Abstract
In the bottom sediments from a number of the Barents Sea sites, including coastal areas of the Novaya Zemlya, Franz Josef Land, and Svalbard archipelagos, sulphate reduction rates were measured and the phylogenetic composition of sulphate-reducing bacterial (SRB) communities was analysed for the first time. Molecular genetic analysis of the sequences of the 16S rRNA and dsrB genes (the latter encodes the β-subunit of dissimilatory (bi)sulphite reductase) revealed significant differences in the composition of bacterial communities in different sampling stations and sediment horizons of the Barents Sea depending on the physicochemical conditions. The major bacteria involved in reduction of sulphur compounds in Arctic marine bottom sediments belonged to Desulfobulbaceae, Desulfobacteraceae, Desulfovibrionaceae, Desulfuromonadaceae, and Desulfarculaceae families, as well as to uncultured clades SAR324 and Sva0485. Desulfobulbaceae and Desulfuromonadaceae predominated in the oxidised (Eh = 154–226 mV) upper layers of the sediments (up to 9% and 5.9% from all reads of the 16S rRNA gene sequences in the sample, correspondingly), while in deeper, more reduced layers (Eh = −210 to −105 mV) the share of Desulfobacteraceae in the SRB community was also significant (up to 5%). The highest relative abundance of members of Desulfarculaceae family (3.1%) was revealed in reduced layers of sandy-clayey sediments from the Barents Sea area affected by currents of transformed (mixed, with changed physicochemical characteristics) Atlantic waters.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Barents Sea is an open marine system, where large amounts of dispersed organic matter (OM) from the northern Atlantic and Arctic oceans interchange, mainly precipitating on the sea bottom (Stiansen et al. 2009; Lind et al. 2018). Transversal structure of the Barents Sea summer waters along the Kola meridian demonstrates that warm Atlantic waters almost completely fill the southern part of the sea, forming the central and northern branches of the Nordkapp Current (Fig. 1). Cold waters inflowing from the Arctic Ocean via the Franz-Victoria and St. Anna trenches prevail in the northern part of the Barents Sea; thus, deep-water trenches are the key areas for water exchange between the Arctic basin and the Barents Sea (Mityaev et al. 2018). The estimated averaged gross production for most of the Barents Sea area including coastal waters is 82–174 g C m−2 per year (Makarevich 2012), with the largest contribution of the Atlantic area (Dalpadado et al. 2014). An increased organic matter concentration in the near-bottom water layers can be explained by deep currents, gravitational processes and bottom topography, providing the presence of a nepheloid layer in the Barents Sea almost everywhere (Politova et al. 2019).
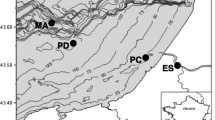
Sampling stations (67th cruise of the RV Akademik Mstislav Keldysh, August-October 2016) and water circulation in the Barents Sea: Atlantic waters—red (grey for black and white version) lines, transformed waters of Atlantic origin—black dotted lines, Barents Sea waters—black dashed lines, Arctic waters—blue (black thick for black and white version) lines, Norwegian and Murmansk coastal currents—green (black thin for black and white version) lines (Stiansen et al. 2009)
Organic material resulting from primary production precipitates to the sea bottom, where microorganisms of the bottom sediments mineralised most of it (Wollast 1991). Significant variation in the redox potential of these sediments, from oxidised surface horizons to strongly reduced deeper ones, provides diverse ecological niches for a broad range of microorganisms. Investigation of drill samples from the Arctic bottom sediments revealed bacterial 16S rRNA genes in all horizons, while genetic material of the Archaea domain members were found only in strongly reduced sulphide-enriched horizons; microbial communities of each horizon were phylogenetically diverse and interacted closely with communities of the neighbouring horizons with noticeably different physicochemical characteristics (Forschner et al. 2009).
The products of fermentative metabolism act as electron donors for bacteria responsible for the terminal phase of OM oxidation in Svalbard coastal sediments (Finke and Jørgensen 2008; Robador et al. 2009; Glombitza et al. 2015). Sulphates and metal oxides are the most important electron acceptors in the anoxic horizons of marine coastal bottom sediments (Thamdrup 2000). In the upper horizons (0–2 cm) of Arctic sediments, sulphate reduction and iron reduction are responsible for mineralization of 75 and 25%, respectively, while in deeper horizons (5–9 cm) sulphate reduction is the almost exclusively process of mineralization (Jensen et al. 2003; Vandieken et al. 2006a; Finke et al. 2007). Most measurements of sulphate reduction rate (SRR) in the Barents Sea were carried out for the samples from the bottom sediments of Svalbard fjords in the course of investigation of the OM mineralization pathways and of the temperature effect on sulphate reduction (Arnosti et al. 1998; Nickel et al. 2008; Sawicka et al. 2012; Robador et al. 2015).
Sulphate-reducing bacteria (SRB), which help to mineralize over 50% of organic carbon resulting from pelagic primary production, play the key role in the carbon and sulphur global biogeochemical cycles in the ocean (Jørgensen 1982; Müller et al. 2018). SRB are a phylogenetically heterogeneous group (Muyzer and Stams 2008) comprising the microorganisms capable of obtaining energy by anaerobic oxidation of molecular hydrogen or low molecular weight organic compounds (which are often fermentation products), using sulphate as a final electron acceptor. Sulphate-reducing bacteria may be subdivided into two separate groups according to the utilization of organic compounds. Lactate, formate, and propionate are the typical substrates for oxidation to acetate by SRB with an incomplete tricarboxylic acid cycle; in most ecosystems, including Arctic marine sediments, it is mainly carried out by members of the genera Desulfovibrio and Desulfobulbus (Glombitza et al. 2015). Most members of the genera Desulfobacter, Desulfobacterium, Desulfococcus, Desulfosarcina can oxidise organic substrates completely (Knoblauch et al. 1999a; Glombitza et al. 2015).
One of the first molecular studies on the composition of microbial communities in Svalbard coastal bottom sediments revealed the presence of ~ 30% of potentially psychrophilic microorganisms, most of which were sulphate-reducing bacteria (Sahm et al. 1999). Several psychrophilic SRB capable of growth even at subzero temperatures have been isolated from Svalbard coastal bottom sediments, including Desulfofrigus oceanense, Desulfofaba gelida, and Desulfotalea psychrophila (Knoblauch et al. 1999b), Desulfotomaculum arcticum (Vandieken et al. 2006c) and Desulfoconvexum algidum (Könneke et al. 2013). Psychrophilic SRB (Desulfovibrio arcticus) were also found in the Arctic cryopegs, saline water-saturated horizons located at various depths in permafrost soils (Pecheritsyna et al. 2012). The results of several studies (Jørgensen et al. 1990; Thamdrup et al. 1994; Sagemann et al. 1998; Pimenov et al. 2000) showed the rate of sulphate reduction in northern seas’ bottom sediments to be comparable with the rates of this process in moderate latitudes. Temperature of the Arctic bottom sediments affects the competition for substrates between SRB phylogenetic subgroups, some of which contain psychrophilic species and have very high rates of metabolism at low temperatures (Knoblauch et al. 1999a).
The data on sulphate reduction rates in the Barents Sea bottom sediments are presently scarce (Arnosti et al. 1998; Savvichev et al. 2009; Robador et al. 2015), while there is almost no information on the phylogenetic composition of sulphate-reducing microbial communities from Arctic seas. Thus, the aim of our study was to carry out in-depth investigation of the phylogenetic diversity in sulphate-reducing bacterial communities from the Barents Sea bottom sediments collected at hydrochemically different sediment horizons and sampling stations, including those at the coastal areas of the Novaya Zemlya, Franz Josef Land, and Svalbard archipelagos.
Materials and methods
Sampling of the bottom sediments
Samples of the Barents Sea bottom sediments and near-bottom water were collected during the 67th cruise of RV Akademik Mstislav Keldysh (August–October 2016) using a Mini Muc K/MT 410 multi-corer (KUM, Germany) and a large-diameter geological tube (Shirshov Institute of Oceanology, Russia). Samplings were carried out at the sites of hydrocarbon deposits in the Pechora Sea (the south-eastern part of the Barents Sea) near Vaygach Island (station 5407); at the Shtokman gas condensate deposit (stations 5412 and 5419); in the North Barents Sea Deep (station 5421); at the Novaya Zemlya coasts in the Russkaya Gavan' Bay (station 5424) and in the Western Novaya Zemlya Trench (station 5415); at the Franz Josef Land coasts in the Cambridge Strait (strait entrance at station 5453, deep-water area at station 5454, and in the Dezhnev Bay at station 5456); at the Svalbard coast in Storfjorden (station 5441); and in the Medvezhinskii Trench (station 5431)—see Fig. 1 and Table 1 (Politova et al. 2018, 2019).
Determination of organic carbon and rates of sulphate reduction
Initial description of the sedimentary cores (0–26-cm layer) and samplings for determination of SO42− and organic carbon (Corg) contents, as well as measurements of sulphate reduction rates and redox potential (Eh), were carried out in the laboratory on the ship’s board. After sampling, the sediments were frozen inboard at −18 °C. Prior to analysis in stationary laboratories, the samples were defrosted and dried at 50 °C. Corg contents in the bottom sediments were determined by dry incineration using AN-7560 carbon analyser (GZIP, Belarus) with the accuracy 3–6 rel.% in the Shirshov Institute of Oceanology, Russia.
