Abstract
House dust mites (HDMs) including Dermatophagoides spp. are an important cause of respiratory allergies. However, their relationship with microorganisms in house dust has not been fully elucidated. Here, we characterized bacteria and fungi associated with HDMs in house dust samples collected in 107 homes in Korea by using DNA barcode sequencing of bacterial 16S rRNA gene, fungal internal transcribed spacer 2 (ITS2) region, and arthropod cytochrome c oxidase I (COI) gene. Our inter-kingdom co-occurrence network analysis and/or indicator species analysis identified that HDMs were positively related with a xerophilic fungus Wallemia, mycoparasitic fungi such as Cystobasidium, and some human skin-related bacterial and fungal genera, and they were negatively related with the hygrophilous fungus Cephalotrichum. Overall, our study has succeeded in adding novel insights into HDM-related bacteria and fungi in the house dust ecosystem, and in confirming the historically recognized fact that HDMs are associated with xerophilic fungi such as Wallemia. Understanding the microbial ecology in house dust is thought to be important for elucidating the etiology of human diseases including allergies, and our study revealed baseline information of house dust ecology in relation to HDMs. The findings could be useful from a perspective of human health.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
House dust mites (HDMs) belong to the family Pyroglyphidae and inhabit human dwellings, with examples including Dermatophagoides farinae, Dermatophagoides pteronyssinus, and Euroglyphus maynei. Many of them are cosmopolitan and distributed worldwide (Colloff 2009; Thomas 2010). In Korea, they are frequently detected in house dust, with D. farinae being reported as the most predominant species followed by D. pteronyssinus (Ree et al. 1997). HDMs are known to be allergenic and more than 60 allergens of Dermatophagoides spp. are reported by the WHO/IUIS Allergen Nomenclature Sub-committee (http://allergen.org/) as of December 2021. Moreover, a recent proteogenomic study identified additional allergen-like proteins (Erban et al. 2020b). HDM allergens are associated with allergic rhinitis and asthma, and exposure to them is known to exacerbate asthma in children who are sensitized to them (Kanchongkittiphon et al. 2015). According to one estimate, 1–2% of the world’s population and 50% of asthma patients are sensitized to HDM allergens (Caldert al. 2015). In Korea, HDM is the most common type of aeroallergens to which about 30–40% of the patients with allergic rhinitis and/or asthma are sensitized (Park et al. 2019).
House dust mites inhabit human dwellings. The indoor reservoirs of HDMs include carpets, upholstered furniture, mattresses, bedding, and settled house dust (Portnoy et al. 2013). These indoor microenvironments serve not only as habitats for HDMs, but also for other arthropods (Bertone et al. 2016; Madden et al. 2016), such as domestic cockroaches (Bernton and Brown 1964; Kang et al. 1979). Additionally, human dwellings serve as habitats for microorganisms such as bacteria (Jeon et al. 2013; Thompson et al. 2021) and fungi (Adams et al. 2013; An and Yamamoto 2016). Consequently, a unique ecosystem is formed in house dust, which is a mixture of human dander, animal hair and dander, pollen, plant debris, food debris, soil and fiber, and can be the source of nutrients for them (van Bronswijk 1981).
The co-occurrence of HDMs and microorganisms in house dust has been investigated historically by culture and microscopic observation (Sinha et al. 1970; Lustgraaf 1978b; Rijckaert et al. 1981), but more recently by DNA sequencing (Loo et al. 2018). By analyzing HDMs in culture, the associations between HDMs and microorganisms (bacteria and fungi) have been investigated in vitro, both inside (intestine) and outside (environment) of the mites (van Bronswijk and Sinha 1973; Lustgraaf 1978a; Oh et al. 1986; Hay et al. 1992; Hay et al. 1993; Hubert et al. 2019; Klimov et al. 2019; Nesvorna et al. 2021). Overall, these studies revealed that fungi in small quantity are beneficial to HDMs in that fungi digest lipids and proteins in house dust and facilitate the absorption of nutrients by HDMs and that HDMs utilize fungi as a food source (Van Asselt 1999; Acevedo et al. 2019). However, large amounts of fungi are known to be detrimental because the dense colonization of fungi is thought to impede the movement of HDMs and/or produce toxins that are harmful to HDMs (Van Asselt 1999; Acevedo et al. 2019). Taxonomically, xerophilic fungi, such as Wallemia sebi and Aspergillus penicillioides, are known to be related to xerotolerant HDMs (Lustgraaf 1978a; Lustgraaf 1978b; Van Asselt 1999). However, there is still a lack of detailed information on taxonomical composition and diversity of bacteria and fungi related to HDMs, especially in actual dust systems in homes (i.e., in vivo).
Here, we aimed to characterize bacteria and fungi associated with HDMs in house dust samples collected from homes in Korea. We used DNA barcode sequencing of bacterial 16S rRNA gene, fungal internal transcribed spacer 2 (ITS2) region, and arthropod cytochrome c oxidase I (COI) gene to detect and identify these organisms. HDMs were detected and identified by DNA barcode sequencing of the arthropod COI gene. The use of culture-independent high-throughput DNA barcoding has the advantage of being able to detect rare species and species that are difficult to culture (Shinohara et al. 2021), and allows characterization of the full spectrum of bacterial and fungal microbiomes associated with HDMs in house dust. Based on the results of DNA metabarcoding, we identified bacteria and fungi associated with HDMs in house dust by statistical methods such as network analysis and indicator species analysis.
Materials and methods
Sample collection and preparation
A total of 107 settled house dust samples in homes in Seoul city and Gyeonggi province of South Korea were collected by a field technician during April and May of 2021. The descriptive statistics of sampled homes are summarized in Table S1. Each of the dust samples was collected from the vacuum cleaner of the resident and enclosed into a sterilized zipper bags and transported to a laboratory. A sieve with 150 µm stainless mesh was used to remove large particles from each sample. Prior to each sieving, the mesh was washed with tap water using a scourer, rinsed with soapy water, deionized water, and acetone in that order, and dried. More than 1.2 g of each sieved dust sample was obtained and stored in a sterilized vial and kept at −20 °C until the subsequent experiment.
DNA extraction
The genomic DNA was extracted from 0.025 g of each sieved dust sample using AccuPrep Stool DNA Extraction Kit (Bioneer, Daejeon, Korea). For DNA extraction, 0.025 g of each sieved dust sample, 20 µl of proteinase K, 400 µl of SL buffer from the extraction kit, 0.3 g of 0.1 mm diameter glass beads, and 0.1 g of 0.3 mm diameter glass beads were added into sterilized 2-ml tube and homogenized by a beadbeater-24 (Biospec Products, Bartlesville, OK, USA). After homogenization, the genomic DNA was eluted into 50 µl of TE (10 mM Tris–HCl, 1 mM EDTA, pH 8.0) by the kit’s protocol. The extracted DNA was kept at −80 °C until the following experiment.
