Abstract
Hydrogen-uptake (Hup) activity is implicated in the mitigation of energy losses associated with the biological nitrogen fixation process, and has been related to productivity increases in some legume hosts. However, in common bean (Phaseolus vulgaris L.) the expression of hydrogenase is rare. In this study an 18-kb hup gene cluster from Rhizobium leguminosarum bv. viciae encoding a NiFe hydrogenase was successfully transferred to three common bean rhizobial strains lacking hydrogenase activity (Hup−) but symbiotically very effective and used in commercial inoculants in Brazil: one strain originally from Colombia (Rhizobium tropici CIAT 899), and two strains from Brazil (R. tropici H 12 and Rhizobium freirei PRF 81). The inclusion of NiCl2 in the nutrient solution did not increase hydrogenase activity, indicating that common bean plants allow efficient nickel provision for hydrogenase synthesis in the bacteroids. The symbiotic performance—evaluated by nodulation, plant growth, N accumulation and seed production—of wild-type and Hup+ derivative strains was compared in experiments performed with cultivar Carioca under greenhouse conditions, in sterile substrate and in non-sterile soil. Statistically significant increases in one or more parameters were observed for all three Hup+ derivatives when compared to the respective wild-type strain. Differences were found mainly with the Brazilian strains, reaching impressive increases in nodule efficiency and seed total N content. The results highlight the potential of using Rhizobium Hup+ strains for the design of more energy-efficient inoculants for the common bean crop.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biological nitrogen fixation (BNF) is a key biological process for Earth’s N balance. Reduction of atmospheric nitrogen (N2) to ammonia is mediated by the enzyme nitrogenase, in a process that produces hydrogen gas (H2) as an energy-rich obligate by-product that consumes at least 25% of the energy supplied to the nitrogenase. The H2 produced usually diffuses into the surrounding soil, implying in loss of energy that otherwise could be directed to the reduction of N2, thereby affecting the energy efficiency of BNF, and with a great impact on the symbioses of rhizobia with several legumes (Hungria and Neves 1987; Neves and Hungria 1987; Hungria and Ruschel 1989; Baginsky et al. 2005; Imperial et al. 2006; Annan et al. 2012). Some rhizobial strains possess genes coding for another enzyme, a hydrogen-uptake NiFe hydrogenase—Hup—that recycles part of the energy (ATP) spent in the BNF process through the oxidation of the released H2 (Ruiz-Argüeso et al. 1979; Hungria and Neves 1987; Neves and Hungria 1987; Dong and Layzell 2001; McLearn and Dong 2002). Soybean (Glycine max (L.) Merr.), lupines (Lupinus spp.), and cowpea (Vigna unguiculata L.) are among the most well-known legumes capable of forming symbiotic relationships with rhizobia that possess functional hydrogenase-uptake genes (Hungria et al. 1989; Souza et al. 1999; Baginsky et al. 2005; Brito et al. 2005; Annan et al. 2012).
After hydrogen-uptake capacity was demonstrated in legume nodules, symbioses with rhizobia lacking hydrogenase-uptake systems have been viewed as energetically inefficient. However, the superiority of symbiotic performance in rhizobia possessing functional hydrogenases (called H2-uptake positive or Hup+ strains) over those lacking the enzyme (Hup−) reported by some authors (e.g. Schubert and Evans 1976; Albrecht et al. 1979; Hanus et al. 1981; Hungria et al. 1989; Baginsky et al. 2005), has not been conclusive according to others (Drevon et al. 1982; Golding and Dong 2010).
Partnership between rhizobia and common bean (Phaseolus vulgaris L.) is crucial, as this legume represents a basic protein source in many countries of South and Central America, Asia and Africa. However, there are concerns about the capacity of BNF to support common bean plant growth and yield; in general, limitations have been attributed to plant breeding programs focused on N-fertilizer, to highly competitive but poorly effective indigenous population of rhizobia in soils, and to high susceptibility to diseases and environmental stresses (Graham 1981; Buttery et al. 1987; Hungria and Vargas 2000; Dwivedi et al. 2015). Nevertheless, increases in nodulation, BNF and grain yield have been observed when selected elite commercial strains are used as inoculants in Brazil (Hungria et al. 2000, 2003, 2013), even when the strains are Hup− (Ormeño-Orrillo et al. 2012). Besides, it is worth mentioning that it has been long suggested that inoculation with Hup+ rhizobial strains could help to optimize the BNF process with common beans (Hungria and Neves 1987; Neves and Hungria 1987; Hungria and Ruschel 1989).
In this study, Hup+ derivatives of three very effective but Hup−Rhizobium strains used in commercial inoculants for the common bean crop in Brazil were obtained. Afterwards, the symbiotic performance of wild-type and engineered strains was compared in cultivar Carioca, considered as a host genotype with high capacity for BNF.