Sulphate reduction rates (SRR) were determined by radioisotope analysis using 35S-sulphate. Immediately after hauling on board, the sample (3 cm3) of a bottom sediment from a corresponding horizon was collected into a cut-off 5-ml plastic syringe and sealed with a gas-tight butyl rubber stopper. After injecting 0.2 ml of 35S-SO42− solution (10 µCi), the samples were incubated at 1–3 °C for 1–2 days. After incubation, the samples were fixed with 1 ml of 2 M KOH before transportation to the stationary laboratory. The samples were then treated as described previously (Pimenov and Bonch-Osmolovskaya 2006). Pore waters were separated by centrifugation for 10 min at 8000 g in a TsUM-1 centrifuge (Russia). Sulphate concentrations in pore water samples were measured using a Stayer ion chromatograph (Russia).
The sediment samples with the highest sulphate reduction rates were used for molecular studies of phylogenetic composition of microbial communities and for isolation of SRB enrichment cultures.
All SRR and organic carbon measurements were carried out in three replicates.
SRB enrichment cultures
Initial enrichment cultures were obtained by injecting 1 cm3 of the corresponding bottom sediment sample into 18-ml Hungate test tube with 9 ml of anoxic liquid Widdel lactate/sulphate medium for marine SRB (Widdel and Bak 1992). The samples used for SRB enrichments originated from four stations located at the south-east (station 5407, 0–1.5 cm) and north of the Novaya Zemlya (station 5424, 5–7 cm), also at the west of the Franz Josef Land (stations 5454, 0–3 cm and 5456, 6–10 cm). Enrichments were maintained by inoculations using sterile syringes; the inoculum volume was 10%. The enrichment cultures were incubated for 5–25 days at 20 °C in the dark.
Growth of the enrichments was monitored by light microscopy and by colorimetric sulphide determination with N,N-dimethyl-p-phenylenediamine (Trüper and Schlegel 1964).
Isolation of total DNA
Total DNA was isolated from 0.25 g of corresponding bottom sediment samples using the MO BIO’s PowerSoil DNA Isolation Kit (Qiagen, Netherlands), according to the manufacturer's protocol. The isolation of total DNA from the SRB enrichment cultures was carried out using the Genomic DNA Purification Kit (Thermo Fischer Scientific, USA). The concentration and purity of DNA samples were estimated spectrophotometrically at λ 260 and 280 nm on NanoDrop 2000C (Thermo Fisher Scientific, USA).
PCR analysis of the dsrB and 16S rRNA genes
The DSRp2060F and DSR4R oligonucleotide PCR primers (Geets et al. 2006) specific to the dsrB gene encoding the β-subunit of dissimilatory (bi)sulphite reductase, an essential enzyme for sulphate reduction, were used for SRB detection in the Barents Sea bottom sediments and to obtain the amplicons for the subsequent DGGE analysis. Preliminary assessment of SRB phylogenetic diversity was carried out by PCR with oligonucleotide primers specific to the 16S rRNA genes of six major SRB subgroups—DFM140/DFM842, DBB121/DBB1237, DBM169/DBM1006, DSB127/DSB1273, DCC305/DCC1165, DSV230/DSV838 (Daly et al. 2000; Korneeva et al. 2015).
PCR was carried out using a GeneAmp PCR System 9700 amplifier (Applied Biosystems, USA). The reaction mixture (25 µl) contained the following: ~ 25 ng template DNA; 2.0 mM MgCl2; 400 µM dNTP (Thermo Fischer Scientific, USA); 500 nM of each primer (Syntol, Russia); and 2.5 U Taq DNA polymerase (Syntol, Russia). Amplification was carried out as follows: 95 °C for 5 min; 35 cycles of 94 °C for 1 min, corresponding Ta for 1 min, 72 °C for 1 min; and 72 °C for 10 min.
Nested PCR was used for detection of minor SRB members in the obtained enrichment cultures. This method provides for higher sensitivity and amplification specificity. Nested PCR was carried out in two steps. First, fragments of bacterial 16S rRNA genes were amplified using total DNA from enrichments as a template, and pA + pH’ oligonucleotide primers (Edwards et al. 1989; Dar et al. 2005) under the following conditions: 95 °C for 5 min; 35 cycles of 94 °C for 1 min, 37 °C for 1 min, 72 °C for 6 min; and 72 °C for 10 min. Then, fragments of the 16S rRNA genes of six major SRB subgroups were amplified with corresponding primers using the product of the first step as a template. Amplification products were analysed using 110 V electrophoresis in 1% agarose gel and 1 × TAE buffer.
DGGE and sequencing of the dsrB gene fragments
Products of amplification (with the use of total DNA from SRB enrichments as template) of the dsrB gene fragments were separated by denaturing gradient gel electrophoresis (DGGE). To improve DGGE separation of the DNA fragments, the sequence of the forward primer DSRp2060F contained a GC-rich fragment of 40 bp at the 5′-end (Muyzer et al. 1993). Amplification products were applied to 7% (v/v) polyacrylamide gel (acrylamide–N,N’-methylenebisacrylamide, 37.5:1) with acrylamide concentration gradient of 35–65% in 0.5× TAE buffer (pH 7.4). Denaturing agents used for the preparation of a 100% solution were 7 M urea and 40% deionised formamide (Merck, Germany). Ammonium persulphate (50 μl of 10% APS per 10 ml solution) and TEMED (10 μl per 10 ml solution) were used as an initiator and a catalyst for gel polymerization, respectively (Green et al. 2010).
DGGE was carried out for 8 h at 200 V and 60 °C in a DCode Universal Mutation Detection System (Bio-Rad, USA). After electrophoresis, the gel was washed with distilled water and stained with SYBR®Gold (Thermo Fisher Scientific, USA) for 40 min in the dark. The stained gel was visualised on a transilluminator; the separate bands were excised, transferred into test tubes with 20 µl of sterile distilled water, and incubated for 16 h at 4 °C for elution of the DNA fragments from the polyacrylamide gel.
PCR was carried out using the DNA fragments eluted from the gel as templates and the primer pair DSRp2060F (5′-CAACATCGTTCATACCCAGGG-3′) and DSR4R (5′-GTGTAGCAGTTACCGCA-3′) in order to reamplify the dsrB fragments, which were then purified from 1% agarose gel using DNA Gel Extraction Kit (Thermo Fisher Scientific, USA). Nucleotide sequences of the dsrB gene fragments were determined by Sanger sequencing with the BigDye Terminator v. 3.1 Cycle Sequencing Kit in an ABI PRISM 3730 automatic DNA analyser (Thermo Fisher Scientific, USA).
All PCR and DGGE experiments were carried out in three replicates.
High-throughput sequencing of the 16S rRNA gene fragments
PCR amplification of the 16S rRNA gene fragments containing the V3–V4 variable regions was carried out using the universal, covering Bacteria and Archaea, primers PRK341F (5′-CCTACGGGRBGCASCAG-3′) and PRK806R (5′-GGACTACYVGGGTATCTAAT-3′) (Takai, Horikoshi 2000; Yu et al. 2005). The following program was used: 96 °C for 2 min, followed by 30 cycles of 96 °C for 30 s, 53 °C for 30 s, and 72 °C for 60 s, and a final elongation at 72 °C for 10 min. PCR fragments were then sequenced on a GS FLX genome analyser (Roche, Switzerland) according to the Titanium protocol using the GS FLX Titanium Sequencing Kit XL+ . Creation of the library, its amplification, and sequencing were carried out according to the relevant Roche protocols.
Phylogenetic analysis
Similarity between the translated amino acid sequences encoded by the dsrB gene and the corresponding GenBank sequences (about 125 amino acids each) was determined after aligning by the MUSCLE (Edgar 2004), using also the ExPASy and the BLASTX software. Phylogenetic tree on the dsrB gene was constructed with the Maximum Likelihood method implemented in the MEGA-X software (Kumar et al. 2016), using homologically close reference sequences from the GenBank.
Reads of the 16S rRNA gene fragments starting with the forward primer were selected and trimmed to the same length of 250 bp, using Mothur v.1.35.1 (Schloss et al. 2009). All the subsequent operational taxonomic unit (OTU) analysis was done with Usearch v.11 (Edgar 2010). Low-quality reads were filtered (fastq_maxee = 1.00) and high-quality reads were clustered into OTUs at 97% identity level. At the clustering stage, chimera and singleton sequences were removed by the Usearch algorithm. Then all reads, including low-quality ones and singletons, were mapped to OTU representative sequences at 97% global identity level to determine OTU relative abundance for each sample. OTU taxonomic identification was performed using the SINA online alignment and classification platform, and the Silva v.1.2.11 database with default parameters (Pruesse et al. 2012; Quast et al. 2013), and searching for close sequences in GenBank using the BLAST software. When a sequence with more than 95% similarity with the 16S rRNA gene of the described microorganism was detected, OTU was assigned to the corresponding genus.