DNA sequencing
Three DNA barcode markers targeting arthropods, bacteria, or fungi were PCR amplified separately. For arthropods, the cytochrome c oxidase I (COI) gene was amplified with primers BF1 and BR2 (Elbrecht and Leese 2017). For bacteria, the 16S rRNA gene was amplified with primers Bakt_341F and Bakt_805R (Herlemann et al. 2011). For fungi, the internal transcribed spacer 2 (ITS2) region was amplified with primers fITS7 and ITS4 (White et al. 1990; Ihrmark et al. 2012). For the tagged sequencing on a MiSeq platform (Illumina, San Diego, CA, USA), the primers with the adapter sequences were used.
Each PCR reaction mixture (25 µl) contained 12.5 µl of the 2× PCR Solution Premix Taq (Takara Bio, Otsu, Shiga, Japan), and 1 µl of extracted DNA. The concentrations of primers were 0.5 µM for arthropods, 1 µM for bacteria, and 0.33 µM for fungi. For arthropods, the PCR thermal cycle was 94 °C for 3 min for initial denaturation, followed by 35 cycles of 94 °C for 30 s, 50 °C for 30 s, and 65 °C for 90 s, and completed at 65 °C for 5 min for final elongation. For bacteria and fungi, the PCR thermal cycle was 95 °C for 5 min for initial denaturation, followed by 35 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 60 s, and completed at 72 °C for 10 min for final elongation. After amplification, the PCR products were purified by AMPure XP beads (Beckman Coulter, Brea, CA, USA). After purification, the index PCR was conducted with the Nextera XT Index Kit v.2 (Illumina) against each of the purified PCR product. For the index PCR, the thermal cycle was 95 °C for 3 min for initial denaturation, followed by 8 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s, and completed at 72 °C for 5 min for final elongation.
The indexed PCR products were purified by AMPure XP beads (Beckman Coulter) and quantified by Quant-iT PicoGreen ds DNA reagent kit (Life Technologies, Carlsbad, CA, USA). The quantified samples were normalized, pooled with the internal control PhiX (30%), and injected to the v.3 600 cycle-kit reagent cartridge (Illumina) for 2× 300 bp paired-end sequencing on an Illumina MiSeq system.
Taxonomic assignment
The sequencing data processing method for bacterial libraries was described in our previous paper (Woo and Yamamoto 2020). Briefly, USEARCH v.11.0.667 (Edgar 2010) was used for quality trimming and taxonomic assignment. Low-quality reads with >1.0 expected errors and/or those with lengths <200 bp were excluded. From the resultant reads, unique sequences were identified. Using the UNOISE algorithm (Edgar and Flyvbjerg 2015), chimeric reads were also removed, and zero-radius OTUs (ZOTUs) were identified. The SINTAX algorithm (Edgar 2016) was used for bacterial taxonomic assignment of each ZOTU against the RDP training set v.18 (rdp_16s_v18.fa.gz) with a cutoff confidence value 0.8.
For arthropod libraries, taxonomic assignments were performed in mothur v.1.48.0 (Schloss et al. 2009). First, we constructed a custom reference database of arthropod COI gene sequences. In total, 6,037,295 arthropod COI gene sequences were downloaded from the Barcode of life data system (BOLD) v.4 on July 28, 2021, from which, using the pcr.seqs command, those amplifiable by the arthropod primer pair used in this study were incorporated into our database. The sequences containing ambiguous reads were removed using the screen.seqs command. In total, 75,526 sequences were obtained and 22 human mitochondrial DNA sequences (Table S2) were added and incorporated into our custom database. Second, the obtained arthropod COI gene sequence reads were taxonomically assigned against our custom database. The forward and reverse reads of each sequence were combined with the make.contigs command. For combined reads, the screen.seqs command was used with criteria of 250 min-length, 450 max-length, and 0 max ambiguous reads to eliminate low-quality reads. Then, unique sequences were identified using the unique.seqs command. The identified unique sequences were aligned to our custom database with the align.seqs command. From the aligned sequences, gaps were removed using the degap.seqs command, and sequences amplifiable with our primer pair were selected using the screen.seqs command. Subsequently, the unique sequences were screened for chimeric reads using chimera.vsearch command and removed. The remaining reads were taxonomically assigned against our custom database. The reads assigned to human mitochondrial DNA were removed using the remove.lineage command, and amplicon sequence variants (ASVs) were identified with the classify.otu command for the remaining reads.
For fungal libraries, our previously reported method (An and Yamamoto 2016) was used. Briefly, the reads after quality trimming were taxonomically assigned using the BLAST 2.12.0+ against the reference database (sh_general_release_dynamic_s_10.05.2021.fasta) downloaded from UNITE (Nilsson et al. 2018; Kõljalg et al. 2020) from which sequences of ambiguous species were removed. Specifically, the reference database initially contained 98,090 sequences, but we excluded sequences of ambiguous species (denoted as ‘Ascomycota_sp’) to prepare the database containing only 42,881 unambiguous sequences. After taxonomic assignment, FHiTHINGS (Dannemiller et al. 2014) v.1.5 was used to summarize the number of sequences at each taxonomic rank with a cutoff of 0.001.
Diversity analysis
The obtained high-quality sequence reads were analyzed using the phyloseq package (McMurdie and Holmes 2013), vegan package (Oksanen et al. 2017), and R v.4.2.1. Prior to performing diversity analysis, bacterial ZOTUs classified as Streptophyta (green plants) or Zea (maize) at the genus level were excluded. After the exclusion, bacterial and arthropod ZOTUs and fungal reads that have the same taxonomy were merged by the tax_glom function in phyloseq package. Then, bacterial, fungal, and arthropod libraries were rarefied into 1,100, 4,700, and 400 reads, respectively. Based on the rarefied libraries, the estimate_richness function in phyloseq package was used to measure α diversity indices of bacterial, fungal, and arthropod libraries. Furthermore, β diversity analysis was conducted based on the rarefied libraries with the Jaccard similarity coefficient and Bray–Curtis dissimilarity. To compare dissimilarities, the adonis2 function in vegan package was used for performing permutational multivariate analysis of variance (PERMANOVA).
Network analysis
We constructed an inter-kingdom co-occurrence network at the genus level for fungi, bacteria and arthropods using the SParse InversE Covariance estimation for Ecological ASsociation Inference (SPIEC-EASI) package v.1.1.2 in R (https://github.com/zdk123/SpiecEasi/) (Kurtz et al. 2015). All the count tables were filtered prior to running SpiecEasi to include only genera found in at least 20% of samples. The Meinshausen-Buhlmann (MB) method for neighborhood selection with a lambda minimum ratio of 1e-2 and nlambda of 40 was used for the network inference. Network plots were displayed using Gephi software (https://github.com/gephi/gephi).
