Abstract
Circular RNAs function as important regulators in the pathogenesis of human cancers, including nasopharyngeal carcinoma (NPC). We aimed to explore the functions of circ_0028007 in NPC development. Quantitative real-time polymerase chain reaction assay was employed for the levels of circ_0028007, NUAK family kinase 1, microRNA-656-3p (miR-656-3p), and E74 like ETS transcription factor 2 (ELF2). RNase R assay was used to verify the feature of circ_0028007. Cell Counting Kit-8 assay and colony formation assay were performed to assess cell growth. Wound-healing assay and transwell assay were adopted to analyze cell migration and invasion. Tube formation assay was conducted for cell angiogenic capacity. Flow cytometry analysis was performed for cell apoptosis. Western blot assay was conducted for protein levels. Compared to normal tissues and cells, circ_0028007 level was elevated in NPC tissues and cells. Knockdown of circ_0028007 repressed NPC cell growth, migration, invasion, and angiogenesis, facilitated apoptosis in vitro and blocked tumor growth in vivo. Moreover, circ_0028007 silencing could regulate the AMP-activated protein kinase/mammalian target of rapamycin pathway in NPC cells. Circ_0028007 promoted the malignant behaviors of NPC cells via acting as miR-656-3p sponge. In addition, ELF2 was demonstrated to be the target gene of miR-656-3p. MiR-656-3p overexpression restrained NPC cell malignant phenotypes, while ELF2 elevation reversed the effects. Circ_0028007 contributed to the progression of NPC by decoying miR-656-3p and elevating ELF2. The findings might provide potential targets for NPC therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nasopharyngeal carcinoma (NPC) is a tumor that occurs in nasopharyngeal mucosa, and has a high incidence in China (Wei et al. 2010; Chen et al. 2019a, b). Presently, with the progress of radiotherapy technology, the treatment of NPC has achieved good results (Perri et al. 2019). However, due to the insidious nasopharyngeal location and complex early symptoms, most NPC patients have deteriorated to the advanced stage by the time of seeking medical treatment, and local recurrence and distant metastasis are common in advanced patients, so the prognosis of NPC patients is very dismal (Petersson 2015; Lee et al. 2019). Hence, elucidating the mechanism of NPC progression and finding better strategies for NPC treatment are very crucial.
Circular RNAs (circRNAs) are derived from reverse splicing of protein-coding genes and play an important role in indicating pathophysiological changes within organisms (Jeck et al. 2014). Furthermore, circRNAs were able to serve as the sponge for microRNAs (miRNAs) to modulate gene expression, thereby altering the biological processes (Panda 2018; Yin et al. 2019). Currently, the regulatory roles of circRNAs in tumor progression have gradually been identified, including NPC. For example, circ_0081534 aggravated the malignancy of NPC by altering miR-508-5p/FN1 axis (Li et al. 2020). CircSERPINA3 sponged miR-944 to elevate MDM2 expression, thereby facilitating NPC cell growth and invasion (Liu et al. 2020a, b). Circ-ZNF609 promoted NPC growth and motility by decoying miR-150-5p (Zhu et al. 2019). As a member of circRNA, circ_0028007 has been demonstrated to promote cell migration and improve chemoresistance in NPC cells (Qiongna et al. 2020). Even so, the exact roles of circ_0028007 in NPC development are largely unclear.
MiRNAs are short non-coding RNAs with about ~ 22 nucleotides, which play vital roles in malignant tumor progression (Peng et al. 2016). Up to date, multiple miRNAs, such as miR-96-5p (Luo et al. 2020), miR-34a (Jiang et al. 2020), miR-204 (Zong et al. 2020), and miR-18a (Mai et al. 2019), have been confirmed to be related to the progression of NPC by altering tumor cell viability, apoptosis, invasion and migration. Moreover, Tian et al. declared that miR-656-3p took part in regulating NPC cell malignant phenotypes by lncRNA SNHG8/miR-656-3p/SATB1 axis (Tian et al. 2020). E74 like ETS transcription factor 2 (ELF2) is a member of the ELF subfamily and functions as a promoter in NPC development (Liu et al. 2017). Through bioinformatics analysis, miR-656-3p was found to possess the complementary sequences of circ_0028007 and ELF2. However, the relationships among circ_0028007, miR-656-3p, and ELF2 are not reported.
In this present work, the expression profiles of circ_0028007, miR-656-3p, and ELF2 in NPC were detected. Moreover, the roles and related mechanisms of circ_0028007 were clarified.
Materials and Methods
Tissues Acquisition
Thirty-six NPC patients (including 22 patients at stages I and II and 14 patients at stage III) who underwent surgical treatment at An'kang Central Hospital were enrolled in our study. The tumor tissues and adjacent non-tumor tissues were harvested from the patients after the research was approved by the Ethics Committee of An'kang Central Hospital and written informed consents were offered by all patients. The study was conducted in accordance with the Declaration of Helsinki. The samples were preserved at − 80 °C. All patients did not receive any treatment before this research. The following patients were excluded: (1) complicated with severe liver and kidney injury; (2) intolerance to surgery; (3) a history of cancer; (4) in pregnancy. The clinicopathological characteristics of NPC patients were shown in Table 1.
Cell Culture
Normal nasopharyngeal epithelial cell line (NP69) and three NPC cell lines (SUNE-1, C666-1 and 5-8F) were purchased from MingZhoubio. (Ningbo, China). These cells were cultivated at 37 °C in RPMI 1640 medium (Sigma-Aldrich, St. Louis, MO, USA) added with 10% fetal bovine serum (FBS; Sigma-Aldrich) and 1% penicillin–streptomycin (Sigma-Aldrich) in the condition of 5% CO2.
