Abstract
The amyloid precursor like protein-1 (APLP1) belongs to the amyloid precursor protein family that also includes the amyloid precursor protein (APP) and the amyloid precursor like protein-2 (APLP2). Though the three proteins share similar structures and undergo the same cleavage processing by α-, β- and γ-secretases, APLP1 shows divergent subcellular localization from that of APP and APLP2, and thus, may perform distinct roles in vivo. While extensive studies have been focused on APP, which is implicated in the pathogenesis of Alzheimer’s disease, the functions of APLP1 remain largely elusive. Here we report that the expression of APLP1 in Drosophila induces cell death and produces developmental defects in wing and thorax. This function of APLP1 depends on the transcription factor dFoxO, as the depletion of dFoxO abrogates APLP1-induced cell death and adult defects. Consistently, APLP1 up-regulates the transcription of dFoxO target hid and reaper-two well known pro-apoptotic genes. Thus, the present study provides the first in vivo evidence that APLP1 is able to induce cell death, and that FoxO is a crucial downstream mediator of APLP1’s activity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Human amyloid precursor like protein-1 (APLP1) belongs to a protein family which also contains amyloid precursor protein (APP) and amyloid precursor like protein-2 (APLP2) [1–5].The three proteins share similar structures with conserved N- and C-terminal domains, and can be cleaved by the same α-, β- and γ-secretases [6–10], implying that they might perform similar functions in development. Previous studies revealed that single knock-out mice of APP, APLP1 or APLP2 produced only subtle phenotypes, while double knock-out APP/APLP2 or APLP1/APLP2 mice died shortly after birth, implying a possible functional redundancy among the proteins [11–13].However, other studies suggest that these proteins may bear diverse functions rather than simply compensating for each other [14–16]. In particular, APLP1 exhibits distinct expression pattern and subcellular localization from that of APP and APLP2. While APP and APLP2 are ubiquitously expressed and predominantly distributed in intracellular compartments including the endosomes, ER and Golgi apparatus [5], APLP1 specifically concentrates in the nervous system but also shows a weak signal in other tissues such as heart, lung, liver and kidney of E15 mouse embryos observed by in situ hybridization [17]. However, the functions of APLP1 in the non-neuronal tissues remain unknown. APLP1 mainly localizes to the plasma membrane [5], indicating APLP1 may perform divergent in vivo functions from that of APP and APLP2. Though all APP family members are able to generate the intracellular domains (ICDs) through cleavage by γ- and ε-secretases [18–22], ICDs derived from APP and APLP2, but not APLP1, interact with Tip60 and Fe65 to localize to the spherical nuclear AFT complexes, where they regulate target genes expression [23, 24].
APLP1 is shown to be transcriptionally regulated by p53 tumor suppressor [25], and forms oligomers on the plasma membrane via its E2 domain [26]. APLP1 is found in the senile plaques of Alzheimer’s disease (AD) patients’ brains [27, 28], suggesting a possible role of APLP1 in the progression of the disease. Consistent with this, APLP1 undergoes the processing by β- and γ-secretases to form short Aβ-like peptides [29], which is also reported to be altered in Down’s Syndrome [30]. Besides, APLP1 is shown to regulate synapse formation, neuronal differentiation and synaptic plasticity [1, 4, 31–34]. However, compared with APP, the in vivo functions of APLP1 have remained largely unknown.
Drosophila has been used as a powerful model organism to investigate the in vivo functions and underlining mechanisms of human genes. Here in this report, we introduced APLP1 into Drosophila and found that expression of APLP1 induced dramatical cell death and developmental defects in various tissues. In addition, APLP1-induced cell death is mediated by the transcription factor dFoxO. Consistently, APLP1 up-regulates the expression of dFoxO target genes hid and reaper. To our knowledge, this study provides the first in vivo evidence that APLP1 triggers cell death in development.
Materials and methods
Drosophila strains
UAS-APLP1, UAS-APLP2 and UAS-APPLsd [35] were kind gifts from Dr. Merders. dfoxO △94 and dfoxO 21 [36] were kind gifts from Dr. Partridge. ptc-Gal4, en-Gal4, pnr-Gal4, sd-Gal4, UAS-GFP, UAS-dfoxO-IR#1, UAS-dfoxO-IR#2, hid-LacZ [37] were previously described. UAS-LacZ, dpp-Gal4, reaper-LacZ, APPL-Gal4, UAS-p35 and UAS-DIAP1 were obtained from the Bloomington Stock center.
AO staining
Wing discs were dissected from the 3rd instar larvae in 1 % PBS buffer and stained for acridine orange as described [38]. Each genotype was dissected with 20 discs for statistics.
Light image
Freshly eclosed flies of indicated genotypes were collected and immediately frozen in −80 °C. Wings were dissected and mounted on the slide in the alcohol/glycerol (1:1) medium, and flies were mounted on the 1 % agarose plate in the alcohol/glycerol medium. Light images of wings were collected with Olympus microscope BX51, and light images of thoraxes were collected with OLYMPUS stereo microscope SZX16.
