Abstract
The Cerrado biome in the Sete Cidades National Park, an Ecological Reserve in Northeastern Brazil, has conserved its native biodiversity and presents a variety of plants found in other savannas in Brazil. Despite this finding the soil microbial diversity and community structure are poorly understood. Therefore, we described soil bacterial diversity and distribution along a savanna vegetation gradient taking into account the prevailing environmental factors. The bacterial composition was retrieved by sequencing a fragment of the 16S ribosomal RNA gene. The bacterial operational taxonomic units (OTUs) were assigned to 37 different phyla, 96 classes, and 83 genera. At the phylum level, a core comprised by Proteobacteria, Acidobacteria, Actinobacteria, Firmicutes, Verrucomicrobia and Planctomycetes, was detected in all areas of Cerrado. ‘Cerrado stricto sensu’ and ‘Cerradao’ share more similarities between edaphic properties and vegetation and also present more similar bacterial communities, while ‘Floresta decidual’ and ‘Campo graminoide’ show the largest environmental differences and also more distinct bacterial communities. Proteobacteria (26%), Acidobacteria (21%) and Actinobacteria (21%) were the most abundant phyla within the four areas. All the samples present similar bacteria richness (alpha diversity) and the observed differences among them (beta diversity) were more related to the abundance of specific taxon OTUs compared to their presence or absence. Total organic C, N and P are the main abiotic factors structuring the bacterial communities. In summary, our findings show the bacterial community structure was clearly different across the Cerrado gradient, but that these environments share a bacterial phylum-core comprising Proteobacteria, Acidobacteria, Actinobacteria, Verrucomicrobia and Planctomycetes with other Brazilian savannas.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The Brazilian Cerrado (Tropical Savanna), as the second largest Brazilian biome after the Amazon rainforest, is an ecosystem that contains a high biodiversity (Forzza et al. 2010). The biome occupies a quarter of the Brazilian territory and extends across almost the entire country, making it one of the most biodiverse savannas in the world (Myers et al. 2000). In the Cerrado from the Northeast of Brazil, the Federal Government established the Sete Cidades National Park, which covers 6221 ha and aims to study and protect the region’s biodiversity (IBDF 1979). In this National Park, the Cerrado comprises a gradient of vegetation from ‘Campo graminoide’ (continuous grass stratum), ‘Cerrado stricto sensu’ (grass, shrubs, low trees and woody stratum) ‘Cerradao’ (woody stratum with varying density of shrubs and trees) to ‘Floresta decidual’ (trees stratum) (Coutinho 1978).
Although other regions of Brazil also present similar gradients of Cerrado vegetation, such as the Southeast and Central plateau (Castro et al. 1999), Castro et al. (1998) hypothesized that the Cerrado from Sete Cidades National Park could be a hotspot of biodiversity and could have different patterns of plant diversity. Therefore, a Long-Term Ecological Program (PELD) was initiated in 1994 to study the plant diversity within Sete Cidades National Park, and the major finding was that different plant diversities exist across a gradient of that Cerrado (Castro et al. 1998, 1999).
Despite the importance of Cerrado biodiversity, inventories of soil microbial diversity using high-throughput sequencing technologies are scarce and concentrated in Southeast and Central plateau Cerrado (Araujo et al. 2012; Rampelotto et al. 2013; Castro et al. 2016). Therefore, information about soil microbial diversity across the Cerrado in the Northeast of Brazil is still lacking.
Microbes are the largest reservoir of biodiversity in natural forests and are involved in nutrient transformations that maintain the forest ecosystem (DeMandal et al. 2015). Obtaining further information regarding soil microbial diversity is important because soil microorganisms are involved in vital ecosystem functions (Rodrigues et al. 2013). Sequencing 16S rRNA genes that have been recovered from the environment is the most frequently used molecular method for evaluating bacterial communities and has been widely used to describe bacterial diversity in different ecosystems (Zhang and Xu 2008; Mishra et al. 2014).
