Abstract
The main objective of this paper is to analyze the prognostic and immunological value of CDC45 in kidney renal clear cell carcinoma (KIRC) using single-cell and bulk RNA-sequencing approaches. The expression of CDC45 in KIRC was evaluated by the HPA database, the TCGA-KIRC dataset and verified by PCR analysis and single-cell RNA-sequencing. The ability of CDC45 to independently predict prognosis in KIRC was confirmed by univariate/multivariate regression analysis. Gene set enrichment analysis (GSEA) was employed to explore CDC45-related pathways in KIRC. In addition, Relationships between CDC45 and immunity were also examined. Elevated CDC45 expression in KIRC was demonstrated at mRNA and protein levels. The results of the correlation analysis showed that as CDC45 expression increased, so did the histological grade, clinical stage, and TNM stage of the patients (p < 0.05). Univariate/multivariate regression analysis suggested CDC45 as an independent prognostic factor for KIRC. Seven pathways related to CDC45 were screened through GSEA. Meanwhile, we found that CDC45 was correlated with tumor mutational burden (TMB) and microsatellite instability (MSI) but not tumor neoantigen burden (TNB). Regarding immunity, CDC45 exhibited correlations with the tumor microenvironment, immune cell infiltration, and immune checkpoints. Besides, low CDC45 expression was shown to be associated with a better response to immunotherapy. Single-cell RNA-sequencing revealed that CDC45 was differently expressed in T cells (p < 0.05). CDC45 showed potential as a prognostic biomarker and therapeutic target for KIRC. Meanwhile, the CDC45 low expression group was more sensitive to immunotherapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In urology, renal cell carcinoma (RCC) was a common form of cancer, second only to bladder cancer in terms of incidence. Mortality from RCC was rising, with an increasing annual incidence globally (Hsieh et al. 2017). Kidney renal clear cell carcinoma (KIRC), the most common type, accounted for 70–85% of cases (Escudier et al. 2019). With the improvement of medical technology and the popularization of medical checkups, the early diagnosis rate of KIRC had improved. Despite this, in 30% of cases, cancer was still in an advanced stage. The prognosis for advanced KIRC was poor, with a 5-year survival rate as low as 11.7% (Siegel et al. 2017). Research studies had shown that the incidence of postoperative metastases in KIRC patients was 30% or more, and neither postoperative nor preoperative metastases had a very good survival rate. Metastatic KIRC featured a 5-year survival rate of only 10% (Wang et al. 2018). As such, the search for novel biomarkers and therapeutic targets was crucial to improving KIRC patient outcomes.
It had been reported that cell-division cycle proteins (CDC) were commonly found in cells that were actively in the process of division, but were downregulated during cell quiescence, differentiation, or senescence (Kingsbury et al. 2005). CDC45 was originally discovered during genetic screening for yeast cell cycle mutants (Hennessy et al. 1991). Later studies revealed that CDC45 was one of the essential factors in the process of DNA replication (Hardy 1997; Zou et al. 1997). Since then, numerous studies had demonstrated that CDC45 was necessary for the entire process of DNA replication, including the establishment of the initiation complex, the unhealing of chromosomes at the replication fork, and the orderly synthesis of DNA (Pacek and Walter 2004). In the field of oncology, CDC45 had also been extensively studied. It had been reported that CDC45 was commonly highly expressed in various malignant tumors such as breast cancer, cervical cancer, lung cancer, and glioblastoma (Tomita et al. 2011), and that CDC45 may be a marker for the proliferation of various malignant tumors (Pollok et al. 2007). However, the research on the mechanism of CDC45 in the field of KIRC is not thorough. Recently, the immune microenvironment in cancer patients has received extensive attention. Its complexity and diversity have affected the occurrence and development of cancer, providing a new idea for clinical immunotherapy. Therefore, here we examined the prognosis of KIRC in relation to CDC45 and further investigated the associations between CDC45 and immunity and immunotherapy.
Materials and Methods
Data Collection
The Cancer Genome Atlas (TCGA; http://cancergenome.nih.gov/) provided the CDC45 expression matrix and clinical data for 539 KIRC and 72 normal kidney samples. Following this, incomplete or missing data were eliminated, and the CDC45 gene expression profile and corresponding clinical information were used for further analysis. All Analyses were conducted using R software (https://www.r-project.org/). The different expression levels of CDC45 mRNA were conducted by the R package "limma". In addition, we took |log2 fold change (FC)|≥ 1, and adjusted p-value < 0.05 as the definition of cut-off criteria.