Results and discussion
Physicochemical analysis of the Barents Sea bottom sediments
The Holocene bottom sediments were represented by grey aleuro-pelitic clayey hydrotroilite silts with inclusions of detritus and a characteristic upper (0–3 cm) reddish oxidised layer. Small (up to 1 cm) Fe–Mn crusts occurred at stations 5412, 5415, 5421, 5431, 5453, and 5454. Sulphate concentration in the bottom sediments was within the range 20.8–29.7 μM (Table 1). In the Pechora Sea (the south-eastern part of the Barents Sea), the sediments were coarse-grained, represented by sand-aleurite silts (station 5407) with the lowest Corg content (0.2%). Average Corg content in the upper layers of the Barents Sea bottom sediments was 1.4% (n = 10), compared to the average of 1.1% (n = 90) for the whole sea area (Nemirovskaya 2020), while the highest values defined in our work (2.6%) were typical of predominantly pelitic silts of the deep-water part of the North Barents Sea Trench (Table 1). Reduced conditions with active sulphate reduction and degradation of plankton biomass precipitating from the water column usually result in Corg content decreasing from the surface horizon (0–3 cm) of sediments to deeper ones (4–15 cm) (Nemirovskaya 2020).
Station 5407 (Pechora Sea) is under predominant influence of the Coastal Current (seasonal fluctuation: S = 30–34.5‰; T = −1.8–8 °C) (Ingvaldsen and Loeng 2009), which transfers desalinated waters from the North and White seas and is located in the area of hydrocarbon deposits, gas seeps, and relics of permafrost sediments. In spite of predominant coarse-grained bottom sediments with the lowest Corg, reducing conditions developed in these sediments from the horizon of 1.5 cm (Fig. 2).
Redox potential (a), sulphate concentration (b) and sulphate reduction rate (c) in the bottom sediments at sampling stations in the Barents Sea: – (large black triangle) − station 5407, − (black square) − station 5412, − (small black triangle) − station 5424, − × − station 5441, − ж − station 5454, − (black circle) − station 5456
The Barents-Kara oil-and-gas province is represented by sand-clay Mesozoic (Triassic-Jurassic-Cretaceous) and Cenozoic sediments (Margulis 2008). These areas (stations 5412 and 5419) are affected by the waters of the Barents Sea and transformed Atlantic waters (seasonal fluctuation: S = 34.5–35; T = −1.5–5 °C) (Ingvaldsen and Loeng 2009). In such surface sediment horizons with relatively low Corg concentrations (1.5%), the composition of the hydrocarbon molecular markers indicates their enrichment with components of thermally mature endogenic organic matter.
High production of the Barents Sea is caused by mixing of the relatively warm Atlantic waters with the cold Arctic ones, which results in formation of the Polar front at the border of the Barents Sea waters (seasonal fluctuation: S = 34.5–35‰; T = −1.5–5 °C) with Arctic waters (seasonal fluctuation: S = 32–34.8‰; T = < 0 °C) (Ingvaldsen and Loeng 2009). In this region, on the Perseus Plateau station 5421 is located, which is characterised by high rate of sediment accumulation and the highest Corg value for the surface bottom sediments (2.6%).
Station 5424 is located in the area of the Russkaya Gavan' Bay dominated by discharge of a freshwater terrigenic flow from the Shokal'skogo Glacier, one of the outlets of the ice sheet on the Severny Island of the Novaya Zemlya archipelago. Content of organic matter in the bottom sediments at this station was low, just 0.31%.
Station 5415 is located in the Western Novaya Zemlya Trench, where the coastal Novaya Zemlya Current flows (seasonal fluctuation: S = 33–34.7‰; T = −1.8–6 °C), which is significantly affected by transformed highly productive Atlantic waters (Barents Sea waters: seasonal fluctuation S = 34.5–35‰; T = −1.5–5 °C) and the drift of Novaya Zemlya glaciers (Ingvaldsen and Loeng 2009). Organic carbon content in the bottom sediments here was above the average values for the Barents Sea (Corg = 2.2%).
Station 5431 is located in the Medvezhinskii Trench at the border with Atlantic waters (seasonal fluctuation S = > 35‰; T = > 3 °C) and transformed Arctic waters (seasonal fluctuation S = < 34.4‰; T = 1–3 °C) (Ingvaldsen and Loeng 2009). The content of organic carbon in the sediments here (Corg = 1.4%) corresponds to the average values for the Barents Sea (Nemirovskaya 2020).
Station 5441 is located in Storfjorden at the western part of the Barents Sea continental outskirts, at the boundary between oceanic and continental crust (Gabrielsen et al. 1990). This area is characterised by wide occurrence of cold Arctic methane seeps of endogenous origin and is affected by the flow off Svalbard glaciers (Åström et al. 2016). Organic carbon content in the bottom sediments (Corg = 1.6%) was slightly higher than the average values for the Barents Sea (Nemirovskaya 2020).
The Cambridge Strait is located between the Alexandra Land and Prince George Land islands (Franz Josef Land archipelago), where ice sheets are well developed. Arctic waters dominate in this area (seasonal fluctuation: S = 32–34.8‰; T = < 0 °C), although Atlantic modified waters (seasonal fluctuation S = 34.8–34.9‰; 0 °C < T < 1,5 °C) penetrates into the Barents Sea via the Franz Josef Trench (along the western entrance to the Cambridge Strait with station 5453), arriving into the Arctic basin via the Fram Strait (Ingvaldsen and Loeng 2009). Station 5456 (Corg = 1.5%) is located at the fracturing zone of the Nagursky Fault near Dezhnev Cape, while station 5454 (Corg = 1.4%) is located in the deep-water part of the Cambridge Strait.
Sulphate reduction in the Barents Sea bottom sediments
The lowest rates of sulphate reduction were revealed at station 5431 in the Medvezhinskii Trench bottom sediments (0.6–1.5 nmol S cm−3 day−1), at Svalbard coast at station 5441 (0.7–1.3 nmol S cm−3 day−1), and especially at the entrance to the Cambridge Strait at station 5453 (0.2–0.3 nmol S cm−3 day−1). In the bottom sediments of the central Barents Sea area (stations 5412, 5419, and 5421) sulphate reduction rates (SRR) varied from 2.1 to 6.5 nmol S cm−3 day−1, with higher values in the Shtokman deposit area at station 5412, up to 6.5 nmol S cm−3 day−1 in the 12–14-cm sediment horizon.
The higher SRR were revealed in the Pechora Sea bottom sediments at station 5407 (7.8–10.4 nmol S cm−3 day−1), as well as in the bays affected by glacier movement: near Shokal'skogo Glacier at the Russkaya Gavan' Bay, Novaya Zemlya coast (station 5424, 3.0–20.1 nmol S cm−3 day−1), near the Polyarnykh Letchikov Glacier in the deep-water part of the Cambrigde Strait, Franz Josef Land coast (station 5454, 8.2–27.4 nmol S cm−3 day−1) and in the Dezhnev Bay (station 5456, 5.2–47.2 nmol S cm−3 day−1).
It should be noted, that most of the previous measurements of SRR in the Barents Sea were carried out for the samples from the bottom sediments of Svalbard fjords in the course of investigation of the OM mineralization pathways and of the temperature effect on SRB activity. These rates varied from 4–53 to 200–350 nmol S cm−3 day−1 (Arnosti et al. 1998; Sagemann et al. 1998; Nickel et al. 2008; Sawicka et al. 2012; Robador et al. 2015). The SRR at the east coastline of Svalbard were low because of higher Fe(III) concentrations, a substrate for a competitive reducing process, in the bottom sediments, versus the west coastline area (Vandieken et al. 2006a). The rates of sulphate reduction in the sediments of the central Barents Sea area varied from 1.1 to 16.8 nmol S cm−3 day−1, while in the sediments of the continental slope SRR were much lower, from 0.08 to 0.34 nmol S cm−3 day−1 (Savvichev et al. 2000, 2009). Our results obtained for the bottom sediments from a number of the Barents Sea sites, including coastal areas of the Novaya Zemlya, Franz Josef Land, and Svalbard archipelagos, are within this range and give clear evidence for the presence of active SRB cells in the investigated sediments.
Metabolic activity of SRB cells in bottom sediments near the Svalbard coast (Smeerenburgfjorden) at a water depth 218 m was previously shown to change along the sediment depth profile, with the highest sulphate reduction rate detected at the subsurface, in the upper 2–6 cm. The maximum amount of SRB cells in these sediments was at the 2.25 cm horizon—up to 5.2 × 108 SRB ml−1 (Ravenschlag et al. 2000). In the present work, the highest sulphate reduction rates were found in deeper horizons, at the horizons of 5–10 cm and more (see Table 1). It should be noted that in some cases the SRB distribution profile in marine Arctic sediments varies significantly due to the action of tides and bioturbation caused by marine invertebrates. The physical structure of the bottom sediments, the content of pore water as well as organic matter contents and metal concentrations also have large effects on abundance of marine microorganisms and their metabolic activities (Ravenschlag et al. 2001; Hop et al. 2002).
PCR detection of sulphate-reducing bacteria in enrichment cultures from the Barents Sea bottom sediments
From samples of bottom sediments taken at stations 5407, 5424, 5454 and 5456 off the coast of the Novaya Zemlya and Franz Josef Land archipelagoes, actively growing and producing H2S enrichment cultures were obtained on the Widdel lactate/sulphate medium for marine SRB. Microscopic examination of fixed and fuchsin-stained cell preparations from the SRB enrichment cultures revealed that the main part of cells in all preparations was represented by rods of different lengths, whereas vibrios, spirillae and cocci were less common.