RNA Extraction, RNase R Treatment, and Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR) Assay
TRIzol reagent (Beyotime, Shanghai, China) was employed to extract total RNA. Total RNA was exposed to 3 U/μg RNase R (Epicenter Biotechnologies, Madison, WI, USA) for 15 min. Then PrimeScript™ RT reagent Kit (Takara, Dalian, China) or miRNA 1st Strand cDNA Synthesis Kit (Vazyme, Nanjing, China) was adopted to generate cDNAs. QRT-PCR was done through the usage of BeyoFast™ SYBR Green qPCR Mix (Beyotime) and specific primers (GeneCopoeia, Guangzhou, China). The relative expression was computed with the 2−ΔΔCt way and normalized to GAPDH or U6. The primers were as follows: circ_0028007: (F: 5′-TTTCGTTGGAAACTGATTTTTG-3′ and R: 5′-GCTGGCATATTCCATGATGA-3′); NUAK family kinase 1 (NUAK1): (F: 5′-AAGGCACCTACGGCAAAGTC-3′ and R: 5′-GTCTGATGTGAACCATGTCTTGT-3′); miR-656-3p: (F: 5′-ACACTCCAGCTGGGAATATTATACAGTCA-3′ and R: 5′-CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAGAGGUUG-3′); ELF2: (F: 5′-AAACTGTAGTGGAGGTGTCAACT-3′ and R: 5′-CATGGCTATCTGGTGATGTTGG-3′); GAPDH: (F: 5′-GAAGGTGAAGGTCGGAGTC-3′ and R: 5′-GAAGATGGTGATGGGATTTC-3′); U6: (F: 5′-CTCGCTTCGGCAGCACA-3′ and R: 5′-AACGCTTCACGAATTTGCGT-3′).
Cell Transfection
Specific small interfering RNA against circ_0028007 (si-circ_0028007; 5′-TTGTTCTCAAACACAGTTCAT-3′) and si-NC, short hairpin RNA against circ_0028007 (sh-circ_0028007; 5′-CTTTGTTCTCAAACACAGTTC-3′) and sh-NC, the mimics of miR-656-3p (miR-656-3p; 5′-AAUAUUAUACAGUCAACCUCU-3′) and scramble mimics (miR-NC), the inhibitors of miR-656-3p (anti-miR-656-3p; 5′-AGAGGTTGACTGTATAATATT-3′) and anti-miR-NC, ELF2 overexpression vector (ELF2), and control vector (vector) were provided by GeneCopoeia. Then SUNE-1 and 5-8F cells were plated into 6-well plates and cell transfection was done utilizing Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA).
Cell Growth Determination
Cell growth was evaluated by Cell Counting Kit-8 (CCK-8) experiments and colony formation test. For CCK-8 assay, SUNE-1 and 5-8F cells were added into 96-well plates with 1 × 104 cells per well. After cultivation for 0, 24, 48, and 72 h, 10 μL CCK-8 solution (Sigma-Aldrich) was supplemented into each well and incubated for another 4 h. The absorption at 450 nm was measured utilizing a microplate reader (Bio-Rad, Hercules, CA, USA).
For the colony formation test, SUNE-1 and 5-8F cells were incubated for 14 days in 6-well plates. Next, the colonies were fixed in paraformaldehyde (Sangon, Shanghai, China) and dyed with 0.1% crystal violet (Sangon). Ultimately, the colonies were imaged and counted.
Wound-Healing Assay
In brief, the transfected cells (1 × 104 cells) were seeded into 6-well plates and cultured until 90% confluence. The scratches were then created with a pipette tip. The scratch width was recorded at 0 and 24 h. The migration rate was estimated via the formula: (width at 0 h-width at 24 h)/width at 0 h.
Transwell Assay
The migration and invasion were evaluated utilizing the Transwell chambers (BD Bioscience, San Jose, CA, USA) coated without and with Matrigel (BD Bioscience), respectively. In brief, the transfected SUNE-1 and 5-8F cells (3 × 104) suspended in serum-free medium were plated into the top chamber and 600 μL complete culture medium was filled into the lower chamber. After 24 h, the migrated/invaded cells were stained with 0.1% crystal violet (Sangon). The results were counted under a microscope (100×; Olympus, Tokyo, Japan).
Tube Formation Assay
The angiogenic capacity of HUVECs (Procell, Wuhan, China) was analyzed by tube formation assay. HUVECs were seeded into 96-well plates that contained dissolved Matrigel matrix (BD Bioscience). Then the media from SUNE-1 and 5-8F cells with various transfection were harvested and added into the well. After 8 h of incubation at 37 °C, the branches in each well were photographed and counted under a microscope (100×; Olympus).
Flow Cytometry Analysis
The SUNE-1 and 5-8F cells with various transfection were suspended in binding buffer. Next, Annexin V-fluorescein isothiocyanate (FITC; Vazyme) and propidium iodide (PI; Vazyme) were used to stain cells. The results were examined with FACScan® flow cytometry (BD Bioscience). All procedures were in line with the instructions of Annexin V-FITC/PI Apoptosis Detection Kit (Vazyme).