Detection of CDC45 Expression in KIRC and Enrichment of Functions
The “limma” was applied to detect differently expressed genes (DEGs) between tumors and adjacent normal tissues, including the CDC45 gene in KIRC. CDC45 expression at the protein level was verified by immunohistochemical methods applying the human protein atlas (HPA, http://www.proteinatlas.org/).
Collection of Clinical Samples and Quantitative Real‑Time PCR (qRT‑PCR)
To increase the credibility of the data, we collected eight pairs of tissues from patients with KIRC who underwent nephrectomy at the Affiliated Hospital of Nantong University to further validate CDC45 expression at the mRNA level. The details of these eight patients were as follows: age 50 to 80 years; weight 55 to 85 kg; three males and five females; T stage T1-T4; N stage N0-N1; and M stage all M0. RNA was extracted from freshly frozen KIRC samples using the TRIzol reagent (Life Technology, USA). In accordance with the instructions, reverse transcription was performed on RNA. Finally, CDC45 expression was detected with the help of the SYBR Green reagent (Vazyme, Nanjing, China). GAPDH served as a standardized control for KIRC specimens. Relative CDC45 expression was calculated using the 2−ΔΔCt method. The above data analysis was based on GraphPad Prism 8.0.1 and a t-test. The corresponding primers in this study were: CDC45 (F: 5’-TGAGTATGACCTCCGCCTGG-3’, R: 5’-CCATGCACAGACCACAGCTT-3’) and GAPDH (F: 5’-CAGGAGGCATTGCTGATGAT-3’, R: 5’-GAAGGCTGGGGCTCATTT-3’).
Univariate/Multivariate Cox Hazard Regression Analysis
To further investigate the effect of CDC45 on overall survival (OS), we used the R package to perform univariate/multivariate regression analysis on KIRC patients in the TCGA database to determine whether CDC45 was an independent factor associated with OS by eight clinical factors (race, sex, age, grading, T, N, M, stage). And we presented it in the form of a forest plot using the R language.
Gene Set Enrichment Analysis (GSEA)
The TCGA-KIRC samples were stratified into high- and low-CDC45 groups. Following this, the potential mechanism of CDC45 in KIRC was explored by GSEA where p < 0.05 and FDR q value < 0.25 were regarded as the cut-off criteria (Hänzelmann et al. 2013).
Correlation Analysis of CDC45 with Microsatellite Instability (MSI), Tumor Mutational Burden (TMB) and Tumor Neoantigen Burden (TNB)
To further analyze the possible key genes in KIRC, we constructed and visualized PPI networks using the STRING database and Cytoscape software. Moreover, to examine the association between CDC45 expression and MSI, TMB, and TNB, we took advantage of the Sangerbox website (http://www.sange rbox.com/tool), while setting the threshold to less than 0.05 (Cai et al. 2020; Li et al. 2021; Liu et al. 2021a).
Immunotherapy Response Prediction and Correlations between CDC45 and Immunity
Based on the ESTIMATE algorithm and CDC45 expression matrix, the ImmuneScore, StromalScore, and ESTIMATEScore were calculated (Yu et al. 2022). TIMER was employed to examine the relationships between CDC45 and immune cell infiltration in KIRC (TIMER, https://cistrome.shinyapps.io/timer/) (Chen et al. 2021). Correlation analysis examined the CDC45-immune checkpoint and cell associations. These correlation analyses were done with the help of the Sangerbox tool. Immunotherapy outcomes were calculated by uploading the tumor expression matrix to the TIDE (tumor immune dysfunction and exclusion) database (http://tide.dfci.harvard.edu) and TIGER (Tumor Immunotherapy Gene Expression Resource) database (http://tiger.canceromics.org/) (Chen et al. 2022; Jiang et al. 2018).
Detection of CDC45 Expression in Various Cells in the Tumor Microenvironment by Single-Cell Sequencing
The TISCH (http://tisch.comp-genomics.org) database integrated single-cell sequencing data for 27 cancers, providing gene expression visualization at the single-cell level (Sun et al. 2021). This study used the TISCH database to explore the single-cell level expression of CDC45 in KIRC in the GSE159115 dataset.
