Abstract
YPK9/YOR291W of Saccharomyces cerevisiae encodes a vacuolar membrane protein. Previous research has suggested that Ypk9p is similar to the yeast P5-type ATPase Spf1p and that it plays a role in the sequestration of heavy metals. In addition, bioinformatics analysis has suggested that Ypk9p is a homolog of human ATP13A2, which encodes a protein of the subfamily of P5 ATPases. However, no specific function of Ypk9p has been described to date. In this study, we found, for the first time, that YPK9 is involved in the oxidative stress response and modulation of the replicative lifespan (RLS). We found that YPK9 deficiency confers sensitivity to the oxidative stress inducer hydrogen peroxide accompanied by increased intracellular ROS levels, decreased mitochondrial membrane potential, abnormal mitochondrial function, and increased incidence of early apoptosis in budding yeast. More importantly, YPK9 deficiency can lead to a shortened RLS. In addition, we found that overexpression of the catalase-encoding gene CTA1 can reverse the phenotypic abnormalities of the ypk9Δ yeast strain. Collectively, these findings highlight the involvement of Ypk9p in the oxidative stress response and modulation of RLS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
YPK9/YOR291W of Saccharomyces cerevisiae (S. cerevisiae) encodes a protein localized at the vacuolar membrane. Previous studies have reported that Ypk9p is similar to the yeast P5 ATPase Spf1p (Gitler et al. 2009; Sørensen et al. 2018). In addition, bioinformatics analysis has suggested that among proteins, Ypk9p has the greatest homology to human ATP13A2, which encodes a protein of the subfamily of P5 ATPases (Kühlbrandt 2004; Li et al. 2021; Palmgren and Nissen 2011).
P-type ATPases form a large superfamily of cation and lipid pumps that carry out many fundamental biological processes, including generation of membrane potential for muscle contraction and removal of toxic ions from cells (Kühlbrandt 2004; Tadini-Buoninsegni 2020). Based on sequence homology, the P-type ATPase family is divided into five distinct subfamilies, which are referred to as types P1–P5 (Kühlbrandt 2004; Palmgren and Axelsen 1998; Palmgren and Nissen 2011). Interestingly, P1–P4 ATPases have specific substrates; for example, P2C ATPases are Na+/K+ and H+/K+ pumps in animals, and P2D ATPases are Na+ pumps in fungi (Palmgren and Nissen 2011). However, substrate specificities and biological roles have not been elucidated for any P5 pumps (Palmgren and Nissen 2011).
Previous studies in yeast have reported that deletion of YPK9 leads to sensitivity to certain metals (Gitler et al. 2009; Heins-Marroquin et al. 2019; Schmidt et al. 2009), implicating YPK9 in the transport or regulation of metal cations. Mutations in the ATP13A2 gene are involved in the pathogenesis of neurodegenerative diseases such as Parkinson’s disease (Kong et al. 2014; Spataro et al. 2019), Kufor-Rakeb syndrome (Kett et al. 2015; Ramirez et al. 2006) and neuronal ceroid lipofuscinosis (Bras et al. 2012; Estrada-Cuzcano et al. 2017), implying a potential role of ATP13A2 in senescence; however, the underlying mechanism is largely unknown.
In this paper, we provide the first evidence that YPK9 deficiency leads to sensitivity to oxidative stress, increased intracellular reactive oxygen species (ROS) levels, abnormal mitochondrial function, increased incidence of early apoptosis and shortened replicative lifespan (RLS) in budding yeast. These observations highlight the involvement of Ypk9p in the oxidative stress response and modulation of RLS.
Materials and methods
Yeast strains and plasmids
All S. cerevisiae strains used in this paper (listed in Table 1) were derivatives of the wild-type BY4742 strain.
To generate a YPK9 deletion strain, we used the plasmid pRS306 with a URA3 selection marker as a template and amplified the deletion cassette by polymerase chain reaction (PCR) (Baudin et al. 1993). The primers were 5′-AGCCCAGACTTACTGATAGATCTTGCATATACTCCGGTAAGATTGTACTG.AGAGTGCAC-3′ and 5′ - CATGGTACTTGTACACATACATAGATAAAAATCTTTGCTCTGTGCGGTATTTCACACCG-3′. The PCR product was transformed into BY4742 yeast cells with the standard lithium acetate (LiAc) method, and then the YPK9 gene sequence fragment was replaced via homologous recombination (Jamshad and Darby 2012). The transformants were selected on selective plates (SD-URA). The positive clones were confirmed by PCR.
To generate the CTA1 overexpression plasmid pAUR123CTA1, the yeast CTA1 ORF was amplified from wild-type yeast genomic DNA using PCR. The forward primer (5′-ATAGTCGACATGTCGAAATTGGGACAAGA-3′) contained a Sal I site, and the reverse primer (5′-CGCTCTAGATCAAAATTTGGAGTTACTCG-3′) contained a Xba I site (Zhao et al. 2018). The PCR products were cloned into the empty plasmid pAUR123. The recombinant plasmid pAUR123CTA1 was transformed into the E. coli strain DH5α. DNA sequencing of the recombinant plasmids was performed by Sangon (Shanghai, China).
The plasmid pAUR123CTA1 was transformed into ypk9Δ yeast cells with the standard LiAc method to generate ypk9Δ CTA1 overexpression yeast strain (ypk9Δ CTA1 OX). The transformants were screened on YPD medium plates containing 0.2 g/ml aureobasidin A (AbA) at 30 °C for 2 days. The positive clones were confirmed by PCR.
Culture conditions
For all experiments, the strains were removed from storage at − 80 °C, thawed, inoculated in YPD solid medium, and incubated in a thermostatically controlled incubator at 30 °C for 2 to 3 days. YPD medium containing 1% (w/v) yeast extract, 2% (w/v) peptone, and 2% (w/v) dextrose glucose was used for yeast cell culture. The solid YPD plates contained 2% (w/v) agar. The selective SD-URA plates contained 0.67% (w/v) yeast nitrogen base, 2% (w/v) glucose, 2% (w/v) agar and 0.77 g/l URA dropout amino acid mixture.
Growth curve determination
The growth rates were detected with a Bioscreen C instrument (Growth Curves, USA). First, single colonies were inoculated into YPD medium and grown overnight at 30 °C with shaking. Second, the overnight cell cultures were adjusted to an optical density at 600 nm (OD600) of 0.1 by dilution with YPD medium or H2O2-containing YPD medium in culture plates. The strains were then cultured at 30 °C with shaking, and the OD600 was automatically measured every 2 h for 2–3 days until cell growth plateaued (Jasnos et al. 2005). The results represent the averages from three separate experiments. The Friedman test was used for analysis, and a p value less than 0.05 was considered to indicate statistical significance.
Spot assay
Single colonies of the tested yeast strains were inoculated into YPD medium and grown overnight at 30 °C with shaking. The overnight cell cultures were adjusted to an optical density at 600 nm (OD600) of 0.1. The cells were then diluted with sterile PBS in a fivefold series, and 5 μl of each dilution was spotted onto solid agar plates with or without the stress agent. The plates were incubated at 30 °C, and images were taken 48 h after plating.
Real-time quantitative PCR (RT-qPCR)
Briefly, exponential-phase cells were pelleted and washed twice with PBS after they had been treated with or without 3 mM H2O2 for 1 h. Total RNA was then extracted by using a Yeast RNA Kit (Omega Biotek, USA) according to the manufacturer’s instructions. First-strand cDNA was synthesized using a FastKing RT Kit (with gDNase) (Tiangen, China), and RT-qPCR was performed in a LightCycler 480 instrument with the SYBR Green method. The relative expression of the target genes was quantified relative to the expression of the housekeeping gene PRP8. The gene-specific primers for RT-qPCR are listed in Tables 2 and 3. The results represent the averages from three independent experiments. Student’s t test was used for analysis, and a p-value less than 0.05 was considered to indicate statistical significance.