Indicator species analysis
The indicator species analysis was performed to identify bacterial and fungal genera significantly associated with HDMs (Dermatophagoides spp.). Specifically, the samples were divided into ‘High HDM’ or ‘Low HDM’ groups using the median value of the relative abundance of HDMs obtained by analysis of arthropods as a threshold. Bacterial and fungal genera that were disproportionately high in relative abundance in ‘High HDM’ or ‘Low HDM’ groups were identified using the ‘multipatt’ function (with ‘r.g’ function and 9,999 permutations) in the R package ‘indicspecies’ (De Caceres and Legendre 2009) in R v.4.2.1.
Results
Sequencing statistics
From a total of 107 settled house dust samples, 94 arthropod libraries, 103 bacterial libraries, and 103 fungal libraries were prepared. Some samples (13 samples for arthropods, and four samples for bacteria and fungi) were not PCR amplified, so the libraries could not be constructed. A total of 3,792,537 high-quality sequence reads consisting of 631,800 reads of arthropods, 1,301,958 reads of bacteria, and 1,858,779 reads of fungi were obtained (Table S3).
Arthropods
House dust mites of the genus Dermatophagoides were detected from 86 of 94 dust samples (91.5%) analyzed (Fig. 1 A). The mean relative abundance of Dermatophagoides in all analyzed samples was 28.7%. In total, 981 ASVs were assigned to Dermatophagoides or Tyrophagus, and centroid sequences of 35 sequence clusters from these ASVs are shown in Supplementary Data 1. Tyrophagus (mold mite) was detected from 43 of 94 dust samples (45.7%) analyzed. The mean relative abundance of Tyrophagus in all analyzed samples was 2.4%. It is known that eight species of mites are found in dust in South Korea, including three major species: D. farinae, D. pteronyssinus, and Tyrophagus putrescentiae (Jeong et al. 2012). Dermatophagoides farinae is known to be the most dominant (65.3% of the total mite individuals), followed by D. pteronyssinus (20.6%) and T. putrescentiae (6.5%) (Ree et al. 1997). In our study, both Dermatophagoides and Tyrophagus were detected. The misidentification of other mite species as Dermatophagoides or Tyrophagus is unlikely as the sequenced COI regions are dissimilar among the mite species inhabiting Korea (Fig. S1).
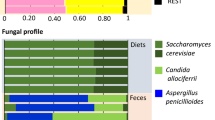
Relative abundance of A arthropod genera identified based on the COI gene, B bacterial genera identified based on 16S rRNA gene, and C fungal genera identified based on the ITS2 region. For each organism, the top-30 most abundant genera are shown. The tree shows the log-transformed Euclidean distance of the compositional similarity of the genera shown
The measured relative abundance of Dermatophagoides ranged from 0 to 96.6%, and the median and mean values were 15.8% and 28.7%, respectively. Based on its relative abundance, the samples were divided into groups of first (Q1), second (Q2), third (Q3), and fourth (Q4) quartiles (Fig. 2) for the subsequent α diversity analysis. In the β diversity and indicator species analyses, the group combining Q1 and Q2 was named the ‘High HDM’ group, and the group combining Q3 and Q4 was named the ‘Low HDM’ group.
Cumulative distribution of the relative abundance of the number of sequences of house dust mites (HDMs) (Dermatophagoides spp.) to the number of sequences of all detected arthropods. The samples are classified into groups of first (Q1), second (Q2), third (Q3), and fourth (Q4) quartiles depending on their relative abundance for subsequent analyses
Bacteria
The abundant bacteria identified from settled house dust appears to be related with the human microbiome, especially skin, mucous, and oral cavity. Specifically, the dominant genera and their mean relative abundances were Staphylococcus (23%), Corynebacterium (14%), Streptococcus (9%), Lactobacillus (8%), and Cutibacterium (4%). The top-30 bacterial genera and their abundance are shown in Fig. 1B.
Fungi
Among fungi, Malassezia, which is the resident yeast on human and animal skin, was the most abundant genus throughout the samples with 21% of mean relative abundance. Saccharomyces, which is the genus containing yeasts, was also abundant throughout the sample and its mean relative abundance was 20%. Lentinula and Pleurotus, the well-known edible mushrooms, each showed 5% of mean relative abundance. The top-30 fungal genera identified from the settled house dust samples are shown in Fig. 1 C.
Alpha diversity
As mentioned, the samples were divided into groups of first (Q1), second (Q2), third (Q3), and fourth (Q4) quartiles based on the relative abundance of HDMs (Dermatophagoides spp.) (Fig. 2). The α diversity indices of bacteria and fungi observed in each group were presented in Fig. S2. There were no significant differences between the groups for all diversity indices and organisms tested (Kruskal-Wallis test: p > 0.05). The descriptive statistics of α diversity indicators of each organism is listed in Table S4, and the rarefaction curves are shown in Fig. S3.
Beta diversity
Beta diversity analysis was performed to compare bacterial and fungal communities between the ‘High HDM’ and ‘Low HDM’ groups (Fig. S4). On both the community structures (Bray–Curtis dissimilarity) and memberships (Jaccard similarity), the effects of the groups appear to be small in both bacteria and fungi (PERMANOVA: r2 < 0.03).
Inter-kingdom network
The inter-kingdom co-occurrence network among fungi, bacteria and arthropods genera in the house dust consisted of 289 nodes (227 fungi, 57 bacteria and five arthropods) and 922 edges (69.85% positive and 30.15% negative co-occurrences) (Fig. 3). We found that HDMs (Dermatophagoides) were associated with five genera, with a positive association with Wallemia (fungi), and negative associations with Faecalimonas (bacteria), Cephalotrichum (fungi), Dinoderus (arthropods) and Lasioderma (arthropods) (Table 1). The complementary cumulative distributions of the network metrics, such as centralities, are shown in Fig. S5. The top 200 cross-taxon edges with the highest observed interactions are listed in Table S5.