Results
Expression and External Verification of CDC45 in KIRC
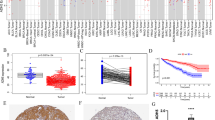
To investigate the expression of CDC45 in tumors and adjacent normal tissues, the expression levels of CDC45 were examined in pan-cancers (Fig. 1A). Meanwhile, we could see that CDC45 expression had a significant increase in tumor tissues than in normal tissues (p < 0.05, Fig. 1B–C). Pairwise boxplot also suggested the same results (p < 0.05, Fig. 1D). One, three, five-year area under the curve (AUC) values of CDC45 in KIRC of OS prognosis were 0.625, 0.592 and 0.621(Fig. 1E). By K-M survival analysis we concluded that elevated CDC45 expression predicted poor OS (Fig. 1F). Immunohistochemical staining of CDC45 downloaded from the HPA database (https: //www.proteinatlas.org/) also clearly revealed that normal tissues had a lower expression of CDC45 than KIRC (Fig. 1G–H). In addition, validation results of PCR at the mRNA level further indicated that CDC45 expression was elevated in KIRC (Fig. 1I).
The expression of CDC45 in KIRC. A Expression of CDC45 in various cancers in TCGA database; B Expression and distribution of genes in human body; C Boxplot of CDC45 expression in normal renal tissue and KIRC tissue; D Pairwise boxplot of CDC45 expression in normal renal tissue and KIRC tissue; E ROC curve of CDC45 and its AUCs of 1-, 3-, and 5-year survival. F K–M survival analysis of CDC45. G–H Immunohistochemical staining of CDC45 in HPA database. I Verification of CDC45 expression at tissue level by PCR, T represented KIRC tissue; N represented adjacent normal tissue. * p < 0.05, ** p < 0.01, *** p < 0.001
Relationships between CDC45 and Clinicopathologic Features
Logistic analysis was used to screen the clinicopathologic data of TCGA-KIRC patients to determine the relationships between five clinical features and CDC45. Elevated CDC45 expression was significantly correlated with higher grade (p = 0.00082), stage (p = 3.6E-05), T (p = 2.9E-06), M (p = 1.4E-06) and N (p = 4.6E-05) (Fig. 2). Considering the above findings, we speculated that elevated CDC45 expression was associated with the progression of KIRC.
Prognostic Analysis of CDC45 in KIRC
Univariate regression analysis reflected that there was a significant association between OS and age, grade, stage, T, M, CDC45 expression (all p < 0.05; Fig. 3A and Table 1). Multivariate regression analysis reflected that age, grade, stage, and CDC45 expression of KIRC were correlated with OS in KIRC (all p < 0.05; Fig. 3B and Table 1). The p values of CDC45 in the above two analyses were significant, which further indicated that CDC45 may be an independent prognostic factor for KIRC.
Identification of CDC45-Related Signaling Pathways
To further explore pathways associated with CDC45, GSEA assays were taken between high- and low-CDC45 matrices, and seven signaling pathways, including CELL CYCLE, CHEMOKINE, JAK STAT, NOD LIKE RECEPTOR, p53, T CELL RECEPTOR SIGNALING PATHWAY, PRIMARY IMMUNODEFICIENCY were identified by setting the p < 0.05 and FDR q value < 0.25 (Fig. 4 and Table 2). These findings might provide insights into CDC45-mediated KIRC pathogenesis.
Enrichment plots from gene set enrichment analysis (GSEA). A CELL CYCLE; B CHEMOKINE signaling pathway; C JAK STAT signaling pathway; D NOD LIKE RECEPTOR signaling pathway; E p53 signaling pathway; F PRIMARY IMMUNODEFICIENCY; G T CELL RECEPTOR signaling pathway; H seven CDC45-related signaling pathways
Associations between CDC45 and PPI, TMB, TNB, MSI in KIRC
We constructed a PPI network through the String database (https://string-db.org/) to search for genes in KIRC that may be closely related to CDC45 (Szklarczyk et al. 2011). A total of ten genes closely related to CDC45 were finally obtained (GINS2, GINS4, WDHD1, POLE2, MCM2, MCM4, MCM6, MCM5, MCM7, POLA1, Fig. 5A). Meanwhile, we evaluated the relationships between CDC45 and MSI, TMB, TNB through the Sangerbox website with a threshold of p < 0.05. According to the findings, CDC45 was significantly connected with MSI as well as TMB but not TNB in KIRC (Fig. 5B–D).
