Abstract
Objectives
To develop a convenient chemical transformation mediated CRISPR/Cas9 (CT-CRISPR/Cas9) system for genome editing in Escherichia coli.
Results
Here, we have constructed a CT-CRISPR/Cas9 system, which can precisely edit bacterial genome (replacing, deleting, inserting or point mutating a target gene) through chemical transformation. Compared with the traditional electroporation mediated CRISPR/Cas9 (ET-CRISPR/Cas9) system, genome editing with the CT-CRISPR/Cas9 system is much cheaper and simpler. In the CT-CRISPR/Cas9 system, we observed efficient genome editing on LB-agar plates. The CT-CRISPR/Cas9 system has successfully modified the target gene with the editing template flanked by short homologous DNA fragments (~ 50 bp) which were designed in primers. We used the lab-made CaCl2 solution to perform the CT-CRISPR/Cas9 experiment and successfully edited the genome of E. coli. Potential application of the CT-CRISPR/Cas9 system in high-throughput genome editing was evaluated in two E. coli strains by using a multiwell plate.
Conclusions
Our work provides a simple and cheap genome-editing method, that is expected to be widely applied as a routine genetic engineering method.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The CRISPR/Cas system, an RNA-guided immune system identified in bacteria and archaea, is consisted of the clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated proteins (Cas) (Barrangou et al. 2007; Brouns et al. 2008; Marraffini and Sontheimer 2008; Barrangou and Marraffini 2014; Marraffini 2015). Until now, seven different types of CRISPR/Cas systems have been characterized (Sorek et al. 2013; Wright et al. 2016; Gootenberg et al. 2017). Among them, the type II CRISPR/Cas9 system, consisted of a mature CRISPR RNA (crRNA), a trans-activating crRNA (tracrRNA) and an endonuclease Cas9 which generates double strand breaks (DSBs), has been most widely applied in editing genomes of prokaryocytes and eukaryocytes (Cong et al. 2013; Mojica and Montoliu 2016; Wang et al. 2016; Heidari et al. 2017; Komor et al. 2017). For example, the CRISPR/Cas9 system is able to generate DSBs close to a protospacer adjacent motif (PAM), making the sequence near DSBs editable through either non-homologous end joining (NHEJ) or homologous recombination (HR) in the presence of template DNA for repairing and small guiding RNA (sgRNA) for guiding Cas9 to the target loci in the genome (Wang et al. 2016).
Genome editing with electroporation mediated CRISPR/Cas9 (ET-CRISPR/Cas9) system has been developed (Jiang et al. 2013, 2015; Li et al. 2015; Pyne et al. 2015; Su et al. 2016; Zhao et al. 2016) and validated in large scale in E. coli (Zerbini et al. 2017). The ET-CRISPR/Cas9 system has been well applied in metabolic engineering of E. coli (Heo et al. 2017; Zhu et al. 2017). Because a commercial electroporator and disposable electroporation cuvettes are required for ET-CRISPR/Cas9 system, it is quite expensive to create mutants through this method. The cost of ET-CRISPR/Cas9 limits the application of genome editing. Chemical transformation, which only needs cheap standard-buffers made in-lab, has been widely used as a routine experiment in many labs. However, to our knowledge, chemical transformation mediated CRISPR/Cas9 system (CT-CRISPR/Cas9) has not been established. Here, we report a CT-CRISPR/Cas9 system for genome editing. In this new genome editing system, template DNA is delivered into the chemically competent E. coli cell by heat shock. High-throughput genome editing is useful in many fields (e.g. metabolic engineering and synthetic biology). Although ET-CRISPR system mediated high-throughput genome editing is available, a low-cost genome editing method is still in demand. In this study, we established a CT-CRISPR method, that can edit bacterial genome by simple and cheap means. We anticipate that this method would facilitate the application of CRISPR/Cas9 genome editing in a broader scope.