Immunohistochemistry
3rd instar larvae of indicated genotypes were collected and dissected in 1 % PBS buffer. And the antibody staining of imaginal discs was conducted as previously described [39]. The following antibodies were used: rabbit anti-cleaved caspase-3 (1:400, Cell Signaling and Technology), anti-rabbit-Alexa (1:1,000, Cell Signaling and Technology).
X-gal staining
Wing discs were dissected from the 3rd instar larvae in 1 % PBS buffer and stained for β-galactosidase activity as described [40].
Statistical analysis
For the statistics of anterior cross vein (acv), wings from freshly eclosed virgins were dissected and the presence of acv was counted. 20 wings were examined for each genotype. For the statistics of scutellum size, freshly eclosed virgins were collected and photoed, the area of the scutellum was measured with the software Cellsens of Olympus. Each genotype was tested with 10 flies.
Results
APLP1 induces caspase-dependent cell death in Drosophila
To characterize the in vivo functions of APLP1 in development, we expressed APLP1 in multiple tissues of Drosophila. Targeted expression of APLP1 along the anterior/posterior (A/P) compartment boundary in 3rd instar wing discs driven by the patched-Gal4 (ptc-Gal4) driver (Fig. 1a, b) [37, 41] initiated extensive cell death in the ptc domain, as revealed by acridine orange (AO) staining (Fig. 1d, g), compared with the ptc-Gal4 control (Fig. 1c, g), indicating the expression of APLP1 is sufficient to induce cell death in Drosophila. To examine whether APLP1-induced cell death is caspase dependent, we performed immunostaining against the cleaved caspase-3. Expression of APLP1 initiated strong cleaved caspase-3 staining along the A/P compartment boundary (Fig. 1i), compared with the ptc-Gal4 control (Fig. 1h), suggesting APLP1 induces caspase activation in Drosophila. Consistent with this finding, expression of p35, a viral protein that inhibits effector caspases [42], or the Drosophila inhibitor of apoptosis protein1 (DIAP1) [43, 44], suppressed APLP1-induced cell death and cleaved caspase-3 activation (Fig. 1e–g, j, k).Together, these data suggest that APLP1 induces caspase-dependent cell death in Drosophila.
APLP1 induces caspase-dependent cell death in Drosophila. Fluorescent images of GFP expression (a, b), or acridine orange staining (c–f), or anti-cleaved caspase-3 staining (h–k) of wing discs from 3rd instar larvae are shown. ptc-Gal4 was used as a control (c, h), or to drive the expression of GFP (a, b) or APLP1 (d–f, i–k). APLP1-induced cell death (d) and cleaved caspase-3 activation (i) are suppressed by the expression of p35 (e, j) or DIAP1 (f, k). g is the statistical analysis of acridine orange-positive cells in figures c–f. ***P ≤ 0.001. Genotypes: ptc-Gal4 UAS-GFP/+ (a, b); ptc-Gal4/+ (c, h); ptc-Gal4/+; UAS-APLP1/+ (d, i); ptc-Gal4/+; UAS-APLP1/UAS-p35 (e, j); ptc-Gal4/+; UAS-APLP1/UAS-DIAP1(f, k)
To further examine the role of APLP1 in regulating cell death in wing development, we used engrailed-Gal4 (en-Gal4) to drive APLP1’s expression in the posterior compartment of wing discs (Fig. 2a), and observed evident cell death as compared with the en-Gal4 control (Fig. 2b–d). Interestingly, a loss of acv phenotype in the posterior compartment was noticed in wings of APLP1-expressing flies, but not those of en-Gal4 controls or the ones that express GFP (Fig. 2e–h), indicating APLP1 affects vein formation in Drosophila. Similarly, ptc-Gal4 driven expression of APLP1 resulted in strong cell death (Fig. 1d, g and Fig. 3c, h) and the complete loss of the acv in adult wings (Fig. 4c, f), while the ptc-Gal4 control (Fig. 4a, f) or expression of GFP (Fig. 4b, f) failed to generate such phenotype, further suggesting APLP1 affects vein development. Furthermore, expression of APLP1 driven by dpp-Gal4 along the A/P boundary (Fig. S1a) triggered strong cell death in wing discs (Fig. S1c, d), and generated the loss-of-acv phenotype in adult wings (Fig. S1g, h), compared with the controls (Fig. S1b, d, e, f, h).Finally, expression of APLP1 driven by sd-Gal4 (Fig. S2a) provoked extensive cell death in the wing pouch (Fig. S2c) and produced a blistered wing phenotype (Fig. S3b, d), compared with the controls (Fig. S2b, S3a).