.For all total DNA samples isolated from obtained SRB enrichment cultures, positive PCR signals indicated the presence of dsrB, one of the key genes for sulphate reduction pathway.
Direct PCR analysis of the 16S rRNA gene fragments specific for six major SRB subgroups revealed members of subgroup 5 (Desulfococcus-Desulfonema-Desulfosarcina) in enrichments from the bottom sediments of Franz Josef Land (station 5454 at the Polyarnykh Letchikov Glacier, 0–3 cm horizon) and of subgroup 6 (Desulfovibrio-Desulfomicrobium) at station 5456 (6–10 cm) in the Dezhnev Bay. SRB of subgroup 6 were also detected in enrichments from the bottom sediments of two stations (5407 and 5424, 0–1.5 and 5–7-cm horizons, correspondingly) to the south and to the north of the Novaya Zemlya archipelago. No SRBs of subgroups 1–4 were revealed in the obtained enrichment cultures (Table 2).
Nested PCR was carried out in order to detect minor SRB in the enrichment cultures. Analysis of the results showed the presence of SRB subgroups 1 (Desulfotomaculum), 2 (Desulfobulbus) and 4 (Desulfobacter) in enrichments from the bottom sediments collected to the south of Novaya Zemlya (station 5407 close to Vaygach Island near the continent). Nested PCR also revealed SRB subgroup 5 (Desulfococcus-Desulfonema-Desulfosarcina) in enrichments from the sediments collected at both stations (5407 and 5424) to the south and to the north of Novaya Zemlya and to the west of Franz Josef Land (station 5456, Dezhnev Bay). Nested PCR detected 16S rRNA gene fragments belonged to SRB subgroup 6 (Desulfovibrio-Desulfomicrobium) at station 5454 at the Franz Josef Land—see Table 2.
PCR analysis revealed no members of SRB subgroup 3 (Desulfobacterium) in all enrichment cultures from the Barents Sea bottom sediments, while SRB subgroups 1 and 2 (Desulfotomaculum and Desulfobulbus, respectively) were detected only in the enrichments from the sediments of station 5407 at the Novaya Zemlya southern coast by sensitive nested PCR.
It should be noted that members of SRB subgroup 6 (Desulfovibrio-Desulfomicrobium), which are considered the most widespread ones in marine ecosystems, were revealed by direct PCR in enrichment cultures from the bottom sediments collected at both stations (5407 and 5424) to the south and to the north of Novaya Zemlya and at station 5456 near Franz Josef Land. Nested PCR detected SRB subgroup 5 (Desulfococcus-Desulfonema-Desulfosarcina) in enrichments from the bottom sediments of all four investigated stations, which was in agreement with the earlier studies of the Svalbard coastal bottom sediments by fluorescent in situ hybridization and rRNA blot hybridization, revealed predominance of the Desulfosarcina-Desulfococcus of the family Desulfobacteraceae (up to 73% of all SRB). This fact partially confirms their abundance in seas and metabolic versatility in respect of electron donors (Ravenschlag et al. 2000).
Phylogenetic analysis of sulphate-reducing bacteria in enrichment cultures from the Barents Sea bottom sediments based on the dsrB gene
Analysis of five translated amino acid sequences encoded by the dsrB gene revealed that in total DNA isolated from the SRB enrichments, obtained from the surface horizon (0–1.5 cm) of the oxidised bottom sediments at station 5407 to the south-east of Novaya Zemlya, they exhibited the highest similarity to the dsrB sequences of Desulfatiferula (95.2%), Desulfobulbus (91.2%), Desulfopila (88.8%) genera, and uncultured Desulfofustis spp. (86.4%).
In the oxidised upper horizon (0–3 cm) of the bottom sediments collected in the area of Franz Josef Land at station 5454 (639 m depth) west of the archipelago, the highest dsrB gene translated sequence similarity (four sequences were obtained) was observed for members of Desulfatiferula genus (93.6%) too, but also with Desulfovermiculus halophilus (93.2%), Desulfocurvus vexinensis (92.8%), and uncultured SRB isolated from the hypersaline sediments of the Great Salt Lake in Utah, USA (93.1%).
As for reduced bottom sediments, in the SRB enrichments from 5–7-cm sediment horizon collected at station 5424 to the north of Novaya Zemlya archipelago in the Russkaya Gavan' Bay (depth 176 m), five translated amino acid sequences encoded by dsrB were similar to those from Desulfobulbus (96.0%), Desulfatiferula (95.2%), Desulfoconvexum (92.8%), Desulfopila (90.4%), and Desulforhopalus (89.9%) genera. Interestingly, Desulfoconvexum algidum, the only known member of Desulfoconvexum genus (Desulfobacteraceae family) is a psychrophilic SRB isolated from the bottom sediments at the Svalbard north-western coast, which is capable of chemolithoautotrophic growth on hydrogen as an electron donor (Könneke et al. 2013).
In the SRB enrichments from reduced bottom sediments (6–10-cm horizon) collected at station 5456 south of Franz Josef Land in Dezhnev Bay near the Lunnyi Kupol Glacier (depth 606 m), three translated amino acid sequences encoded by dsrB were similar to those from Desulfovibrio frigidus (93.6%), as well as from Desulfomicrobium norvegicum (91.5%) and Desulfosarcina variabilis (84.5%). D. frigidus was isolated from cold sediments at Svalbard western coast and is able to grow on a broad range of electron donors (hydrogen, formate, and lactate) and acceptors (sulphate, sulphite, thiosulphate, and elemental sulphur) (Vandieken et al. 2006b). The relevant sequences distributed among the major detected SRB families (Desulfobacteraceae, Desulfobulbaceae, and Desulfovibrionaceae) are designated on the phylogenetic tree (Fig. 3) by the numbers of corresponding sampling stations.
Phylogenetic tree of sulphate-reducing bacteria enrichment cultures from the bottom sediments of the Barents Sea, obtained by comparative analysis of the translated amino acid sequences encoded by the dsrB gene. The numbers next to each node represent a measure of support for the node (given as percentages where 100% represent maximal support). The scale bar corresponds to 5% of the calculated sequence divergence
As can be seen from the above data, dsrB gene fragments belonged to Desulfatiferula genus were detected in the oxidized upper bottom sediments (stations 5407 and 5454) as well as in some reduced lower bottom sediments (station 5424). Representatives of Desulfobulbus and Desulfopila genera existed in the oxidized (station 5407) and reduced (station 5424) bottom sediments near Novaya Zemlya archipelago. Only oxidized upper bottom sediments contains dsrB gene fragments specific to Desulfofustis spp. (station 5407 at Novaya Zemlya), Desulfovermiculus spp. and Desulfocurvus spp. (station 5454 at Franz Josef Land), while variety of SRB genera in the reduced lower bottom sediments was somewhat higher—including SRB closed to Desulfoconvexum, Desulforhopalus (station 5424 at Novaya Zemlya), Desulfovibrio, Desulfomicrobium and Desulfosarcina (station 5456 at Franz Josef Land).
Occurrence of the dsrB gene fragments with the highest similarity to the relevant fragments in genomes of members of Desulfobacteraceae and Desulfovibrionaceae families was shown previously for a number of other marine sediment ecosystems, including the meromictic Black Sea (Bryukhanov et al. 2018). Two presently known members of Desulfatiferula genus (Desulfobacteraceae family) are able to utilize long-chain alkanes and fatty acids as electron donors and are mesophiles, isolated from oil-contaminated bottom sediments of Étang de Berre Lagoon at the Mediterranean coast of France (Cravo-Laureau et al. 2007; Hakil et al. 2014). It should be noted that all known Desulfobulbus spp. (Desulfobulbaceae family) oxidize three-carbon compounds and have a wide lability of metabolism from fermentative to mixotrophic, some species were isolated from desalinated bottom sediments of river estuaries (Oakley et al. 2010). Members of the genus Desulfopila of the same family, including Desulfopila aestuarii and Desulfopila inferna, use a broad spectrum of organic electron donors (and sometimes also hydrogen) and are also found mainly in the coastal areas, such as river estuaries and the littoral zone (Gittel et al. 2010).
Phylogenetic analysis of sulphate-reducing bacterial communities from the Barents Sea bottom sediments based on the 16S rRNA gene profiling
Analysis of DNA fragments isolated and reamplified from DGGE bands during investigation of enrichment cultures is not always able to detect the minor components of microbial communities, and application of dsrB as a genetic marker, although widely used for study the phylogenetic position of SRB (Bagwell et al. 2009), has certain limitations due to horizontal gene transfer (Klein et al. 2001). Therefore, a high-throughput sequencing of the 16S rRNA gene fragments was used for complete qualitative and quantitative analysis of native microbial communities in the Barents Sea bottom sediments.