Methods
Strains, plasmids, primers and growth conditions
The bacterial strains and plasmids used in this study are provided in Table 1. Primers used in this study are provided in Table S1. Plasmids were isolated with SanPrep column plasmid mini-preps kit (Sangon Biotech Co., Ltd). E. coli was grown in the LB medium (1% tryptone, 0.5% yeast extract and 1% NaCl) or on the LB-agar plate supplemented with ampicillin (100 μg/ml), kanamycin (50 μg/ml), spectinomycin (100 μg/ml) or chloramphenicol (50 μg/ml) at 37 °C or 30 °C when necessary.
Plasmid and editing template DNA construction
To construct pTargetF-pMB1 serial plasmids that express sgRNA for guiding Cas9 to the target gene, inverse PCR was performed with pTargetF-pMB1 as the template and a pair of reverse primers containing 20 nucleosides (N20) homologous to the sequence adjacent to the protospacer adjacent motif (PAM) in the target gene (Table S2). The PCR product was treated by Dpn I to degrade the template plasmid before being transformed into chemical competent E. coli cells. Construction of the pTargetF-pMB1 derivative plasmids was confirmed by sequencing.
To construct the template DNA used for genome editing, the DNA fragment homologous to the upstream of the target, the DNA fragment for editing the target and the DNA fragment homologous to the downstream of the target were assembled together through overlapping PCR (Heckman and Pease 2007).
Genome editing with chemical transformation
E. coli strains carrying pCas were grown overnight in 5 mL LB cultures at 30 °C. 1 mL of the culture was transferred to 50 mL LB and incubated with shaking (180 rpm). To induce the expression of the λ-Red recombination system, arabinose (15 mM final concentration) was added to the culture with an OD600 of 0.2–0.3. The chemical competent cells were prepared with either a Competent Cell Preparation Kit (Takara, Co., Ltd) or the CaCl2 solution. Briefly, 50 ml of the culture at an OD600 of 0.45 was precipitated and then washed twice with the solution A or the solution of CaCl2 at different concentrations, followed by resuspension in 500 µL of the solution B or the CaCl2 solution supplemented with 10% glycerol.
For chemical transformation, 50 µL of the competent cell culture was mixed with 1.38 µg of the pTargetF- pMB1 derivative plasmid and 1–3 µg of the editing template DNA. To evaluate the effect of the concentration of Ca2+ on CT-CRISPR, competent cells were prepared with the solution of CaCl2 at concentrations of 0, 25, 50, 100, 200, 300, 400 and 500 mM. To evaluate the effect of the template DNA concentration on editing efficiency, a serial concentrations (0, 0.5, 1, 3, 6 μg) of template DNA was mixed with 1.38 µg of pTargetF-panD. To evaluate the effect of the length of the homology on editing efficiency, the template DNA fragments flanked by homologies of different lengths (0.05, 0.25, 0.5, and 1 kb) were constructed. The mixture of pTargetF-panD and the editing template DNA, together with the chemical competent cell, was heated at 42 °C for 90 s and then rapidly placed on ice for 1–2 min, followed by the addition of 945 µL of preheated LB medium. Cells were recovered at 30 °C for 1 h before being spread onto LB-agar plates supplemented with appropriate antibiotics. To eliminate DNA after transformation in the liquid, DNase I (0.4 U/μl) was added to the transformed liquid cell culture prior to plating. Antibiotic resistance of transformants on plates was examined by replica-plating with toothpicks. Genotypes of transformants were examined by colony PCR, when necessary.
Editing efficiency evaluation
To evaluate the editing efficiency, the chloramphenicol resistance gene flanked by homologous arms was set as the editing template (conferring chloramphenicol resistance, Cmr). Editing efficiency was evaluated through direct plating and replica-plating. For the direct plating method, competent E. coli cells harboring pCas (conferring kanamycin resistance, Kanr) were transformed with pTargetF-panD (conferring spectinomycin resistance, Specr) and the editing template DNA (Cmr). Transformed cells were partially spread onto plates supplemented with appropriate antibiotics. Editing efficiency was determined by the ratio of Cmr Kanr Specr colonies to Kanr Specr colonies. For editing efficiency evaluated by the replica-plating method, the transformed cell culture was spread onto plates supplemented with kanamycin plus spectinomycin, followed by the examination of Cmr colonies from the Kanr Specr colonies by replica-plating with toothpicks. Editing efficiency was also determined by the ratio of Cmr Kanr Specr colonies to Kanr Specr colonies. For both of the two methods, Cmr Kanr Specr colonies were examined by colony PCR to confirm the modification of targeted gene.