Materials and methods
Rhizobial strains and generation of hydrogen-recycling derivative strains
Three strains used in commercial inoculants for the common bean crop in Brazil [Rhizobium tropici CIAT 899 (= SEMIA 4077) and H 12 (= SEMIA 4088), and Rhizobium freirei PRF 81 (= SEMIA 4080) (Hungria et al. 2000, 2003)] were selected as targets to improve their nitrogen fixation efficiency by incorporating the ability to recycle hydrogen evolved by nitrogenase. Analysis of genome sequences (available for two out of the three strains (Ormeño-Orrillo et al. 2012) revealed the presence of DNA regions encoding versions of hydrogenase gene clusters inactivated by relevant deletions. Consistent with this, these strains do no synthesize a functional H2-uptake hydrogenase in bean bacteroids (our unpublished results and Fig. 1a). Also, Southern hybridization using a Rhizobium leguminosarum bv. viciae (Rlv) UPM791 hupS probe on EcoRI-digested genomic DNAs from these strains allowed the detection of faint, single bands on each strain corresponding to high molecular weight restriction fragments (20–23 kb, data not shown). To generate strains with H2-recycling ability, a Tn5 derivative mini-transposon (TnHB100) containing an 18-kb H2-uptake (hup) gene cluster from Rhizobium leguminosarum bv. viciae strain UPM791 was transferred by mating into each rhizobial strain as described before (Báscones et al. 2000). Basically, the mini-transposon was introduced into the different rhizobial strains by conjugation using Escherichia coli S17.1 (pTnHB100) as the donor strain and spectinomycin-resistant transconjugants were checked for the presence of the hydrogenase gene cluster by Southern analysis. Transconjugants arose at a frequency of approximately 10−8 per recipient cell; at least 50% of the spectinomycin-resistant transconjugants showed an additional signal, corresponding to a ca. 4-kb EcoRI restriction fragment following hybridization to the hupS probe (data not shown). Using this procedure, R. tropici CIAT 899H19 and PRF 81H1, and R. freirei H 12H9 derivatives incorporating Rlv UPM791 hydrogenase gene cluster were generated. Bacteroid suspensions from these TnHB100-containing strains were evaluated for expression of hydrogenase activity using a hydrogen electrode, as previously reported (Báscones et al. 2000). Three transconjugants per recipient strain were tested and found to display similar levels of hydrogenase activity, as expected from a transposon in which hydrogenase genes have their own symbiotic NifA-dependent promoter. By the end of the screening one derivative strain was chosen for each one of the three parental strains to proceed with the greenhouse experiments. The expression of hydrogenase was further confirmed by immunological detection of the small subunit of the enzyme in bacteroid cell crude extracts using antiserum raised against R. leguminosarum HupS, as previously described by Brito et al. (1994).
Expression of hydrogenase activity in bacteroids induced by Rhizobium strains in common bean nodules. A hydrogenase activity was measured in suspensions of bacteroids obtained from plants grown with regular nutrient solutions (white bars), or in the same solutions supplemented with Ni2+ (85 μM NiCl2, shaded bars). Data are the average of two replicates, and bars indicate standard error; B Immunological detection of hydrogenase small subunit in bacteroid crude extracts of the indicated R. tropici (Rt) or R. freirei (Rf) strains. Bacteroids were obtained from common bean plants grown as described in panel A. 30 μg of protein obtained from bacteroids of the indicated strains were loaded onto each lane. Immunoblots were revealed with antisera raised against R. leguminosarum HupS (diluted 1:100). The position of relevant molecular weight markers (kDa) is shown at the left of the figure. The horizontal arrow at the right denotes the position of HupS-immunoreactive band
We determined the insertion sites for two of these derivative strains: in the case of R. tropici H12H9, TnHB100 was located within a gene putatively encoding a cellulose synthase and 100% identical to R. tropici CIAT899H19_PC05720; in R. freirei PRF 81H1, the insertion was on gene locus RHSP_05501, putatively encoding a dihydroxyacetone kinase. None of these two insertions are in regions whose mutations could be rationally linked to increases in productivity.
All wild-type and derivative strains are deposited at the “Diazotrophic and Plant Growth Promoting Bacteria Culture Collection of Embrapa Soja” (WFCC Collection # 1213, WDCM Collection # 1054), in Londrina, State of Paraná, Brazil, and at the Centro de Biotecnología y Genómica de Plantas (C.B.G.P.), Universidad Politécnica de Madrid, Spain.