APLP1 induces cell death and morphological defects in wing development. Fluorescent images of GFP expression (a) or acridine orange staining (b, c) of wing discs from 3rd instar larvae and light images of adult wing (e–g) are shown. en-Gal4 was used as a control (b, f), or to drive the expression of GFP (a, e) or APLP1 (c, g). The lower panels are high magnification of the boxed areas in the upper panels. d shows the statistical analysis of acridine orange-positive cells in the boxed areas in b and c, while h shows the statistical analysis of the acv presence in f and g. ***P ≤ 0.001. Genotypes: en-Gal4 UAS-GFP/+ (a, e); en-Gal4/+ (b, f); en-Gal4/+; UAS-APLP1/+ (c, g)
APLP1 induces dFoxO-dependent cell death in wing discs. Fluorescent images of GFP expression (a) or acridine orange staining (b–g) of wing discs from 3rd instar larvae are shown. ptc-Gal4 was used as a control (b), or to drive the expression of GFP (a) or APLP1 (c–g).APLP1-induced cell death is suppressed by two dfoxO mutations (d, e) or expression of a dfoxO RNAi (f), but remains unaffected by the expression of LacZ (g). The lower panels are high magnification of boxed areas in the upper panels. h is the statistical analysis of acridine orange-positive cells in the lower panels of b–g. ***P ≤ 0.001; **, P < 0.01; n.s., not significant. Genotypes: ptc-Gal4 UAS-GFP/+ (a); ptc-Gal4/+ (b); ptc-Gal4/+; UAS-APLP1/+ (c); ptc-Gal4/+; UAS-APLP1/dfoxO 21 (d); ptc-Gal4/+; UAS-APLP1/dfoxO △94 (e); ptc-Gal4/+; UAS-APLP1/UAS-dfoxO-IR#1 (f); ptc-Gal4/+; UAS-APLP1/UAS-LacZ (g)
APLP1 induces dFoxO-dependent loss-of-acv phenotype in adult wings. Light images of adult wings are shown (a–e). ptc-Gal4 was used as a control (a), or to drive the expression of GFP (b) or APLP1 (c–e). Expression of a dfoxO RNAi (d), but not LacZ (e), partially suppressed APLP1-induced loss-of-acv phenotype. The lower panels are high magnification of boxed areas (showing acv) in the upper panels. f is the statistical analysis of the acv presence in a–e. ***P ≤ 0.001. Genotypes ptc-Gal4/+ (a); ptc-Gal4 UAS-GFP/+ (b); ptc-Gal4/+; UAS-APLP1/+ (c); ptc-Gal4/+; UAS-APLP1/UAS-dfoxO-IR#1 (d); ptc-Gal4/+; UAS-APLP1/UAS-LacZ (e)
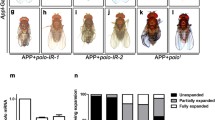
To check if expression of APLP1 could induce cell death in other tissues, we expressed APLP1 in the thorax by pnr-Gal4, and observed a reduced scutellum phenotype (Fig. 5c, i) resulted from enhanced cell death in the notum tips of wing discs (Fig. S4b, d). As negative controls, neither pnr-Gal4 nor expression of GFP was able to induce cell death in wing discs and scutellum defect in adult flies (Fig. 5a, b, i; Fig. S4a, d). In addition, expression of APLP1 under the control of the pan-neuron elav-Gal4 driver (Fig. S5a) [37] provoked neuronal cell death in the ventral nerve cord (Fig. S5b–d), while expression of APLP1 in the eyediscs driven by GMR-Gal4 (Fig. S5e) induced strong cell death posterior to the morphogenetic furrow (Fig. S5f–h). Taken together, these results indicate that APLP1 induces cell death in a non-tissue specific manner in Drosophila.
Loss of dfoxO suppresses APLP1-induced small scutellum phenotype. Light images of Drosophila adult thoraxes are shown. Compared with the pnr-Gal4 control (a), expression of APLP1 (c) but not GFP (b) induced a small scutellum phenotype, which was suppressed by two dfoxO mutations (e, f) or expression of two independent dfoxO RNAi (g, h), but not that of LacZ (d). i is the statistical analysis of the size of scutellum in a–h. ***P ≤ 0.001; n.s. not significant. Genotypes: pnr-Gal4/+ (a); pnr-Gal4 UAS-GFP/+ (b); pnr-Gal4 UAS-APLP1/+ (c); pnr-Gal4 UAS-APLP1/UAS-LacZ (d); pnr-Gal4 UAS-APLP1/dfoxO 21 (e); pnr-Gal4 UAS-APLP1/dfoxO △94 (f); pnr-Gal4 UAS-APLP1/UAS-dfoxO-IR#1 (g); pnr-Gal4 UAS-APLP1/UAS-dfoxO-IR#2 (h)
Together, the above data indicate that APLP1 induces cell death and developmental defects in Drosophila. Consistent with our observation, the depletion of APLP1 in cultured neuroblastoma cells reduced stress-induced apoptosis while overexpression of APLP1 slightly enhanced stress-induced apoptosis [25]. However, expression of APLP1 alone failed to trigger apoptosis in these cells [25], suggesting APLP1 induces cell death in a cell context dependent manner. Therefore, our data not only demonstrate for the first time that APLP1 by itself is sufficient to induce cell death, but also provide the first in vivo evidence for this function of APLP1.
APLP1 belongs to a protein family which also contains amyloid precursor like protein-2 (APLP2) [1–5]. To check if APLP2 can induce cell death and produce similar phenotypes as that of APLP1, we also expressed APLP2 under the control of various Gal4 drivers in Drosophila. Interestingly, ptc>APLP2 promoted cell death along the A/P boundary in the wing disc (Fig. S6a–c) and loss of acv in the adult wing (Fig. S6d-f), sd>APLP2 triggered strong cell death in the wing pouch (Fig. S7a, b) that resulted in small blistered wings (Fig. S7c, d), and pnr>APLP2 produced dramatically diminished scutella (Fig. S7e, f). Thus, APLP2 is able to recapitulate the cell death phenotype of APLP1 in Drosophila.