Yeast petite mutation assay
Petite mutants were identified by the 2,3,5-triphenyltetrazolium chloride (TTC) overlay technique. First, exponential-phase cells were harvested after they had been treated with or without 3 mM H2O2 for 1 h. An aliquot was diluted in PBS and spread onto YPD plates. The plates were incubated at 30 °C for 2 days until colonies formed and then overlain with TTC agar. In this assay, yeast colonies formed by respiration-deficient yeast mutants were white, whereas colonies formed by normal yeast were red (Cho et al. 1998; Laskowski 1954).
Detection of generated ROS
The 2′,7′-dichlorofluorescein diacetate (DCFH-DA) (Beyotime, China) method was used to measure intracellular ROS levels. When DCFH-DA is taken up by cells, it is de-esterified into the ionized free acid DCFH. DCFH is then further oxidized into fluorescent DCF by intracellular ROS (Eruslanov and Kusmartsev 2010). For the ROS generation assay, exponential-phase cells were harvested and washed twice with PBS after they had been treated with or without 3 mM H2O2 for 1 h. Then, the cells were incubated with 5 M DCFH-DA at 30 °C in the dark for 1 h. The cells were again harvested and washed three times with precooled PBS to remove free reagent, and the mean green fluorescence intensity was evaluated by flow cytometry (BD FACSCanto II, USA). The results represent the averages from three separate experiments. Student’s t test was used for analysis, and a p value less than 0.05 was considered to indicate statistical significance.
Detection of MMP
Rhodamine 123 (Rh123) (Beyotime, China) was used as a fluorescent probe to detect MMP (Ludovico et al. 2001). Briefly, yeast cells in the exponential phase (treated with or without 3 mM H2O2 for 1 h) were harvested, washed three times with PBS, and then incubated in liquid YPD medium containing 10 μM Rh123 for 60 min at 30 °C in the dark. Next, the cells were washed three times with PBS, and the fluorescence intensity was analyzed using a flow cytometer (BD FACSCanto II, USA). The results represent the averages from three separate experiments. Student’s t test was used for analysis, and a p value less than 0.05 was considered to indicate statistical significance.
Apoptotic marker assay
Apoptosis was measured using fluorescein isothiocyanate (FITC)-annexin V and caspase-3 activity assays as previously described.
Exposure of phosphatidylserine on the outer cell surface is an early event in apoptosis. Phosphatidylserine normally resides in the inner leaflets of cells facing the cytoplasm but is rapidly translocated to the outer leaflets of the cells facing the surface at the early stage of apoptosis. Surface exposure to phosphatidylserine can be specifically detected by staining with fluorescent FITC-labeled annexin V (Herker et al. 2004).
Briefly, yeast cells in the exponential phase (treated with or without 3 mM H2O2 for 1 h) were harvested, and the cell walls were then digested with zymolyase in a buffer containing 0.5 mM MgCl2, 1.2 M sorbitol, and 35 mM H3PO4 at pH 6.8 and 30 °C. After the cell walls were digested, the cells were washed with PBS. Next, the protoplasts were resuspended in 500 μl of annexin V binding solution, incubated with FITC-conjugated Annexin V for 15 min at room temperature, and then analyzed by flow cytometry (BD FACSCanto II, USA).
Caspase-3 is a key enzyme in apoptosis (Nicholson et al. 1995; Qi et al. 2020). The activity of yeast caspase-3 was determined as described previously (Rona et al. 2015) with a Caspase 3 Activity Assay Kit (Beyotime, China) Briefly, yeast cells in the exponential phase (treated with or without 3 mM H2O2 for 1 h) were harvested and then resuspended in cell lysis buffer (50 mM Tris–HCl, pH 8.0; 50 mM KCl; 2 M citrate; 10% glycerol; and 1 mM PMSF). The cells were then broken with glass beads by vigorous shaking, and the supernatants were used to determine caspase-3 activity according to the manufacturer’s instructions. All experiments were performed at 4 °C.
The results represent the averages from three separate experiments, Student’s t tests were used for analysis, and a p value less than 0.05 was considered to indicate statistical significance.
Catalase activity assay
To determine the intracellular catalase activity, a single yeast colony was inoculated into 5 ml of YPD and grown to the exponential phase at 30 °C. Next, the yeast cells were harvested and suspended in precooled RIPA lysis buffer containing protease inhibitors. The cells were then broken with glass beads by vigorous shaking, and the supernatant was collected by centrifugation at 4 °C. The protein concentration was determined using a Bradford Protein Assay Kit (Beyotime, China). Catalase activity was quantified using commercial assay kits (Beyotime, China).
Determination of RLS
The RLS assay was performed as previously described to count the total number of daughter cells generated by individual mother cells by using an optical microscope equipped with a micromanipulator (Postnikoff and Harkness 2014). All lifespan experiments were carried out on YPD plates. Statistical significance was calculated with the Wilcoxon rank-sum test, and p < 0.05 was considered to indicate statistical significance.
DAPI staining
Staining of exponential phase cells by 4′,6-diamidino-2-phenilindole (DAPI) was carried out as follows. Cells were fixed with 70% (v/v) ethanol for 30 min at room temperature. Then, the cells were washed once with NS buffer (20 mM Tris–HCl pH 7.6, 0.25 M sucrose, 1 mM EDTA, 1 mM MgCl2, 0.1 mM ZnSO4, 0.1 mM CaCl2, 0.8 mM PMSF, 0.05% 2-mercaptoethanol) and resuspended in NS buffer containing 0.5 μg/ml DAPI for 5 min (Massardo et al. 2000). The cells were then dropped on a glass slide, squashed lightly, and observed under UV excitation (Leica DMi8, Germany).
Results and discussion
ypk9Δ yeast cells are sensitive to H2O2
The generation of yeast mutants knocked out for a certain gene and their employment to explore the response to distinct environmental stresses constitute an ideal methodology for discovering gene functions (Santos and Riezman 2012). H2O2 is a widely used radical-generating reagent that can elicit harmful effects on cells, including enhanced DNA damage, oxidative stress and aging (Janero et al. 1991; Popa et al. 2010).
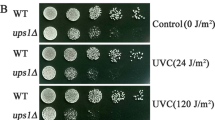
To explore the potential role of YPK9 in the oxidative stress response, the ypk9Δ mutant was generated by PCR-mediated gene disruption. Specifically, the YPK9 gene was replaced with the selectable marker URA3 in the haploid BY4742 background. Then, the ypk9Δ mutant was subjected to oxidative stress induced by H2O2. We found that there was no significant difference between the wild-type strain and the ypk9Δ strain under unstressed conditions. When stressed with 3 mM H2O2, the ypk9Δ strain exhibited obvious growth defects, and the growth defects were more severe under the high (3.5 mM) H2O2 concentration (Fig. 1A). The hypersensitivity to H2O2 of the ypk9Δ cells was further confirmed by the results of the spot assay, the ypk9Δ cells exhibited defective growth compared with wild-type cells (Fig. 1B). In addition, the ypk9Δ cells also exhibited hypersensitivity to oxidative stressor tert-butyl hydroperoxide (TBHP) and cube hydroperoxide (CHP) (data not shown). These observations strongly suggest a role of YPK9 in the oxidative stress response.
ypk9-deleted yeast cells are sensitive to H2O2. The growth curves (A) of the wild-type strain and ypk9Δ strain were assessed with a Bioscreen C machine under 3 mM H2O2 and 3.5 mM H2O2 stressed and unstressed conditions. The results represent the averages from three separate experiments, and a p-value less than 0.05 was considered to indicate statistical significance. (B) Wild-type and ypk9Δ cells were fivefold serially diluted, spotted onto YPD plates (left) and YPD plates containing 3 mM H2O2 (right) and then incubated at 30 °C until colonies formed
YPK9 deficiency induces changes in the mRNA expression levels of oxidative stress-related genes
The growth defects of the ypk9Δ strain under H2O2-stressed conditions implied that the intracellular oxidation and antioxidation systems were imbalanced. Therefore, we monitored the changes in the expression patterns of oxidative stress-related genes in the ypk9Δ strain under H2O2-stressed and unstressed conditions (Jamieson 1998). These genes included superoxide dismutase genes (SOD1, SOD2), catalase genes (CTA1, CTT1), glutathione peroxidase genes (GPX1, GPX2, GPX3), the thioredoxin-2 gene (TRX2), a methionine-s-sulfoxide reductase gene (MXR1), a thioredoxin peroxidase gene (TSA1), a cytoplasmic and mitochondrial glutathione oxidoreductase gene (GLR1), glutathione synthase genes (GSH1, GSH2), and transcription factor genes (YAP1 and SKN7).