Phylogenetic analysis of the composition of microbial communities was carried out for 12 sediment samples with high sulphate reduction rates: stations 5407 (0–1.5 and 10–14-cm horizons), 5412 (0–2 and 10–14-cm horizons), 5424 (0–2, 5–7, and 19–24-cm horizons), 5441 (0–2 and 6–8-cm horizons), and 5454 (0–3, 7–9, and 8–22-cm horizons). About 6000–20,000 sequences of the 16S rRNA gene fragments were obtained for each sample. It was demonstrated, that the most numerous subgroups of sulphate- and sulphur-reducing bacteria of Deltaproteobacteria class in the investigated bottom sediments belonged to Desulfobulbaceae and Desulfobacteraceae families (Desulfobacterales order), Desulfuromonadaceae family (Desulfuromonadales order), Desulfarculaceae family (Desulfarculales order), Desulfovibrionaceae family (Desulfovibrionales order), and two uncultured clades, SAR324 and Sva0485 (Fig. 4).
Distribution of the main subgroups of sulphate/sulphur-reducing bacteria (at the family level) in the bottom sediments of the Barents Sea based on the results of high-throughput sequencing of the 16S rRNA gene fragments. The X-axis shows the numbers of sampling stations and horizons (in cm) of bottom sediments, and the Y-axis indicates the abundance (in %) relative to all reads of the 16S rRNA gene sequences in the sample, i.e. percentages of SRB subgroups from the entire microbial community
Clade SAR324 comprises widespread oceanic Deltaproteobacteria preferring the biotopes with low O2 content and possessing versatile metabolism (chemolithotrophy, heterotrophic growth on C1 compounds, and alkane oxidation). The SAR324 dsrA gene clustered phylogenetically in a bootstrap supported clade that is distinct from the common sulphur-oxidizing and sulphate-reducing types (Sheik et al. 2014). For clade Sva0485, which comprises tentative sulphate/iron-reducing Deltaproteobacteria, detected to date in acidic mine ecosystems, deep-water hydrothermal sulphide smokers, and iron-containing lake sediments, a new taxon, Acidulodesulfobacterales order, has been proposed (Tan et al. 2019).
At station 5407 (south-east of Novaya Zemlya archipelago, near Vaygach Island, depth 47 m) with the lowest Corg content (0.2%), which is a subject to the effects of the Coastal Current carrying desalinated seawater and of the Pechora catchment basin, most numerous SRB in the surface oxidised sediments (0–1.5-cm horizon, Eh = 190 mV) belonged to Desulfobulbaceae (5.4% of all reads in the sample), Desulfuromonadaceae (3.2%), Desulfobacteraceae (1.8%) families, and to SAR324 clade (0.5% of all reads)—see Fig. 4. At the deeper, strongly reduced horizon (10–14 cm, Eh = −160 mV) the SRB community composition was different, with predominance of members of the Desulfobacteraceae family (4.4% of all reads in the sample), while the shares of Desulfobulbaceae and Desulfuromonadaceae decreased to 1.9% and 1.6%, respectively. SRB of Desulfarculaceae family (1.2%) and Sva0485 clade (0.9%) were also revealed. Within Desulfobacteraceae family, the closest similarity of the obtained 16S rRNA gene sequences was to those of uncultured SEEP-SRB1 and Sva0081 clades, as well as to Desulfosarcina genus.
The samples collected at station 5412 (west of Novaya Zemlya archipelago, area of the Shtokman gas condensate deposit, depth 267 m), showed no pronounced differences in the composition of the SRB communities in similar sediment horizons (0–2 and 10–14 cm), in spite of Corg content being 7 times higher (Fig. 4). It should be noted, however, that compared to the coastal station 5407, in the upper oxidised bottom sediments (Eh = 226 mV) at station 5412, relative abundance of SRB cells was 2.8 lower, although their phylogenetic composition was similar (% of all reads in the sample): Desulfobulbaceae (1.7%), Desulfuromonadaceae (0.9%), Desulfobacteraceae (0.8%), and SAR324 clade (0.4%). In the deeper, reduced sediments of station 5412 (10–14 cm, Eh = −175 mV), the quantitative and qualitative composition of the SRB community also did not differ significantly from that of station 5407 (Fig. 4), apart from 2.5 times higher relative abundance of Desulfarculaceae and especially a significant decrease in relative abundance of Desulfuromonadaceae family (from 1.6% to 0.08%). It looking as follows: Desulfobacteraceae (5% of all reads in the sample), Desulfarculaceae (3.1%), Desulfobulbaceae (2.4%), and Sva0485 clade (0.6%).
In the bottom sediments of station 5424, together with station 5407 located at the Novaya Zemlya coast, although at its northern part (Russkaya Gavan' Bay, 176 m depth) and is affected by freshwater inflow from the Shokal'skii Glacier, the composition of the SRB community was interesting. Similar to station 5407, the sediments at station 5424 had low organic matter content (0.3%) and was located in the area affected by coastal currents. In the oxidised upper sediments (0–2 cm, Eh = 208 mV) of station 5424, members of Desulfuromonadaceae family were the most abundant group (5.9% of all reads in the sample), rather than Desulfobulbaceae predominant at stations 5407 and 5412. Other subgroups were Desulfobulbaceae (2.4%), Desulfobacteraceae (1.2%), and SAR324 clade (0.3%). Deeper reduced horizons (5–7 and 19–24 cm, Eh = −147 and −210 mV) of station 5424 differed from the similar horizon (10–14 cm) at station 5407 in predominance of Desulfobulbaceae family (especially in the 19–24-cm horizon, where SRB relative abundance was 15% of all reads, the highest value among all collected sediments) and in almost complete absence of Desulfoarculaceae members. Phylogenetic composition of predominant bacteria reducing sulphur compounds in the horizons 5–7 and 19–24 cm of station 5424 was as follows: Desulfobulbaceae (3.9% and 9%), Desulfobacteraceae (3.2% and 2.5%), and Desulfuromonadaceae (1.9% and 2.8%)—see Fig. 4. Apart from numerous uncultured forms, members of Desulforhopalus and Desulfocapsa genera were revealed within the Desulfobulbaceae family, while the most common members of Desulfobacteraceae family belonged to Desulfobacula and Desulfoconvexum genera, as well as to uncultured Sva0081 and SEEP-SRB1 clades.
In the oxidised surface sediments (0–2-cm horizon, Eh = 154 mV) of station 5441 at the Svalbard southern coast, which are affected by glacial flows and contain relatively high Corg content (similar to station 5412), SRB relative abundance was relatively low (not exceeding 5%). The quantitative phylogenetic composition of the microbial community reducing sulfur compounds was relatively similar to that of the surface sediments of stations 5407 and 5412: Desulfobulbaceae (1.7% of all reads in the sample), Desulfobacteraceae (1.5%), Desulfuromonadales (1.1%), and SAR324 and Sva0485 clades—0.4% and 0.2%, respectively. Compared to stations 5407 and 5412, reduced sediments (horizon 6–8 cm, Eh = −150 mV) of station 5441 had higher relative abundance of members of Desulfobulbaceae family (4.9% of all reads in the sample) and lower relative abundance of Desulfoarculaceae (Fig. 4). While most Desulfobulbaceae from this horizon belonged to uncultured lineages, some were identified at the genus level (Desulfocapsa and Desulfobulbus); within Desulfobacteraceae family, Desulfofaba, Desulfococcus, and Desulfatirhabdium genera were revealed, as well as SEEP-SRB1 and Sva0081 clades. Low amounts of the 16S rRNA gene sequences exhibiting the highest similarity to those of Desulfatiglans spp. (Desulfarculaceae family, Desulfarculales order) were also detected.
To compare the obtained data on phylogenetic composition of SRB communities with the previous research of Arctic marine sediments it should be noted that only a few data is known up to date. For instance, several psychrophilic SRB have been isolated from Svalbard coastal bottom sediments, including the representatives of Desulfofrigus, Desulfofaba, Desulfotalea (Knoblauch et al. 1999b), Desulfotomaculum (Vandieken et al. 2006c) and Desulfoconvexum (Könneke et al. 2013) genera. As it is mentioned above, Desulfofaba spp. and Desulfoconvexum spp. were revealed by 16S rRNA analysis in the oxidised surface sediments (0–2-cm horizon) of station 5441 at the Svalbard southern coast and in the reduced bottom sediments (5–7-cm horizon) of station 5424 to the north of Novaya Zemlya correspondingly.
The bottom sediments collected at the deep-water (639 m) station 5454 west of the Franz Josef Land, affected by the flow from the Polyarnykh Letchikov Glacier had the SRB phylogenetic composition differing significantly from that of other sampling stations. Thus, the surface (0–3 cm, Eh = 147 mV) oxidised sediments showed the lowest relative abundance of sulphate- and sulphur-reducing bacteria among the all collected Barents Sea sediment samples (2.5% of all reads in the sample). Members of Desulfoarculaceae family and SVA0485 clade were not detected, and relative abundance of Desulfobulbaceae was very low (only 0.3% of all reads), while Desulfuromonadales formed the majority (1.5% of all reads in the sample) (Fig. 4). While, unlike the similar horizon of other stations, the deeper horizon (7–9 cm) of station 5454 was also oxidised (Eh = 125 mV), the rate of sulphate reduction there was very high (27.4 nmol S cm−3 day−1). In this sample, members of Desulfobulbaceae family, which was the most numerous SRB subgroup in the majority of the studied samples, were almost absent; relative abundance of Desulfuromonadales was also very low; the share of uncultured SAR324 clade was significant (over 1.3% of all reads in the sample). In the deep reduced horizons (18–22 cm, Eh = −105 mV) of station 5454 sediments, SRB relative abundance was lower than in similar horizons of other stations, although the phylogenetic composition of the microbial community was similar, except for the high share of SAR324 clade (Fig. 4): SAR324 (2.1% of all reads in the sample), Desulfobacteraceae (2%), Desulfobulbaceae (1.5%), Desulfuromonadales (1.1%), Sva0485 (0.2%), and Desulfoarculaceae (0.2%).