In Drosophila, there is only one APP family member named amyloid precursor protein-like (APPL) protein [45], which plays a vital role in regulating the development of the Drosophila nervous system [46]. APPL regulates the synaptic growth [47], axon transport [48, 49] and is also a modulator of wnt PCP signaling [50]. Expression of APPL in the thorax resulted in loss of thoracic macrochaetes and a reduction in scutellum size [51], reminiscent of the thorax phenotypes of pnr>APLP1 in this study (Fig. 5c). Furthermore, expression of APPL in the wing disc induced caspase dependent cell death and produced a loss-of-acv phenotype [51], which is similar to that of ptc>APLP1 (Fig. 4c). To examine whether APPL could trigger cell death in the nervous system, we expressed APPL in the ventral nerve cord (Fig. S8a), and observed mild cell death compared with the control (Fig. S8b–d), suggesting APPL could induce neuronal cell death in Drosophila.
Previous work showed that expression of human APP or Drosophila APPL induced cell death in Drosophila [37]. This study indicated a similar function for human APLP1 and APLP2. Thus, the roles in regulating cell death have been conserved by all APP family members.
Loss of dFoxO suppresses APLP1-induced cell death in Drosophila
To investigate the mechanism underlying APLP1-induced cell death, we called specific attention to the transcription factor dFoxO encoding the Drosophila homolog of mammalian FoxO3a, based on the following reasons. Firstly, both APLP1 and FoxO3a play pivotal roles in regulating stress-induced cell death [25, 52]. Secondly, both APLP1 and FoxO3a are required for p53-dependent apoptosis [25, 53]. Thirdly, both APLP1 and dFoxO are sufficient to induce cell death in Drosophila [54]. Finally, we previously noted that expression of dFoxO or human FoxO3a driven by ptc-Gal4 recapitulated the loss-of-acv wing phenotype of ptc>APLP1 (Fig. S9) [37].
To check whether dFoxO is required for APLP1-induced cell death, we expressed APLP1 in Drosophila with depleted dfoxO expression. In support of the hypothesis, we found that ptc>APLP1 induced cell death in wing discs was significantly suppressed by mutations in dfoxO(Fig. 3d, e, h) or RNAi-mediated knocking down of dfoxO (Fig. 3f, h), but not by the expression of LacZ (Fig. 3g, h). Consistently, ptc>APLP1 induced loss-of-acv phenotype in adult wings was also suppressed by loss of dfoxO, but remained unaffected by the expression of LacZ (Fig. 4c–f). Furthermore, sd>APLP1 triggered blistered wing phenotype was also suppressed in dfoxO mutants (Fig. S3c, d).
Consistent with the results obtained in the wing, we found that pnr>APLP1 induced cell death in the notum area of wing discs and the reduced scutellum phenotype in adult were considerably suppressed by mutations or RNAi-mediated down-regulation of dfoxO (Fig. 5e–i, Fig. S4c, d), but remained unchanged by the expression of LacZ (Fig. 5d, i).
Taken together, these data suggest that dFoxO is required for APLP1-induced cell death and developmental defects in the Drosophila wing and thorax.
APLP1 up-regulates dFoxO target gene expression
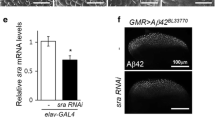
The above results highlight a role of dFoxO in mediating APLP1-induced cell death in Drosophila, which suggests that APLP1 may activated FoxO to initiate the downstream cell death machinery. To examine whether APLP1 activates dFoxO in vivo, we checked the expression of the pro-apoptotic genes hid and reaper (rpr), the well documented transcriptional targets of dFoxO [37, 54, 55]. Indeed, the expression of APLP1 in the wing pouch driven by sd-Gal4 resulted in dramatically up-regulated transcription of hid and rpr, as monitored by a hid-lacZ and a rpr-LacZ reporters, respectively (Fig. 6). Hence, the data demonstrate that APLP1 is able to activate dFoxO invivo, which provides a mechanism for APLP1-induced cell death in Drosophila. Yet, it remains unknown how APLP1 activates dFoxO in Drosophila, and whether APLP1 could trigger FoxO3a activation in mammalian cells. Further investigation will be required to address these important questions. Consistent with the observation that APLP2 phenocopied APLP1 in regulating cell death, expression of APLP2 can also activate dFoxO target genes hid and rpr transcription (Fig. S10). Collectively, these data indicate amyloid precursor like proteins are sufficient to activate FoxO mediated cell death machinery.