The mRNA expression levels of TRX2, MXR1, YAP1, CTA1, and GPX1 were decreased, while those of GSH1 and GPX2 were increased in ypk9Δ cells compared to wild-type control cells under unstressed conditions. After stress with 3 mM H2O2, most of the investigated genes were upregulated in the ypk9Δ strain, such as GPX3, TSA1, GSH1, SKN7, CTA1, GLR1, GSH1, SOD1, GPX2, CTT1, and SOD2 (Fig. 2). We hypothesize that the upregulation of antioxidative gene expression in ypk9Δ cells under stressed conditions may be a compensatory response to oxidative stress, suggesting that the cellular redox status is altered in ypk9Δ cells.
Oxidative stress-related gene expression pattern of ypk9Δ yeast cells. The relative mRNA expression levels of SOD1, SOD2, CTA1, CTT1, GPX1, GPX2, GPX3, TRX2, MXR1, TSA1, GLR1, GSH1, GSH2, YAP1 and SKN7 in the wild-type strain and ypk9Δ strain were measured by RT-qPCR under 3 mM H2O2-stressed and unstressed conditions. All data are expressed as the fold-changes relative to the levels in unstressed wild-type cells, which were set to 1. Differences between groups were analyzed by Student’s t test, and a p-value less than 0.05 was considered to indicate statistical significance. *p < 0.05, unstressed ypk9Δ vs. unstressed wild-type; **p < 0.01, unstressed ypk9Δ vs. unstressed wild-type; Δ indicates p < 0.05, stressed ypk9Δ vs. stressed wild-type; ΔΔ indicates p < 0.01, stressed ypk9Δ vs. stressed wild-type; ΔΔΔ indicates p < 0.001, stressed ypk9Δ vs. stressed wild-type
Notably, the catalase-encoding gene CTA1, which can catalyze the degradation of H2O2 in cells into H2O and O2 (Jamieson 1998), was downregulated in the ypk9Δ strain compared to the wild-type control strain under unstressed conditions, implying a relationship between YPK9 and CTA1 in the oxidative stress response.
In addition, we found that MXR1, a methionine-s-sulfoxide reductase involved in the oxidative stress response and RLS regulation (Kaya et al. 2010; Moskovitz et al. 1997), was also downregulated in the ypk9Δ strain compared to the wild-type control strain under unstressed conditions. Interestingly, a synthetic genetic array analysis showed that MXR1 has a negative genetic interaction with YPK9 (Costanzo et al. 2016), and our observation highlights the potential role of YPK9 in RLS regulation.
YPK9 deficiency increases ROS levels, decreases MMP and results in a high rate of respiration-deficient mutants under H2O2 treatment
H2O2 can induce the production of intracellular ROS that subsequently attack cellular components, including lipids, proteins, and DNA (Janero et al. 1991; Popa et al. 2010). Therefore, we measured the intracellular ROS levels by the DCFH-DA method. As expected, we found that ROS levels were significantly higher in ypk9Δ cells than in wild-type cells under both unstressed and stressed conditions, indicating the occurrence of intracellular oxidative stress in ypk9Δ cells (Fig. 3A).
Increased ROS production and (A) decreased MMP (B) in ypk9Δ yeast cells. Wild-type yeast cells and ypk9Δ yeast cells in the exponential phase were stressed or not stressed with 3 mM H2O2 for 1 h and then subjected to ROS or MMP assays. The results are shown as the mean relative fluorescence intensities from three separate experiments. ypk9Δ yeast cells formed more petite colonies than wild-type when treated with 3 mM H2O2 (C). Differences between groups were analyzed by Student’s t test, and a p value greater than 0.05 was considered to indicate statistical significance. *p < 0.05, **p < 0.01, and ***p < 0.001. (D) Mitochondrial and nuclear DNA stained with DAPI. 1, indicates nuclear DNA, 2, indicates Mitochondrial DNA, 3, indicates cytoplasm. a–d: fluorescent image; e–h: gray image. A bar represents 2.5 μM. (E) The electrophoretogram of the PCR product of CYTB gene
Intracellular ROS can cause damage to DNA, especially mitochondrial DNA (mtDNA) (Kurihara et al. 2012). Damage to mtDNA causes electron transport chain (ETC) dysfunction, generates oxidative stress, impairs MMP regulation and leads to abnormal MMP (Lenaz and Genova 2012).
Given the significant increases in intracellular ROS levels, we next determined whether YPK9 deficiency also affected mitochondrial function. We observed that MMP was not significantly different between wild-type cells and ypk9Δ cells under unstressed conditions; however, after stress with H2O2, MMP of the ypk9Δ yeast strain was lower than that of the wild-type control strain (Fig. 3B).
Generally, under normal physiological conditions, the mitochondrial respiratory system maintains MMP at an adequate level to guarantee intracellular ATP synthesis (Larsen et al. 2012; Moon et al. 2016). The decrease in MMP suggested that mitochondrial respiratory dysfunction occurred in ypk9Δ yeast cells. Therefore, the proportions of respiration-deficient petite cells (result from the loss/mutation of nuclear-encoded proteins, which are essential for the mitochondrial respiration capacity, or from mtDNA mutation or loss) were determined by the TTC overlay technique. Respiring colonies could reduce TTC to a red color, while those with reduced respiratory function are unable to reduce TTC and remain white.
We found that under unstressed conditions, the proportion of petite yeast cells did not differ between the wild-type yeast strain and the ypk9Δ yeast strain. However, after stress with 3 mM H2O2, the ratio of petite yeast cells in the ypk9Δ yeast strain was significantly greater than that in the wild-type control strain (Fig. 3C), suggesting that mitochondrial respiratory dysfunction was serious under stressed conditions in ypk9Δ yeast cells.
There are two kinds of specific types of petite cells: rho0 cells, where the mtDNA has been completely lost, and rho– cells, where there are one or more mutations in the mtDNA that prevent respiratory metabolism, but some residual mtDNA remains in the cell (Ferguson and von Borstel 1992). Therefore, we further determined that the petite mutants of ypk9Δ are rho0 or rho– by the DAPI staining method. Four randomly selected petite colonies (white colonies on the TTC agar plate) of ypk9Δ cells were cultivated in YPD medium to the exponential phase, and the cells were harvested for DAPI staining as previously described (Massardo et al. 2000).
We found no obvious mitochondrial bodies in these ypk9Δ petite cells (only a few cells could be stained with one or two mitochondrial bodies, and these mitochondrial bodies had very weak fluorescence), while obvious mitochondrial bodies could be detected in the wild-type and ypk9Δ cells (red colonies on the TTC agar plate), and the mitochondrial DNA appeared as small beads on strings, as previously described (Massardo et al. 2000). We note that the petite cells of the wild-type were stained with more mitochondrial bodies than ypk9 petite cells (Fig. 3D). In addition, we extracted the total genomic DNA of the 4 ypk9 petite cells and used gene-specific PCR primers (5′-AGAAGTGTAGTAAGAGGT-3′ and 5′-TCAATAGTAGAGATAACAGG-3′) to amplify the fragment (194 bp) of the CYTB gene, which is located in the mitochondrial genome of yeast (Tzagoloff et al. 1976). The electrophoretogram of the PCR product showed that all 4 ypk9Δ petite cells had the specific expected product of wild-type (Fig. 3 E). Both the DAPI staining and PCR results indicated that the ypk9Δ petite cells were rho− not rho0. These findings raise the possibility that YPK9 is involved in stabilizing mitochondria.