Genome editing with multiwell plate
In each well of the 12-well plate, 30 µL of the competent cell culture was mixed with 1.38 µg of the pTargetF-pMB1 derivative plasmid and 3 µg of the corresponding editing template DNA unless otherwise indicated. The mixture was heat shocked at 42 °C for 90 s with a metal bath, followed by ice bath for 1 to 2 min and addition of 170 µL of preheated LB medium. Transformed cells were recovered at 30 °C for 1 h, before being spread onto LB agar plates supplemented with appropriate antibiotics (refer to the supplemental materials for procedures of genome editing with multiwell plate).
Results
Establishment of the CT-CRISPR/Cas9 genome editing system
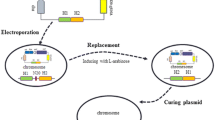
We constructed the CT-CRISPR/Cas9 genome editing system which delivered editing template DNA simply by heat shocking. In this system, the editing template DNA was acquired by overlapping PCR and the sgRNA was expressed by the pTargetF-pMB1 derivative plasmid which had been generated by PCR amplification with a pair of reverse primers (Fig. 1). Chemical competent cells were prepared with a Competent Cell Preparation Kit. The edited genome was obtained after co-transfer of the editing template DNA and the pTargetF-pMB1 derivative plasmid (Fig. 1). To evaluate the CT-CRISPR/Cas9 editing system, the chloramphenicol resistance gene (cat) flanked with the left and the right homologies (~ 1 kb) of crp was constructed through overlapping PCR (Fig. S1). The PCR product was used as the editing template DNA, that was expected to replace the chromosomal gene crp with the cat gene, yielding chloramphenicol resistance (Cmr) colonies. Colony PCR with the Cmr colonies as the template showed that the loci of crp was replaced by cat, in accordance with the prediction (Fig. S2). In a similar way, another E. coli chromosomal gene panD was successfully edited with the CT-CRISPR/Cas9 system (Figs. S3, S4).
Procedures of chemical transformation mediated CRISPR/Cas9 (CT-CRISPR/Cas9). Template DNA flanked by homologous regions was constructed by overlapping PCR and co-transformed with the pTargetF-pMB1 derivative plasmid containing sgRNA to chemical competent cells expressing Cas9 and λ Red recombinase, in which the target gene is editable with the template DNA
Genome editing efficiency of CT-CRISPR/Cas9 was evaluated by calculating the ratio of transformants with the target gene replaced by a cat gene (making the transformant confer chloramphenicol resistance) to the total transformants. Two methods, direct plating and replica-plating (see Materials and Methods for details), were used to evaluate genome editing efficiency. Direct plating method has been commonly used (Jiang et al. 2015; Zhao et al. 2016). In this method, Cmr transformants were screened on plates supplemented with chloramphenicol by plating transformed liquid cell culture. While in the replica-plating method, the transformed liquid cell culture was plated on LB-agar plates without chloramphenicol and Cmr transformants were screened out by replica-plating. Our results showed that CT-CRISPR/Cas9 achieved genome editing efficiencies of 19.0% ± 6.9% and 95.7% ± 1.45% respectively, evaluated by the direct plating and replica-plating methods.