Discussion
Amyloid pecursor like protein-1(APLP1) is amammalian paralog of amyloid precursor protein (APP). While APP has been extensively studied for its involvement in the Alzheimer’s disease, few studies have been directed to APLP1 and its in vivo functions remain largely unknown. In the present study, we investigated the in vivo functions of APLP1 using Drosophila as a model organism. We found that ectopic expression of APLP1 induced cell death and developmental defects in the nervous and non-nervous system. Our genetic study characterized the transcription factor dFoxO as a critical downstream factor that mediates APLP1’s activity, for the depletion of dFoxO significantly suppressed APLP1-induced cell death in larval discs and associated phenotypes in adults. Further study confirmed that APLP1 was able to up-regulate the transcription of dFoxO target genes hid and reaper.
APLP1 was reported to function mainly in the nervous system, as high expression level of APLP1 was detected in the developing central and peripheral nervous systems, yeta weak expression signal of APLP1 was also observed in organs like heart, lung, liver and kidney in mouse embryos [17], implying a role of APLP1 in the development of non-neuronal tissues. Consistent with this explanation, RNAi mediated knockdown of APLP1 in WI-38 and MCF7 cells dramatically reduced the proliferation of these cells [25]. In the present study, we showed that expression of APLP1 could induce cell death and developmental defects in both neuronal and non-neuronal systems in Drosophila, and thus, providing further evidence for the function of APLP1 in non-neuronal cells.
Previous studies showed that loss of APLP1 diminishes stress induced apoptosis in neuroblastoma cells, whereas ectopic expression of APLP1 moderately enhances cell death upon stress stimulation [25]. However, expression of APLP1 alone is not sufficient to induce neuroblastoma cell death [25], suggesting APLP1 induces cell death in a context dependent manner. Our data not only demonstrate for the first time that APLP1 by itself is sufficient to induce cell death, but also provide the first in vivo evidence for this function of APLP1. APLP1 was reported to be a direct transcriptional target of the p53 tumor suppressor [25], which suggests a possible involvement of APLP1 in p53-induced cell death. p53 was known to interact with the transcriptional factor FoxO [56–58], and MDM2 was reported to act downstream of p53 to promote FoxO ubiquitination and degradation [58]. In the present study, we showed that FoxO mediates APLP1-induced cell death. The exact relationship between APLP1, FoxO and p53 in cell death will require further investigation. Overall, this study highlights a novel function of APLP1 in promoting FoxO-mediated cell death in vivo, which will shed light on the role of APLP1 in mammalian cells.
References
Beckman M, Iverfeldt K (1997) Increased gene expression of beta-amyloid precursor protein and its homologues APLP1 and APLP2 in human neuroblastoma cells in response to retinoic acid. Neurosci Lett 221(2–3):73–76
Kuan YH et al (2006) PAT1a modulates intracellular transport and processing of amyloid precursor protein (APP), APLP1, and APLP2. J Biol Chem 281(52):40114–40123
Lenkkeri U et al (1998) Structure of the human amyloid-precursor-like protein gene APLP1 at 19q13.1. Hum Genet 102(2):192–196
Adlerz L et al (2003) Accumulation of the amyloid precursor-like protein APLP2 and reduction of APLP1 in retinoic acid-differentiated human neuroblastoma cells upon curcumin-induced neurite retraction. Brain Res Mol Brain Res 119(1):62–72
Kaden D et al (2009) Subcellular localization and dimerization of APLP1 are strikingly different from APP and APLP2. J Cell Sci 122(Pt 3):368–377
Li Q, Sudhof TC (2004) Cleavage of amyloid-beta precursor protein and amyloid-beta precursor-like protein by BACE 1. J Biol Chem 279(11):10542–10550
Kume H, Maruyama K, Kametani F (2004) Intracellular domain generation of amyloid precursor protein by epsilon-cleavage depends on C-terminal fragment by alpha-secretase cleavage. Int J Mol Med 13(1):121–125
Pastorino L et al (2004) BACE (beta-secretase) modulates the processing of APLP2 in vivo. Mol Cell Neurosci 25(4):642–649
Eggert S et al (2004) The proteolytic processing of the amyloid precursor protein gene family members APLP-1 and APLP-2 involves alpha-, beta-, gamma-, and epsilon-like cleavages: modulation of APLP-1 processing by n-glycosylation. J Biol Chem 279(18):18146–18156
Walsh DM et al (2003) gamma-Secretase cleavage and binding to FE65 regulate the nuclear translocation of the intracellular C-terminal domain (ICD) of the APP family of proteins. Biochemistry 42(22):6664–6673