Western Blot Assay
Total protein was obtained with RIPA buffer (Beyotime) and examined with BCA Protein Quantification Kit (Vazyme). Then sodium dodecyl sulfate–polyacrylamide gel (Sangon) electrophoresis was conducted to separate the protein. The separated proteins were then blotted to polyvinylidene difluoride membranes (Sangon). Thereafter, the membranes were blocked in 5% defatted milk and kept with primary antibodies overnight at 4 °C and secondary antibody (ab97023; 1:10,000; Abcam, Cambridge, MA, USA) for 2 h at room temperature. At last, the protein blots were visualized with the ECL kit (Vazyme). The primary antibodies including phospho-AMP-activated protein kinase (p-AMPK; ab92701; 1:1000; Abcam), AMPK (ab32407; 1:1000; Abcam), p-mammalian target of rapamycin (mTOR) (ab84400; 1:1000; Abcam), mTOR (ab32028; 1:1000; Abcam), ELF2 (ab225958; 1:1000; Abcam), and GAPDH (ab37168; 1:10,000; Abcam). The measurements were repeated 3 times.
Subcellular Fractionation Location Assay
The PARIS Kit (Invitrogen) was employed to separate the cytoplasm and nucleus of SUNE-1 and 5-8F cells strictly following the instructions. The level of circ_0028007 was examined with the aforementioned qRT-PCR assay. GAPDH and U6 served as the controls for cytoplasmic transcript and nuclear transcript, respectively.
Dual-Luciferase Reporter Assay
The fragments of wild type (WT) circ_0028007 or ELF2 3′UTR including miR-656-3p binding sequences were cloned into pmirGLO plasmid (Promega, Madison, WI, USA) to establish WT-circ_0028007 and WT-ELF2 3′UTR. The mutant vectors (MUT-circ_0028007 and MUT-ELF2 3′UTR) were generated by mutating the binding sequences of miR-656-3p. Then the luciferase vectors and miR-656-3p or miR-NC were transfected into NPC cells and incubated for 48 h. The luciferase activity was measured with Dual-Luciferase Reporter Assay Kit (Promega).
Murine Xenograft Model
The BALB/c nude mice were acquired from Beijing Vital River Laboratory Animal Technology Co., Ltd. (Beijing, China) and divided into 2 groups (n = 6). The SUNE-1 cells with sh-circ_0028007 or sh-NC stably transfection were constructed by lentivirus transfection by GeneCopoeia and then injected into the mice. Tumor volume was examined once a week for 4 weeks and evaluated with the formula: Length × Width2 × 0.5. After 4 weeks, the mice were euthanized. The collected tumors were weighted and saved at − 80 °C until further use. The animal study was permitted by the Ethics Committee of Animal Research of An'kang Central Hospital and performed in accordance with the Guidelines for Care and Use of Laboratory Animals of "National Institutes of Health".
Statistical Analysis
All experiments were conducted triple times. The obtained data were evaluated by GraphPad Prism 7 (GraphPad Inc., La Jolla, CA, USA) and presented as mean ± standard deviation. The significance was evaluated by Student’s t-test or one-way analysis of variance. It was considered as significant if P < 0.05.
Results
Circ_0028007 was Increased in NPC Tissues and Cells
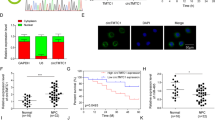
To investigate the function of circ_0028007 in NPC development, the expression of circ_0028007 in NPC tissues was firstly determined by qRT-PCR assay. As a result, compared to normal tissues, circ_0028007 level was increased in NPC tissues (Fig. 1A). Moreover, circ_0028007 level was positively correlated with the clinical pathological stage of NPC (Fig. 1B). Circ_0028007 level in NPC patients was also related to lymph node metastasis and tumor size (Table 1). Then the expression of circ_0028007 in NPC cells was detected. The results showed that circ_0028007 was highly expressed in SUNE-1, C666-1, and 5-8F cells compared to NP69 cells (Fig. 1C). In addition, we found that circ_0028007 was resistant to RNase R treatment, while NUAL1 mRNA was evidently digested by RNase R, indicating that circ_0028007 was stable than linear transcript (Fig. 1D). These results suggested the potential role of circ_0028007 in NPC.
Expression of circ_0028007 in NPC tissues and cells. A The expression of circ_0028007 in NPC tissues and normal tissues was detected by qRT-PCR assay. B The expression of circ_0028007 in NPC tissues at different clinical stages was examined by qRT-PCR assay. C The expression of circ_0028007 in NP69, SUNE-1, C666-1, and 5-8F cells was examined by qRT-PCR assay. D After total RNA in SUNE-1 and 5-8F cells was treated with or without RNase R, the levels of circ_0028007 and NUAK1 mRNA were detected by qRT-PCR assay. *P < 0.05
Silencing of circ_0028007 Repressed Cell Growth, Migration, Invasion, and Angiogenesis and Promoted Apoptosis in NPC Cells
As shown in Fig. 2A, si-circ_0028007 transfection conspicuously decreased circ_0028007 expression in SUNE-1 and 5-8F cells, but did not affect NUAK1 mRNA expression. CCK-8 assay indicated that circ_0028007 knockdown suppressed the viability of SUNE-1 and 5-8F cells compared to si-NC control groups (Fig. 2B, C). The results of colony formation assay showed that the colony formation capacity of SUNE-1 and 5-8F cells was repressed by the downregulation of circ_0028007 (Fig. 2D). Wound-healing assay exhibited that circ_0028007 silencing blocked the migration of SUNE-1 and 5-8F cells in contrast to si-NC control groups (Fig. 2E). As illustrated by transwell assay, circ_0028007 interference inhibited the abilities of SUNE-1 and 5-8F cells to migrate and invade in comparison with control groups (Fig. 2F). Tube formation assay indicated that circ_0028007 silencing repressed tube formation ability relative to si-NC groups (Fig. 2G). Moreover, flow cytometry analysis presented that circ_0028007 knockdown facilitated the apoptosis of SUNE-1 and 5-8F cells compared to si-NC control groups (Fig. 2H). AMPK/mTOR pathway has been reported to play a vital role in tumor cell growth (Faubert et al. 2015). Thus, we detected the function of circ_0028007 in AMPK/mTOR pathway by detecting the levels of p-AMPK/AMPK and p-mTOR/mTOR in SUNE-1 and 5-8F cells. The results exhibited that circ_0028007 silencing increased p-AMPK/AMPK ratio and decreased p-mTOR/mTOR ratio in SUNE-1 and 5-8F cells (Fig. 2I, J). Collectively, circ_0028007 knockdown suppressed the malignant behaviors of NPC cells.