Greenhouse experiments in sterile substrate
Wild-type and Hup+ derivative strains were individually grown in modified-yeast mannitol medium (YM) (Hungria et al. 2016), for 5 days, in the dark, at 28 °C. Cultures were adjusted to 109 cells mL−1 by optical density, based on growth curves previously obtained for each strain. Seeds of common bean cultivar Carioca, a good N2-fixing genotype (e.g. Hungria et al. 2000), were surface sterilized and three seeds were sown in each sterilized Leonard jar containing sterile substrate composed of sand and vermiculite (1:2, v/v), as described before (Hungria et al. 2016). Each seed received 1 mL of the corresponding rhizobial culture. Two treatments were included as controls: non-inoculated plants without and with N [applied as 80 mg of N (KNO3) plant−1 week−1]. All jars received sterile N-free nutrient solution (Hungria and Araujo 1994) whenever necessary.
The experiment was performed under greenhouse conditions, with photoperiod of 12/12 h (day/night) and temperature averages of 28.1/21.3 °C (day/night). After 5 days, plants were thinned to two per pot. The experiment was performed in a complete randomized block design with eight replicates.
At flowering, 38 days after emergence (DAE), plants were harvested. Shoots were separated from roots and placed in a forced-air dryer at 65 °C until constant weight was obtained (approximately 72 h). Roots were carefully washed and also placed in the dryer. After 72 h in an air-forced drier at 50 °C nodules were removed from roots and dried again. Nodulation (nodule number and dry weight), shoot dry weight and shoot N concentration (Kjeldahl-based analyzer, Tecator, Sweden) were determined.
Greenhouse experiments in pots containing non-sterile soil
Rhizobial inoculants and seed surface sterilization were performed as described in the previous experiment with sterile substrate. The experiment was performed in pots of 5-kg capacity containing 4 kg of soil. The indigenous population of common bean rhizobia in the soil was estimated as 104 cells g−1 of soil by the most probable number (MPN) counting technique with common bean plants of cultivar Carioca, and the statistical tables of Hungria and Araujo (1994). Soil chemical properties were evaluated as described by Hungria et al. (2013), and the main values were as follows: pH in CaCl2 of 5.3; N, 0.18 g kg−1; C, 2.35 g kg−1, P, 8.2 mg kg−1.
Five seeds were sown per pot and each received 1 mL of inoculant containing 109 cells mL−1 of the corresponding strain. As in the experiment with sterile substrate, non-inoculated controls with and without N-mineral (the same amount as under sterile conditions) were included. The experiment was performed in a complete randomized block design with eight replicates. After 5 days, plants were thinned to two plants per pot. Growth conditions, including light, temperature and nutrient solution were as described in the greenhouse experiment with sterile substrate.
At late maturity stage, plants were harvested to evaluate seed production parameters. Pods were collected, opened and the number of seeds per plant was evaluated. After drying (72 h in an air-forced drier at 50 °C), seed dry weight was determined, following the evaluation of seed N concentration (Kjeldahl-based analyzer, Tecator, Sweden).
Statistical analysis of the greenhouse data
Prior to the statistical analyses data were checked for normality of errors and homogeneity of variances. One-way general linear model ANOVA was used, and when significant difference among treatments was detected, Tukey’s Test was employed for a posteriori multiple comparison of the treatment means. Differences were considered significant at p < 0.05 and p < 0.01. Statistical analyses were performed with the software SAS® 9.4 (SAS Institute, North Caroline, USA).
Results
A mini-transposon (TnHB100) containing an 18-kb H2-uptake (hup) gene cluster from R. leguminosarum bv. viciae (Rlv) strain UPM791 was successfully transferred into three common bean elite rhizobial strains used in commercial inoculants in Brazil, R. tropici strains CIAT 899 and H 12, and R. freirei PRF 81. All three Hup+ derivatives exhibited high levels of hydrogenase activity in common bean bacteroids, suggesting efficient recycling of the H2 evolved by nitrogenase in nodules (Fig. 1a).
R. leguminosarum bv. phaseoli is also a symbiont of common bean, and as in the genetically closely related R. leguminosarum bv. viciae it has been shown that nickel availability to the host plant pea (Pisum sativum L.) limits hydrogenase, by affecting the processing of the hydrogenase structural subunits (Brito et al. 1994), we investigated the effects of including NiCl2 in the nutrient solution. We did not observe any increase in the level of activity of the hydrogenase, indicating that common bean plants allow efficient nickel provision to bacteroids (Fig. 1a). In one case (R. tropici PRF 81H1), the addition of nickel was detrimental for activity, suggesting higher sensitivity to nickel toxicity in this strain.
By using antiserum raised against Rlv HupS (Brito et al. 1994) we verified that immunoblots of extracts from bacteroids from the engineered strains, and not those from wild-type strains, showed immunereactive bands of an estimated size compatible with that of the hydrogenase small subunit (ca. 35 kDa, Fig. 1b).