Inter-kingdom co-occurrence network of fungi, bacteria and arthropods genera in settled house dust samples constructed using the SpiecEasi method. Nodes represent individual genera, and edges represent significant positive (green edges) and negative (red edges) associations. The size of each node is proportional to the number of connections (degree). The organisms associated with house dust mites (Dermatophagoides) are highlighted with thick lines. The weight values of top 200 cross-taxon edges with the highest observed interactions are listed in Table S5
Indicator genera
The indicator species analysis revealed several bacterial and fungal genera that were found in the ‘High HDM’ and ‘Low HDM’ groups. Four bacterial genera were found to be disproportionately abundant in the ‘High HDM’ group (Table 2). No indicator bacterial genus was found in the ‘Low HDM’ group. Similarly, nine fungal genera were found to be disproportionately abundant in the ‘High HDM’ group, with examples including Coniothyrium, Cystobasidium, Torulaspora, Trichophyton, and Zygosaccharomyces (Table 3). Conversely, 10 fungal genera were found to be disproportionately abundant in the ‘Low HDM’ group.
In Fig. 4, the relative abundances of bacteria and fungi are shown that were found to be associated with HDMs (Dermatophagoides) by the indicator species analysis and/or co-occurrence network analysis.
Relative abundance of A bacteria and B fungi that were found to be associated with house dust mites (HDMs) (Dermatophagoides) by the indicator species analysis and/or co-occurrence network analysis. The genera shown in red and green indicate the genera that were negatively and positively associated with HDMs, respectively. Each column of the heatmaps represents a house dust sample. From left to right, the columns are arranged in descending order based on the relative abundance of HDMs. The tree shows the log-transformed Euclidean distance of the compositional similarity of the genera
Discussion
Using DNA metabarcoding followed by indicator species analysis and/or network analysis, this study identified bacteria and fungi associated with HDMs (Dermatophagoides) in settled house dust in Korea. In some previous studies (Horak et al. 1996; Li et al. 2022), HDMs (number of mites or allergen levels) and microorganisms (bacteria and/or fungi) in house dust have been measured in parallel. In this study, we used DNA metabarcoding to identify arthropods, including HDMs, and microorganisms (both bacteria and fungi) and statistically analyzed their co-occurrences. We discovered that the microorganisms positively related with HDMs included the xerophilic fungi, mycoparasitic fungi and some human skin-related bacteria, and those negatively related included the hygrophilous fungi. These findings were made possible because we could collect a large number of house dust samples (n = 107) to ensure enough statistical power.
Wallemia was found to be positively networked with HDMs. Wallemia is a xerophilic fungus with species such as Wallemia sebi commonly found in house dust (Desroches et al. 2014). The coexistence of Wallemia and HDMs has also been reported by previous studies. For instance, W. sebi has been isolated from HDMs in culture (Hay et al. 1992). A more recent DNA-based study has reported that Wallemia is the predominant fungus in the microbiome of D. pteronyssinus in culture (Hubert et al. 2019). An in vitro study also suggested that the growth of xerophilic fungi on the media stimulated increase in the number of HDMs, although the effect of Wallemia was small compared to other xerophilic fungi, e.g., Aspergillus penicillioides (Lustgraaf 1978a). These studies may suggest that xerophilic fungi, including Wallemia, may be suitable food sources for HDMs. Additionally, field studies have also reported co-detection of xerophilic fungi such as W. sebi and A. penicillioides and HDMs in indoor dust, which have been analyzed by conventional culture and microscopic methods (Lustgraaf 1978b; Rijckaert et al. 1981). Using DNA metabarcoding, our study also revealed that Wallemia and HDMs were networked in house dust ecosystems. The co-occurrence of HDMs and Wallemia seems reasonable because they are both xerotolerant and therefore their ecological preferences may be similar. For instance, HDMs are reported to survive even in an environment maintained at 10% relative humidity (RH), if they are exposed to a high relative humidity (90% RH) for 1.5 h a day (de Boer and Kuller 1997), and 65–75% RH has been reported to be optimal for their population growth (Arlian 1992; Arlian et al. 1998).
Cystobasidium was also relatively abundant and positively associated with HDMs by the indicator species analysis. Cystobasidium is a pink-to-orange colored basidiomycete yeast that is genetically distant but resembles Rhodotorula in appearance (Yurkov et al. 2015), a commonly found indoor yeast (An and Yamamoto 2016). Rhodotorula, previously a polyphyletic genus, used to contain some species of Cystobasidium, but they are now separated (Wang et al. 2015; Yurkov et al. 2015) in response to recent changes in the code of fungal nomenclature (Hawksworth et al. 2011). Cystobasidium spp. have often been isolated from soil (Li et al. 2020). Our study shows that it is not uncommon in house dust in Korea. Cystobasidium is known to be mycoparasitic (Bauer et al. 2006; Wang et al. 2015). One hypothesis that Cystobasidium was positively related with HDMs may be due to its mycoparasitism. Interestingly, mycoparasitism or killer activity against other fungi is also known for Coniothyrium (Campbell 1947), Torulaspora (Rosa-Magri et al. 2011), and Zygosaccharomyces (Weiler and Schmitt 2003), which were also positively related with HDMs. We speculate that these mycoparasitic yeasts or filamentous fungi are beneficial to HDMs as they may prevent from excessive fungal growth, which is known to be detrimental to HDMs (Van Asselt 1999; Acevedo et al. 2019).
Cephalotrichum was identified as a fungus that is negatively related with HDMs by network analysis. The genus Cephalotrichum contains the former synnematous genus Doratomyces (Woudenberg et al. 2017). Doratomyces is reported to have grown in the building envelope that appeared to be in chronically wet conditions (Morey 2011), and is known to occur on wet cellulosic substrates such as wood (Woudenberg et al. 2017). The negative correlation between HDMs and hygrophilous Cephalotrichum appears to be consistent with the known fact that large amounts of fungi are not beneficial to HDMs (Van Asselt 1999; Acevedo et al. 2019). It may be possible to hypothesize that the lack of co-occurrence between xerotolerant HDMs and hygrophilous Cephalotrichum is due to the dense fungal colonization and/or the differences in habitat or ecological preferences.
In addition to fungi, some bacteria have been found to be positively related with HDMs in house dust by indicator species analysis. We speculate that it is due to their relationship with human skin. For instance, Granulicatella, Lautropia, and Stenotrophomonas have been reported to be detected on human skin (Dekio et al. 2007; Gao et al. 2007; Ross et al. 2017). Similarly, Trichophyton was also found to be positively related with HDMs in house dust, and this fungus is also known to be associated with human skin (Wang et al. 2021). It is thought that human skin-related bacteria and fungi likely co-occur with HDMs, as HDMs are known to eat human skin fragments that are abundant in house dust (van Bronswijk 1981). In addition, Neisseria, which is known to be found mainly in the upper respiratory tract as a commensal and/or pathogenic bacterial genus (Weyand 2017), is also found to be positively related with HDMs. Exposure to HDMs is known to affect airway microbiome diversity, and Neisseria is known to be more abundantly detected in patients with allergic rhinitis induced by HDMs (Chiu et al. 2017, 2020). Therefore, one hypothesis we can propose is that exposure to HDMs caused allergic reactions such as rhinitis to increase Neisseria in the airways of the residents, which in turn increased Neisseria in the house dust.