We observed that genome editing efficiency was significantly affected by the concentration of the editing template DNA. Through the direct plating method, the editing efficiency was gradually increased from 0% to the highest level 27.0% ± 0.60% when the concentration of the template DNA was increased from 0 to 3 μg, followed by a slight decrease to 23.4% ±1.7% when 6 μg of the template DNA was provided (Fig. 2). The dosage effect of the editing template DNA on genome editing efficiency was more dramatic when measured by the replica-plating method: the editing efficiency reached 98.7% ±0.6% and 98% ± 2.8% respectively, when 3 μg and 6 μg of the template DNA were provided (Fig. 2). The results also clearly showed that the genome editing efficiency was much higher with the replica-plating method than with the direct-plating method.
Effect of template DNA concentration on editing efficiency. Genome editing efficiency was evaluated through the direct plating method (a) and the replica-plating method (b). Editing efficiency was calculated by dividing the number of Kanr and Specr colonies by the number of Cmr colonies (refers to the Materials and methods for detailed information). Each experiment was performed in triplicate and representative data were shown
Application of the CT-CRISPR/Cas9 system in gene deletion and insertion
Marker-free gene deletion
To construct a scarless mutant with deleted target gene, the editing template DNA was constructed by assembling the two DNA fragments homologous to the upstream and downstream of the target gene panD through overlapping PCR (Fig. S5). Chemical competent cells transformed with the editing template DNA were screened on plates and the corresponding mutants were further confirmed by colony PCR (Fig. S6). Transformation efficiency for marker-free gene deletion was 60 ± 14 CFU/μg and the gene editing efficiency was 100%.
Gene insertion
To insert an exogenous gene into the genome, the template DNA of a chloramphenicol resistant gene cat flanked by two DNA fragments homologous to the upstream and downstream of the target locus (at position of 146443 of E. coli MG1655) was constructed with the One Step Cloning Kit (Fig. S7). Chemical competent cells transformed with the editing template DNA were screened on selective plates and the mutants were confirmed by colony PCR (Fig. S8). Transformation efficiency for gene insertion was 16 ± 3 CFU/μg and the gene editing efficiency was (89 ± 15)%.
CT-CRISPR/Cas9 genome editing with the standard lab-made CaCl2 solution
To further reduce the cost of CT-CRISPR/Cas9 mediated genome editing, we replaced the reagents of the Competent Cell Preparation Kit with the lab-made CaCl2 solution. The mutation (A128C) on rpsL (named rpsL* hereafter) make E. coli resistant to streptomycin (StrepR). CT-CRISPR/Cas9 genome editing was evaluated via transforming the streptomycin-sensitive (StrepS) E. coli cells expressing Cas9 with rpsL* and pTargetF-rpsL. StrepR transformants were detected with an efficiency of ~ 150 CFU/μg when competent cells were prepared with 100 mM Ca2+ (Fig. 3a). To optimize the condition, the effect of CaCl2 concentration on CT-CRISPR/Cas9 mediated genome editing was evaluated. Competent cells were prepared with the solution of CaCl2 at different concentrations (0 mM to 500 mM). We observed the highest transformation efficiency (> 500 CFU/μg) when competent cells were prepared with 200 mM CaCl2 (Fig. 3a).
CT-CRISPR/Cas9 genome editing with lab-made CaCl2 solution. a Competent E. coli cells were prepared with the solution of CaCl2 at concentrations of 0, 25, 50, 100, 200, 300, 400 and 500 mM. 50 μl of the competent cell culture was transformed with 1 μg of the template DNA and pTargetF-rpsL for providing the sgRNA which targets to rpsL. b Competent E. coli cells were prepared with 200 mM CaCl2 solution. The up- and down- stream homology of panD with no selection marker (1 μg) were mixed with 40 μl of the competent cell culture. Transformants were screened on plates supplemented with appropriate antibiotics. The experiment was performed in duplicate and representative data were shown
We applied the CT-CRISPR/Cas9 mediated genome editing in gene scarless deletion. To construct the panD mutant, competent cells expressing the Cas9 system were prepared with the lab-made solution of CaCl2 and transformed with the marker-free up- and down- stream homologue arms. A transformation efficiency of 45 ± 10 CFU/μg was detected in competent cells prepared with 200 mM CaCl2 solution (Fig. 3b). Twenty transformants were examined by colony PCR with primers targeted to the up- and down- stream of panD. The PCR results showed that panD was deleted in all tested transformants.