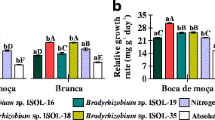
In order to determine the effect of hydrogen recycling in the symbiotic performance, wild-type and Hup+-derivative strains were compared in assays carried out with sterile substrate and also with non-sterile soil as substrate. In the first experiment, performed with sterile substrate, significant increases in parameters related to N2-fixation were associated to the Hup+ trait in one or more of the engineered strains. Differences were observed in nodule number (NN) for R. tropici H 12H9, nodule dry weight (NDW) for R. tropici CIAT 899H19, shoot N concentration (SNC) for CIAT 899H19 and R. freirei PRF 81H1, shoot total N content (STNC) for PRF 81H1 and nodule efficiency (NE) for H 12H9 and PRF 81H1 (Table 1). Remarkable responses were observed mainly with plants inoculated with the Brazilian elite strains H 12H9 and PRF 81H1, with increases in NE of 27.5 and 23.2%, respectively, in comparison to the respective wild-type strains. Noticeable also was the increase in NN of 45.2% with strain R. tropici H 12H9 (Table 1).
In the experiment performed using as substrate non-sterile soil, the Hup+ trait also resulted in statistically significant increases in seed production parameters in plants inoculated with the Brazilian strains R. tropici H 12H9 and R. freirei PRF 81H1, but not with the Colombian strain CIAT 899H19 (Table 2). With both Brazilian strains significant increases were observed in seed dry weight (SeDW) and seed total N content (SeTNC). Impressively, R. tropici H 12H9 Hup+ strain increased SeTNC by 27.1% as compared to the wild type (Table 2).
Discussion
Brazil is amongst the world’s greatest producers and consumers of common bean, and in the country the grains represent the main source of protein for the lower income population; nevertheless, National mean yield is very low when compared to the genetic potential of the cultivars released (SEAB 2017), and a similar situation is reported in other countries. As N is usually the most limiting nutrient in soils used to cultivate common bean in South and Central America, Africa and Asia, improvements in the contribution of BNF could result in great impact and increased grain yields. However, under field conditions it has long been a general idea that the legume establishes inefficient N2-fixing symbioses (Graham 1981; Buttery et al. 1987; van Kessel and Hartley 2000). Besides, it has also been shown that it is feasible to successfully survey elite strains with higher capacity of BNF and competitiveness, greatly improving grain yield (Hungria et al. 2000, 2003; Mulas et al. 2011; Vanlauwe et al. 2019). Intriguingly, although the hydrogen-uptake property has been subject of intensive studies and promises of increased yields in the 1970s and 1980s (e.g. Schubert and Evans 1976; Hungria and Neves 1987), it has fallen into a relative ostracism in the following decades. Yet, several studies have shown that effective hydrogen-uptake rhizobial strains can improve the efficiency of the BNF, plant performance and yield (e.g. Schubert and Evans 1976; Albrecht et al. 1981; Hanus et al. 1981; Hungria and Neves 1987; Hungria et al. 1989; Baginsky et al. 2005).
The hydrogen-uptake activity is variable among rhizobia (Bedmar and Phillips 1984; Bedmar et al. 1984; Golding and Dong 2010; Imperial et al., 2006) and, apparently, it is less frequently observed in rhizobia nodulating temperate legumes (Drevon et al. 1982; Golding and Dong 2010). In addition, although hydrogenase is a bacterial enzyme, it has been noted that the host plant may exert an influence on its expression as reported for common bean (Navarro et al. 1993), lupines (Murillo et al. 1989), lotus (Lotus corniculatus L.; Monza et al. 1997), pea (Bedmar and Phillips 1984) and lentil (Lens culinaris; Brito et al. 2008) symbioses.
In a pioneer study of hydrogenases of common bean microsymbionts, both CIAT 899 and CFN 299 were positive for the hybridization with a probe containing genes of B. japonicum, but both had Hup− phenotypes in nodules of 30-day-old plants (van Berkum et al. 1994). The sequencing of the genomes of R. tropici CIAT 899 and strain PRF 81 (Ormeño-Orrillo et al. 2012), this last one now reclassified as R. freirei (Dall’Agnol et al. 2013), as well as of strain CFN 299 (Ormeño-Orrillo et al. 2016), now reclassified as R. leucaenae (Ribeiro et al. 2012), and also of R. tropici H 12 (data not published) has shown that all harbor truncated genes in the hydrogenase operons. These genetic defects are consistent with the lack of hydrogenase activity observed in the wild-type strains in the present study.