However, no association was found with other abundantly detected skin-related bacteria such as Staphylococcus and Corynebacterium. In addition, no association was found with HDMs with Malassezia and Saccharomyces, the two most abundantly detected fungal genera. These fungal genera are known to be associated with HDMs under laboratory conditions. For example, Malassezia has been suggested to have a persistent symbiotic relationship with HDMs (Nesvorna et al. 2021). Saccharomyces is known to be food for HDMs (Molva et al. 2019). However, in our study, neither network analysis nor indicator species analysis showed a significant relationship between these microorganisms and HDMs. One explanation is that in the actual indoor environment, these microorganisms may be present in much higher abundance than is required as food or as symbionts of HDMs, compared to the controlled laboratory conditions reported by the previous studies. If it is true that the abundances of these genera always far exceed those required for HDM in the actual indoor environment, it would explain that these genera are not the controlling factor of HDM abundance and therefore not correlated with HDMs.
A negative association was found between HDMs and Candida, which seems to contradict the results of previous laboratory-based studies (Hubert et al. 2019; Nesvorna et al. 2021). We do not know the reason. One possibility is that the abnormally high abundances of Candida observed in three samples (samples 26, 96, and 152) biased the overall statistical result. However, more research is needed on this point.
Recent studies have reported more about bacteria inside the mites in culture, and some studies report intracellular bacteria symbiotic to the mites, such as Cardinium (Erban et al. 2020a; Hubert et al. 2021) and Bartonella or Bartonella-like bacteria (Valerio et al. 2005; Hubert et al. 2012; Kim et al. 2018), which are thought to serve as indicators of the mites. In our study, however, neither Cardinium nor Bartonella was detected, which may be due in part to our reference database. In fact, our database lacks a reference Cardinium sequence, but we also expect that it is possible that Cardinium was absent in our samples, as Kim et al. (2018) reported the absence of Cardinium in the Korean D. farinae strain in culture. Our database contained 44 reference sequences of Bartonella (Table S6). To identify the possible bias in our method, we performed an in silico analysis and scrutinized ZOTUs assigned to the order Rhizobiales to which Bartonella belongs (Supplementary Information). We could confirm that our method can unambiguously detect and identify Bartonella. Therefore, we consider that Bartonella was truly absent in our DNA libraries. The absence of Bartonella may be due to the differences in microbiome between the mites. For example, T. putrescentiae is known to have populations with and without endosymbiotic bacteria (Erban et al. 2021). Another study analyzing the microbiome of D. pteronyssinus in culture did not also report on Bartonella (Nesvorna et al. 2021), indicating the absence of Bartonella in their population. Additionally, another explanation is that Bartonella was present but was below detection limit because its abundance was too low compared to other bacteria contained in house dust whose abundance is much higher than the abundance of the mites themselves.
In both bacteria and fungi, HDMs had negligible effects on the community structure Bray–Curtis dissimilarity) and membership (Jaccard similarity) (PERMANOVA: r2 < 0.03), suggesting that the role of HDMs in shaping the characteristic microbial communities in house dust is small. In fact, our network analysis revealed that HDMs (Dermatophagoides) were located peripherally in the network of the house dust ecosystem. The relatively small values of network metrics such as degrees and centralities calculated for Dermatophagoides also support that HDMs play only peripheral roles in shaping the overall microbial communities in house dust.
A caveat of this study is that the quantification of organisms is relative. The measured relative abundance might be affected, especially among species of arthropods. Specifically, even if the absolute abundance is the same, the relative abundance of one species becomes high if the abundance of other species is low. This effect might be enhanced in arthropods in which only a few species were detected, and we suspect that the observed negative correlations of Dermatophagoides to Dinoderus (bamboo powder-post beetles) and Lasioderma (cigarette beetle) might be due to this effect as there seems to be no conceivable biological basis. Furthermore, it should be noted that the inter-kingdom co-occurrence network constructed is based on the relative abundance of organisms quantified within each kingdom. Though the similar approach has been also used by many of other studies (Tipton et al. 2018; Schlatter et al. 2020), the use of methods such as quantitative PCR or metagenomic sequencing that quantifies the abundance of organisms on the same scale across kingdoms may be preferable in the future. Nonetheless, we believe that the network constructed in this study well represents the house dust ecosystems as we could successfully identify the historically well-known positive relationship between xerophilic Wallemia and HDMs (Lustgraaf 1978a; Lustgraaf 1978b; Rijckaert et al. 1981; Hay et al. 1992; Hubert et al. 2019).
In summary, using high-throughput DNA metabarcoding followed by co-occurrence network analysis and indicator species analysis, we identified microorganisms associated with HDMs in house dust. Specifically, we identified that HDMs were positively related with the xerophilic fungus Wallemia, mycoparasitic fungi such as Cystobasidium, and some human skin-related bacterial and fungal genera, and they were negatively related with the hygrophilous fungus Cephalotrichum (syn. Doratomyces). Understanding the microbial ecology of house dust is important for elucidating the etiology of allergies, for example, because the house dust microbiomes are thought to affect human intestinal microbiomes that are thought to be associated with human diseases such as allergies (Shan et al. 2019). In this study, we provided baseline information regarding the HDM-related microorganisms in house dust. The findings could be also useful from a perspective of human health.
Data availability
The raw sequencing data is available on NCBI under the BioProject number PRJNA791469 for arthropods, PRJNA791474 for bacteria, and PRJNA791477 for fungi.