Circ_0028007 knockdown restrained the progression of NPC cells. SUNE-1 and 5-8F cells were transfected with si-NC or si-circ_0028007. A The levels of circ_0028007 and NUAK1 mRNA in SUNE-1 and 5-8F cells were detected by qRT-PCR assay. B and C The viability of SUNE-1 and 5-8F cells was evaluated by CCK-8 assay. D The colony formation capacity of SUNE-1 and 5-8F cells was analyzed by colony formation assay. E The migration of SUNE-1 and 5-8F cells was assessed by wound-healing assay. F The migration and invasion of SUNE-1 and 5-8F cells were investigated by transwell assay. G The tube formation ability of HUVECs was analyzed by tuba formation assay. H The apoptosis of SUNE-1 and 5-8F cells was analyzed by flow cytometry analysis. I and J The protein levels of p-AMPK, AMPK, p-mTOR, and mTOR in SUNE-1 and 5-8F cells were measured by western blot assay. *P < 0.05
Circ_0028007 Sponged miR-656-3p to Regulate miR-656-3p Expression
To explore the underlying mechanism of circ_0028007 in NPC cell development, we analyzed the potential target miRNAs of circ_0028007 using circular RNA interactome (https://circinteractome.nia.nih.gov/) and starbase (https://starbase.sysu.edu.cn/starbase2/index.php). Then we selected 6 miRNAs (miR-1179, miR-326, miR-331-3p, miR-346, miR-496, and miR-656-3p) that targeted by circ_0028007 and investigated the effect of circ_0028007 knockdown on their expression in SUNE-1 and 5-8F cells. The results showed that circ_0028007 knockdown markedly increased miR-656-3p expression than other miRNAs. Thus, we selected miR-656-3p (Fig. S1A, B). The binding sites between miR-656-3p and circ_0028007 were presented in Fig. 3A. Subcellular fractionation location assay showed that circ_0028007 mainly localized in the cytoplasm of SUNE-1 and 5-8F cells, which suggested the combination possibility of circ_0028007 and miR-656-3p in space (Fig. 3B, C). Then miR-656-3p transfection elevated miR-656-3p expression in SUNE-1 and 5-8F cells (Fig. 3D). As demonstrated by dual-luciferase reporter assay, miR-656-3p overexpression repressed the luciferase activity of WT-circ_0028007 but not the luciferase activity of MUT-circ_0028007 in SUNE-1 and 5-8F cells (Fig. 3E, F). These results indicated the interaction between miR-656-3p and circ_0028007. Indeed, there was a reduction in miR-656-3p expression in NPC tissues and cells compared to normal tissues and cells (Fig. 3G, H). Thereafter, anti-miR-656-3p was transfected into SUNE-1 and 5-8F cells to decrease miR-656-3p expression and the results were demonstrated by qRT-PCR assay (Fig. 3I). In addition, our results exhibited that circ_0028007 knockdown led to a noteworthy elevation in miR-656-3p expression in SUNE-1 and 5-8F cells, while anti-miR-656-3p transfection reversed the effect (Fig. 3J, K). Taken together, circ_0028007 functioned as the sponge for miR-656-3p.
Circ_0028007 directly targeted miR-656-3p. A The binding sites between circ_0028007 and miR-656-3p. B and C The expression of circ_0028007 in the cytoplasm and nucleus of SUNE-1 and 5-8F cells was detected by qRT-PCR assay. D After SUNE-1 and 5-8F cells were transfected with miR-656-3p or miR-NC, the expression of miR-656-3p was detected via qRT-PCR assay. E and F The interaction between circ_0028007 and miR-656-3p was analyzed by dual-luciferase reporter assay. G and H The expression of miR-656-3p in NPC tissues, cells and corresponding normal tissues and cells was determined by qRT-PCR assay. I The expression of miR-656-3p in anti-miR-NC or anti-miR-656-3p transfected SUNE-1 and 5-8F cells was examined by qRT-PCR assay. J and K SUNE-1 and 5-8F cells were transfected with si-NC, si-circ_0028007, si-circ_0028007+anti-miR-NC, or si-circ_0028007+anti-miR-656-3p and then the expression of miR-656-3p was detected by qRT-PCR assay. *P < 0.05
Circ_0028007 Repressed NPC Cell Progression by Targeting miR-656-3p
Afterward, the relationship between circ_0028007 and miR-656-3p in the regulation of NPC cell malignant behaviors was investigated. The results of CCK-8 assay and colony formation assay showed that miR-656-3p inhibition abrogated the suppressive roles of circ_0028007 knockdown in cell viability and colony formation in both SUNE-1 and 5-8F cells (Fig. 4A–C). Transwell assay indicated that circ_0028007 silencing repressed the migration and invasion of SUNE-1 and 5-8F cells, while these effects were weakened by decreasing miR-656-3p (Fig. 4D–F). Tube formation assay showed that circ_0028007 knockdown-mediated suppressive role in angiogenesis was ameliorated by reducing miR-656-3p (Fig. 4G). The promotional effect of circ_0028007 knockdown on SUNE-1 and 5-8F cell apoptosis was abated by the inhibition of miR-656-3p (Fig. 4H). Moreover, our results showed that miR-656-3p suppression partly overturned the effects of circ_0028007 knockdown on p-AMPK/AMPK and p-mTOR/mTOR levels in SUNE-1 and 5-8F cells (Fig. 4I, G). All these results suggested that circ_0028007 silencing suppressed NPC cell growth, migration, invasion, and angiogenesis, promoted apoptosis and regulated AMPK/mTOR pathway by sponging miR-656-3p.