von Koch CS et al (1997) Generation of APLP2 KO mice and early postnatal lethality in APLP2/APP double KO mice. Neurobiol Aging 18(6):661–669
Heber S et al (2000) Mice with combined gene knock-outs reveal essential and partially redundant functions of amyloid precursor protein family members. J Neurosci 20(21):7951–7963
Beglopoulos V et al (2004) Reduced beta-amyloid production and increased inflammatory responses in presenilin conditional knock-out mice. J Biol Chem 279(45):46907–46914
Shariati SA, De Strooper B (2013) Redundancy and divergence in the amyloid precursor protein family. FEBS Lett 587(13):2036–2045
Scheinfeld MH, Matsuda S, D’Adamio L (2003) JNK-interacting protein-1 promotes transcription of A beta protein precursor but not A beta precursor-like proteins, mechanistically different than Fe65. Proc Natl Acad Sci USA 100(4):1729–1734
Gersbacher MT et al (2013) Turnover of amyloid precursor protein family members determines their nuclear signaling capability. PLoS One 8(7):e69363
Lorent K et al (1995) Expression in mouse embryos and in adult mouse brain of three members of the amyloid precursor protein family, of the alpha-2-macroglobulin receptor/low density lipoprotein receptor-related protein and of its ligands apolipoprotein E, lipoprotein lipase, alpha-2-macroglobulin and the 40,000 molecular weight receptor-associated protein. Neuroscience 65(4):1009–1025
Dimitrov M et al (2013) Alzheimer’s disease mutations in APP but not gamma-secretase modulators affect epsilon-cleavage-dependent AICD production. Nat Commun 4:2246
Schettini G et al (2010) Phosphorylation of APP-CTF-AICD domains and interaction with adaptor proteins: signal transduction and/or transcriptional role–relevance for Alzheimer pathology. J Neurochem 115(6):1299–1308
Zhang C et al (2007) An AICD-based functional screen to identify APP metabolism regulators. Mol Neurodegener 2:15
Nunan J, Small DH (2002) Proteolytic processing of the amyloid-beta protein precursor of Alzheimer’s disease. Essays Biochem 38:37–49
Scheinfeld MH et al (2002) Processing of beta-amyloid precursor-like protein-1 and -2 by gamma-secretase regulates transcription. J Biol Chem 277(46):44195–44201
Muller T et al (2013) A ternary complex consisting of AICD, FE65, and TIP60 down-regulates Stathmin1. Biochim Biophys Acta 1834(1):387–394
Cao X, Sudhof TC (2001) A transcriptionally [correction of transcriptively] active complex of APP with Fe65 and histone acetyltransferase Tip60. Science 293(5527):115–120
Tang X et al (2007) Amyloid-beta precursor-like protein APLP1 is a novel p53 transcriptional target gene that augments neuroblastoma cell death upon genotoxic stress. Oncogene 26(52):7302–7312
Mayer MC et al (2014) Novel zinc-binding site in the E2 domain regulates amyloid precursor-like protein 1 (APLP1) oligomerization. J Biol Chem 289(27):19019–19030
Bayer TA et al (1997) Amyloid precursor-like protein 1 accumulates in neuritic plaques in Alzheimer’s disease. Acta Neuropathol 94(6):519–524
McNamara MJ et al (1998) Immunohistochemical and in situ analysis of amyloid precursor-like protein-1 and amyloid precursor-like protein-2 expression in Alzheimer disease and aged control brains. Brain Res 804(1):45–51
Yanagida K et al (2009) The 28-amino acid form of an APLP1-derived Abeta-like peptide is a surrogate marker for Abeta42 production in the central nervous system. EMBO Mol Med 1(4):223–235
Portelius E et al (2014) Altered cerebrospinal fluid levels of amyloid beta and amyloid precursor-like protein 1 peptides in Down’s syndrome. Neuromol Med 16(2):510–516
Zheng H et al (1995) beta-Amyloid precursor protein-deficient mice show reactive gliosis and decreased locomotor activity. Cell 81(4):525–531
Guilarte TR et al (2008) Increased APLP1 expression and neurodegeneration in the frontal cortex of manganese-exposed non-human primates. J Neurochem 105(5):1948–1959
Bergmans BA et al (2010) Neurons generated from APP/APLP1/APLP2 triple knockout embryonic stem cells behave normally in vitro and in vivo: lack of evidence for a cell autonomous role of the amyloid precursor protein in neuronal differentiation. Stem Cells 28(3):399–406
Guilarte TR (2010) APLP1, Alzheimer’s-like pathology and neurodegeneration in the frontal cortex of manganese-exposed non-human primates. Neurotoxicology 31(5):572–574
Merdes G et al (2004) Interference of human and Drosophila APP and APP-like proteins with PNS development in Drosophila. EMBO J 23(20):4082–4095
Slack C et al (2011) dFOXO-independent effects of reduced insulin-like signaling in Drosophila. Aging Cell 10(5):735–748
Wang X et al (2014) FoxO mediates APP-induced AICD-dependent cell death. Cell Death Dis 5:e1233
Ma X et al (2013) dUev1a modulates TNF-JNK mediated tumor progression and cell death in Drosophila. Dev Biol 380(2):211–221