Association of CDC45 with KIRC Tumor Microenvironment, Tumor Immune Infiltration, Immune Cell Pathways, and Immune Checkpoint
Based on the results of the correlation analysis between CDC45 and immunity, we found that CDC45 had a strong link to ImmuneScore, ESTIMATEScore, and StromalScore (p < 0.05, Fig. 6A). In terms of tumor immune infiltration, CDC45 was significantly associated with B cell, CD8 + T cell, CD4 + T cell, neutrophil, dendritic and macrophage cell infiltration (all p < 0.001, Fig. 6B). Meanwhile, the data from TCGA showed that CDC45 was associated with immune checkpoints such as PDCD1, PDCD1LG2, TNFSF4, in KIRC (all p < 0.05; Fig. 6C). In a co-expression analysis, CDC45 was significantly correlated with immune cell pathways like activated CD4 T cell, central memory CD8 T cell, monocyte, etc. (all p < 0.05, Fig. 6D). The above results concluded that CDC45 was significantly associated with immunity in four aspects.
CDC45-Related Immunotherapy Response Prediction
Based on our results, CDC45 was differentially expressed across immune subtypes, including C1, C2, C3, C4, and C6 (Thorsson et al. 2018) (Fig. 7A). Among them, CDC45 accounted for the highest proportion in the C3 immune subtype. According to Fig. 7B–D, KIRC patients with high CDC45 expression had lower MSI scores, higher tumor immune dysfunction scores and higher TIDE scores. Accordingly, these patients may have a worse outcome following immunotherapy than those with low CDC45 expression. Meanwhile, we further validated the impact of the relationships between CDC45 and immunotherapy on the prognosis of KIRC through TIGER. It could be seen from Fig. 7E that the prognosis was worse for patients whose levels of CDC45 were high. The whole figure reflected that elevated CDC45 expression was associated with a poorer immune response; thus those patients with elevated CDC45 expression who were not sensitive to immunotherapy naturally had a worse prognosis.
Prediction of CDC45-related immune responses of immunotherapy. A Proportion of high and low expression groups of CDC45 in immune subtypes. B Differences of CDC45 expression in T cell dysfunction scores; C Differences of CDC45 expression in TIDE scores; D Differences of CDC45 expression in MSI; E Survival analysis between high and low CDC45 expression groups. **p < 0.01, ***p < 0.001
Results of Single-Cell Sequencing
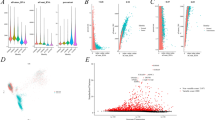
In parallel, we used the TISCH dataset to further validate the expression of CDC45 in the tumor microenvironment of KIRC at the single-cell level. In GSE159115, CDC45 was mainly expressed in CD8 T, Mono/Macro, endothelial, and malignant cells (Figure S1; Fig. 8A). The GSE159115 violin plot suggested that the expression of CDC45 in KIRC CD8 T cells as well as Erythroblasts was significantly different from that of CDC45 in corresponding normal tissue CD8 T cells as well as Erythroblasts (Fig. 8B). In addition, the results of CDC45 in four additional datasets showed significant expression of CDC45 in Tprolif (Fig. 8C). The results of single-cell sequencing showed that CDC45 was mainly expressed in T cells, partially explaining its immune correlations.
CDC45 expression in various cells in the tumor microenvironment by single-cell sequencing. A UMAP plot CDC45 in various cells in the GSE159115 dataset. B Violin plot of CDC45 expression in immune cells and stromal cells of KIRC tissue and normal tissue in the GSE159115 dataset. C The results of the expression of CDC45 in various cells in five datasets. *p < 0.05, **p < 0.01, ***p < 0.001
Discussion
KIRC was a highly heterogeneous urological malignancy, and the biological mechanism of its development was not fully understood. Targeted therapy was the main treatment modality for patients with advanced KIRC, but studies had shown that only 20–40% of patients benefited from it, and most high-risk patients still had a poor prognosis. The search for biomarkers with high specificity and sensitivity was important for targeted individualized therapeutic treatment and prediction of the prognosis of patients with KIRC (Barata and Rini 2017; Li et al. 2019a). Therefore, this study explored the possibility of CDC45 as a potential therapeutic target for KIRC based on bioinformatics.
The results of the paper reflected that CDC45 was highly expressed in a variety of human malignancies and was also upregulated in KIRC tissues. We also verified the expression level of CDC45 in KIRC using HPA and PCR. Clinicopathological correlation analysis showed that as CDC45 expression increased, so did the histological grade, clinical stage, and TNM stage of the patients (p < 0.05). Univariate/multivariate analyses suggested CDC45 as an independent prognostic factor associated with OS in KIRC. Based on the above results, the paper hypothesized that CDC45 may be a potential oncogenic factor of KIRC and was expected to be a potential prognostic biomarker of KIRC. Seven signaling pathways related to CDC45 were obtained by applying GSEA, including CELL CYCLE, CHEMOKINE, JAK STAT, NOD LIKE RECEPTOR, P53, PRIMARY IMMUNODEFICIENCY, T CELL RECEPTOR signaling pathways. In addition, we analyzed and concluded that CDC45 was closely related to the tumor microenvironment, immune cell infiltration, and immune checkpoint. It was further concluded by TIDE that low CDC45 expression was associated with better immunotherapy efficacy.