Simplified CRISPR-Cas9 genome editing system with short homologous region designed in primers
We asked if short homologous sequences, which could be designed in primers, were sufficient for genome editing with CT-CRISPR/Cas9. As a first step, we examined the genome editing efficiencies with shortened DNA fragments homologous to the upstream and downstream of panD. As expected, the editing efficiency dramatically decreased with shortened homologous sequences (Figs. 4, S9). With the direct plating method, no transformants were detected on the LB-agar plates supplemented with chloramphenicol when the homologous fragment was decreased to 250 bp (Figs. 4, S9). However, with the same homologous fragment (Fig. S9), we detected an editing efficiency of 54.6% ± 0.4% by using the replica-plating method (Fig. 4). We further shortened the homologous arm to 50 bp which had been designed in primers (Fig. S9). A mutation efficiency of 5.88% ± 0% was detected (Fig. 4). Successful genome editing with short homologous fragments was confirmed by colony PCR (Fig. S10). With short homologous sequences designed in primers, we also successfully edited other two genes (crp and dsrA) in E. coli BW25113 (Fig. S10). These results demonstrated that the CT-CRISPR/Cas9 genome editing process could be further simplified by designing the homologous region in primers, omitting the procedure for assembling long homologous DNA fragments via overlapping PCR (Fig. 5).
Effect of the length of homologous region on editing efficiency. With template DNA flanking with a serial of homologous sequences of different length (0, 50, 250, 500, 1000 bp), genome editing efficiency was evaluated through the direct plating method and the replica-plating method as described. The experiment was performed in duplicate and representative data were shown
Simplified CT-CRISPR/Cas9 genome editing with short homologous DNA fragments. Short homologous DNA fragment was designed in primers for amplifying the editing template (Step 1). To edit the target loci in the genome, chemical competent cells expressing Cas9 and λ Red recombinase were transformed with the editing template DNA and pTargetF-Pmb1 derivative plasmid (Step 2 and 3). Finally, the plasmid expressing Cas9 were eliminated in the edited cells by increasing the temperature for cell incubation (Step 4)
Application of CT-CRISPR/Cas9 system in high-throughput genome editing
To evaluate potential application of our CT-CRISPR/Cas9 system in high-throughput genome editing, we attempted to edit 6 individual genes (panD, crp, hns, stpA, leuO and dsrA) in 2 different E. coli strains (MG1655 and BW25113) with a 12-well pate (refers to the supplemental materials for detailed experimental procedures). The 6 genes were divided into two groups: 4 genes (panD, crp, hns and stpA) were edited with long homologous DNA sequences and the other 2 genes (leuO and dsrA) were edited with short homologous DNA sequences designed in primers. The delivery of the template DNA was accomplished by using a metal bath. All of the four genes (panD, crp, hns and stpA) were successfully modified with template DNA flanked by long homologous DNA fragments (~ 1 kb) in both of the two E. coli strains in the 12-well plate (Table 2, Fig. S11). With template DNA flanked by short homologous DNA fragments (50 bp) designed in primers, modification of the other two genes (leuO and dsrA) was not successful in the 12-well plate. All edited E. coli strains were confirmed by colony PCR (Table 2, Fig. S11). The results showed potential application of the CT-CRISPR/Cas9 system in high-throughput genome editing.