Igaki T, Pagliarini RA, Xu T (2006) Loss of cell polarity drives tumor growth and invasion through JNK activation in Drosophila. Curr Biol 16(11):1139–1146
Xue L, Noll M (2002) Dual role of the Pax gene paired in accessory gland development of Drosophila. Development 129(2):339–346
Ma X et al (2014) Bendless modulates JNK-mediated cell death and migration in Drosophila. Cell Death Differ 21(3):407–415
Hay BA, Wolff T, Rubin GM (1994) Expression of baculovirus P35 prevents cell death in Drosophila. Development 120(8):2121–2129
Wang SL et al (1999) The Drosophila caspase inhibitor DIAP1 is essential for cell survival and is negatively regulated by HID. Cell 98(4):453–463
Lisi S, Mazzon I, White K (2000) Diverse domains of THREAD/DIAP1 are required to inhibit apoptosis induced by REAPER and HID in Drosophila. Genetics 154(2):669–678
Rosen DR et al (1989) A Drosophila gene encoding a protein resembling the human beta-amyloid protein precursor. Proc Natl Acad Sci USA 86(7):2478–2482
Martin-Morris LE, White K (1990) The Drosophila transcript encoded by the beta-amyloid protein precursor-like gene is restricted to the nervous system. Development 110(1):185–195
Torroja L et al (1999) The Drosophila beta-amyloid precursor protein homolog promotes synapse differentiation at the neuromuscular junction. J Neurosci 19(18):7793–7803
Gunawardena S, Goldstein LS (2001) Disruption of axonal transport and neuronal viability by amyloid precursor protein mutations in Drosophila. Neuron 32(3):389–401
Torroja L et al (1999) Neuronal overexpression of APPL, the Drosophila homologue of the amyloid precursor protein (APP), disrupts axonal transport. Curr Biol 9(9):489–492
Soldano A et al (2013) The Drosophila homologue of the amyloid precursor protein is a conserved modulator of Wnt PCP signaling. PLoS Biol 11(5):e1001562
Kim HJ et al (2007) Drosophila homolog of APP-BP1 (dAPP-BP1) interacts antagonistically with APPL during Drosophila development. Cell Death Differ 14(1):103–115
Kajihara T et al (2006) Differential expression of FOXO1 and FOXO3a confers resistance to oxidative cell death upon endometrial decidualization. Mol Endocrinol 20(10):2444–2455
Bouchard C et al (2007) FoxO transcription factors suppress Myc-driven lymphomagenesis via direct activation of Arf. Genes Dev 21(21):2775–2787
Luo X et al (2007) Foxo and Fos regulate the decision between cell death and survival in response to UV irradiation. EMBO J 26(2):380–390
Siegrist SE et al (2010) Inactivation of both Foxo and reaper promotes long-term adult neurogenesis in Drosophila. Curr Biol 20(7):643–648
Shen J, Tower J (2010) Drosophila foxo acts in males to cause sexual-dimorphism in tissue-specific p53 life span effects. Exp Gerontol 45(2):97–105
You H, Mak TW (2005) Crosstalk between p53 and FOXO transcription factors. Cell Cycle 4(1):37–38
Fu W et al (2009) MDM2 acts downstream of p53 as an E3 ligase to promote FOXO ubiquitination and degradation. J Biol Chem 284(21):13987–14000
Acknowledgments
We thank Dr. Merders, Dr. Partridge and the Bloomington Drosophila Stock Center for fly stocks. This work is supported by the National Basic Research Program of China (973 Program) (2010CB944901, 2011CB943903), National Natural Science Foundation of China (31071294, 31171413, 31371490), the Specialized Research Fund for the Doctoral Program of Higher Education of China (20120072110023), and Shanghai Committee of Science and Technology (09DZ2260100, 14JC1406000).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10495_2015_1097_MOESM1_ESM.docx
Fig. S1 APLP1 induces cell death and defects in wing development. Fluorescent images of GFP expression (a) or acridine orange staining (b, c) of wing discs from 3rd instar larvae and light images of adult wing (e-g) are shown. dpp-Gal4 was used as a control (b,f), or to drive the expression of GFP (a,e) or APLP1 (c, g). The lower panels are high magnification of the boxed areas in the upper panels. d shows the statistical analysis of acridine orange-positive cells in band c, whereas h shows the statistical analysis of the acv presence in f andg.***: P ≤ 0.001. Genotypes: UAS-GFP/+ ; dpp-Gal4/+ (a,e); dpp-Gal4/+ (b, f); dpp-Gal4/UAS-APLP1 (c,g) (DOCX 201 kb)
10495_2015_1097_MOESM2_ESM.docx
Fig. S2 APLP1 induces cell death in the wing pouch area. Fluorescent images of GFP expression (a) or acridine orange staining (b, c) of wing discs from 3rd instar larvae are shown. sd-Gal4 was used as a control (b), or to drive the expression of GFP (a) or APLP1 (c). The lower panels are high magnification of the boxed areas in the upper panels. Genotypes: sd-Gal4/+ ; UAS-GFP/+ (a); sd-Gal4/+ (b);sd-Gal4/+ ; UAS-APLP1/+ (c) (DOCX 159 kb)
10495_2015_1097_MOESM3_ESM.docx