In budding yeast, mtDNA is packaged into protein–DNA complexes that are called mitochondrial nucleoids (mt-nucleoids) (Chen and Butow 2005). More than 20 proteins have been identified as potential components of mt-nucleoids in yeast. These proteins play crucial roles in mitochondrial organization and maintenance. For example, ABF2 deficiency leads to an unstable mitochondrial genome and hypersensitivity to damage from endogenously produced free radicals (Diffley and Stillman 1991; O'Rourke et al. 2002). Therefore, we measured the mRNA expression levels of eleven canonical component proteins of mt-nucleoids (Chen and Butow 2005). The results revealed that most of the genes studied, including genes involved in mtDNA packaging and protection (ABF2), mtDNA assembly (ILV5), mtDNA stability (ACO1 and HSP60), citric acid cycle (KGD1), ATP synthesis (ATP1), mtDNA replication (MIP1), and mtDNA transcription (RPO41), were downregulated in YPK9-deficient cells under unstressed conditions (Fig. 4). We speculate that the downregulated mt-nucleoid gene expression pattern may be another reason for the observed high rates of respiration-deficient petite cells in the ypk9Δ yeast strain under stressed conditions, although we do not know why these mt-nucleoid genes were differentially expressed in YPK9-deficient cells.
Mt-nucleoid genes were differentially expressed in ypk9-deficient cells. The relative mRNA expression levels of ABF2, ACO1, ATP1, HSP60, ILV5, KGD1, KGD2, LPD1, MGM101, MIP1 and RPO41 in the wild-type strain and ypk9Δ strain were measured by RT-qPCR under 3 mM H2O2-stressed and unstressed conditions. All data are expressed as the fold-changes relative to the levels in unstressed wild-type cells, which were set to 1. Differences between groups were analyzed by Student's t test, and a p-value less than 0.05 was considered to indicate statistical significance. *p < 0.05, unstressed ypk9Δ vs. unstressed wild-type; **p < 0.01, unstressed ypk9Δ vs. unstressed wild-type; Δ indicates p < 0.05, stressed ypk9Δ vs. stressed wild-type; ΔΔ indicates p < 0.01, stressed ypk9Δ vs. stressed wild-type; ΔΔΔ indicates p < 0.001, stressed ypk9Δ vs. stressed wild-type
YPK9 deficiency induces early apoptosis in yeast
Cells undergoing apoptosis always exhibit a decrease in MMP, which precedes nuclear signs of apoptosis (Dai et al. 2009; Fehrmann et al. 2013). Given the observed decrease in MMP, apoptosis was evaluated by FITC-labeled annexin V staining and caspase-3 activity assays.
The exposure of phosphatidylserine at the outer cell surface is an early event in apoptosis that is remarkably conserved from yeast to mammals and can be specifically detected by staining with FITC-labeled annexin V (Herker et al. 2004). As expected, the mean FITC fluorescence intensity was stronger in ypk9Δ cells than in wild-type control cells under both unstressed and stressed conditions (Fig. 5A).
YPK9 deficiency leads to early apoptosis. wild-type yeast cells and ypk9Δ yeast cells in the exponential phase were stressed or not stressed with 3 mM H2O2 for 1 h. The cell walls were then digested, and the cells were stained with FITC-labeled annexin V for apoptosis assessment (A). Total yeast protein was extracted and subjected to a caspase activity assay (B). Differences between groups were analyzed by Student's t test, and a p value less than 0.05 was considered to indicate statistical significance. *p < 0.05, and **p < 0.01
In addition, caspase-3, one of the most important caspases, plays a central role in mediating nuclear apoptosis, and its activation is a hallmark and early marker of apoptosis. S. cerevisiae contains a metacaspase (encoded by YCA1 gene) that is similar to mammalian metacaspases (Guaragnella et al. 2006; Uren et al. 2000). As expected, caspase activity was also increased in ypk9Δ yeast cells under both unstressed and stressed conditions (Fig. 5B).
As mentioned above, the observed overproduction of intracellular ROS may have been a causal factor for mitochondrial dysregulation and apoptosis in ypk9Δ cells. Previous research has reported that inhibition of Na+/K+-ATPase (belonging to the P2-type ATPase subfamilies) with ouabain can result in increased ROS levels in cardiac cells through a mechanism that may be mediated by Ras and p42/44 mitogen-activated protein kinases (MAPKs) (Xie et al. 1999). Activation of Na+/K+-ATPase attenuates high glucose-induced H9c2 cell apoptosis by suppressing ROS accumulation and MAPKs (Yan et al. 2016). However, based on sequence homology and structural organization analysis, Ypk9p is similar to P5-type ATPases but not similar to Na+/K+-ATPase (Kühlbrandt 2004; Palmgren and Nissen 2011).
YPK9 deficiency leads to shortened RLS
Accumulating evidence suggests that there is crosstalk between apoptosis and aging in model organisms, including Drosophila melanogaster (D. melanogaster), nematodes and S. cerevisiae (Laun et al. 2008). For example, caloric restriction (CR), a method of lifespan elongation that is effective in many model organisms for aging research, can attenuate age-related apoptosis and extend lifespan (Zhang and Herman 2002). However, the relationship between apoptosis and lifespan remains unclear.
Budding yeast has two different paradigms of aging: RLS, which is defined as the number of divisions an individual yeast cell undergoes before dying (Jazwinski 1990), and chronological life span (CLS), which is defined as the length of time a population of yeast cells remains viable in a nondividing state following nutrient deprivation (MacLean et al. 2001).
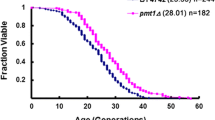
Given the observed early apoptosis of ypk9Δ yeast cells, we monitored RLS and found that the RLS of the ypk9Δ yeast strain (mean RLS = 15, cells = 160) was 32% shorter than that of the wild-type yeast strain (mean RLS = 22, cells = 80) (p < 0.05) (Fig. 6).
YPK9 deficiency leads to shortened RLS. The mean RLS (under 2% glucose conditions) and total number of daughter cells are shown in parentheses; the total number of daughter cells is denoted by “N”. The values were analyzed for statistical significance using the Wilcoxon rank-sum test. p < 0.05 was considered to indicate statistical significance
At least 17 proapoptotic genes and 4 antiapoptotic genes have been reported thus far in budding yeast. Deletion or overexpression of these genes could impact CLS or RLS. The majority of related studies have focused on the connection between apoptosis and chronological aging, and the impact of deleting just a few genes on RLS has been examined (Laun et al. 2012; MacLean et al. 2001; Rockenfeller and Madeo 2008). For example, previous studies have reported that deletion of the antiapoptotic gene MMI1 results in increased RLS (Rinnerthaler et al. 2006). In addition, deletion of DNM1 can extend RLS, possibly by increasing cellular resistance to apoptosis (Scheckhuber et al. 2007). However, very few papers have been published that deal explicitly with the relationship between aging and apoptosis in single-celled organisms, and whether apoptosis or related processes are causes or consequences of aging still needs further research and discussion. On the other hand, apoptosis is well accepted as an anti-aging mechanism in multicellular animals because it can remove damaged cells from tissues and is critical for tissue repair and tissue homeostasis (Schmitt et al. 2007).
Overexpression of CTA1 can reverse the phenotypic abnormalities of the ypk9Δ yeast strain
As mentioned in the previous section, the catalase-encoding gene CTA1 was downregulated in the ypk9Δ strain compared to the wild-type control strain under unstressed conditions. There are two kinds of catalases in budding yeast that catalyze the intracellular degradation of H2O2 into H2O and O2: peroxisomal/mitochondrial catalase A (Cta1p) and cytosolic catalase T (Ctt1p) (Alugoju et al. 2020). Both CTA1 and CTT1 are involved in the oxidative stress response and longevity of S. cerevisiae (Mesquita et al. 2010).