Discussion
In this study, we established a simple and cheap CRISPR/Cas9 mediated genome editing method by delivering editing template DNA via chemical transformation. We have successfully achieved genome editing with both the commercial competent cell preparation kit and the lab-made standard CaCl2 solution. By simply heat shocking, the target loci in the genome can be edited in the chemically competent cell. Our CT-CRISPR/Cas9 genome editing system has been applied in gene replacement, scarless deletion, insertion and multigene editing. In the traditional ET-CRISPR/Cas9 genome editing systems (Jiang et al. 2013, 2015; Li et al. 2015; Pyne et al. 2015; Su et al. 2016; Zhao et al. 2016), the template DNA is delivered into bacteria with an electroporator (~ 2000 USD) and a disposable cuvette (~ 10 USD). For the CT-CRISPR/Cas9 genome editing system, the above expenditure can be saved. With lab-made CaCl2 solution, transformation efficiency for marker-free gene deletion reached 45 ± 10 CFU/µg and the editing efficiency reached as high as 100%. This advantage is extremely useful for large-scale genome editing in metabolic engineering and synthetic biology. We exemplified the potential application of our CT-CRISPR/Cas9 method in high-throughput genome editing by editing multiple genomes in parallel with a 12-well plate. With the template DNA flanked by ~ 1 kb homologous fragments, all of the tested genomes (8/8) were successful edited by the CT-CRISPR/Cas9 system. Although single-gene knockout mutant collection is readily available in E. coli BW25113, our CT-CRISPR/Cas9 gene editing system provides a simple method for constructing mutants in E. coli strains beyond BW25113, by either transferring the mutated gene from the KO mutant collection or de novo construction. Moreover, our CT-CRISPR/Cas9 provides a method for genome editing beyond gene inactivation (i.e. point mutation, gene insertion and double gene inactivation).
To evaluate the editing efficiency of the CT-CRISPR/Cas9 system, we used two different methods: direct plating and replica-plating. With the direct plating method, genome editing should be completed in the liquid culture before being spread on plates; while with the replica-plating method, genome editing could continue on non-selective plates before replica-plating. Compared with the direct plating method, the replica-plating method showed obviously higher genome editing efficiency (Figs. 2, 4).
Our previous work showed plasmid transformation on agar plates (Sun et al. 2006, 2009; Zhang et al. 2012; Sun et al. 2013; Sun 2016, 2018). High genome editing efficiency with the replica-plating method could be resulted from extended period for chemical transformation or genome editing in E. coli cells on the LB-agar plates. To check whether high genome editing via replica-plating was resulted from extended time for transformation with the template DNA on plates, the heat shocked mixture of competent cells and the template DNA was treated by DNase I before spreading it onto LB-agar plates. We observed that, without and with DNase I treatment, the editing efficiencies were 99.3% ± 1.0% and 96.4% ± 0.9% respectively, showing no obvious difference. The results showed that DNase I treatment did not significantly affect editing efficiency, revealing that the increase of genome editing efficiency with the replica-plating method could not be attributed to extended time for DNA transfer on LB-agar plates. Therefore, we conclude that efficient CRISPR/Cas9 mediated genome editing should occur on LB-agar plates. The discovery of genome editing on LB-agar plates would further expand the application scope of the CT-CRISPR/Cas9 system.
Using template DNA flanked by short homologous DNA fragments in primers, instead of long homologous DNA fragments which need to be constructed by time-consuming overlap PCR, would further simplify the parallel genome editing procedure, making it more amenable to high throughput genome-editing with CT-CRISPR/Cas9. Although CRISPR/Cas9 genome editing with short homology in a plasmid has been documented, to our knowledge, unless both RecA and the λ RED recombinase are over-expressed (Zhao et al. 2016), CRISPR/Cas9 genome editing with linear DNA carrying homology no longer than 50 bp has not been documented. Attempts to edit chromosomal gene with short homology designed in primers were not successful (Jiang et al. 2015). We observed that shortening the length of homologous sequences dramatically reduced the CRISPR/Cas9 editing efficiency (Fig. 4). It is noticeable that, we have successfully used our CT-CRISPR/Cas9 system to edit the genome with PCR product amplified with short homologies designed in primers (Figs. 4, S10), while RecA over-expression was omitted. With the multiwell plate, we failed to edit the genome with short homology designed in primers (Table 2). Optimizing the CT-CRISPR/Cas9 editing system in the multiwell plate could make it possible to edit genomes with primers containing short homology in the future.