In R. leguminosarum, hup and hyp genes are organized as hupSLCDEFGHIJKhypABFCDEX and result in a functional hydrogenase (Brito et al. 2005; Imperial et al. 2006). A method of transferring the capacity of oxidation of H2 from R. leguminosarum to other rhizobial strains by the introduction of a mini-transposon has been developed (Báscones et al. 2000), and in this study it was used to transfer these genes to three very effective but Hup− strains used in Brazilian commercial inoculants, CIAT 899, PRF 81 and H 12. The Hup+ phenotype was confirmed, allowing a proper comparison of Hup− and Hup+ symbiotic performances. In all three strains the introduction of the Hup mini-tranposon TnHB100 resulted in high levels of hydrogenase activity in bacteroids, and the activity was not increased in the presence of extra levels of nickel in the plant nutrient solution. Similar results have been previously observed in bacteroids induced by R. etli CFN 42 in common bean of cultivar Negro Jamapa (Brito et al. 2000). In contrast, R. leguminosarum bacteroids induced in pea nodules require high levels of nickel in the nutrient solution to attain maximum levels of hydrogenase activity and enzyme processing levels (Brito et al. 1994). These data indicate that P. vulgaris plants are able to efficiently provide nickel to bacteroids for hydrogenase synthesis.
When symbiotic parameters were compared in the wild-type and Hup+ derivative strains, the results have shown that the Hup+ trait, in general, resulted in significant increases in BNF and seed production parameters. CIAT 899 was isolated in Colombia, while H 12 and PRF 81 are indigenous to Brazil, and the results were more prominent with the Brazilian strains. Outstanding increases were shown in nodule number and nodule efficiency, of 45.2 and 27.5%, respectively, with the H 12-derivative Hup+ strain. Our results with common bean are in agreement with studies comparing Hup+ and Hup− strains in cowpea (Souza et al. 1999; Baginsky et al. 2005) and soybean (Hungria et al. 1989). Most importantly, parameters related to seed production, including seed dry weight and seed total N content were also significantly improved by the Hup+ trait in plants inoculated with the two Brazilian derivative strains.
The synthesis of inorganic N-fertilizers requires large quantities of fossil fuel and its application results in the release of nitrous oxide from soil as well in contamination of water reservoirs; therefore, improving BNF efficiency and reducing the use of N-fertilizers would also help to alleviate the environmental N footprint by decreasing the need for N fertilizers (Dong et al. 2003; Golding and Dong 2010; Hungria et al. 2013; Sá et al. 2017). In this context, improving rhizobial strains for use as inoculants with legume crops by means of properly engineering Hup+ rhizobial strains might result in superior symbiotic performance (Kent et al. 1998; Golding and Dong 2010). In the engineering process described here, strains have incorporated an 18-kb hydrogenase gene cluster obtained from the symbiotic plasmid of R. leguminosarum UPM791, along with a resistance marker (spectinomycin). Removal of this marker is feasible (Báscones et al. 2000), thus resulting in modified strains containing only rhizobial DNA. Our results highlight the biotechnological potential of using Hup+ strains for the improvement of BNF with common bean.
References
Albrecht SL, Maier RJ, Hanus FJ, Russell SA, Emerich DW, Evans HJ (1979) Hydrogenase in Rhizobium japonicum increases nitrogen fixation by nodulated soybeans. Science 203:1255–1257. https://doi.org/10.1126/science.203.4386.1255
Albrecht SL, Okon Y, Lonnquist J, Burris RH (1981) Nitrogen fixation by corn-Azospirillum associations in a temperate climate. Crop Sci 21:301–306. https://doi.org/10.2135/cropsci1981.0011183X002100020024x
Annan H, Golding A-L, Zhao Y, Dong Z (2012) Choice of hydrogen uptake (Hup) status in legume-rhizobia symbioses. Ecol Evol 2:2285–2290. https://doi.org/10.1002/ece3.325
Baginsky C, Brito B, Imperial J, Ruiz Argüeso T, Palacios JM (2005) Symbiotic hydrogenase activity in Bradyrhizobium sp. (Vigna) increases nitrogen content in Vigna unguiculata plants. Appl Environ Microbiol 71:7536–7538. https://doi.org/10.1128/AEM.71.11.7536-7538.2005