We generated the ypk9Δ CTA1 OX strain (in which CTA1 was overexpressed by using a high-copy-number vector, and both RT-qPCR (Fig. 7A) and catalase activity assays (Fig. 7B) confirmed that CTA1 was overexpressed in ypk9Δ yeast cells) and investigated whether overexpression of CTA1 could eliminate defects associated with YPK9 deficiency.
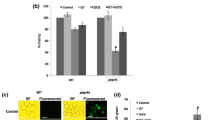
Overexpression of CTA1 eliminates the phenotype associated with YPK9 deficiency. (A) RT-qPCR and (B) catalase activity assays confirmed that CTA1 was overexpressed in ypk9Δ yeast cells. (C) The growth curves of the wild-type strain, the ypk9Δ strain and the ypk9Δ CTA1 OX strain were assessed with a Bioscreen C machine under H2O2-stressed and unstressed conditions. The results represent the averages from three separate experiments. The Friedman test was applied for statistical significance analysis, and a p-value less than 0.05 was considered to indicate statistical significance. (D) The exponential-phase wild-type, ypk9Δ, and ypk9Δ CTA1 OX cells were fivefold serially diluted with PBS and were spotted onto YPD or H2O2-added solid YPD plates, which were kept at 30 °C until colonies formed. Wild-type cells, ypk9Δ cells and ypk9Δ CTA1 OX cells were stressed or not stressed with 3 mM H2O2 for 1 h and then subjected to a ROS assay (E), an MMP assay (F), a caspase activity assay (G) and a FITC-labeled annexin V apoptosis assay (H). The results are displayed as the mean ± SEM value. *p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001. (I) Overexpression of CTA1 restores the RLS of the ypk9Δ strain. The mean RLS and total number of daughter cells are shown in parentheses; the total number of daughter cells is denoted by “N”. The Wilcoxon rank-sum test was applied for statistical significance analysis, and p < 0.05 was considered to indicate statistical significance
We obtained several findings. ①There were no significant differences between the ypk9Δ CTA1 OX and ypk9Δ strains under unstressed conditions, but under H2O2-stressed conditions, ypk9Δ CTA1 OX cells grew faster than ypk9Δ cells (Fig. 7C, D). ② The intracellular ROS levels were lower in ypk9Δ CTA1 OX cells than in ypk9Δ cells under both H2O2-stressed conditions and unstressed conditions (Fig. 7E). ③ MMP was higher in ypk9Δ CTA1 OX yeast cells than in ypk9Δ yeast cells under both H2O2-stressed conditions and unstressed conditions (Fig. 7F). ④ Caspase activity (Fig. 7G) and FITC-labeled annexin V fluorescence intensity (Fig. 7H), which are early markers of apoptosis, were lower in ypk9Δ CTA1 OX cells than in ypk9Δ cells under both H2O2-stressed conditions and unstressed conditions. ⑤ More importantly, the RLS of ypk9Δ CTA1 OX cells was obviously higher than that of ypk9Δ cells and even reached a level similar to that of wild-type cells (Fig. 7I). These findings strongly suggest that overexpression of CTA1 can eliminate defects associated with YPK9 deficiency.
Notably, previous research has reported that CTA1 deletion does not influence the lifespan of wild-type yeast (W303a strain) grown on glucose but decreases the lifespan of wild-type yeast grown on ethanol (Van Zandycke et al. 2002). In contrast, our observations suggest that downregulation of CTA1 expression may play a role in shortening RLS in ypk9Δ cells grown on glucose and imply the existence of an interaction between CTA1 and YPK9.
CR increased the RLS of ypk9Δ
Previous reports have suggested that CR could extend yeast RLS by increasing the rate of yeast mitochondrial respiration (Lin et al. 2002; Schleit et al. 2013). In yeast, mitochondrial function is important for both RLS and CLS (Delaney et al. 2013). CR-mediated pathways could influence mitochondrial segregation and inheritance, nuclear-mitochondrial communication, and mitochondrial quality control, ultimately affecting RLS (Ruetenik and Barrientos 2015).
Altered mitochondrial metabolism is one hallmark of aging (Lin and Beal 2006). The aging process is always accompanied by decreased efficiency of the mitochondrial respiratory chain, reduced MMP and ATP generation, and increased electron leakage (Amigo et al. 2016). Since we observed that YPK9 deficiency leads to mitochondrial dysfunction, including an abnormally expressed mt-nucleoid gene profile, decreased MMP and a high rate of respiration-deficient petite rate under H2O2 treatment, we wished to determine whether CR could extend the RLS of ypk9Δ cells.
Glycerol is a widely used dietary supplement capable of mimicking CR by shifting metabolism away from glycolysis and toward oxidative phosphorylation (Wei et al. 2009). Given this, we determined the RLS of ypk9Δ under the CR conditions (3% glycerol). We found that the RLS of ypk9Δ yeast cells was 19 generations under the CR conditions and was 26.7% higher than the RLS under standard conditions (2% glucose), while, the RLS of wild-type BY4742 cells (23 generations) was not obviously changed under the 3% glycerol conditions (Schleit et al. 2013) (Fig. 8). This result suggested that the shortened RLS of ypk9Δ cells with dysfunctional mitochondria could be rescued by CR. This observation was in accord with previous studies (Schleit et al. 2013) that growth on the nonfermentable carbon source, 3% glycerol, could extend the RLS of phbΔ (with mitochondrial dysfunction and decreased RLS under 2% glucose conditions).
CR increased the RLS of ypk9Δ. The mean RLS (under 3% glycerol conditions) and total number of daughter cells are shown in parentheses; the total number of daughter cells is denoted by “N”. The values were analyzed for statistical significance using the Wilcoxon rank-sum test. p < 0.05 was considered to indicate statistical significance
In conclusion, we have demonstrated, for the first time, that YPK9 deficiency leads to oxidative stress accompanied by abnormal mitochondrial function and an increased incidence of early apoptosis in budding yeast, which may have contributed to the observed shortened RLS of ypk9Δ yeast in this study. More importantly, we found that these senescence-related phenotypes can be eliminated by overexpression of the catalase-encoding gene CTA1. These findings highlight the involvement of Ypk9p in the oxidative stress response and modulation of RLS.
Data availability
The data used to support the findings of this study are available from the corresponding author upon request.