References
Acevedo N, Zakzuk J, Caraballo L (2019) House dust mite allergy under changing environments. Allergy Asthma Immunol Res 11:450–469. https://doi.org/10.4168/aair.2019.11.4.450
Adams RI, Miletto M, Taylor JW, Bruns TD (2013) The diversity and distribution of fungi on residential surfaces. PLoS ONE 8:e78866. https://doi.org/10.1371/journal.pone.0078866
An C, Yamamoto N (2016) Fungal compositions and diversities on indoor surfaces with visible mold growths in residential buildings in the Seoul Capital Area of South Korea. Indoor Air 26:714–723. https://doi.org/10.1111/ina.12261
Arlian LG (1992) Water balance and humidity requirements of house dust mites. Exp Appl Acarol 16:15–35. https://doi.org/10.1007/bf01201490
Arlian LG, Confer PD, Rapp CM, Vyszenski-Moher DL, Chang JC (1998) Population dynamics of the house dust mites Dermatophagoides farinae, D. pteronyssinus, and Euroglyphus maynei (Acari: Pyroglyphidae) at specific relative humidities. J Med Entomol 35:46–53. https://doi.org/10.1093/jmedent/35.1.46
Bauer R, Begerow D, Sampaio JP, Wei M, Oberwinkler F (2006) The simple-septate basidiomycetes: a synopsis. Mycol Prog 5:41–66. https://doi.org/10.1007/s11557-006-0502-0
Bernton HS, Brown H (1964) Insect allergy—Preliminary studies of the cockroach. J Allergy 35:506–513. https://doi.org/10.1016/0021-8707(64)90082-6
Bertone MA, Leong M, Bayless KM, Malow TL, Dunn RR, Trautwein MD (2016) Arthropods of the great indoors: characterizing diversity inside urban and suburban homes. PeerJ 4:e1582. https://doi.org/10.7717/peerj.1582
Calder A, Linneberg A, Kleine-Tebbe J, De Blay F, Hernandez Fernandez de Rojas D, Virchow JC et al (2015) Respiratory allergy caused by house dust mites: What do we really know? J Allergy Clin Immunol 136:38–48. https://doi.org/10.1016/j.jaci.2014.10.012
Campbell WA (1947) A new species of Coniothyrium parasitic on Sclerotia. Mycologia 39:190–195. https://doi.org/10.2307/3755006
Chiu C-Y, Chan Y-L, Tsai Y-S, Chen S-A, Wang C-J, Chen K-F et al (2017) Airway microbial diversity is inversely associated with mite-sensitized rhinitis and asthma in early childhood. Sci Rep 7:1820. https://doi.org/10.1038/s41598-017-02067-7
Chiu C-Y, Chan Y-L, Tsai M-H, Wang C-J, Chiang M-H, Chiu C-C et al (2020) Cross-talk between airway and gut microbiome links to IgE responses to house dust mites in childhood airway allergies. Sci Rep 10:13449. https://doi.org/10.1038/s41598-020-70528-7
Colloff MJ (2009) Dust mites. CSIRO Publishing. & Springer, Melbourne. https://doi.org/10.1007/978-90-481-2224-0
Dannemiller KC, Reeves D, Bibby K, Yamamoto N, Peccia J (2014) Fungal High-throughput Taxonomic Identification tool for use with Next-Generation Sequencing (FHiTINGS). J Basic Microbiol 54:315–321. https://doi.org/10.1002/jobm.201200507
de Boer R, Kuller K (1997) Mattresses as a winter refuge for house-dust mite populations. Allergy 52:299–305. https://doi.org/10.1111/j.1398-9995.1997.tb00994.x
De Caceres M, Legendre P (2009) Associations between species and groups of sites: indices and statistical inference. Ecology 90:3566–3574. https://doi.org/10.1890/08-1823.1
Dekio I, Sakamoto M, Hayashi H, Amagai M, Suematsu M, Benno Y (2007) Characterization of skin microbiota in patients with atopic dermatitis and in normal subjects using 16S rRNA gene-based comprehensive analysis. J Med Microbiol 56:1675–1683. https://doi.org/10.1099/jmm.0.47268-0
Desroches TC, McMullin DR, Miller JD (2014) Extrolites of Wallemia sebi, a very common fungus in the built environment. Indoor Air 24:533–542. https://doi.org/10.1111/ina.12100
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Edgar RC (2016) SINTAX: a simple non-Bayesian taxonomy classifier for 16S and ITS sequences. bioRxiv. https://doi.org/10.1101/074161
Edgar RC, Flyvbjerg H (2015) Error filtering, pair assembly and error correction for next-generation sequencing reads. Bioinformatics 31:3476–3482. https://doi.org/10.1093/bioinformatics/btv401
Elbrecht V, Leese F (2017) Validation and development of COI metabarcoding primers for freshwater macroinvertebrate bioassessment. Front Environ Sci. https://doi.org/10.3389/fenvs.2017.00011
Erban T, Klimov PB, Harant K, Talacko P, Nesvorna M, Hubert J (2021) Label-free proteomic analysis reveals differentially expressed Wolbachia proteins in Tyrophagus putrescentiae: Mite allergens and markers reflecting population-related proteome differences. J Proteom 249:104356. https://doi.org/10.1016/j.jprot.2021.104356
Erban T, Klimov P, Molva V, Hubert J (2020a) Whole genomic sequencing and sex-dependent abundance estimation of Cardinium sp., a common and hyperabundant bacterial endosymbiont of the American house dust mite, Dermatophagoides farinae. Exp Appl Acarol 80:363–380. https://doi.org/10.1007/s10493-020-00475-5
Erban T, Klimov P, Talacko P, Harant K, Hubert J (2020b) Proteogenomics of the house dust mite, Dermatophagoides farinae: Allergen repertoire, accurate allergen identification, isoforms, and sex-biased proteome differences. J Proteom 210:103535. https://doi.org/10.1016/j.jprot.2019.103535
Gao Z, Tseng C-h, Pei Z, Blaser MJ (2007) Molecular analysis of human forearm superficial skin bacterial biota. Proc. Natl. Acad. Sci. USA 104:2927–2932. https://doi.org/10.1073/pnas.0607077104
Hawksworth DL, Crous PW, Redhead SA, Reynolds DR, Samson RA, Seifert KA et al (2011) The Amsterdam Declaration on fungal nomenclature. IMA Fungus 2:105–112. https://doi.org/10.5598/imafungus.2011.02.01.14
Hay DB, Hart BJ, Pearce RB, Kozakiewicz Z, Douglas AE (1992) How relevant are house dust mite-fungal interactions in laboratory culture to the natural dust system? Exp Appl Acarol 16:37–47. https://doi.org/10.1007/bf01201491
Hay DB, Hart BJ, Douglas AE (1993) Effects of the fungus Aspergillus penicillioides on the house dust mite Dermatophagoides pteronyssinus: an experimental re-evaluation. Med Vet Entomol 7:271–274. https://doi.org/10.1111/j.1365-2915.1993.tb00687.x
Herlemann DPR, Labrenz M, Jns K, Bertilsson S, Waniek JJ, Andersson AF (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5:1571–1579. https://doi.org/10.1038/ismej.2011.41