It should be noted that members of Desulfobulbaceae family, which constituted the most abundant SRB subgroup in most horizons of the studied Barents Sea bottom sediments, have been previously detected at the boundary between anoxic sulphide-containing sediments and upper oxidised ones (Pfeffer et al. 2012). The members of Desulfobacteraceae family also occur in marine bottom sediments (Strittmatter et al. 2009). The presence of metabolically active Desulfobacteraceae and Desulfobulbaceae was previously revealed near Axel Heiberg Island at the north of the Canadian Arctic archipelago (Colangelo‐Lillis et al. 2019) and in Arctic deposits of northern Finland, where Desulfobulbus was the predominant genus (Virpiranta et al. 2019). Aceticlastic SRB of Desulfobacteraceae and Desulfobulbaceae families were shown to play a significant role in the degradation of cyanobacterial necromass in marine Arctic sulphide sediments (Müller et al. 2018).
Bacteria able to reduce sulphate and elemental sulphur constituted ~ 10% of all microorganisms in surface oxidised sediments at stations 5407 and 5424, which are located near the Novaya Zemlya archipelago coast, affected by coastal currents and contained the lowest Corg content, with predominance of Desulfobulbaceae and Desulfuromonadaceae. In deeper, reduced sediment horizons of station 5407, Desulfobacteraceae family was the dominant SRB group, while Desulfobulbaceae prevailed in reduced sediments of station 5424, where the rate of sulphate reduction was higher. Slight predominance of Desulfobulbaceae over Desulfobacteraceae was observed also in reduced sediments of station 5441, located at Svalbard archipelago southern coast, affected by both Arctic and Atlantic currents, and characterised by much higher Corg content. The highest relative abundance of members of Desulfarculaceae family was revealed in reduced horizons of sand-clayey sediments of station 5412 in the Barents Sea area affected by the currents of transformed Atlantic waters. In the reduced horizon of sandy-aleurite sediments from the deepest station 5454 (Cambridge Bay, Franz Josef Land, depth 639 m), the lowest rate of sulphate reduction was detected, as well as emergence of the SAR324 clade, members of which, as was mentioned above, possess highly labile metabolism.
Some sulphur-reducing bacteria of the family Desulfuromonadaceae, which was also detected in the bottom sediment samples in present research (Fig. 4), are able to use as electron acceptors, apart from elemental sulphur, also Mn(IV), Fe(III), Co(III), Tc(VII), U(VI) etc. Thus, psychrophilic Fe(III)-reducing bacteria Desulfuromonas svalbardensis and Desulfuromusa ferrireducens have been isolated from marine sediments at Svalbard archipelago coast (Vandieken et al. 2006d) The bottom sediments of Svalbard fjords, which are affected by nearby glaciers, have low content of organic matter, which limits formation of iron sulphides, so that iron is released into the water column. Fe(III)-reducing bacteria of the order Desulfuromonadales were detected recently in the upper 12 cm of these sediments. At a station remote from the coast, transition from the iron cycle microbial communities to sulphate-reducing communities occurred in the sediments already in the ~ 5-cm horizon, which was in agreement with the higher Corg content and higher rate of sulphate reduction (Buongiorno et al. 2019).
The lowest SRB relative abundance, not exceeding 3–4.5% of all microorganisms, was observed here in all oxidised surface sediment horizons, except for two stations at the Novaya Zemlya coast. Detection of anaerobic SRB of Desulfobacterales order in such strongly oxidised conditions (with Eh up to 226 mV) at all stations is of great interest, confirming aerotolerance of many SRB cells inhabiting the oxic-anoxic interface in a number of biotopes and possessing effective antioxidation systems (Brioukhanov et al. 2010).
High relative abundance of uncultured (up to 40%) sulphate- and sulphur-reducing bacteria revealed in the studied Barents Sea bottom sediments indicates considerable prospects for identification, isolation to pure cultures, and in-depth biochemical and genetic investigation of members of new microbial taxa inhabiting the cold Arctic seas, a presently insufficiently studied part of the World Ocean.
Data availability
The datasets generated and analysed during the current study from the 16S rRNA gene sequencing are available in the NCBI Sequence Read Archive (SRA) repository under accession numbers SRX8118116-SRX8118127 and BioSample repository under accession numbers SAMN14599701-SAMN14599712 (BioProject PRJNA624280)—https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA624280. The datasets generated and analysed during the current study from the dsrB gene sequencing are available in the GenBank repository under accession numbers MT806071-MT806087—https://www.ncbi.nlm.nih.gov/nuccore/.
Code availability
Not applicable.
References
Arnosti C, Jørgensen BB, Sagemann J, Thamdrup B (1998) Temperature dependence of microbial degradation of organic matter in marine sediments: polysaccharide hydrolysis, oxygen consumption, and sulfate reduction. Mar Ecol Prog Ser 165:59–70. https://doi.org/10.3354/meps165059
Åström EKL, Carroll ML, Ambrose WG Jr, Carroll J (2016) Arctic cold seeps in marine methane hydrate environments: impacts on shelf microbenthic community structure offshore Svalbard. Mar Ecol Prog Ser 552:1–18. https://doi.org/10.3354/meps11773
Bagwell CE, Formolo M, Ye Q, Yeager CM, Lyons TW, Zhang CL (2009) Direct analysis of sulfate-reducing bacterial communities in gas hydrate-impacted marine sediments by PCR-DGGE. J Basic Microbiol 49:87–92. https://doi.org/10.1002/jobm.200800278
Brioukhanov AL, Durand M-C, Dolla A, Aubert C (2010) Response of Desulfovibrio vulgaris Hildenborough to hydrogen peroxide: enzymatic and transcriptional analyses. FEMS Microbiol Lett 310:175–181. https://doi.org/10.1111/j.1574-6968.2010.02061.x
Bryukhanov AL, Vlasova MA, Malakhova TV, Perevalova AA, Pimenov NV (2018) Phylogenetic diversity of the sulfur cycle bacteria in the bottom sediments of the Chersonesus Bay. Microbiology 87:372–381. https://doi.org/10.1134/S0026261718030025
Buongiorno J, Herbert LC, Wehrmann LM, Michaud AB, Laufer K, Røy H, Jørgensen BB, Szynkiewicz A, Faiia A, Yeager KM, Schindler K, Lloyda KG (2019) Complex microbial communities drive iron and sulfur cycling in Arctic fjord sediments. Appl Environ Microbiol 85:e00949-e1019. https://doi.org/10.1128/AEM.00949-19
Colangelo-Lillis J, Pelikan C, Herbold CW, Altshuler I, Loy A, Whyte LG, Wing BA (2019) Diversity decoupled from sulfur isotope fractionation in a sulfate-reducing microbial community. Geobiology 17:660–675. https://doi.org/10.1111/gbi.12356
Cravo-Laureau C, Labat C, Joulian C, Matheron R, Hirschler-Réa A (2007) Desulfatiferula olefinivorans gen. nov., sp. Nov., a long-chain n-alkene-degrading, sulfate-reducing bacterium. Int J Syst Evol Microbiol 57:2699–2702
Dalpadado P, Arrigo KR, Hjøllo SS, Rey F, Ingvaldsen RB, Sperfeld E, van Dijken GL, Stige LC, Olsen A, Ottersen G (2014) Productivity in the Barents Sea – response to recent climate variability. PLoS ONE 9:e95273. https://doi.org/10.1371/journal.pone.0095273
Daly K, Sharp RJ, McCarthy AJ (2000) Development of oligonucleotide probes and PCR primers for detecting phylogenetic subgroups of sulfate-reducing bacteria. Microbiology (reading) 146:1693–1705. https://doi.org/10.1099/00221287-146-7-1693
Dar SA, Kuenen JG, Muyzer G (2005) Nested PCR-denaturing gradient gel electrophoresis approach to determine the diversity of sulfate-reducing bacteria in complex microbial communities. Appl Environ Microbiol 71:2325–2330. https://doi.org/10.1128/AEM.71.5.2325-2330.2005
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinfo 5:113. https://doi.org/10.1186/1471-2105-5-113
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Edwards U, Rogall T, Blöcker H, Emde M, Böttger EC (1989) Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res 17:7843–7853. https://doi.org/10.1093/nar/17.19.7843