Báscones E, Imperial J, Ruiz-Argueso T, Palacios JM (2000) Generation of new hydrogen-recycling Rhizobiaceae strains by introduction of a novel hup minitransposon. Appl Environ Microbiol 66:4292–4299. https://doi.org/10.1128/aem.66.10.4292-4299.2000
Bedmar EJ, Phillips DA (1984) A transmissible plant shoot factor promotes uptake hydrogenase activity in Rhizobium symbionts. Plant Physiol 75:629–633. https://doi.org/10.1104/pp.75.3.629
Bedmar EJ, Brewin NJ, Phillips DA (1984) Effect of plasmid pIJ1008 from Rhizobium leguminosarum on symbiotic function of Rhizobium meliloti. Appl Environ Microbiol 47:876–878
Brito B, Palacios JM, Hidalgo E, Imperial J, Ruiz-Argüeso T (1994) Nickel availability to pea (Pisum sativum L.) plants limits hydrogenase activity of Rhizobium leguminosarum bv. viciae bacteroids by affecting the processing of the hydrogenase structural subunits. J Bacteriol 176:5297–5303. https://doi.org/10.1128/jb.176.17.5297-5303.1994
Brito B, Monza J, Imperial J, Ruiz-Argüeso T, Palacios JM (2000) Nickel availability and hupSL activation by heterologous regulators limit symbiotic expression of the Rhizobium leguminosarum bv viciae hydrogenase system in Hup− rhizobia. Appl Environ Microbiol 66:937–942. https://doi.org/10.1128/aem.66.3.937-942.2000
Brito B, Baginsky C, Palacios JM, Cabrera E, Ruiz-Argüeso T, Imperial J (2005) Biodiversity of uptake hydrogenase systems from legume endosymbiotic bacteria. Biochem Soc Trans 33:33–35. https://doi.org/10.1042/BST0330033
Brito B, Toffanin A, Prieto RI, Imperial J, Ruiz-Argüeso T, Palacios JM (2008) Host-dependent expression of Rhizobium leguminosarum bv. viciae hydrogenase is controlled at transcriptional and post-transcriptional levels in legume nodules. Mol Plant Microbe Interact 21:597–604. https://doi.org/10.1094/MPMI-21-5-0597
Buttery B, Park S, Findlay W (1987) Growth and yield of white bean (Phaseolus vulgaris L.) in response to nitrogen, phosphorus and potassium fertilizer and to inoculation with Rhizobium. Can J Plant Sci 67(2):425–432. https://doi.org/10.4141/cjps87-061
Dall’Agnol RF, Ribeiro RA, Ormeño-Orrillo E, Rogel MA, Delamuta JRM, Andrade DS, Martínez-Romero E, Hungria M (2013) Rhizobium freirei, a symbiont of Phaseolus vulgaris very effective in fixing nitrogen. Int J Syst Evol Microbiol 63:4167–4173. https://doi.org/10.1099/ijs.0.052928-0
Dong Z, Layzell D (2001) H2 oxidation, O2 uptake and CO2 fixation in hydrogen treated soils. Plant Soil 229:1–12. https://doi.org/10.1023/A:1004810017490
Dong Z, Wu L, Kettlewell B, Caldwell CD, Layzell DB (2003) Hydrogen fertilization of soils—is this a benefit of legumes in rotation? Plant, Cell Environ 26:1875–1879. https://doi.org/10.1046/j.1365-3040.2003.01103.x
Drevon JJ, Frazier L, Russell SA, Evans HJ (1982) Respiratory and nitrogenase activities of soybean nodules formed by hydrogen uptake negative (Hup−) mutant and revertant strains of Rhizobium japonicum characterized by protein patterns. Plant Physiol 70:1341–1346. https://doi.org/10.1104/pp.70.5.1341
Dwivedi SL, Sahrawat KL, Upadhyaya HD, Mengoni A, Galardini M, Bazzicalupo M, Biondi EG, Hungria M, Kaschuk G, Blair MW, Ortiz R (2015) Advances in host plant and Rhizobium genomics to enhance symbiotic nitrogen fixation in grain legumes. In: Sparks DL (ed) Adv Agron 129:1–116. Elsevier Inc, Academic Press. https://doi.org/10.1016/bs.agron.2014.09.001
Golding AL, Dong Z (2010) Hydrogen production by nitrogenase as a potential crop rotation benefit. Environ Chem Lett 8:101–121. https://doi.org/10.1007/s10311-010-0278-y
Graham PH (1981) Some problems of nodulation and symbiotic nitrogen fixation in Phaseolus vulgaris L. Field Crop Res 4:93–112. https://doi.org/10.1016/0378-4290(81)90060-5
Hanus FJ, Albrecht SL, Zablotowicz RM, Emerich DW, Russell SA, Evans HJ (1981) Yield and N content of soybean seed as influenced by Rhizobium japonicum inoculants possessing the hydrogenase characteristic. Agron J 73:368–372. https://doi.org/10.2134/agronj1981.00021962007300020028x