CDC45 had been considered to be an indispensable factor in DNA replication since its discovery. DNA replication was a critical step in the mid-cell phase and required tight control to ensure the accuracy of genomic information. Abnormal replication could lead to genetic instability and cancer (Hills and Diffley 2014). As the study of CDC45 progresses, more and more researchers had explored the expression of CDC45 in tumor cells. A high level of expression of CDC45 was found in different types of tumor cells, and in the study of the mechanism, it was found that CDC45 mainly regulated the abnormal expression of cyclin through cell cycle, which caused tumorigenesis (Hanahan and Weinberg 2000). Sun et al. showed that elevated CDC45 expression was strongly correlated with tumor size and grade, and demonstrated that silencing of CDC45 inhibited tumor growth through cell cycle G1 phase arrest and induction of apoptosis by gene silencing in papillary thyroid cancer (Sun et al. 2017). Li et al. showed that CDC45 was higher in malignant squamous cell carcinoma than in mild precancerous lesions, and the expression level tended to increase with the grade of precancerous lesions from mild, moderate to high (Li et al. 2008). In the exploration of large sample data in the TCGA database, CDC45 was found to be a key tumor-associated gene in cervical cancer (Zhang and Zhao 2016), prostate cancer (Li et al. 2017) and lung cancer (Zhang et al. 2014), and the analysis of prognostic correlation showed that CDC45 was significantly related to poor prognosis in non-small cell lung cancer (Piao et al. 2018) and pancreatic cancer (Haider et al. 2014). In conclusion, tumors were developed in part by CDC45.
To provide insights into potential CDC45-mediated KIRC pathogenesis, we identified seven corresponding pathways through GSEA. Some of these pathways had been extensively studied in the field of oncology. P53 signaling was an important intracellular oncogenic pathway. Lacroix et al. showed that P53 and P21 in this pathway were biologically active after phosphorylation and widely affected the physiopathological processes of tumor cell metabolism, proliferation, cell cycle, and metastasis and invasion (Lacroix et al. 2020). Vennin et al. showed that P53 signaling inhibited epithelial mesenchymal transition in pancreatic cancer cells by regulating transcription of downstream related genes and driving phosphorylation of related oncogenic molecules such as P21 (Vennin et al. 2019). Wang et al. showed that in prostate cancer, CREB1 regulated P53 activity and inhibited angiogenesis in tumor cells (Wang et al. 2019). JAK/STAT pathway was an important pathway for cytokine signaling and was of great interest in tumors. Studies had shown that inhibiting of the IL-6/JAK2/STAT3 signaling pathway inhibited the proliferation of glioma cells and promotes apoptosis (Stanzani et al. 2017; Zhou et al. 2020). Activation of JAK2/STAT3 signaling in pancreatic cancer can lead to tumorigenesis, progression, cancer stem cell maintenance, and treatment resistance (Tyagi et al. 2016). CHMP4C affected the proliferation of lung cancer cells by cell cycle pathway in squamous cell carcinoma of the lung (Liu et al. 2021b). In summary, we speculated that CDC45 may be involved in the progression of KIRC through these pathways, however, this remained to be further verified by subsequent experiments.
Subsequently, to further explore the pathogenesis, we screened ten genes that closely interact with CDC45. MCM2, MCM3, MCM4, MCM5, MCM6, MCM7 as key proteins in the initiation of DNA replication played a crucial role in gene stabilization (Neves and Kwok 2017; Wang et al. 2020). Studies had shown that aberrant expression of MCM2-7 caused failure of DNA replication leading to tumorigenesis (Pruitt et al. 2007; Wu et al. 2018). As GINS2, GINS4, which were also involved in the DNA replication process, previous studies had reported that they were abnormally expressed in many tumor tissues including KIRC, affecting tumor development. In hepatocellular carcinoma, up-regulation of GINS2 expression suggested a poorer prognosis and GINS2 may influence the extent of immune cell infiltration and thus the tumor microenvironment and ultimately altered the immune response (Li et al. 2022). However, the mechanisms involved were not yet clear. The study by Jin et al. concluded that GINS4 was associated with immune cell infiltration in esophageal squamous cell carcinoma and may play a role in the immune system (Jin et al. 2022). Taken together, the current state of research described above suggested that CDC45 may play an indispensable role in KIRC through these genes however the exact mechanism remained to be explored.