Soil bacterial communities are variable across spatial scales, and several factors, such as soil physicochemical properties and plant diversity, drive the distribution of microorganisms in the environment (Bru et al. 2011). Specifically, plant diversity influences the bacterial community both directly through carbon sources in plant litter and indirectly through its influence on soil physicochemical properties (Bardgett and Shine 1999). Previous studies in Brazilian Central Cerrado have assessed the microbial community structure along the gradient of vegetation and revealed significant influences of vegetation and physicochemical properties (Araujo et al. 2012; Rampelotto et al. 2013; Castro et al. 2016). Araújo et al. (2012) found that bacterial communities from ‘Cerrado Denso’ and ‘Cerrado sensu stricto’ grouped together and are distinct from those of ‘Campo Sujo’ and ‘Mata de Galeria’. Those authors found that the dominant bacterial phylum in all Cerrado habitats was Acidobacteria. Castro et al. (2016) showed that changes in bacterial, archaeal, and fungal community structures in Central Cerrado are strongly correlated with seasonal patterns of soil water uptake. Rampelotto et al. (2013) observed relevant differences in the abundance and structure of bacterial communities in Cerrado soils under different land use systems. Taking into account the peculiarities of the edaphoclimatic conditions and the Cerrado plant composition and diversity found within the Sete Cidades National Park (Castro et al. 1998, 1999), we hypothesize that these environmental variations in soils are reflected in differences in the bacterial community composition.
Materials and methods
Study area
The study was conducted within Sete Cidades National Park (PNSC) (04°02′–08′S and 41°40′–45′W), located in the northeastern state of Piauí, Brazil (Fig. 1). The park covers an area of 6221 ha. The climate is sub-humid with two distinct seasons (wet and dry) during the year, and annual average of temperature at 25 °C. The area has an annual average rainfall of 1558 mm distributed in February, March and April.
We evaluated preserved sites (with 1000 m2 each one), belonging to the long-term ecological program (PELD-CNPq) from the Brazilian government, across a gradient of different Cerrado formations ranging from ‘Campo graminoide’, ‘Cerrado stricto sensu’, ‘Cerradao’ to ‘Floresta decidual’ (Table 1). Basically, ‘Campo graminoide’ is covered by a continuous grass stratum; ‘Cerrado stricto sensu’ is covered by grass, shrubs, low trees and woody stratum; ‘Cerradao’ is covered by woody stratum with varying density of shrubs and trees; and ‘Floresta decidual’ is covered by trees (Coutinho 1978).
Soil sampling and chemical analysis
Each site was divided in three transects (replication) where soil samples were collected at 0–20 cm depth (three points per transect) in March (wet season), 2014. All soil samples were immediately stored in sealed plastic bags and transported in an icebox to the laboratory. A portion of the soil samples was stored in bags and kept at −20 °C for DNA analysis and another portion was air-dried, sieved through a 2 mm screen and homogenized for chemical analyses.
Soil chemical properties were determined and measured using standard laboratory protocols (Tedesco et al. 1995). Soil pH was determined in a 1:2.5 soil/water extract. Available P and exchangeable K+ were extracted using Mehlich-1 extraction method and determined by colorimetry and photometry, respectively (Table 2). Total organic C (TOC) and total N were determined by the wet combustion method using a mixture of potassium dichromate and sulfuric acid under heating (Yeomans and Bremner 1988).
DNA extraction and library preparation
Soil DNA was extracted from 0.5 g (total humid weight) of soil using the PowerLyzer PowerSoil DNA Isolation Kit (MoBIO Laboratories, Carlsbad, CA, USA), according to the manufacturer’s instructions. The DNA extraction was performed in triplicate for each soil sample. The quality and concentration of the extracted DNA was estimated using a Nanodrop 1000 (Thermo Scientific, Waltham, MA, USA).
The amplicon library of the 16S rRNA gene V4 region was prepared as previously described (Illumina 2013), using the region-specific primers (515F/806R) (Caporaso et al. 2011). First step amplification comprised 25 μL reaction containing the following: 14.8 μL of nuclease-free water (Certified Nuclease-free, Promega, Madison, WI, USA), 2.5 μL of 10× High Fidelity PCR Buffer (Invitrogen, Carlsbad, CA, USA), 1.0 μL of 50 mM MgSO4, 0.5 μL of each primer (10 μM concentration, 200 pM final concentration), 1.0 unit of Platinum Taq polymerase High Fidelity (Invitrogen, Carlsbad, CA, USA), and 4.0 μL of template DNA (10 ng). The conditions for PCR were as follows: 94 °C for 4 min to denature the DNA, with 25 cycles at 94 °C for 45 s, 60 °C for 60 s, and 72 °C for 2 min, with a final extension of 10 min at 72 °C. In the second step, a unique pair of Illumina Nextera XT indexes (Illumina, San Diego, CA) was added to both ends of the amplified products. Each 50 μL reaction contained the following: 23.5 uL of nuclease-free water (Certified Nuclease-free, Promega, Madison, WI, USA), 5.0 uL of 10× High Fidelity PCR Buffer (Invitrogen, Carlsbad, CA, USA), 4.8 uL of 25 mm MgSO4, 1.5 uL of dNTP (10 mm each), 5.0 uL of each Nextera XT index (Illumina, San Diego, CA, USA), 1.0 unit of Platinum Taq polymerase High Fidelity (Invitrogen, Carlsbad, CA, USA), and 5.0 uL of each product from previous PCR. The conditions for this second round PCR were as follows: 95 °C for 3 min to denature the DNA, with eight cycles at 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 min, with a final extension of 5 min at 72 °C.