Circ_0028007 knockdown suppressed NPC cell malignancy behaviors by sponging miR-656-3p. SUNE-1 and 5-8F cells were transfected with si-NC, si-circ_0028007, si-circ_0028007+anti-miR-NC, or si-circ_0028007+anti-miR-656-3p. A–C The viability and colony formation of SUNE-1 and 5-8F cells were assessed by CCK-8 assay and colony formation assay, respectively. D–F The migration and invasion of SUNE-1 and 5-8F cells were evaluated by wound-healing assay and transwell assay. G The tube formation capacity was assessed by tube formation assay. H The apoptosis of SUNE-1 and 5-8F cells was analyzed by flow cytometry analysis. I and J The protein levels of p-AMPK, AMPK, p-mTOR, and mTOR in SUNE-1 and 5-8F cells were measured through western blot assay. *P < 0.05
MiR-656-3p Directly Targeted ELF2
In order to further explore the related mechanism of circ_0028007/miR-656-3p in NPC cell development, bioinformatics software targetscan (http://www.targetscan.org/vert_72//) was adopted to analyze the target gene of miR-656-3p. Then we selected 6 genes (MDK, MARCH5, ELF2, LPAR1, MUAK1, and KLF9) that were reported to be related to NPC progression. Moreover, our results showed that miR-656-3p overexpression decreased ELF2 expression in SUNE-1 and 5-8F cells. Thus, we selected ELF2 for further experiments (Fig. S1C, D). The complementary sequences between ELF2 and miR-656-3p were exhibited in Fig. 5A. Then dual-luciferase reporter assay further verified the interaction between miR-656-3p and ELF2 for miR-656-3p overexpression suppressed the luciferase activity of WT-ELF2 3’UTR but not the luciferase activity of MUT-ELF2 3’UTR in both SUNE-1 and 5-8F cells (Fig. 5B, C). As expected, the mRNA and protein levels of ELF2 were drastically enhanced in tumor tissues compared to normal tissues (Fig. 5D, E). Compared to NP69 cells, the protein level of ELF2 was increased in SUNE-1, C666-1, and 5-8F cells (Fig. 5F). Next, the protein level of ELF2 in SUNE-1 and 5-8F cells was increased by transfecting ELF2 overexpression vector into SUNE-1 and 5-8F cells (Fig. 5G). Additionally, our results presented that miR-656-3p overexpression reduced the protein level of ELF2 in SUNE-1 and 5-8F cells, with ELF2 elevation rescued the impacts (Fig. 5H, I). In summary, miR-656-3p negatively altered ELF2 expression by targeting ELF2.
ELF2 was the target gene of miR-656-3p. A The complementary sequences between miR-656-3p and ELF2. B and C The combination between miR-656-3p and ELF2 was demonstrated by dual-luciferase reporter assay. D and E The mRNA and protein levels of ELF2 in NPC tissues and normal tissues were examined by qRT-PCR assay and western blot assay, respectively. F The protein level of ELF2 in NP69, SUNE-1, C666-1, and 5-8F cells was measured by western blot assay. G The protein level of ELF2 in ELF2 or vector transfected SUNE-1 and 5-8F cells was measured through western blot assay. H and I SUNE-1 and 5-8F cells were transfected with miR-NC, miR-656-3p, miR-656-3p+vector, or miR-656-3p+ELF2 and then ELF2 protein level was detected by western blot assay. *P < 0.05
Overexpression of miR-656-3p Hampered NPC Cell Malignant Phenotypes by Targeting ELF2
To clarify the association between miR-656-3p and ELF2 in the regulation of NPC cell progression, miR-NC, miR-656-3p, miR-656-3p+vector, or miR-656-3p+ELF2 was transfected into SUNE-1 and 5-8F cells. As illustrated by CCK-8 assay, colony formation assay, wound-healing assay, and transwell assay, miR-656-3p overexpression inhibited the viability, colony formation, migration, invasion, and tube formation of SUNE-1 and 5-8F cells, while these effects were abolished by increasing ELF2 (Fig. 6A–G). The results of flow cytometry analysis indicated that miR-656-3p elevation facilitated cell apoptosis in SUNE-1 and 5-8F cells, with ELF2 upregulation abrogated this impact (Fig. 6H). Furthermore, we found that miR-656-3p overexpression increased the level of p-AMPK/AMPK and decreased the level of p-mTOR/mTOR in SUNE-1 and 5-8F cells, whereas these impacts were weakened by increasing ELF2 (Fig. 6I, J). Taken together, miR-656-3p decelerated NPC cell progression by interacting with ELF2.