Fig. S3 Loss of dfoxO suppresses APLP1-induced blistered wing phenotype. (a-c) Light images of adult wings are shown. Compared with the sd-Gal4 control (a), expression of APLP1 produced a blistered wing phenotype (b), which was suppressed in heterozygous dfoxO mutants (c). The red arrow indicates a blister on the wing. d is the statistical analysis of the presence of blistered wing.***, P ≤ 0.001. Genotypes: sd-Gal4/+ (a); sd-Gal4/+ ;UAS-APLP1/+ (b); sd-Gal4/+ ;UAS-APLP1/dfoxO △94 (c) (DOCX 155 kb)
10495_2015_1097_MOESM4_ESM.docx
Fig.S4 APLP1 induces dFoxO-mediated cell death in the notum. Fluorescent images of acridine orange staining of notum tips of the wing discs from 3rd instar larvae are shown(a-c). Compared with the pnr-Gal4 control (a), expression of APLP1 resulted in enhanced cell death in the notum (b), which was suppressed by the expression of a dfoxO RNAi (c). The lower panels are high magnification of the boxed areas in the upper panels. d is the statistical analysis of acridine orange-positive cells in a-c.***, P ≤ 0.001; **, P ≤ 0.01. Genotypes: pnr-Gal4/+ (a); pnr-Gal4/UAS-APLP1 (b); pnr-Gal4 UAS-APLP1/UAS-dfoxO-IR#1 (c) (DOCX 118 kb)
10495_2015_1097_MOESM5_ESM.docx
Fig. S5 APLP1 induces cell death in the nervous system. Fluorescent images of GFP expression (a, e) or acridine orange staining (b, c, f, g) of the ventral nervecord (VNC) or eye disc from 3rd instar larvae are shown. elav-Gal4 and GMR-Gal4 were used as controls(b, f), or to drive the expression of GFP (a, e) or APLP1 (c, g). d shows statistical analysis of AO positive cells in b and c.**, P < 0.01. h shows statistical analysis of AO positive cells in f and g.***: P ≤ 0.001. Genotypes: elav-Gal4/+ ; UAS-GFP/+ (a); elav-Gal4/+ (b);elav-Gal4/+ ; UAS-APLP1/+ (c); UAS-GFP/+ ; GMR-Gal4/+ (e); GMR-Gal4/+ (f); GMR-Gal4/UAS-APLP1 (g) (DOCX 177 kb)
10495_2015_1097_MOESM6_ESM.docx
Fig. S6 APLP2 induces cell death and defects in wing development. Acridine orange staining (a, b) of wing discs from 3rd instar larvae and light images of adult wing (d,e) are shown. ptc-Gal4 was used as a control (a,d), or to drive the expression of APLP2 (b, e). The lower panels are high magnification of the boxed areas in the upper panels. c shows the statistical analysis of acridine orange-positive cells in a and b. f shows the statistical analysis of theacv presence in dande.***: P ≤ 0.001. Genotypes: ptc-Gal4/+ (a, d); ptc-Gal4/UAS-APLP2(b,e) (DOCX 275 kb)
10495_2015_1097_MOESM7_ESM.docx
Fig. S7 APLP2 induces cell death and defects in the wing and thorax. Acridine orange staining (a, b) of wing discs from 3rd instar larvae and light images of adult wing (c, d) or thorax (e, f) are shown. sd-Gal4 (a, c) or pnr-Gal4 (e) was used as controls, or to drive the expression of APLP2(b, d, f). The lower panels are high magnification of the boxed areas in the upper panels. Genotypes: sd-Gal4/+ (a, c); sd-Gal4/+ ; UAS-APLP2/+ (b, d); pnr-Gal4/+ (e); UAS-APLP2/+ ; pnr-Gal4/+ (f) (DOCX 251 kb)
10495_2015_1097_MOESM8_ESM.docx
Fig. S8 APPL induces cell death in the nervous system. Fluorescent images of RFP expression (a) or acridine orange staining (b, c) of the ventral nervecord from 3rd instar larvae are shown. APPL-Gal4 was used as a control (b), or to drive the expression of RFP (a) or APPLsd (c). The dashed white box indicates the ventral nerve cord middle line. The lower panels are high magnification of the boxed areas in the upper panels. d shows the statistical analysis of AO positive cells in b and c.**, P < 0.01. Genotypes: APPL-Gal4/+ ; UAS-RFP/+ (a); APPL-Gal4/+ (b);APPL-Gal4/+ ; UAS-APPLsd/+ (c) (DOCX 191 kb)
10495_2015_1097_MOESM9_ESM.docx
Fig. S9 Expression of FoxO proteins produces the loss-of-acv phenotype. (a-c) Light images of adult wings are shown. Compared with the ptc-Gal4 control (a), expression of dFoxO or hFoxO3a resulted in the loss-of-acv phenotype (b, c). The lower panels are high magnification of the boxed areas in the upper panels. d shows the statistical analysis of the presence of acv in a-c.***, P ≤ 0.001. Genotypes: ptc-Gal4/+ (a); ptc-Gal4/+ ;UAS-dFoxO/+ (b); ptc-Gal4/+ ;UAS-hFoxO3a/+ (c) (DOCX 220 kb)
10495_2015_1097_MOESM10_ESM.docx
Fig. S10 Expression of APLP2 activates the transcription of FoxO target genes. Light images of Drosophila 3rd instar wing discs are shown. X-Gal staining of hid-LacZ and reaper-LacZ reporters in the wing pouch (a, c) are dramatically up-regulated by the expression of APLP2 (b, d) (DOCX 190 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Ma, Y., Zhao, Y. et al. APLP1 promotes dFoxO-dependent cell death in Drosophila . Apoptosis 20, 778–786 (2015). https://doi.org/10.1007/s10495-015-1097-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10495-015-1097-1