Finke N, Jørgensen BB (2008) Response of fermentation and sulfate reduction to experimental temperature changes in temperate and Arctic marine sediments. ISME J 2:815–829. https://doi.org/10.1038/ISMEJ.2008.20
Finke N, Vandieken V, Jørgensen BB (2007) Acetate, lactate, propionate, and isobutyrate as electron donors for iron and sulfate reduction in Arctic marine sediments, Svalbard. FEMS Microbiol Ecol 59:10–22. https://doi.org/10.1111/j.1574-6941.2006.00214.x
Forschner SR, Sheffer R, Rowley DC, Smith DC (2009) Microbial diversity in Cenozoic sediments recovered from the Lomonosov Ridge in the Central Arctic Basin. Environ Microbiol 11:630–639. https://doi.org/10.1111/j.1462-2920.2008.01834.x
Gabrielsen RH, Faerseth RB, Jensen LN, Kalheim JE, Riis F (1990) Structural elements of the Norwegian continental shelf. Pt. 1. The Barents Sea Region. Norwegian Petroleum Directorate Bulletin 6:33
Geets J, Borremans B, Diels L, Springael D, Vangronsveld J, van der Lelie D, Vanbroekhoven K (2006) DsrB gene-based DGGE for community and diversity surveys of sulfate-reducing bacteria. J Microbiol Meth 66:194–205. https://doi.org/10.1016/j.mimet.2005.11.002
Gittel A, Seidel M, Kuever J, Galushko AS, Cypionka H, Könneke M (2010) Desulfopila inferna sp. nov., a sulfate-reducing bacterium isolated from the subsurface of a tidal sand-flat. Int J Syst Evol Microbiol 60:1626–1630. https://doi.org/10.1099/ijs.0.015644-0
Glombitza C, Jaussi M, Røy H, Seidenkrantz MS, Lomstein BA, Jørgensen BB (2015) Formate, acetate, and propionate as substrates for sulfate reduction in sub-arctic sediments of Southwest Greenland. Front Microbiol 6:1–14. https://doi.org/10.3389/fmicb.2015.00846
Green SJ, Leigh MB, Neufeld JD (2010) Denaturing gradient gel electrophoresis (DGGE) for microbial community analysis. In: Timmis KN (eds) Handbook of hydrocarbon and lipid microbiology. Springer, Berlin, Heidelberg, pp 4137–4158. https://doi.org/10.1007/978-3-540-77587-4_323
Hakil F, Amin-Ali O, Hirschler-Réa A, Mollex D, Grossi V, Duran R, Matheron R, Cravo-Laureau C (2014) Desulfatiferula berrensis sp. nov., a n-alkene-degrading sulfate-reducing bacterium isolated from estuarine sediments. Int J Syst Evol Microbiol 64:540–544. https://doi.org/10.1099/ijs.0.057174-0
Hop H, Pearson T, Nøst Hegseth E, Kovacs KM, Wiencke C, Kwasniewski S, Eiane K, Mehlum F, Gulliksen B, Wlodarska-Kowalczuk M, Lydersen C, Weslawski JM, Cochrane S (2002) The marine ecosystem of Kongsfjorden, Svalbard. Polar Res 21:167–208. https://doi.org/10.3402/polar.v21i1.6480
Ingvaldsen R, Loeng H (2009) Physical oceanography. In: Sakshaug E, Johnsen G, Kovacs K (eds) Ecosystem Barents Sea. Tapir Academic Press, Trondheim, pp 33–64
Jensen MM, Thamdrup B, Rysgaard S, Holmer M, Fossing H (2003) Rates and regulation of microbial iron reduction in sediments of the Baltic-North Sea transition. Biogeochemistry 65:295–317. https://doi.org/10.1023/A:1026261303494
Jørgensen BB (1982) Mineralization of organic matter in the sea bed – the role of sulphate reduction. Nature 296:643–645. https://doi.org/10.1038/296643a0
Jørgensen BB, Bang M, Blackburn TH (1990) Anaerobic mineralization in marine sediments from the Baltic Sea-North Sea transition. Mar Ecol Prog Ser 59:39–54. https://doi.org/10.3354/meps059039
Klein M, Friedrich M, Roger AJ, Hugenholtz P, Fishbain S, Abicht H, Blackall LL, Stahl DA, Wagner M (2001) Multiple lateral transfers of dissimilatory sulfite reductase genes between major lineages of sulfate-reducing prokaryotes. J Bacteriol 183:6028–6035. https://doi.org/10.1128/JB.183.20.6028-6035.2001
Knoblauch C, Jørgensen BB, Harder J (1999a) Community size and metabolic rates of psychrophilic sulfate-reducing bacteria in Arctic marine sediments. Appl Environ Microbiol 65:4230–4233. https://doi.org/10.1128/AEM.65.9.4230-4233.1999
Knoblauch C, Sahm K, Jørgensen BB (1999b) Psychrophilic sulfate-reducing bacteria isolated from permanently cold Arctic marine sediments: description of Desulfofrigus oceanense gen. nov., sp. nov., Desulfofrigus fragile sp. nov., Desulfofaba gelida gen. nov., sp. nov., Desulfotalea psychrophila gen. nov., sp. nov. and Desulfotalea arctica sp. nov. Int J Syst Evol Microbiol 49:1631–1643. https://doi.org/10.1099/00207713-49-4-1631
Könneke M, Kuever J, Galushko A, Jørgensen BB (2013) Desulfoconvexum algidum gen. nov., sp. nov., a psychrophilic sulfate-reducing bacterium isolated from a permanently cold marine sediment. Int J Syst Evol Microbiol 63:959–964. https://doi.org/10.1099/ijs.0.043703-0
Korneeva VA, Pimenov NV, Krek AV, Tourova TP, Bryukhanov AL (2015) Sulfate-reducing bacterial communities in the water column of the Gdansk Deep (Baltic Sea). Microbiology 84:297–306. https://doi.org/10.1134/S002626171502006X
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lind S, Ingvaldsen RB, Furevik T (2018) Arctic warming hotspot in the northern Barents Sea linked to declining sea-ice import. Nat Clim Chang 8:634–639. https://doi.org/10.1038/s41558-018-0205-y
Makarevich PR, Druzhkova EI, Larionov V (2012) Primary production of the Barents Sea. Bulletin of MSTU 15:786–793 (in Russian). https://doi.org/10.5772/37512
Margulis EA (2008) Factors of the formation of the unique Shtokman-Ludlovsky gas accumulation point in the Barents Sea. Oil and Gas Geology. Theory Pract 3:1–15 ((in Russian))
Mityaev MV, Gerasimova MV, Pavlova LG, Druzhkova EI (2018) Lateral flows of suspended matter in the Kola Meridian section. In: Proceedings of the Kola scientific centre of the russian academy of sciences 9:109–117 (in Russian). https://doi.org/10.25702/KSC.2307-5252.2018-9-4-109-117
Müller AL, Pelikan C, de Rezende JR, Wasmund K, Putz M, Glombitza C, Kjeldsen KU, Jørgensen BB, Loy A (2018) Bacterial interactions during sequential degradation of cyanobacterial necromass in a sulfidic arctic marine sediment. Environ Microbiol 20:2927–2940. https://doi.org/10.1111/1462-2920.14297
Muyzer G, Stams AJM (2008) The ecology and biotechnology of sulphate-reducing bacteria. Nat Rev Microbiol 6:441–454. https://doi.org/10.1038/nrmicro1892
Muyzer G, de Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700. https://doi.org/10.1128/AEM.59.3.695-700.1993
Nemirovskaya IA (2020) Hydrocarbons in the waters and bottom sediments of the Barents Sea. Geochemistry 65:679–692 (in Russian). https://doi.org/10.31857/S0016752520070079
Nickel M, Vandieken V, Brüchert V, Jørgensen BB (2008) Microbial Mn(IV) and Fe(III) reduction in northern Barents Sea sediments under different conditions of ice cover and organic carbon deposition. Deep-Sea Res II 55:2390–2398. https://doi.org/10.1016/j.dsr2.2008.05.003
Oakley BB, Carbonero F, van der Gast CJ, Hawkins RJ, Purdy KJ (2010) Evolutionary divergence and biogeography of sympatric niche-differentiated bacterial populations. ISME J 4:488–497. https://doi.org/10.1038/ismej.2009.146
Pecheritsyna SA, Rivkina EM, Akimov VN, Shcherbakova VA (2012) Desulfovibrio arcticus sp. nov., a psychrotolerant sulfate-reducing bacterium from a cryopeg. Int J Syst Evol Microbiol 62:33–37. https://doi.org/10.1099/ijs.0.021451-0
Pfeffer C, Larsen S, Song J, Dong M, Besenbacher F, Meyer RL, Kjeldsen KU, Schreiber L, Gorby YA, El-Naggar MY, Leung KM, Schramm A, Risgaard-Petersen N, Nielsen LP (2012) Filamentous bacteria transport electrons over centimetre distances. Nature 491:218–221. https://doi.org/10.1038/nature11586
Pimenov NV, Bonch-Osmolovskaya EA (2006) In situ activity studies in thermal environments. Meth Microbiol 35:29–53. https://doi.org/10.1016/S0580-9517(08)70005-9
Pimenov NV, Savvichev AS, Rusanov II, Lein AYu, Ivanov MV (2000) Microbiological processes of the carbon and sulfur cycles at cold methane seeps of the North Atlantic. Microbiology 69:709–720. https://doi.org/10.1023/A:1026666527034
Politova NV, Novigatsky AN, Kozina NV, Terpugova SA (2018) Multidisciplinary research in the Barents Sea on cruise 67th of the R/V Akademik Mstislav Keldysh. Oceanology 58:499–501. https://doi.org/10.1134/S0001437018030153
Politova NV, Kravchishina MD, Novigatsky AN, Lokhov AS (2019) Dispersed sedimentary matter of the Barents Sea. Oceanology 59:697–714. https://doi.org/10.1134/S0001437019050151
Pruesse E, Peplies J, Glöckner FO (2012) SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28:1823–1829. https://doi.org/10.1093/bioinformatics/bts252