In conclusion, we have established a simple and cheap CT-CRISPR/Cas9 system for genome editing in E. coli. The application of the CT-CRISPR9 genome editing method will largely facilitate the application of genome editing. We anticipate that our CT-CRISPR9 genome editing method would become a routine experiment in lab.
References
Barrangou R, Marraffini LA (2014) CRISPR-Cas systems: prokaryotes upgrade to adaptive immunity. Mol Cell 54:234–244. https://doi.org/10.1016/j.molcel.2014.03.011
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712. https://doi.org/10.1126/science.1138140
Brouns SJ, Jore MM, Lundgren M, Westra ER, Slijkhuis RJ, Snijders AP, Dickman MJ, Makarova KS, Koonin EV, van der Oost J (2008) Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321:960–964. https://doi.org/10.1126/science.1159689
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823. https://doi.org/10.1126/science.1231143
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97:6640–6645. https://doi.org/10.1073/pnas.120163297
Gootenberg JS, Abudayyeh OO, Lee JW, Essletzbichler P, Dy AJ, Joung J, Verdine V, Donghia N, Daringer NM, Freije CA, Myhrvold C, Bhattacharyya RP, Livny J, Regev A, Koonin EV, Hung DT, Sabeti PC, Collins JJ, Zhang F (2017) Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 356:438–442. https://doi.org/10.1126/science.aam9321
Heckman KL, Pease LR (2007) Gene splicing and mutagenesis by PCR-driven overlap extension. Nat Protoc 2:924–932. https://doi.org/10.1038/nprot.2007.132
Heidari R, Shaw DM, Elger BS (2017) CRISPR and the rebirth of synthetic biology. Sci Eng Ethics 23:351–363. https://doi.org/10.1007/s11948-016-9768-z
Heo MJ, Jung HM, Um J, Lee SW, Oh MK (2017) Controlling citrate synthase expression by CRISPR/Cas9 genome editing for n-butanol production in Escherichia coli ACS. Synth Biol 6:182–189. https://doi.org/10.1021/acssynbio.6b00134
Jiang W, Bikard D, Cox D, Zhang F, Marraffini LA (2013) RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat Biotechnol 31:233–239. https://doi.org/10.1038/nbt.2508
Jiang Y, Chen B, Duan C, Sun B, Yang J, Yang S (2015) Multigene editing in the Escherichia coli genome via the CRISPR-Cas9 system. Appl Environ Microbiol 81:2506–2514. https://doi.org/10.1128/aem.04023-14
Komor AC, Badran AH, Liu DR (2017) CRISPR-based technologies for the manipulation of eukaryotic genomes. Cell 168:20–36. https://doi.org/10.1016/j.cell.2016.10.044
Li Y, Lin Z, Huang C, Zhang Y, Wang Z, Tang YJ, Chen T, Zhao X (2015) Metabolic engineering of Escherichia coli using CRISPR-Cas9 meditated genome editing. Metab Eng 31:13–21. https://doi.org/10.1016/j.ymben.2015.06.006
Marraffini LA (2015) CRISPR-Cas immunity in prokaryotes. Nature 526:55–61. https://doi.org/10.1038/nature15386
Marraffini LA, Sontheimer EJ (2008) CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322:1843–1845. https://doi.org/10.1126/science.1165771
Mojica FJ, Montoliu L (2016) On the origin of CRISPR-Cas technology: from prokaryotes to mammals. Trends Microbiol 24:811–820. https://doi.org/10.1016/j.tim.2016.06.005
Pyne ME, Moo-Young M, Chung DA, Chou CP (2015) Coupling the CRISPR/Cas9 system with lambda red recombineering enables simplified chromosomal gene replacement in Escherichia coli. Appl Environ Microbiol 81:5103–5114. https://doi.org/10.1128/aem.01248-15