ELF2 overexpression reversed the effects of miR-656-3p on NPC cell malignant behaviors. SUNE-1 and 5-8F cells were transfected with miR-NC, miR-656-3p, miR-656-3p+vector, or miR-656-3p+ELF2. A–C The viability and colony formation of SUNE-1 and 5-8F cells were tested by CCK-8 assay and colony formation assay, respectively. D–F The migration and invasion of SUNE-1 and 5-8F cells were assessed by wound-healing assay and transwell assay. G The tube formation was assessed by tube formation assay. H The apoptosis of SUNE-1 and 5-8F cells was analyzed by flow cytometry analysis. I and J The protein levels of p-AMPK, AMPK, p-mTOR, and mTOR in SUNE-1 and 5-8F cells were examined by western blot assay. *P < 0.05
Circ_0028007 Knockdown Suppressed ELF2 Expression by Sponging miR-656-3p
Based on the above results, we further explored the relationships among circ_0028007, miR-656-3p, and ELF2. As shown in Fig. 7A–D, circ_0028007 silencing markedly reduced the mRNA and protein levels of ELF2 in SUNE-1 and 5-8F cells, while these effects were ameliorated by the inhibition of miR-656-3p. These results suggested that circ_0028007 positively modulated ELF2 expression in NPC cells by targeting miR-656-3p.
Circ_0028007 directly targeted miR-656-3p to alter ELF2 expression. A–D SUNE-1 and 5-8F cells were transfected with si-NC, si-circ_0028007, si-circ_0028007+anti-miR-NC, or si-circ_0028007+anti-miR-656-3p, and then the mRNA and protein levels of ELF2 were detected by qRT-PCR assay and western blot assay, respectively. *P < 0.05
Silencing of circ_0028007 Hampered Tumor Formation In Vivo
To explore the function of circ_0028007 in tumor growth in vivo, sh-circ_0028007 or sh-NC transfected SUNE-1 cells were injected into nude mice to establish murine xenograft models. The results showed that the mice with circ_0028007 silencing possessed reduced tumor volume and tumor weight compared to control groups (Fig. 8A–C). Moreover, the levels of circ_0028007 and ELF2 mRNA were decreased and the level of mIR-656-3p was increased in the tumor tissues harvested from sh-circ_0028007 groups compared to sh-NC control groups (Fig. 8D). Western blot assay showed that the levels of ELF2 protein and p-mTOR/mTOR were reduced and the level of p-AMPK/AMPK was increased in the tumor tissues of sh-circ_0028007 groups compared to sh-NC control groups (Fig. 8E). To summarize, circ_0028007 knockdown delayed tumor growth in vivo.
Circ_0028007 contributed to tumorigenesis in vivo. A Tumor volume was examined every week. B The solid tumors were presented. C Tumor weight was examined after 4 weeks. D The levels of circ_0028007, miR-656-3p, and ELF2 mRNA in the collected tumors were detected by qRT-PCR assay. E The levels of ELF2 protein, p-AMPK/AMPK, and p-mTOR/mTOR in the collected tumors were measured by western blot assay. *P < 0.05
Discussion
Recently, the involvement of circRNA/miRNA/mRNA regulatory axis in the tumorigenesis has been widely demonstrated (Rong et al. 2017). For example, circZMYM2 promoted pancreatic cancer development by regulating miR-335-5p/JMJD2C axis (An et al. 2018). Circ-SFMBT2/miR-182-5p/CREB1 axis aggravated the malignancy of gastric cancer (Sun et al. 2018). In the present study, a new regulatory axis of circ_0028007/miR-656-3p/ELF2 was discovered in NPC progression.
In NPC, the abnormal expression of circRNAs has been verified to play a crucial role in tumor development. For instance, circ_0008450 silencing restrained NPC cell motility and growth and enhanced apoptosis by decoying miR-577 to reduce CXCL9 (Wei et al. 2019). Circ_001193 functioned as a tumor contributor in NPC by adsorbing miR-496 (Ke et al. 2020). Although circ_0028007 was reported to be upregulated in NPC and contribute to tumor cell motility (Qiongna et al. 2020), the roles of circ_0028007 in NPC are largely unknown. Herein, circ_0028007 level was elevated in NPC tissues and cells. Interference of circ_0028007 restrained the viability, colony formation, migration, and invasion, and facilitated apoptosis of NPC cells in vitro. Angiogenesis is the product of the growth of new blood vessels from the original blood vessels and is related to the growth progression of solid tumors (Ramjiawan et al. 2017). Moreover, angiogenesis has been demonstrated to play a promotional role in NPC development (Lu et al. 2018; Chen et al. 2020). In the present study, we observed that circ_0028007 deficiency markedly suppressed angiogenesis in NPC cells. Besides, it has been documented that AMPK/mTOR pathway plays a critical role in tumorigenesis, including NPC (Faubert et al. 2015; Murugan 2019; Liu et al. 2020a, b). Therefore, we explored whether AMPK/mTOR pathway was involved in circ_0028007-mediated NPC cell progression. As a result, circ_0028007 knockdown enhanced p-AMPK/AMPK level and reduced p-mTOR/mTOR level in NPC cells, suggesting that circ_0028007 could modulate AMPK/mTOR pathway. In addition, in vivo experiments indicated that circ_0028007 deficiency blocked tumor formation in vivo. All the findings demonstrated the oncogenic function of circ_0028007 in NPC and facilitated our understanding on the pathogenesis of NPC.