Hungria M, Araujo RS (eds) (1994) Manual de Métodos Empregados em Estudos de Microbiologia Agrícola. EMBRAPA-SPI, Brasília, Brazil. ISSN 0101-9716. https://doi.org/10.13140/RG.2.1.2663.4727
Hungria M, Neves MCP (1987) Cultivar and Rhizobium strain effect on nitrogen fixation and transport in Phaseolus vulgaris L. Plant Soil 103:111–121. https://doi.org/10.1007/BF02370675
Hungria M, Ruschel AP (1989) Acetylene reduction, hydrogen evolution and nodule respiration in Phaseolus vulgaris. Biol Fertil Soils 7:351–358. https://doi.org/10.1007/BF00257832
Hungria M, Vargas MAT (2000) Environmental factors affecting N2 fixation in grain legumes in the tropics, with an emphasis on Brazil. Field Crop Res 65:151–164. https://doi.org/10.1016/S0378-4290(99)00084-2
Hungria M, Neves MCP, Döbereiner J (1989) Relative efficiency, ureide transport and harvest index in soybeans inoculated with isogenic HUP mutants of Bradyrhizobium japonicum. Biol Fertil Soils 7:325–329. https://doi.org/10.1007/BF00257827
Hungria M, Andrade DS, Chueire LMO, Probanza A, Guttierrez-Mañero FJ, Megías M (2000) Isolation and characterization of new efficient and competitive bean (Phaseolus vulgaris L.) rhizobia from Brazil. Soil Biol Biochem 32:1515–1528. https://doi.org/10.1016/S0038-0717(00)00063-8
Hungria M, Campo RJ, Mendes IC (2003) Benefits of inoculation of the common bean (Phaseolus vulgaris) crop with efficient and competitive Rhizobium tropici strains. Biol Fertil Soils 39:88–93. https://doi.org/10.1007/s00374-003-0682-6
Hungria M, Nogueira MA, Araujo RS (2013) Co-inoculation of soybeans and common beans with rhizobia and azospirilla: strategies to improve sustainability. Biol Fertil Soils 49:791–801. https://doi.org/10.1007/s00374-012-0771-5
Hungria M, O’Hara GW, Zilli JE, Araujo RS, Deaker R, Howieson JG (2016) Isolation and growth of rhizobia. In: Howieson JG, Dilworth MJ (eds) Working with Rhizobia. ACIAR, Canberra, pp 39–60. ISBN 978 1 925436 18 1
Imperial J, Palacios JM, Brito B, Rey I, Cabrera E, Ureta AC, Ruiz-Argüeso T (2006) El sistema hidrogenasa de las bacterias que nodulan leguminosas. In: Bedmar EJ, González J, Lluch C, Rodelas B (eds) Fijación de Nitrógeno: Fundamentos y Aplicaciones. SEFIN, Granada, pp 35–44. ISBN 84-611-1198-5
Kent AD, Wojtasiak ML, Robleto AE, Triplett EW (1998) A transposable partitioning locus used to stabilize plasmid-borne hydrogen oxidation and trifolitoxin production genes in a Sinorhizobium strain. Appl Environ Microbiol 64:1657–1662
McLearn N, Dong Z (2002) Microbial nature of the hydrogen-oxidizing agent in hydrogen-treated soil. Biol Fertil Soils 35:465–469. https://doi.org/10.1007/s00374-002-0495-z
Monza J, Diaz P, Borsani O, Ruiz-Argüeso T, Palacios JM (1997) Evaluation and improvement of the energy efficiency of nitrogen fixation in Lotus corniculatus nodules induced by Rhizobium loti strains indigenous to Uruguay. World J Microbiol Biotechnol 13:565–571. https://doi.org/10.1023/A:1018573527503
Mulas D, García-Fraile P, Carro L, Ramírez-Bahena M-H, Casquero P, Velázquez E, González-Andrés F (2011) Distribution and efficiency of Rhizobium leguminosarum strains nodulating Phaseolus vulgaris in Northern Spanish soils: selection of native strains that replace conventional N fertilization. Soil Biol Biochem 43(11):2283–2293. https://doi.org/10.1016/j.soilbio.2011.07.018
Murillo J, Villa A, Chamber M, Ruiz-Argüeso T (1989) Occurrence of H2-uptake hydrogenases in Bradyrhizobium sp. (Lupinus) and their expression in nodules of Lupinus spp. and Ornithopus compressus. Plant Physiol 89:78–85. https://doi.org/10.1104/pp.89.1.7
Navarro RB, Vargas AAT, Schröder EC, van Berkum P (1993) Uptake hydrogenase (Hup) in common bean (Phaseolus vulgaris) symbioses. Appl Environ Microbiol 59:4161–4165
Neves MCP, Hungria M (1987) The physiology of nitrogen fixation in tropical grain legumes. CRC Crit Rev Plant Sci 6:267–321. https://doi.org/10.1080/07352688709382252
Ormeño-Orrillo E, Menna P, Almeida LGP, Ollero FJ, Nicolás MF, Rodrigues EP, Nakatami AS, Batista JSS, Chueire LMO, Souza RC, Vasconcelos ATR, Megías M, Hungria M, Martínez-Romero E (2012) Genomic basis of broad host range and environmental adaptability of Rhizobium tropici CIAT 899 and Rhizobium sp. PRF 81 which are used in inoculants for common bean (Phaseolus vulgaris L.). BMC Genom 13:735. https://doi.org/10.1186/1471-2164-13-735