As far as immunity was concerned, CDC45 showed significant associations with immune cell infiltration and immune checkpoints (p < 0.05). As part of immune surveillance, the immune system identifies, kills and removes mutated cells in the body before tumors form. Tumor cells evade immune surveillance by using immune checkpoints to masquerade as normal cells in the body. Blocking the immune checkpoint pathway to prevent tumor cells from masquerading as normal components of the body was an effective way to achieve anti-tumor immunity (Li et al. 2019b). In this paper, we found that CDC45 was associated with immune checkpoints, suggesting that through the regulation of particular immune checkpoint genes, CDC45 may regulate tumor immune patterns. And it provided a new idea for the future immunotherapy of KIRC.
In general, there were some highlights in our study. First, we used the TCGA database to identify the elevated expression of CDC45 in KIRC and further verified its expression by PCR. Second, we further determined the relationships between CDC45 and immunity through single-cell and bulk RNA-sequencing and predicted the response to immunotherapy in KIRC patients. However, there were still some shortcomings in this paper. First of all, we did not experimentally verify CDC45 expression at the protein level. Last but not least, whether the pathways associated with CDC45 in this paper affected the progression of KIRC remained to be verified by further experiments.
Conclusion
In summary, this paper validated increased CDC45 expression in KIRC by bioinformatics analysis and demonstrated its potential as a prognostic biomarker of KIRC. In addition, CDC45 correlated with the level of immune cell infiltration in KIRC, suggesting that CDC45 may play a significant role in controlling the immune microenvironment of KIRC, but its specific mechanism still needed to be further investigated.
Data Availability
The RNA-sequencing data and corresponding clinical information were downloaded from the Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov/). Immunohistochemical results of CDC45 were downloaded from Human Protein Atlas (HPA) online website (http://www.proteinatlas.org/).
References
Barata PC, Rini BI (2017) Treatment of renal cell carcinoma: current status and future directions. CA Cancer J Clin 67(6):507–524
Cai X, Qiu W, Qian M, Feng S, Peng C, Zhang J, Wang Y, Wang Y (2020) A Candidate Prognostic Biomarker Complement Factor I Promotes Malignant Progression in Glioma. Front Cell Develop Biol 8:615970
Chen F, Fan Y, Hou J, Liu B, Zhang B, Shang Y, Chang Y, Cao P, Tan K (2021) Integrated analysis identifies TfR1 as a prognostic biomarker which correlates with immune infiltration in breast cancer. Aging 13(17):21671–21699
Chen Z, Luo Z, Zhang D, Li H, Liu X, Zhu K, Zhang H, Wang Z, Zhou P, Ren J et al (2022) TIGER: a web portal of tumor immunotherapy gene expression resource. Geno Proteom Bioinform. ISSN: S1672-0229. https://doi.org/10.1016/j.gpb.2022.08.004
Escudier B, Porta C, Schmidinger M, Rioux-Leclercq N, Bex A, Khoo V, Grünwald V, Gillessen S, Horwich A (2019) Renal cell carcinoma: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up†. Ann Oncol off J Eur Soc Med Oncol 30(5):706–720
Haider S, Wang J, Nagano A, Desai A, Arumugam P, Dumartin L, Fitzgibbon J, Hagemann T, Marshall JF, Kocher HM et al (2014) A multi-gene signature predicts outcome in patients with pancreatic ductal adenocarcinoma. Gen Med 6(12):105
Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100(1):57–70
Hänzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14:7
Hardy CF (1997) Identification of Cdc45p, an essential factor required for DNA replication. Gene 187(2):239–246
Hennessy KM, Lee A, Chen E, Botstein D (1991) A group of interacting yeast DNA replication genes. Genes Dev 5(6):958–969
Hills SA, Diffley JF (2014) DNA replication and oncogene-induced replicative stress. Curr Biol CB 24(10):R435-444