Abbreviations
- AbA:
-
Aureobasidin A
- ATP:
-
Adenosine triphosphate
- CHP:
-
Cube hydroperoxide
- CLS:
-
Chronological life span
- CR:
-
Caloric restriction
- DAPI:
-
4′,6-Diamidino-2-phenilindole
- DCFH-DA:
-
2′,7′-Dichlorofluorescein diacetate
- D. melanogaster :
-
Drosophila melanogaster
- ETC:
-
Electron transport chain
- FITC:
-
Fluorescein isothiocyanate
- H2O2 :
-
Hydrogen peroxide
- LiAc:
-
Lithium acetate
- MMP:
-
Mitochondrial membrane potential
- mtDNA:
-
Mitochondrial DNA
- mt-nucleoids:
-
Mitochondrial nucleoids
- OD:
-
Optical density
- PCR:
-
Polymerase chain reaction
- Rh123:
-
Rhodamine 123
- RLS:
-
Replicative lifespan
- ROS:
-
Reactive oxygen species
- RT-qPCR:
-
Real-time quantitative PCR
- S. cerevisiae :
-
Saccharomyces cerevisiae
- TBHP:
-
Tert-butyl hydroperoxide
- TTC:
-
2,3,5-Triphenyltetrazolium chloride
- WT:
-
Wild-type
References
Alugoju P, Periyasamy L, Dyavaiah M (2020) Protective effect of quercetin in combination with caloric restriction against oxidative stress-induced cell death of Saccharomyces cerevisiae cells. Lett Appl Microbiol 71(3):272–279. https://doi.org/10.1111/lam.13313
Amigo I, da Cunha FM, Forni MF, Garcia-Neto W, Kakimoto PA, Luévano-Martínez LA, Macedo F, Menezes-Filho SL, Peloggia J, Kowaltowski AJ (2016) Mitochondrial form, function and signalling in aging. Biochem J 473(20):3421–3449. https://doi.org/10.1042/bcj20160451
Baudin A, Ozier-Kalogeropoulos O, Denouel A, Lacroute F, Cullin C (1993) A simple and efficient method for direct gene deletion in Saccharomyces cerevisiae. Nucleic Acids Res 21(14):3329–3330. https://doi.org/10.1093/nar/21.14.3329
Bras J, Verloes A, Schneider SA, Mole SE, Guerreiro RJ (2012) Mutation of the parkinsonism gene ATP13A2 causes neuronal ceroid-lipofuscinosis. Hum Mol Genet 21(12):2646–2650. https://doi.org/10.1093/hmg/dds089
Chen XJ, Butow RA (2005) The organization and inheritance of the mitochondrial genome. Nat Rev Genet 6(11):815–825. https://doi.org/10.1038/nrg1708
Cho JH, Ha SJ, Kao LR, Megraw TL, Chae CB (1998) A novel DNA-binding protein bound to the mitochondrial inner membrane restores the null mutation of mitochondrial histone Abf2p in Saccharomyces cerevisiae. Mol Cell Biol 18(10):5712–5723. https://doi.org/10.1128/mcb.18.10.5712
Costanzo M, VanderSluis B, Koch EN, Baryshnikova A, Pons C, Tan G, Wang W, Usaj M, Hanchard J, Lee SD, Pelechano V, Styles EB, Billmann M, van Leeuwen J, van Dyk N, Lin ZY, Kuzmin E, Nelson J, Piotrowski JS, Srikumar T, Bahr S, Chen Y, Deshpande R, Kurat CF, Li SC, Li Z, Usaj MM, Okada H, Pascoe N, San Luis BJ, Sharifpoor S, Shuteriqi E, Simpkins SW, Snider J, Suresh HG, Tan Y, Zhu H, Malod-Dognin N, Janjic V, Przulj N, Troyanskaya OG, Stagljar I, Xia T, Ohya Y, Gingras AC, Raught B, Boutros M, Steinmetz LM, Moore CL, Rosebrock AP, Caudy AA, Myers CL, Andrews B, Boone C (2016) A global genetic interaction network maps a wiring diagram of cellular function. Science 353(6306):e1420. https://doi.org/10.1126/science.aaf1420
Dai S, Hu S, Wu C (2009) Apoptotic effect of sonodynamic therapy mediated by hematoporphyrin monomethyl ether on C6 glioma cells in vitro. Acta Neurochir 151(12):1655–1661. https://doi.org/10.1007/s00701-009-0456-5
Delaney JR, Murakami C, Chou A, Carr D, Schleit J, Sutphin GL, An EH, Castanza AS, Fletcher M, Goswami S, Higgins S, Holmberg M, Hui J, Jelic M, Jeong KS, Kim JR, Klum S, Liao E, Lin MS, Lo W, Miller H, Moller R, Peng ZJ, Pollard T, Pradeep P, Pruett D, Rai D, Ros V, Schuster A, Singh M, Spector BL, Wende HV, Wang AM, Wasko BM, Olsen B, Kaeberlein M (2013) Dietary restriction and mitochondrial function link replicative and chronological aging in Saccharomyces cerevisiae. Exp Gerontol 48(10):1006–1013. https://doi.org/10.1016/j.exger.2012.12.001
Diffley JF, Stillman B (1991) A close relative of the nuclear, chromosomal high-mobility group protein HMG1 in yeast mitochondria. Proc Natl Acad Sci USA 88(17):7864–7868. https://doi.org/10.1073/pnas.88.17.7864
Eruslanov E, Kusmartsev S (2010) Identification of ROS using oxidized DCFDA and flow-cytometry. Methods Mol Biol 594:57–72. https://doi.org/10.1007/978-1-60761-411-1_4
Estrada-Cuzcano A, Martin S, Chamova T, Synofzik M, Timmann D, Holemans T, Andreeva A, Reichbauer J, De Rycke R, Chang DI, van Veen S, Samuel J, Schöls L, Pöppel T, Mollerup Sørensen D, Asselbergh B, Klein C, Zuchner S, Jordanova A, Vangheluwe P, Tournev I, Schüle R (2017) Loss-of-function mutations in the ATP13A2/PARK9 gene cause complicated hereditary spastic paraplegia (SPG78). Brain 140(2):287–305. https://doi.org/10.1093/brain/aww307
Fehrmann S, Paoletti C, Goulev Y, Ungureanu A, Aguilaniu H, Charvin G (2013) Aging yeast cells undergo a sharp entry into senescence unrelated to the loss of mitochondrial membrane potential. Cell Rep 5(6):1589–1599. https://doi.org/10.1016/j.celrep.2013.11.013
Ferguson LR, von Borstel RC (1992) Induction of the cytoplasmic “petite” mutation by chemical and physical agents in Saccharomyces cerevisiae. Mutat Res 265(1):103–148. https://doi.org/10.1016/0027-5107(92)90042-z
Gitler AD, Chesi A, Geddie ML, Strathearn KE, Hamamichi S, Hill KJ, Caldwell KA, Caldwell GA, Cooper AA, Rochet JC, Lindquist S (2009) Alpha-synuclein is part of a diverse and highly conserved interaction network that includes PARK9 and manganese toxicity. Nat Genet 41(3):308–315. https://doi.org/10.1038/ng.300
Guaragnella N, Pereira C, Sousa MJ, Antonacci L, Passarella S, Côrte-Real M, Marra E, Giannattasio S (2006) YCA1 participates in the acetic acid induced yeast programmed cell death also in a manner unrelated to its caspase-like activity. FEBS Lett 580(30):6880–6884. https://doi.org/10.1016/j.febslet.2006.11.050
Heins-Marroquin U, Jung PP, Cordero-Maldonado ML, Crawford AD, Linster CL (2019) Phenotypic assays in yeast and zebrafish reveal drugs that rescue ATP13A2 deficiency. Brain Commun 1(1):fcz019. https://doi.org/10.1093/braincomms/fcz019
Herker E, Jungwirth H, Lehmann KA, Maldener C, Fröhlich KU, Wissing S, Büttner S, Fehr M, Sigrist S, Madeo F (2004) Chronological aging leads to apoptosis in yeast. J Cell Biol 164(4):501–507. https://doi.org/10.1083/jcb.200310014
Jamieson DJ (1998) Oxidative stress responses of the yeast Saccharomyces cerevisiae. Yeast 14(16):1511–1527. https://doi.org/10.1002/(sici)1097-0061(199812)14:16%3c1511::Aid-yea356%3e3.0.Co;2-s
Jamshad M, Darby RA (2012) Yeast transformation to generate high-yielding clones. Methods Mol Biol 866:57–63. https://doi.org/10.1007/978-1-61779-770-5_6
Janero DR, Hreniuk D, Sharif HM (1991) Hydrogen peroxide-induced oxidative stress to the mammalian heart-muscle cell (cardiomyocyte): lethal peroxidative membrane injury. J Cell Physiol 149(3):347–364. https://doi.org/10.1002/jcp.1041490302