Sorek R, Lawrence CM, Wiedenheft B (2013) CRISPR-mediated adaptive immune systems in bacteria and archaea. Annu Rev Biochem 82:237–266. https://doi.org/10.1146/annurev-biochem-072911-172315
Su T, Liu F, Gu P, Jin H, Chang Y, Wang Q, Liang Q, Qi Q (2016) A CRISPR-Cas9 assisted non-homologous end-joining strategy for one-step engineering of bacterial genome. Sci Rep 6:37895. https://doi.org/10.1038/srep37895
Sun D (2016) Two different routes for double-stranded DNA transfer in natural and artificial transformation of Escherichia coli. Biochem Biophys Res Commun 471:213–218. https://doi.org/10.1016/j.bbrc.2016.01.137
Sun D (2018) Pull in and push out: mechanisms of horizontal gene transfer in bacteria. Front Microbiol. https://doi.org/10.3389/fmicb.2018.02154
Sun D, Zhang Y, Mei Y, Jiang H, Xie Z, Liu H, Chen X, Shen P (2006) Escherichia coli is naturally transformable in a novel transformation system. FEMS Microbiol Lett 265:249–255. https://doi.org/10.1111/j.1574-6968.2006.00503.x
Sun D, Zhang X, Wang L, Prudhomme M, Xie Z, Martin B, Claverys JP (2009) Transforming DNA uptake gene orthologs do not mediate spontaneous plasmid transformation in Escherichia coli. J Bacteriol 191:713–719. https://doi.org/10.1128/JB.01130-08
Sun D, Wang B, Zhu L, Chen M, Zhan L (2013) Block and boost DNA transfer: opposite roles of OmpA in natural and artificial transformation of Escherichia coli. PLoS ONE 8:e59019. https://doi.org/10.1371/journal.pone.0059019
Wang H, La Russa M, Qi LS (2016) CRISPR/Cas9 in genome editing and beyond. Annu Rev Biochem 85:227–264. https://doi.org/10.1146/annurev-biochem-060815-014607
Wright AV, Nunez JK, Doudna JA (2016) Biology and applications of CRISPR systems: harnessing nature’s toolbox for genome engineering. Cell 164:29–44. https://doi.org/10.1016/j.cell.2015.12.035
Zerbini F, Zanella I, Fraccascia D, Konig E, Irene C, Frattini LF, Tomasi M, Fantappie L, Ganfini L, Caproni E, Parri M, Grandi A, Grandi G (2017) Large scale validation of an efficient CRISPR/Cas-based multi gene editing protocol in Escherichia coli. Microb Cell Fact 16:68. https://doi.org/10.1186/s12934-017-0681-1
Zhang Y, Shi C, Yu J, Ren J, Sun D (2012) RpoS regulates a novel type of plasmid DNA transfer in Escherichia coli. PLoS ONE 7:e33514. https://doi.org/10.1371/journal.pone.0033514
Zhao D, Yuan S, Xiong B, Sun H, Ye L, Li J, Zhang X, Bi C (2016) Development of a fast and easy method for Escherichia coli genome editing with CRISPR/Cas9. Microb Cell Fact 15:205. https://doi.org/10.1186/s12934-016-0605-5
Zhu X, Zhao D, Qiu H, Fan F, Man S, Bi C, Zhang X (2017) The CRISPR/Cas9-facilitated multiplex pathway optimization (CFPO) technique and its application to improve the Escherichia coli xylose utilization pathway. Metab Eng 43:37–45. https://doi.org/10.1016/j.ymben.2017.08.003
Acknowledgements
We acknowledge Dr. Sheng Yang and Dr. Junjie Yang for kindly donating plasmids pCas and pTargetF-pMB1. This research was supported by National Natural Science Foundation of China under Grant No. 31670084 and Zhejiang Provincial Natural Science Foundation of China under Grant No. LY16C010003.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sun, D., Wang, L., Mao, X. et al. Chemical transformation mediated CRISPR/Cas9 genome editing in Escherichia coli. Biotechnol Lett 41, 293–303 (2019). https://doi.org/10.1007/s10529-018-02639-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-018-02639-1