Hsieh JJ, Purdue MP, Signoretti S, Swanton C, Albiges L, Schmidinger M, Heng DY, Larkin J, Ficarra V (2017) Renal cell carcinoma. Nat Rev Dis Primers 3:17009
Jiang P, Gu S, Pan D, Fu J, Sahu A, Hu X, Li Z, Traugh N, Bu X, Li B et al (2018) Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med 24(10):1550–1558
Jin D, Yuan L, Li F, Wang S, Mao Y (2022) GINS4 might be a novel prognostic immune-related biomarker of not only esophageal squamous cell carcinoma and other cancers. BMC Med Genomics 15(1):75
Kingsbury SR, Loddo M, Fanshawe T, Obermann EC, Prevost AT, Stoeber K, Williams GH (2005) Repression of DNA replication licensing in quiescence is independent of geminin and may define the cell cycle state of progenitor cells. Exp Cell Res 309(1):56–67
Lacroix M, Riscal R, Arena G, Linares LK, Le Cam L (2020) Metabolic functions of the tumor suppressor p53: Implications in normal physiology, metabolic disorders, and cancer. Molec Metabol 33:2–22
Li JN, Feng CJ, Lu YJ, Li HJ, Tu Z, Liao GQ, Liang C (2008) mRNA expression of the DNA replication-initiation proteins in epithelial dysplasia and squamous cell carcinoma of the tongue. BMC Cancer 8:395
Li HY, Jin N, Han YP, Jin XF (2017) Pathway crosstalk analysis in prostate cancer based on protein-protein network data. Neoplasma 64(1):22–31
Li QK, Pavlovich CP, Zhang H, Kinsinger CR, Chan DW (2019a) Challenges and opportunities in the proteomic characterization of clear cell renal cell carcinoma (ccRCC): A critical step towards the personalized care of renal cancers. Semin Cancer Biol 55:8–15
Li B, Chan HL, Chen P (2019b) Immune Checkpoint Inhibitors: Basics and Challenges. Curr Med Chem 26(17):3009–3025
Li D, Liu S, Xu J, Chen L, Xu C, Chen F, Xu Z, Zhang Y, Xia S, Shao Y et al (2021) Ferroptosis-related gene CHAC1 is a valid indicator for the poor prognosis of kidney renal clear cell carcinoma. J Cell Mol Med 25(7):3610–3621
Li Z, Song G, Guo D, Zhou Z, Qiu C, Xiao C, Wang X, Wang Y (2022) Identification of GINS2 prognostic potential and involvement in immune cell infiltration in hepatocellular carcinoma. J Cancer 13(2):610–622
Liu S, Wang Y, Miao C, Xing Q, Wang Z (2021a) High expression of CDCA7 predicts poor prognosis for clear cell renal cell carcinoma and explores its associations with immunity. Cancer Cell Int 21(1):140
Liu B, Guo S, Li GH, Liu Y, Liu XZ, Yue JB, Guo HY (2021b) CHMP4C regulates lung squamous carcinogenesis and progression through cell cycle pathway. J Thorac Dis 13(8):4762–4774
Neves H, Kwok HF (2017) In sickness and in health: The many roles of the minichromosome maintenance proteins. Biochim Biophys Acta 1868(1):295–308
Pacek M, Walter JC (2004) A requirement for MCM7 and Cdc45 in chromosome unwinding during eukaryotic DNA replication. EMBO J 23(18):3667–3676
Piao J, Sun J, Yang Y, Jin T, Chen L, Lin Z (2018) Target gene screening and evaluation of prognostic values in non-small cell lung cancers by bioinformatics analysis. Gene 647:306–311
Pollok S, Bauerschmidt C, Sänger J, Nasheuer HP, Grosse F (2007) Human Cdc45 is a proliferation-associated antigen. FEBS J 274(14):3669–3684
Pruitt SC, Bailey KJ, Freeland A (2007) Reduced Mcm2 expression results in severe stem/progenitor cell deficiency and cancer. Stem Cells (dayton, Ohio) 25(12):3121–3132
Siegel RL, Miller KD, Jemal A (2017) Cancer Statistics, 2017. CA Cancer J Clin 67(1):7–30
Stanzani E, Martínez-Soler F, Mateos TM, Vidal N, Villanueva A, Pujana MA, Serra-Musach J, de la Iglesia N, Giménez-Bonafé P, Tortosa A (2017) Radioresistance of mesenchymal glioblastoma initiating cells correlates with patient outcome and is associated with activation of inflammatory program. Oncotarget 8(43):73640–73653
Sun J, Shi R, Zhao S, Li X, Lu S, Bu H, Ma X (2017) Cell division cycle 45 promotes papillary thyroid cancer progression via regulating cell cycle. Tumour Biol J Int Soc Oncodevelop Biol Med 39(5):1010428317705342
Sun D, Wang J, Han Y, Dong X, Ge J, Zheng R, Shi X, Wang B, Li Z, Ren P et al (2021) TISCH: a comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment. Nucleic Acids Res 49(D1):D1420-d1430