Jasnos L, Sliwa P, Korona R (2005) Resolution and repeatability of phenotypic assays by automated growth curve analysis in yeast and bacteria. Anal Biochem 344(1):138–140. https://doi.org/10.1016/j.ab.2005.04.034
Jazwinski SM (1990) Aging and senescence of the budding yeast Saccharomyces cerevisiae. Mol Microbiol 4(3):337–343. https://doi.org/10.1111/j.1365-2958.1990.tb00601.x
Kaya A, Koc A, Lee BC, Fomenko DE, Rederstorff M, Krol A, Lescure A, Gladyshev VN (2010) Compartmentalization and regulation of mitochondrial function by methionine sulfoxide reductases in yeast. Biochemistry 49(39):8618–8625. https://doi.org/10.1021/bi100908v
Kett LR, Stiller B, Bernath MM, Tasset I, Blesa J, Jackson-Lewis V, Chan RB, Zhou B, Di Paolo G, Przedborski S, Cuervo AM, Dauer WT (2015) α-Synuclein-independent histopathological and motor deficits in mice lacking the endolysosomal Parkinsonism protein Atp13a2. J Neurosci 35(14):5724–5742. https://doi.org/10.1523/jneurosci.0632-14.2015
Kong SM, Chan BK, Park JS, Hill KJ, Aitken JB, Cottle L, Farghaian H, Cole AR, Lay PA, Sue CM, Cooper AA (2014) Parkinson’s disease-linked human PARK9/ATP13A2 maintains zinc homeostasis and promotes α-Synuclein externalization via exosomes. Hum Mol Genet 23(11):2816–2833. https://doi.org/10.1093/hmg/ddu099
Kühlbrandt W (2004) Biology, structure and mechanism of P-type ATPases. Nat Rev Mol Cell Biol 5(4):282–295. https://doi.org/10.1038/nrm1354
Kurihara Y, Kanki T, Aoki Y, Hirota Y, Saigusa T, Uchiumi T, Kang D (2012) Mitophagy plays an essential role in reducing mitochondrial production of reactive oxygen species and mutation of mitochondrial DNA by maintaining mitochondrial quantity and quality in yeast. J Biol Chem 287(5):3265–3272. https://doi.org/10.1074/jbc.M111.280156
Larsen AK, Malinska D, Koszela-Piotrowska I, Parhamifar L, Hunter AC, Moghimi SM (2012) Polyethylenimine-mediated impairment of mitochondrial membrane potential, respiration and membrane integrity: implications for nucleic acid delivery and gene therapy. Mitochondrion 12(1):162–168. https://doi.org/10.1016/j.mito.2011.08.013
Laskowski W (1954) Induction, par le chlorure de tétrazolium, de la mutation “petite colonie” chez la levure. Heredity 8(1):79–88. https://doi.org/10.1038/hdy.1954.6
Laun P, Heeren G, Rinnerthaler M, Rid R, Kössler S, Koller L, Breitenbach M (2008) Senescence and apoptosis in yeast mother cell-specific aging and in higher cells: a short review. Biochim Biophys Acta 1783(7):1328–1334. https://doi.org/10.1016/j.bbamcr.2008.02.008
Laun P, Büttner S, Rinnerthaler M, Burhans WC, Breitenbach M (2012) Yeast aging and apoptosis. Subcell Biochem 57:207–232. https://doi.org/10.1007/978-94-007-2561-4_10
Lenaz G, Genova ML (2012) Supramolecular organisation of the mitochondrial respiratory chain: a new challenge for the mechanism and control of oxidative phosphorylation. Adv Exp Med Biol 748:107–144. https://doi.org/10.1007/978-1-4614-3573-0_5
Li P, Wang K, Salustros N, Grønberg C, Gourdon P (2021) Structure and transport mechanism of P5B-ATPases. Nat Commun 12(1):3973. https://doi.org/10.1038/s41467-021-24148-y
Lin MT, Beal MF (2006) Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature 443(7113):787–795. https://doi.org/10.1038/nature05292
Lin SJ, Kaeberlein M, Andalis AA, Sturtz LA, Defossez PA, Culotta VC, Fink GR, Guarente L (2002) Calorie restriction extends Saccharomyces cerevisiae lifespan by increasing respiration. Nature 418(6895):344–348
Ludovico P, Sansonetty F, Côrte-Real M (2001) Assessment of mitochondrial membrane potential in yeast cell populations by flow cytometry. Microbiology 147(Pt 12):3335–3343. https://doi.org/10.1099/00221287-147-12-3335
MacLean M, Harris N, Piper PW (2001) Chronological lifespan of stationary phase yeast cells; a model for investigating the factors that might influence the ageing of postmitotic tissues in higher organisms. Yeast 18(6):499–509. https://doi.org/10.1002/yea.701
Massardo DR, Zweifel SG, Gunge N, Miyakawa I, Sando N, Del Giudice A, Wolf K, Del Giudice L (2000) Use of lycorine and DAPI staining in Saccharomyces cerevisiae to differentiate between rho0 and rho- cells in a cce1/delta cce1 nuclear background. Can J Microbiol 46(11):1058–1065. https://doi.org/10.1139/w00-096
Mesquita A, Weinberger M, Silva A, Sampaio-Marques B, Almeida B, Leão C, Costa V, Rodrigues F, Burhans WC, Ludovico P (2010) Caloric restriction or catalase inactivation extends yeast chronological lifespan by inducing H2O2 and superoxide dismutase activity. Proc Natl Acad Sci USA 107(34):15123–15128. https://doi.org/10.1073/pnas.1004432107
Moon SH, Lee SJ, Jung KH, Quach CH, Park JW, Lee JH, Cho YS, Lee KH (2016) Troglitazone stimulates cancer cell uptake of 18F-FDG by suppressing mitochondrial respiration and augments sensitivity to glucose restriction. J Nucl Med 57(1):129–135. https://doi.org/10.2967/jnumed.115.162016
Moskovitz J, Berlett BS, Poston JM, Stadtman ER (1997) The yeast peptide-methionine sulfoxide reductase functions as an antioxidant in vivo. Proc Natl Acad Sci USA 94(18):9585–9589. https://doi.org/10.1073/pnas.94.18.9585
Nicholson DW, Ali A, Thornberry NA, Vaillancourt JP, Ding CK, Gallant M, Gareau Y, Griffin PR, Labelle M, Lazebnik YA (1995) Identification and inhibition of the ICE/CED-3 protease necessary for mammalian apoptosis. Nature 376(6535):37
O’Rourke TW, Doudican NA, Mackereth MD, Doetsch PW, Shadel GS (2002) Mitochondrial dysfunction due to oxidative mitochondrial DNA damage is reduced through cooperative actions of diverse proteins. Mol Cell Biol 22(12):4086–4093. https://doi.org/10.1128/mcb.22.12.4086-4093.2002
Palmgren MG, Axelsen KB (1998) Evolution of P-type ATPases. Biochim Biophys Acta 1365(1–2):37–45. https://doi.org/10.1016/s0005-2728(98)00041-3
Palmgren MG, Nissen P (2011) P-type ATPases. Annu Rev Biophys 40:243–266. https://doi.org/10.1146/annurev.biophys.093008.131331
Popa CV, Dumitru I, Ruta LL, Danet AF, Farcasanu IC (2010) Exogenous oxidative stress induces Ca2+ release in the yeast Saccharomyces cerevisiae. Febs j 277(19):4027–4038. https://doi.org/10.1111/j.1742-4658.2010.07794.x
Postnikoff SD, Harkness TA (2014) Replicative and chronological life-span assays. Methods Mol Biol 1163:223–227. https://doi.org/10.1007/978-1-4939-0799-1_17
Qi G, Sun D, Tian Y, Xu C, Zhang Y, Wang D, Ma K, Xu S, Jin Y (2020) Fast activation and tracing of caspase-3 involved cell apoptosis by combined electrostimulation and smart signal-amplified SERS nanoprobes. Anal Chem 92(11):7861–7868. https://doi.org/10.1021/acs.analchem.0c01114