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
Ravenschlag K, Sahm K, Knoblauch C, Jørgensen BB, Amann R (2000) Community structure, cellular rRNA content, and activity of sulfate-reducing bacteria in marine arctic sediments. Appl Environ Microbiol 66:3592–3602. https://doi.org/10.1128/aem.66.8.3592-3602.2000
Ravenschlag K, Sahm K, Amann R (2001) Quantitative molecular analysis of the microbial community in marine Arctic sediments (Svalbard). Appl Environ Microbiol 67:387–395. https://doi.org/10.1128/AEM.67.1.387-395.2001
Robador A, Brüchert V, Jørgensen BB (2009) The impact of temperature change on the activity and community composition of sulfate-reducing bacteria in arctic versus temperate marine sediments. Environ Microbiol 11:1692–1703. https://doi.org/10.1111/j.1462-2920.2009.01896.x
Robador A, Müller AL, Sawicka JE, Berry D, Hubert CRJ, Loy A, Jørgensen BB, Brüchert V (2015) Activity and community structures of sulfate-reducing microorganisms in polar, temperate and tropical marine sediments. ISME J 10:796–809. https://doi.org/10.1038/ismej.2015.157
Sagemann J, Jørgensen BB, Greeff O (1998) Temperature dependence and rates of sulfate reduction in cold sediments of Svalbard, Arctic Ocean. Geomicrobiol J 15:85–100. https://doi.org/10.1080/01490459809378067
Sahm K, Knoblauch C, Amann R (1999) Phylogenetic affiliation and quantification of psychrophilic sulfate-reducing isolates in marine arctic sediments. Appl Environ Microbiol 65:3976–3981. https://doi.org/10.1128/AEM.65.9.3976-3981.1999
Savvichev AS, Rusanov II, Pimenov NV, Mitskevich IN, Bairamov IT, Lein AYu, Ivanov MV (2000) Microbiological explorations in the northern part of the Barents Sea in early winter. Microbiology 69:698–708. https://doi.org/10.1023/A:1026614510196
Savvichev AS, Demidenko NA, Rusanov II, Zakharova EE, Veslopolova EF, Afonina I, Ankudinova I, Pimenov NV, Ivanov MV (2009) Study of the microbial processes in the water column and bottom sediments of the Dolgaya-Vostochnaya Bay (Barents Sea) before construction of the Northern tidal power plant. Microbiology 78:798–801. https://doi.org/10.1134/S0026261709060186
Sawicka JE, Jørgensen BB, Bruchert V (2012) Temperature characteristics of bacterial sulfate reduction in continental shelf and slope sediments. Biogeosciences 9:3425–3435. https://doi.org/10.5194/bg-9-3425-2012
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Sheik CS, Jain S, Dick GJ (2014) Metabolic flexibility of enigmatic SAR324 revealed through metagenomics and metatranscriptomics. Environ Microbiol 16:304–317. https://doi.org/10.1111/1462-2920.12165
Stiansen JE, Korneev O, Titov O, Arneberg P, Filin A, Hansen JR, Høines Å, Marasaev S (2009) Joint Norwegian-Russian environmental status 2008. Report on the Barents Sea ecosystem. Part II – complete report. IMR/PINRO Joint Report Series 3:375
Strittmatter AW, Liesegang H, Rabus R, Decker I, Amann J, Andres S, Henne A, Fricke WF, Martinez-Arias R, Bartels D, Goesmann A, Krause L, Pühler A, Klenk HP, Richter M, Schüler M, Glöckner FO, Meyerdierks A, Gottschalk G, Amann R (2009) Genome sequence of Desulfobacterium autotrophicum HRM2, a marine sulfate reducer oxidizing organic carbon completely to carbon dioxide. Environ Microbiol 11:1038–1055. https://doi.org/10.1111/j.1462-2920.2008.01825.x
Takai K, Horikoshi K (2000) Rapid detection and quantification of members of the archaeal community by quantitative PCR using fluorogenic probes. Appl Environ Microbiol 66:5066–5072. https://doi.org/10.1128/aem.66.11.5066-5072.2000
Tan S, Liu J, Fang Y, Hedlund BP, Lian Z-H, Huang L-Y, Li J-T, Huang L-N, Li W-J, Jiang H-C, Dong H-L, Shu W-S (2019) Insights into ecological role of a new deltaproteobacterial order Candidatus Acidulodesulfobacterales by metagenomics and metatranscriptomics. ISME J 13:2044–2057. https://doi.org/10.1038/s41396-019-0415-y
Thamdrup B (2000) Bacterial manganese and iron reduction in aquatic sediments. In: Schink B (ed) Advances in microbial ecology, vol. 16. Springer, Boston, pp 41–84. https://doi.org/10.1007/978-1-4615-4187-5_2
Thamdrup B, Fossing H, Jørgensen BB (1994) Manganese, iron and sulfur cycling in a coastal marine sediment, Aarhus Bay, Denmark. Geochim Cosmochim Acta 58:5115–5129. https://doi.org/10.1016/0016-7037(94)90298-4
Trüper HG, Schlegel HG (1964) Sulfur metabolism in Thiorhodaceae. I. Quantitative measurements in growing cells of Cromatium okenii. Antonie Van Leeuwenhoek 30:225–238. https://doi.org/10.1007/BF02046728
Vandieken V, Finke N, Jørgensen BB (2006a) Pathways of carbon oxidation in an Arctic fjord sediment (Svalbard) and isolation of psychrophilic and psychrotolerant Fe(III)-reducing bacteria. Mar Ecol Prog Ser 322:29–41. https://doi.org/10.3354/meps322029
Vandieken V, Knoblauch C, Jørgensen BB (2006b) Desulfovibrio frigidus sp. nov. and Desulfovibrio ferrireducens sp. nov., psychrotolerant bacteria isolated from Arctic fjord sediments (Svalbard) with the ability to reduce Fe(III). Int J Syst Evol Microbiol 56:681–685. https://doi.org/10.1099/ijs.0.64057-0
Vandieken V, Knoblauch C, Jørgensen BB (2006c) Desulfotomaculum arcticum sp. nov., a novel spore-forming, moderately thermophilic, sulfate-reducing bacterium isolated from a permanently cold fjord sediment of Svalbard. Int J Syst Evol Microbiol 56:687–690. https://doi.org/10.1099/ijs.0.64058-0
Vandieken V, Mußmann M, Niemann H, Jørgensen BB (2006d) Desulfuromonas svalbardensis sp. nov. and Desulfuromusa ferrireducens sp. nov., psychrophilic, Fe(III)-reducing bacteria isolated from Arctic sediments. Svalbard Int J Syst Evol Microbiol 56:1133–1139. https://doi.org/10.1099/ijs.0.63639-0
Virpiranta H, Taskila S, Leiviskä T, Rämö J, Tanskanen J (2019) Development of a process for microbial sulfate reduction in cold mining waters – cold acclimation of bacterial consortia from an Arctic mining district. Environ Pollut 252:281–288. https://doi.org/10.1016/j.envpol.2019.05.087
Widdel F, Bak F (1992) Gram-negative mesophilic sulfate-reducing bacteria. In: Balows A, Trüper HG, Dworkin M, Harder W, Schleifer K-H (eds) The prokaryotes, 2nd edn. Springer-Verlag, New York, pp 3352–3378
Wollast R (1991) The coastal organic carbon cycle: fluxes, sources, and sinks. In: Mantoura RFC, Martin J-M, Wollast R (eds) Ocean margin processes in global change. Wiley, New York, pp 365–381
Yu Y, Lee C, Kim J, Hwang S (2005) Group-specific primer and probe sets to detect methanogenic communities using quantitative real-time polymerase chain reaction. Biotechnol Bioeng 89:670–679. https://doi.org/10.1002/bit.20347
Funding
Determination of organic carbon was carried out at the expense of the Russian Science Foundation (Grant Number 20-17-00157). Primary treatment of the samples and high-throughput sequencing of the 16S rRNA gene fragments were supported by the Russian Science Foundation also. Analysis of the geological data, radioisotope measurements of sulfate reduction rates, PCR analysis and sequencing of the dsrB gene fragments were supported by the Ministry of Science and Higher Education of the Russian Federation. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
ALB—designed study, performed research, analysed data, wrote the most part of the paper; VVK—designed study, performed research, analysed data, wrote the paper; IIR—performed research, analysed data; ANN—performed research, analysed data, wrote the paper; TAK—performed research, analysed data; NVP—performed research, analysed data; NVR—designed study; NVP—designed study.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare that are relevant to the content of this article.
Ethical approval
This article does not contain any studies with human participants, animals or their biological material performed by any of the authors.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Brioukhanov, A.L., Kadnikov, V.V., Rusanov, I.I. et al. Phylogenetic diversity in sulphate-reducing bacterial communities from oxidised and reduced bottom sediments of the Barents Sea. Antonie van Leeuwenhoek 115, 801–820 (2022). https://doi.org/10.1007/s10482-022-01733-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-022-01733-9