Szklarczyk D, Franceschini A, Kuhn M, Simonovic M, Roth A, Minguez P, Doerks T, Stark M, Muller J, Bork P et al (2011) The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucl Acids Res 39:561–568
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, Porta-Pardo E, Gao GF, Plaisier CL, Eddy JA et al (2018) The Immune Landscape of Cancer. Immunity 48(4):812-830.e814
Tomita Y, Imai K, Senju S, Irie A, Inoue M, Hayashida Y, Shiraishi K, Mori T, Daigo Y, Tsunoda T et al (2011) A novel tumor-associated antigen, cell division cycle 45-like can induce cytotoxic T-lymphocytes reactive to tumor cells. Cancer Sci 102(4):697–705
Tyagi N, Marimuthu S, Bhardwaj A, Deshmukh SK, Srivastava SK, Singh AP, McClellan S, Carter JE, Singh S (2016) p-21 activated kinase 4 (PAK4) maintains stem cell-like phenotypes in pancreatic cancer cells through activation of STAT3 signaling. Cancer Lett 370(2):260–267
Vennin C, Mélénec P, Rouet R, Nobis M, Cazet AS, Murphy KJ, Herrmann D, Reed DA, Lucas MC, Warren SC et al (2019) CAF hierarchy driven by pancreatic cancer cell p53-status creates a pro-metastatic and chemoresistant environment via perlecan. Nat Commun 10(1):3637
Wang H, Peng R, Wang J, Qin Z, Xue L (2018) Circulating microRNAs as potential cancer biomarkers: the advantage and disadvantage. Clin Epigenetics 10:59
Wang Z, Zhao Y, An Z, Li W (2019) Molecular Links Between Angiogenesis and Neuroendocrine Phenotypes in Prostate Cancer Progression. Front Oncol 9:1491
Wang Y, Chen H, Zhang J, Cheng ASL, Yu J, To KF, Kang W (2020) MCM family in gastrointestinal cancer and other malignancies: From functional characterization to clinical implication. Biochim Biophys Acta 1874(2):188415
Wu W, Wang X, Shan C, Li Y, Li F (2018) Minichromosome maintenance protein 2 correlates with the malignant status and regulates proliferation and cell cycle in lung squamous cell carcinoma. Onco Targets Ther 11:5025–5034
Yu Y, Huang Y, Li C, Ou S, Xu C, Kang Z (2022) Clinical value of M1 macrophage-related genes identification in bladder urothelial carcinoma and in vitro validation. Front Genet 13:1047004
Zhang YX, Zhao YL (2016) Pathogenic Network Analysis Predicts Candidate Genes for Cervical Cancer. Comput Math Methods Med 2016:3186051
Zhang W, Gong W, Ai H, Tang J, Shen C (2014) Gene expression analysis of lung adenocarcinoma and matched adjacent non-tumor lung tissue. Tumori 100(3):338–345
Zhou J, Jiang Y, Zhao J, Zhang H, Fu J, Luo P, Ma Y, Zou D, Gao H, Hu J et al (2020) Dp44mT, an iron chelator, suppresses growth and induces apoptosis via RORA-mediated NDRG2-IL6/JAK2/STAT3 signaling in glioma. Cell Oncol (dordr) 43(3):461–475
Zou L, Mitchell J, Stillman B (1997) CDC45, a novel yeast gene that functions with the origin recognition complex and Mcm proteins in initiation of DNA replication. Mol Cell Biol 17(2):553–563
Acknowledgements
We would like to thank the researchers and study participants for their contributions.
Funding
This work was supported by Nantong Science and Technology Planning Project: JC2021183.
Author information
Authors and Affiliations
Contributions
XY.Z and JH.Z and Y.W: manuscript writing/editing/revision; JH.Z: data collection or management; X.W: data analysis; QW.X: protocol/project development; BY.Z: data analysis and manuscript revision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics Approval and Consent to Participate
The study was approved by the Institutional Research Ethics Committees of Affiliated Hospital of Nantong University and written informed consent was obtained.
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, X., Zhou, J., Wang, Y. et al. Elevated CDC45 Expression Predicts Poorer Overall Survival Prognoses and Worse Immune Responses for Kidney Renal Clear Cell Carcinoma via Single-Cell and Bulk RNA-Sequencing. Biochem Genet 62, 1502–1520 (2024). https://doi.org/10.1007/s10528-023-10500-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-023-10500-y