Ramirez A, Heimbach A, Gründemann J, Stiller B, Hampshire D, Cid LP, Goebel I, Mubaidin AF, Wriekat AL, Roeper J, Al-Din A, Hillmer AM, Karsak M, Liss B, Woods CG, Behrens MI, Kubisch C (2006) Hereditary parkinsonism with dementia is caused by mutations in ATP13A2, encoding a lysosomal type 5 P-type ATPase. Nat Genet 38(10):1184–1191. https://doi.org/10.1038/ng1884
Rinnerthaler M, Jarolim S, Heeren G, Palle E, Perju S, Klinger H, Bogengruber E, Madeo F, Braun RJ, Breitenbach-Koller L, Breitenbach M, Laun P (2006) MMI1 (YKL056c, TMA19), the yeast orthologue of the translationally controlled tumor protein (TCTP) has apoptotic functions and interacts with both microtubules and mitochondria. Biochim Biophys Acta 1757(5–6):631–638. https://doi.org/10.1016/j.bbabio.2006.05.022
Rockenfeller P, Madeo F (2008) Apoptotic death of ageing yeast. Exp Gerontol 43(10):876–881. https://doi.org/10.1016/j.exger.2008.08.044
Rona G, Herdeiro R, Mathias CJ, Torres FA, Pereira MD, Eleutherio E (2015) CTT1 overexpression increases life span of calorie-restricted Saccharomyces cerevisiae deficient in Sod1. Biogerontology 16(3):343–351. https://doi.org/10.1007/s10522-015-9550-7
Ruetenik A, Barrientos A (2015) Dietary restriction, mitochondrial function and aging: from yeast to humans. Biochim Biophys Acta 1847(11):1434–1447. https://doi.org/10.1016/j.bbabio.2015.05.005
Santos AX, Riezman H (2012) Yeast as a model system for studying lipid homeostasis and function. FEBS Lett 586(18):2858–2867. https://doi.org/10.1016/j.febslet.2012.07.033
Scheckhuber CQ, Erjavec N, Tinazli A, Hamann A, Nyström T, Osiewacz HD (2007) Reducing mitochondrial fission results in increased life span and fitness of two fungal ageing models. Nat Cell Biol 9(1):99–105. https://doi.org/10.1038/ncb1524
Schleit J, Johnson SC, Bennett CF, Simko M, Trongtham N, Castanza A, Hsieh EJ, Moller RM, Wasko BM, Delaney JR, Sutphin GL, Carr D, Murakami CJ, Tocchi A, Xian B, Chen W, Yu T, Goswami S, Higgins S, Holmberg M, Jeong KS, Kim JR, Klum S, Liao E, Lin MS, Lo W, Miller H, Olsen B, Peng ZJ, Pollard T, Pradeep P, Pruett D, Rai D, Ros V, Singh M, Spector BL, Vander Wende H, An EH, Fletcher M, Jelic M, Rabinovitch PS, MacCoss MJ, Han JD, Kennedy BK, Kaeberlein M (2013) Molecular mechanisms underlying genotype-dependent responses to dietary restriction. Aging Cell 12(6):1050–1061. https://doi.org/10.1111/acel.12130
Schmidt K, Wolfe DM, Stiller B, Pearce DA (2009) Cd2+, Mn2+, Ni2+ and Se2+ toxicity to Saccharomyces cerevisiae lacking YPK9p the orthologue of human ATP13A2. Biochem Biophys Res Commun 383(2):198–202. https://doi.org/10.1016/j.bbrc.2009.03.151
Schmitt E, Paquet C, Beauchemin M, Bertrand R (2007) DNA-damage response network at the crossroads of cell-cycle checkpoints, cellular senescence and apoptosis. J Zhejiang Univ Sci B 8(6):377–397. https://doi.org/10.1631/jzus.2007.B0377
Sørensen DM, Holemans T, van Veen S, Martin S, Arslan T, Haagendahl IW, Holen HW, Hamouda NN, Eggermont J, Palmgren M, Vangheluwe P (2018) Parkinson disease related ATP13A2 evolved early in animal evolution. PLoS ONE 13(3):e0193228. https://doi.org/10.1371/journal.pone.0193228
Spataro R, Kousi M, Farhan SMK, Willer JR, Ross JP, Dion PA, Rouleau GA, Daly MJ, Neale BM, La Bella V, Katsanis N (2019) Mutations in ATP13A2 (PARK9) are associated with an amyotrophic lateral sclerosis-like phenotype, implicating this locus in further phenotypic expansion. Hum Genomics 13(1):19. https://doi.org/10.1186/s40246-019-0203-9
Tadini-Buoninsegni F (2020) Protein adsorption on solid supported membranes: monitoring the transport activity of P-Type ATPases. Molecules 25(18):4167. https://doi.org/10.3390/molecules25184167
Tzagoloff A, Akai A, Foury F (1976) Assembly of the mitochondrial membrane system XVI. Modified form of the ATPase proteolipid in oligomycin-resistant mutants of Saccharomyces cerevisiae. FEBS Lett 65(3):391–395
Uren AG, O’Rourke K, Aravind LA, Pisabarro MT, Seshagiri S, Koonin EV, Dixit VM (2000) Identification of paracaspases and metacaspases: two ancient families of caspase-like proteins, one of which plays a key role in MALT lymphoma. Mol Cell 6(4):961–967. https://doi.org/10.1016/s1097-2765(00)00094-0
Van Zandycke SM, Sohier PJ, Smart KA (2002) The impact of catalase expression on the replicative lifespan of Saccharomyces cerevisiae. Mech Ageing Dev 123(4):365–373. https://doi.org/10.1016/s0047-6374(01)00382-7
Wei M, Fabrizio P, Madia F, Hu J, Ge H, Li LM, Longo VD (2009) Tor1/Sch9-regulated carbon source substitution is as effective as calorie restriction in life span extension. PLoS Genet 5(5):e1000467. https://doi.org/10.1371/journal.pgen.1000467
Xie Z, Kometiani P, Liu J, Li J, Shapiro JI, Askari A (1999) Intracellular reactive oxygen species mediate the linkage of Na+/K+-ATPase to hypertrophy and its marker genes in cardiac myocytes. J Biol Chem 274(27):19323–19328. https://doi.org/10.1074/jbc.274.27.19323
Yan X, Xun M, Li J, Wu L, Dou X, Zheng J (2016) Activation of Na+/K+-ATPase attenuates high glucose-induced H9c2 cell apoptosis via suppressing ROS accumulation and MAPKs activities by DRm217. Acta Biochim Biophys Sin 48(10):883–893. https://doi.org/10.1093/abbs/gmw079
Zhang Y, Herman B (2002) Ageing and apoptosis. Mech Ageing Dev 123(4):245–260. https://doi.org/10.1016/s0047-6374(01)00349-9
Zhao W, Zhou T, Zheng HZ, Qiu KP, Cui HJ, Yu H, Liu XG (2018) Yeast polyubiquitin gene UBI4 deficiency leads to early induction of apoptosis and shortened replicative lifespan. Cell Stress Chaperones 23(4):527–537. https://doi.org/10.1007/s12192-017-0860-3
Acknowledgements
This study was supported by the China National Natural Science Foundation (Grant Nos. 31701050, 81671399, 81971329), the Natural Science Foundation of Guangdong Province (Grant No. 2018A0303070020), the Doctoral Startup Project of Guangdong medical university (Grant No. B2019039) and the Guangdong Medical Research Foundation (Grant No. 2020489). The authors are grateful to Matt Kaeberlein and Brian K. Kennedy for technical assistance.
Author information
Authors and Affiliations
Contributions
Xinguang Liu and Jie Ruan designed the experiments. Fang Guo, Lingyue Kong, Jiaxin Liu, Xiaoshan Hong, Zhiwen Jiang, Haochang Song and Xiaojing Cui performed experiments. Wei Zhao wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhao, W., Guo, F., Kong, L. et al. Yeast YPK9 deficiency results in shortened replicative lifespan and sensitivity to hydrogen peroxide. Biogerontology 22, 547–563 (2021). https://doi.org/10.1007/s10522-021-09935-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10522-021-09935-w