Horak B, Dutkiewicz J, Solarz K (1996) Microflora and acarofauna of bed dust from homes in Upper Silesia, Poland. Ann Allergy Asthma Immunol 76:41–50. https://doi.org/10.1016/S1081-1206(10)63405-9
Hubert J, Kopeck Perotti MA, Nesvorna, Braig HR, Ságová-Marecková et al (2012) Detection and identification of species-specific bacteria associated with synanthropic mites. Microb Ecol 63:919–928. https://doi.org/10.1007/s00248-011-9969-6
Hubert J, Nesvorna M, Kopecky J, Erban T, Klimov P (2019) Population and culture age influence the microbiome profiles of house dust mites. Microb Ecol 77:1048–1066. https://doi.org/10.1007/s00248-018-1294-x
Hubert J, Nesvorna M, Klimov PB, Erban T, Sopko B, Dowd SE et al (2021) Interactions of the intracellular bacterium Cardinium with its host, the house dust mite Dermatophagoides farinae, based on gene expression data. mSystems 6:e00916-21. https://doi.org/10.1128/mSystems.00916-21
Ihrmark K, Ber ITM, Cruz-Martinez K, Friberg H, Kubartova A, Schenck J et al (2012) New primers to amplify the fungal ITS2 region – evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol Ecol 82:666–677. https://doi.org/10.1111/j.1574-6941.2012.01437.x
Jeon Y-S, Chun J, Kim B-S (2013) Identification of household bacterial community and analysis of species shared with human microbiome. Curr Microbiol 67:557–563. https://doi.org/10.1007/s00284-013-0401-y
Jeong KY, Park J-W, Hong C-S (2012) House dust mite allergy in Korea: The most important inhalant allergen in current and future. Allergy Asthma Immunol Res 4:313–325. https://doi.org/10.4168/aair.2012.4.6.313
Kanchongkittiphon W, Mendell MJ, Gaffin JM, Wang G, Phipatanakul W (2015) Indoor environmental exposures and exacerbation of asthma: an update to the 2000 review by the Institute of Medicine. Environ Health Perspect 123:6–20. https://doi.org/10.1289/ehp.1307922
Kang B, Vellody D, Homburger H, Yunginger JW (1979) Cockroach cause of allergic asthma: Its specificity and immunologic profile. J Allergy Clin Immunol 63:80–86. https://doi.org/10.1016/0091-6749(79)90196-9
Kim JY, Yi M-H, Hwang Y, Lee JY, Lee I-Y, Yong D et al (2018) 16S rRNA profiling of the Dermatophagoides farinae core microbiome: Enterococcus and Bartonella. Clin Exp Allergy 48:607–610. https://doi.org/10.1111/cea.13104
Kõljalg U, Nilsson HR, Schigel D, Tedersoo L, Larsson K-H, May TW et al (2020) The taxon hypothesis paradigm—On the unambiguous detection and communication of taxa. Microorganisms 8:1910. https://doi.org/10.3390/microorganisms8121910
Klimov P, Molva V, Nesvorna M, Pekar S, Shcherbachenko E, Erban T et al (2019) Dynamics of the microbial community during growth of the house dust mite Dermatophagoides farinae in culture. FEMS Microbiol Ecol 95:fiz153. https://doi.org/10.1093/femsec/fiz153
Kurtz ZD, Mr CL, Miraldi ER, Littman DR, Blaser MJ, Bonneau RA (2015) Sparse and compositionally robust inference of microbial ecological networks. PLoS Comput Biol 11:e1004226. https://doi.org/10.1371/journal.pcbi.1004226
Li AH, Yuan FX, Groenewald M, Bensch K, Yurkov AM, Li K et al (2020) Diversity and phylogeny of basidiomycetous yeasts from plant leaves and soil: Proposal of two new orders, three new families, eight new genera and one hundred and seven new species. Stud Mycol 96:17–140. https://doi.org/10.1016/j.simyco.2020.01.002
Li Z, Zheng N, An Q, Li X, Sun S, Zhang W et al (2022) Impact of environmental factors and bacterial interactions on dust mite allergens in different indoor dust. Sci Total Environ 844:157177. https://doi.org/10.1016/j.scitotenv.2022.157177
Loo EXL, Chew LJM, Zulkifli AB, Ta LDH, Kuo IC, Goh A et al (2018) Comparison of microbiota and allergen profile in house dust from homes of allergic and non-allergic subjects- results from the GUSTO study. World Allergy Organ J 11:37. https://doi.org/10.1186/s40413-018-0212-5
Lustgraaf Bvd (1978a) Ecological relationships between xerophilic fungi and house-dust mites (Acarida: Pyroglyphidae). Oecologia 33:351–359. https://doi.org/10.1007/bf00348118
Lustgraaf Bvd (1978b) Seasonal abundance of xerophilic fungi and house-dust mites (Acarida: Pyroglyphidae) in mattress dust. Oecologia 36:81–91. https://doi.org/10.1007/bf00344573
Madden AA, Barberan A, Bertone MA, Menninger HL, Dunn RR, Fierer N (2016) The diversity of arthropods in homes across the United States as determined by environmental DNA analyses. Mol Ecol 25:6214–6224. https://doi.org/10.1111/mec.13900
McMurdie PJ, Holmes S (2013) phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8:e61217. https://doi.org/10.1371/journal.pone.0061217
Molva V, Nesvorna M, Hubert J (2019) Feeding interactions between microorganisms and the house dust mites Dermatophagoides pteronyssinus and Dermatophagoides farinae (Astigmata: Pyroglyphidae). J. Med. Entomol. 56:1669–1677. https://doi.org/10.1093/jme/tjz089
Morey PR (2011) Remediation and control of microbial growth in problem buildings. In: Flannigan B, Samson RA, Miller JD (eds) Microorganisms in home and indoor work environments: Diversity, health impacts, investigation and control, 2 edn. CRC Press, Boca Raton, FL, USA, pp 125–144
Nesvorna M, Pekar S, Shcherbachenko E, Molva V, Erban T, Green SJ et al (2021) Microbiome variation during culture growth of the European house dust mite, Dermatophagoides pteronyssinus. FEMS Microbiol Ecol 97:fiab039. https://doi.org/10.1093/femsec/fiab039
Nilsson RH, Larsson K-H, Taylor AFS, Bengtsson-Palme J, Jeppesen TS, Schigel D et al (2018) The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res 47:D259–D264. https://doi.org/10.1093/nar/gky1022
Oh H, Ishii A, Tongu Y, Itano K (1986) Microorganisms associated with the house-dust mite, Dermatophagoides. Jpn J Sanit Zool 37:229–235. https://doi.org/10.7601/mez.37.229
Oksanen FJ, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D et al (2017) vegan: Community Ecology Package. R package version 2.4-4
Park SC, Hwang CS, Chung HJ, Purev M, Al Sharhan SS, Cho H-J et al (2019) Geographic and demographic variations of inhalant allergen sensitization in Koreans and non-Koreans. Allergol Int 68:68–76. https://doi.org/10.1016/j.alit.2018.07.005