Ormeño-Orrillo E, Gomes DF, del Cerro P, Vasconcelos ATR, Canchaya C, Almeida LGP, Mercante FM, Ollero FJ, Megías M, Hungria M (2016) Genome of Rhizobium leucaenae strains CFN 299T and CPAO 29.8: searching for genes related to a successful symbiotic performance under stressful conditions. BMC Genom 17:534. https://doi.org/10.1186/s12864-016-2859-z
Ribeiro RA, Rogel MA, López-López A, Ormeño-Orrillo E, Barcellos FG, Martínez J, Thompson FL, Martìnez-Romero E, Hungria M (2012) Reclassification of Rhizobium tropici type A strains as Rhizobium leucaenae sp. nov. Int J System Evol Microbiol 62(5):1180–1185. https://doi.org/10.1099/ijs.0.032912-0
Ruiz-Argüeso T, Emerich DW, Evans HJ (1979) Hydrogenase system in legume nodules: a mechanism of providing nitrogenase with energy and protection from oxygen damage. Biochem Biophys Res Commun 86:259–264. https://doi.org/10.1016/0006-291x(79)90860-x
Sá JCM, Lal R, Cerri CC, Lorenz K, Hungria M, Carvalho PCC (2017) Low-carbon agriculture in South America to mitigate global climate change and advance food security. Environ Int 98:102–112. https://doi.org/10.1016/j.envint.2016.10.020
Schubert KR, Evans HJ (1976) Hydrogen evolution: a major factor affecting the efficiency of nitrogen fixation in nodulated symbionts. Proc Natl Acad Sci USA 73:1207–1211. https://doi.org/10.1073/pnas.73.4.1207
SEAB (Secretaria de Estado da Agricultura e do Abastecimento DERAL - Departamento de Economia Rural) (2017) Feijão - Análise da Conjuntura Agropecuária. http://www.agricultura.pr.gov.br/arquivos/File/deral/Prognosticos/2018/_feijao_2017_18.pdf. Accessed 22 Apr 2018
Souza AA, Burity HA, Figueiredo MVB, Da Silva MLRB, Melotto M, Tsai SM (1999) Eficiência simbiótica de estirpes Hup+, HupHR e Hup− de Bradyrhizobium japonicum e Bradyrhizobium elkanii em cultivares de caupi. Pesq Agropec Bras 34:1925–1931. https://doi.org/10.1590/S0100-204X1999001000020
van Berkum P, Navarro RB, Vargas AAT (1994) Classification of the uptake hydrogenase-positive (Hup+) bean rhizobia as Rhizobium tropici. Appl Environ Microbiol 60:554–561. https://aem.asm.org/content/60/2/554
van Kessel C, Hartley C (2000) Agricultural management of grain legumes; has it led to an increase in nitrogen fixation? Field Crops Res 65:165–181. https://doi.org/10.1016/S0378-4290(99)00085-4
Vanlauwe B, Hungria M, Kanampiu F, Giller KE (2019) The role of legumes in the sustainable intensification of African smallholder agriculture: lessons learnt and challenges for the future. Agric Ecosyst Environ 284:106583. https://doi.org/10.1016/j.agee.2019.106583
Acknowledgements
A.R. Torres acknowledges a fellowship from CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico, Brazil), and is currently a post-doctoral researcher at Kansas State University. M. Hungria is also a research fellow of CNPq.
Funding
The study was supported by Comunidad de Madrid research project P-AMB-321-0505 (to T.R.A) and by UPM (Convocatoria Proyectos con América Latina, to B.B.). Today the Brazilian group is funded by the INCT-Plant-Growth Promoting Microorganisms for Agricultural Sustainability and Environmental Responsibility (CNPq 465133/2014-2, Fundação Araucária-STI, CAPES).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: TR-A, JI, JMP, MH. Performed the experiments: ART, BB, JI, JMP, TR-A, MH. Analyzed the data: ART, BB, JI, JMP, TR-A, MH. Contributed reagents/materials/analysis tools: TR-A, MH. Wrote the paper: ART, JI, JMP, IAC, TR-A, MH. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval and consent to participate
The authors declare no ethical conflicts; authors declare that they have consented to participate in the manuscript and publish it.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Torres, A.R., Brito, B., Imperial, J. et al. Hydrogen-uptake genes improve symbiotic efficiency in common beans (Phaseolus vulgaris L.). Antonie van Leeuwenhoek 113, 687–696 (2020). https://doi.org/10.1007/s10482-019-01381-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-019-01381-6