Subsequently, the underlying mechanism of circ_0028007 in regulating NPC development was explored. The results exhibited that circ_0028007 functioned as the sponge for miR-656-3p, which directly targeted ELF2. MiR-656-3p has been verified to act as a tumor inhibitor in non-small cell lung cancer via binding to AKT1 (Chen et al. 2019a, b). Moreover, miR-656-3p was weakly expressed in NPC and regulated NPC cell growth and motility by targeting SATB1 (Tian et al. 2020). Similarly, it was found that miR-656-3p was reduced in NPC. Upregulation of miR-656-3p suppressed the malignant behaviors of NPC cells, increased p-AMPK/AMPK level, and decreased p-mTOR/mTOR level. Of note, downregulation of miR-656-3p reversed circ_0028007 knockdown-mediated effects on cell growth, migration, invasion, angiogenesis, apoptosis, and AMPK/mTOR pathway in NPC cells, suggesting that circ_0028007 regulated NPC cell progression by decoying miR-656-3p. This study was the first to explore the relation between circ_0028007 and miR-656-3p. Additionally, the association between miR-656-3p and ELF2 in NPC malignancy was initially investigated in this research, though previous studies showed that ELF2 could be targeted by miR-188 and miR-134-5p to influence NPC progression (Liu et al. 2017; Li et al. 2020). Herein, we observed that ELF2 overexpression rescued the influences of miR-656-3p on NPC cell malignant behaviors and AMPK/mTOR pathway.
In summary, circ_0028007 was increased in NPC and promoted NPC cell growth, migration, invasion, and angiogenesis by altering miR-656-3p/ELF2 axis. Moreover, circ_0028007/miR-656-3p/ELF2 axis participated in regulating AMPK/mTOR pathway. These outcomes might offer novel ideas for the therapeutic targets of NPC treatment.
References
An Y, Cai H, Zhang Y, Liu S, Duan Y, Sun D, Chen X, He X (2018) circZMYM2 competed endogenously with miR-335-5p to regulate JMJD2C in pancreatic cancer. Cell Physiol Biochem 51:2224–2236. https://doi.org/10.1159/000495868
Chen T, Qin S, Gu Y, Pan H, Bian D (2019a) Long non-coding RNA NORAD promotes the occurrence and development of non-small cell lung cancer by adsorbing MiR-656-3p. Mol Genet Genomic Med 7:e757. https://doi.org/10.1002/mgg3.757
Chen YP, Chan ATC, Le QT, Blanchard P, Sun Y, Ma J (2019b) Nasopharyngeal carcinoma. Lancet 394:64–80. https://doi.org/10.1016/S0140-6736(19)30956-0
Chen S, Lv L, Zhan Z, Wang X, You Z, Luo X, You H (2020) Silencing of long noncoding RNA SRRM2-AS exerts suppressive effects on angiogenesis in nasopharyngeal carcinoma via activating MYLK-mediated cGMP-PKG signaling pathway. J Cell Physiol 235:7757–7768. https://doi.org/10.1002/jcp.29382
Faubert B, Vincent EE, Poffenberger MC, Jones RG (2015) The AMP-activated protein kinase (AMPK) and cancer: many faces of a metabolic regulator. Cancer Lett 356:165–170. https://doi.org/10.1016/j.canlet.2014.01.018
Jeck WR, Sharpless NE (2014) Detecting and characterizing circular RNAs. Nat Biotechnol 32:453–461. https://doi.org/10.1038/nbt.2890
Jiang C, Cheng Z, Jiang T, Xu Y, Wang B (2020) MicroRNA-34a inhibits cell invasion and epithelial-mesenchymal transition via targeting AXL/PI3K/AKT/Snail signaling in nasopharyngeal carcinoma. Genes Genomics 42:971–978. https://doi.org/10.1007/s13258-020-00963-3
Ke KJ, Wang CY, Li P, Huang XW, Zhang SL, Li X, Cui XF (2020) Hsa_circ_001193 regulates proliferation and apoptosis of nasopharyngeal carcinoma cells through targeting mir-496. Eur Rev Med Pharmacol Sci 24:3069–3076. https://doi.org/10.26355/eurrev_202003_20671
Lee AWM, Ng WT, Chan JYW, Corry J, Makitie A, Mendenhall WM, Rinaldo A, Rodrigo JP, Saba NF, Strojan P, Suarez C, Vermorken JB, Yom SS, Ferlito A (2019) Management of locally recurrent nasopharyngeal carcinoma. Cancer Treat Rev 79:101890. https://doi.org/10.1016/j.ctrv.2019.101890
Li M, Li Y, Yu M (2020) CircRNA ZNF609 knockdown suppresses cell growth via modulating miR-188/ELF2 axis in nasopharyngeal carcinoma. Onco Targets Ther 13:2399–2409. https://doi.org/10.2147/OTT.S234230
Liu Y, Tao Z, Qu J, Zhou X, Zhang C (2017) Long non-coding RNA PCAT7 regulates ELF2 signaling through inhibition of miR-134-5p in nasopharyngeal carcinoma. Biochem Biophys Res Commun 491:374–381. https://doi.org/10.1016/j.bbrc.2017.07.093
Liu R, Zhou M, Zhang P, Zhao Y, Zhang Y (2020a) Cell proliferation and invasion is promoted by circSERPINA3 in nasopharyngeal carcinoma by regulating miR-944/MDM2 axis. J Cancer 11:3910–3918. https://doi.org/10.7150/jca.42799
Liu Y, Qi X, Zhao Z, Song D, Wang L, Zhai Q, Zhang X, Zhao P, Xiang X (2020b) TIPE1-mediated autophagy suppression promotes nasopharyngeal carcinoma cell proliferation via the AMPK/mTOR signalling pathway. J Cell Mol Med 24:9135–9144. https://doi.org/10.1111/jcmm.15550