Portnoy J, Miller JD, Williams PB, Chew GL, Miller JD, Zaitoun F et al (2013) Environmental assessment and exposure control of dust mites: a practice parameter. Ann Allergy Asthma Immunol 111:465–507. https://doi.org/10.1016/j.anai.2013.09.018
Ree HI, Jeon SH, Lee IY, Hong CS, Lee DK (1997) Fauna and geographical distribution of house dust mites in Korea. Korean J Parasitol 35:9–17. https://doi.org/10.3347/kjp.1997.35.1.9
Rijckaert G, van Bronswijk JEMH, Linskens HF (1981) House-dust community (Fungi, mites) in different climatic regions. Oecologia 48:183–185. https://doi.org/10.1007/bf00347961
Rosa-Magri MM, Tauk-Tornisielo SM, Ceccato-Antonini SR (2011) Bioprospection of yeasts as biocontrol agents against phytopathogenic molds. Brazil Arch Biol Technol 54:1–5. https://doi.org/10.1590/S1516-89132011000100001
Ross AA, Doxey AC, Neufeld JD, Sharpton T (2017) The skin microbiome of cohabiting couples. mSystems 2:e00043-17. https://doi.org/10.1128/mSystems.00043-17
Schlatter DC, Yin C, Hulbert S, Paulitz TC, Stabb EV (2020) Core rhizosphere microbiomes of dryland wheat are influenced by location and land use history. Appl Environ Microbiol 86:e02135-19. https://doi.org/10.1128/AEM.02135-19
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al (2009) Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Shan Y, Wu W, Fan W, Haahtela T, Zhang G (2019) House dust microbiome and human health risks. Int Microbiol 22:297–304. https://doi.org/10.1007/s10123-019-00057-5
Shinohara N, Woo C, Yamamoto N, Hashimoto K, Yoshida-Ohuchi H, Kawakami Y (2021) Comparison of DNA sequencing and morphological identification techniques to characterize environmental fungal communities. Sci Rep 11:2633. https://doi.org/10.1038/s41598-021-81996-w
Sinha RN, van Bronswijk JE, Wallace HA (1970) House dust allergy, mites and their fungal associations. Can Med Assoc J 103:300–301
Thomas WR (2010) Geography of house dust mite allergens. Asian Pac J Allergy Immunol 28:211–224
Thompson JR, Argyraki A, Bashton M, Bramwell L, Crown M, Hursthouse AS et al (2021) Bacterial diversity in house dust: Characterization of a core indoor microbiome. Front Environ Sci 9:754657. https://doi.org/10.3389/fenvs.2021.754657
Tipton L, Mr CL, Kurtz ZD, Huang L, Kleerup E, Morris A et al (2018) Fungi stabilize connectivity in the lung and skin microbial ecosystems. Microbiome 6:12. https://doi.org/10.1186/s40168-017-0393-0
Valerio CR, Murray P, Arlian LG, Slater JE (2005) Bacterial 16S ribosomal DNA in house dust mite cultures. J Allergy Clin Immunol 116:1296–1300. https://doi.org/10.1016/j.jaci.2005.09.046
Van Asselt L (1999) Review: Interactions between domestic mites and fungi. Indoor Built Environ 8:216–220. https://doi.org/10.1177/1420326X9900800402
van Bronswijk JEMH (1981) House dust biology for allergists, acarologists and mycologists. NIB Publishers, Zeist, The Netherlands. https://doi.org/10.1093/ee/2.1.142
van Bronswijk JEMH, Sinha RN (1973) Role of fungi in the survival of Dermatophagoides (Acarina: Pyroglyphidae) in house-dust environment. Environ Entomol 2:142–145. https://doi.org/10.1093/ee/2.1.142
Wang QM, Yurkov AM, G M, Lumbsch HT, Leavitt SD, Groenewald M et al (2015) Phylogenetic classification of yeasts and related taxa within Pucciniomycotina. Stud Mycol 81:149–189. https://doi.org/10.1016/j.simyco.2015.12.002
Wang R, Huang C, Zhang Y, Li R (2021) Invasive dermatophyte infection: A systematic review. Mycoses 64:340–348. https://doi.org/10.1111/myc.13212
Weiler F, Schmitt MJ (2003) Zygocin, a secreted antifungal toxin of the yeast Zygosaccharomyces bailii, and its effect on sensitive fungal cells. FEMS Yeast Res 3:69–76. https://doi.org/10.1111/j.1567-1364.2003.tb00140.x
Weyand NJ (2017) Neisseria models of infection and persistence in the upper respiratory tract. Pathogens and Disease 75:ftx031. https://doi.org/10.1093/femspd/ftx031
White T, Bruns T, Lee S, Taylor J, Innis M, Gelfand D et al (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols. Academic Press, New York, pp 315–322
Woo C, Yamamoto N (2020) Falling bacterial communities from the atmosphere. Environ Microbiome 15:22. https://doi.org/10.1186/s40793-020-00369-4
Woudenberg JHC, Sandoval-Denis M, Houbraken J, Seifert KA, Samson RA (2017) Cephalotrichum and related synnematous fungi with notes on species from the built environment. Stud Mycol 88:137–159. https://doi.org/10.1016/j.simyco.2017.09.001
Yurkov AM, Kachalkin AV, Daniel HM, Groenewald M, Libkind D, de Garcia V et al (2015) Two yeast species Cystobasidium psychroaquaticum f.a sp. nov. and Cystobasidium rietchieii f.a sp. nov. isolated from natural environments, and the transfer of Rhodotorula minuta clade members to the genus Cystobasidium. Antonie van Leeuwenhoek 107:173–185. https://doi.org/10.1007/s10482-014-0315-0
Acknowledgements
We would like to thank the residents of the homes who kindly permitted us to collect house dust samples.
Funding
This project was funded by Dyson Korea.
Author information
Authors and Affiliations
Contributions
Conceptualization, KL and NY; Data curation, CW; Formal analysis, CW, PK, SKL, and NY; Funding acquisition, KL and NY; Investigation, CW, MIUB, and DK; Project administration, JYP; Resources, KD, KL, and NY; Supervision, JYP, KL, and NY; Validation, NY; Visualization, CW and NY; Writing – original draft, CW, PK, and SKL, and NY; Writing – review and editing, CW, KD, and NY. All the authors have read and approved the contents of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the funder (Dyson Korea) has played no role in designing and performing the research.
Ethical approval
This study was approved by the Institutional Review Board of Seoul National University with the approval number 2103/004 - 001.
Informed consent
Written informed consent was obtained from all participants who provided house dust samples in this study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Woo, C., Bhuiyan, M.I.U., Kim, D. et al. DNA metabarcoding-based study on bacteria and fungi associated with house dust mites (Dermatophagoides spp.) in settled house dust. Exp Appl Acarol 88, 329–347 (2022). https://doi.org/10.1007/s10493-022-00755-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10493-022-00755-2