myotonic Dystrophy type 1 locus: implications for therapeutic genome editing. Mol Ther 25:24–43
Wang X, Raghavan A, Chen T, Qiao L, Zhang Y, Ding Q, Musunuru K (2016) CRISPR-Cas9 targeting of PCSK9 in human hepatocytes in vivo-brief report. Arterioscler Thromb Vasc Biol 36:783–786
Wang L, Yi F, Lina F, Yang J, Wang S, Wang Z, Suzuki K, Sun L, Xiuling X, Yang Y, Qiao J, Belmonte JCI, Yang Z, Yuan Y, Jing Q, Liu GH (2017) CRISPR/Cas9-mediated targeted gene correction in amyotrophic lateral sclerosis patient iPSCs. Protein Cell 8:365–378
Xie C, Zhang YP, Song L, Qi W, Jialu H, Danbo L, Yang Z, Zhang J, Xiao J, Zhou B, Du JL, Jing N, Liu Y, Wang Y, Li BL, Song BL, Yan Y (2016) Genome editing with CRISPR/Cas9 in postnatal mice corrects PRKAG2 cardiac syndrome. Cell Res 26:1099
Yang Y, Wang L, Bell P, McMenamin D, He Z, White J, Yu H, Xu C, Morizono H, Musunuru K, Batshaw ML, Wilson JM (2016) A dual AAV system enables the Cas9-mediated correction of a metabolic liver disease in newborn mice. Nat Biotechnol 34:334–338
Yin H, Song CQ, Dorkin JR, Zhu LJ, Li Y, Wu Q, Park A, Yang J, Suresh S, Bizhanova A, Gupta A, Bolukbasi MF, Walsh S, Bogorad RL, Gao G, Weng Z, Dong Y, Koteliansky V, Wolfe SA, Langer R, Xue W, Anderson DG (2016) Therapeutic genome editing by combined viral and non-viral delivery of CRISPR system components in vivo. Nat Biotechnol 34:328–333
Zhu P, Furen W, Mosenson J, Zhang H, He TC, Wen-Shu W (2017) CRISPR/Cas9-mediated genome editing corrects Dystrophin mutation in skeletal muscle stem cells in a mouse model of muscle dystrophy. Mol Ther Nucl Acids 7:31–41
Zych AO, Bajor M, Zagozdzon R (2018) Application of genome editing techniques in immunology. Arch Immunol Ther Exp (Warsz). https://doi.org/10.1007/s00005-018-0504-z
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mollanoori, H., Teimourian, S. Therapeutic applications of CRISPR/Cas9 system in gene therapy. Biotechnol Lett 40, 907–914 (2018). https://doi.org/10.1007/s10529-018-2555-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-018-2555-y