Lu J, Liu QH, Wang F, Tan JJ, Deng YQ, Peng XH, Liu X, Zhang B, Xu X, Li XP (2018) Exosomal miR-9 inhibits angiogenesis by targeting MDK and regulating PDK/AKT pathway in nasopharyngeal carcinoma. J Exp Clin Cancer Res 37:147. https://doi.org/10.1186/s13046-018-0814-3
Luo X, He X, Liu X, Zhong L, Hu W (2020) miR-96-5p suppresses the progression of nasopharyngeal carcinoma by targeting CDK1. Onco Targets Ther 13:7467–7477. https://doi.org/10.2147/OTT.S248338
Mai S, Xiao R, Shi L, Zhou X, Yang T, Zhang M, Weng N, Zhao X, Wang R, Liu J, Sun R, Qin H, Wang H (2019) MicroRNA-18a promotes cancer progression through SMG1 suppression and mTOR pathway activation in nasopharyngeal carcinoma. Cell Death Dis 10:819. https://doi.org/10.1038/s41419-019-2060-9
Murugan AK (2019) mTOR: role in cancer, metastasis and drug resistance. Semin Cancer Biol 59:92–111. https://doi.org/10.1016/j.semcancer.2019.07.003
Panda AC (2018) Circular RNAs Act as miRNA sponges. Adv Exp Med Biol 1087:67–79. https://doi.org/10.1007/978-981-13-1426-1_6
Peng Y, Croce CM (2016) The role of microRNAs in human cancer. Signal Transduct Target Ther 1:15004. https://doi.org/10.1038/sigtrans.2015.4
Perri F, Della Vittoria Scarpati G, Caponigro F, Ionna F, Longo F, Buonopane S, Muto P, Di Marzo M, Pisconti S, Solla R (2019) Management of recurrent nasopharyngeal carcinoma: current perspectives. Onco Targets Ther 12:1583–1591. https://doi.org/10.2147/OTT.S188148
Petersson F (2015) Nasopharyngeal carcinoma: a review. Semin Diagn Pathol 32:54–73. https://doi.org/10.1053/j.semdp.2015.02.021
Qiongna D, Jiafeng Z, Yalin H, Ping H, Chuan Z, Xiaojie J, Miaomiao Z, Yiting S, Hui Z (2020) Implication of hsa_circ_0028007 in reinforcing migration, invasion, and chemo-tolerance of nasopharyngeal carcinoma cells. J Clin Lab Anal 34:e23409. https://doi.org/10.1002/jcla.23409
Ramjiawan RR, Griffioen AW, Duda DG (2017) Anti-angiogenesis for cancer revisited: is there a role for combinations with immunotherapy? Angiogenesis 20:185–204. https://doi.org/10.1007/s10456-017-9552-y
Rong D, Sun H, Li Z, Liu S, Dong C, Fu K, Tang W, Cao H (2017) An emerging function of circRNA-miRNAs-mRNA axis in human diseases. Oncotarget 8:73271–73281. https://doi.org/10.18632/oncotarget.19154
Sun H, Xi P, Sun Z, Wang Q, Zhu B, Zhou J, Jin H, Zheng W, Tang W, Cao H, Cao X (2018) Circ-SFMBT2 promotes the proliferation of gastric cancer cells through sponging miR-182-5p to enhance CREB1 expression. Cancer Manage Res 10:5725–5734. https://doi.org/10.2147/CMAR.S172592
Tian X, Liu Y, Wang Z, Wu S (2020) lncRNA SNHG8 promotes aggressive behaviors of nasopharyngeal carcinoma via regulating miR-656-3p/SATB1 axis. Biomed Pharmacother 131:110564. https://doi.org/10.1016/j.biopha.2020.110564
Wei KR, Yu YL, Yang YY, Ji MF, Yu BH, Liang ZH, Reng X (2010) Epidemiological trends of nasopharyngeal carcinoma in China. Asian Pac J Cancer Prev 11:29–32
Wei H, Liu D, Sun J, Mao Y, Zhao L, Zhu W, Xu G, Gao Z (2019) Circular RNA circ_0008450 upregulates CXCL9 expression by targeting miR-577 to regulate cell proliferation and invasion in nasopharyngeal carcinoma. Exp Mol Pathol 110:104288. https://doi.org/10.1016/j.yexmp.2019.104288
Yin Y, Long J, He Q, Li Y, Liao Y, He P, Zhu W (2019) Emerging roles of circRNA in formation and progression of cancer. J Cancer 10:5015–5021. https://doi.org/10.7150/jca.30828
Zhu L, Liu Y, Yang Y, Mao XM, Yin ZD (2019) CircRNA ZNF609 promotes growth and metastasis of nasopharyngeal carcinoma by competing with microRNA-150-5p. Eur Rev Med Pharmacol Sci 23:2817–2826. https://doi.org/10.26355/eurrev_201904_17558
Zong G, Han J, Yue Z, Liu Y, Cui Z, Shi L (2020) Downregulation of miR-204 facilitates the progression of nasopharyngeal carcinoma by targeting CXCR4 through NF-kappaB signaling pathway. J BUON 25:1098–1104
Acknowledgements
None.
Funding
None.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10528_2022_10205_MOESM1_ESM.tif
Supplementary file1 (TIF 519 KB)—The selection of miR-656-3p and ELF2. (A and B) The expression of miR-1179, miR-326, miR-331-3p, miR-346, miR-496 and miR-656-3p in SUNE-1 and 5-8F cells transfected with si-NC or si-circ_0028007 was detected by qRT-PCR. (C and D) The mRNA levels of MDK, MARCH5, ELF2, LPAR1, MUAK1 and KLF9 in SUNE-1 and 5-8F cells transfected with miR-NC or miR-656-3p were detected by qRT-PCR. *P<0.05.
Rights and permissions
About this article
Cite this article
Ma, X., Li, Y. Circ_0028007 Aggravates the Malignancy of Nasopharyngeal Carcinoma by Regulating miR-656-3p/ELF2 Axis. Biochem Genet 60, 2069–2086 (2022). https://doi.org/10.1007/s10528-022-10205-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-022-10205-8