After indexing, the PCR products were cleaned up using Agencourt AMPure XP—PCR purification beads (Beckman Coulter, Brea, CA, USA), according to the manufacturer’s manual, and quantified using the dsDNA BR assay Kit (Invitrogen, Carlsbad, CA, USA) on a Qubit 2.0 fluorometer (Invitrogen, Carlsbad, CA, USA). Once quantified, different volumes of each library were pooled into a single tube such that each sample was represented equally. After quantification, the molarity of the pool was determined and diluted to 2 nM, denatured, and then diluted to a final concentration of 8.0 pM with a 20% PhiX (Illumina, San Diego, CA, USA) spike for loading into the Illumina MiSeq sequencing machine (Illumina, San Diego, CA, USA).
Processing and analysis of sequencing data
Sequence data were processed using QIIME (Caporaso et al. 2010a) following the UPARSE standard pipeline (Edgar 2013) according to Brazilian Microbiome Project (Pylro et al. 2014, 2016) to produce an OTU table and a set of representative sequences. Briefly, the reads were truncated at 240 bp and quality-filtered using a maximum expected error value of 0.5. Pre-filtered reads were dereplicated and singletons were removed and filtered for additional chimeras using the RDP_gold database using USEARCH 7.0 (Edgar et al. 2011). These sequences were clustered into OTUs at a 97% similarity cutoff following the UPARSE pipeline. After clustering, the sequences were aligned and taxonomically classified against the Greengenes database (version 13.8) (DeSantis et al. 2006) using the PyNAST algorithm (Caporaso et al. 2010b). The sequences were submitted to the NCBI Sequence Read Archive under the number SRP091586.
The strategy of rarefaction (random sub-sampling) was used to normalize to 7000 sequences per sample and evaluate the sequencing effort. Good’s index was calculated to estimate the coverage reached using the rarefaction level chosen. Bacterial diversity was evaluated using observed operational taxonomic units (OTUs), Faith’s PD (phylogenetic diversity), Chao1 and Shannon as alpha diversity metrics. Beta diversity matrices were generated using weighted and unweighted UniFrac phylogenetic distances (Lozupone and Knight 2005) and evaluated by bi-dimensional Principal Coordinates Analysis (PCoA).
The relationships between environmental variables and community OTUs were assessed using redundancy analysis (RDA). Environmental variables were log (x + 1) transformed, except pH. Previously, “decorana” function was performed in order to choose the best ordination model, and this analysis supported the use of RDA. A preliminary RDA (“rda” function) was performed with all available environmental variables: Moisture, TOC, N, P, K, CEC, and pH. Significant environmental variables (p < 0.05) selected with “ordistep” function and confirmed by “anova.cca” were retained to final RDA. Variation inflation factor (VIF) (“vif.cca” function) demonstrated that there was not collinearity among the explanatory variables (VIF < 10). Marginal significance of the remaining variables was tested with 999 permutations (“Anova.cca” function). The adjusted R values (“RsquareAdj” function) were computed to obtain the explanatory power of the final RDA. All analyses were carried out in the R environment (https://www.r-project.org/) using vegan package (Oksanen et al. 2015).
Results
The physicochemical properties of the soils are presented in Table 2. The acidity appeared as the most striking feature of these soils. Moisture and TOC were the most contrasting variables between ‘Campo graminoide’ and ‘Floresta decidual’, whereas ‘Campo graminoide’, ‘Cerrado stricto sensu’ and ‘Cerradao’ shared similar contents of N, P, K, and CEC. In general, ‘Cerrado stricto sensu’ and ‘Cerradao’ were more similar to each other, while ‘Campo graminoide’ and ‘Floresta decidual’ were different from all the others.
Regarding the composition and distribution of bacterial communities, a total of 977,191 reads were obtained from 36 soil samples and were clustered in 4721 OTUs. The analysis of the sequences showed a total of 37 phyla, 96 classes, and 83 genera within the four Cerrado areas. Nine most abundant phyla (Proteobacteria, Acidobacteria, Actinobacteria, Firmicutes, Verrucomicrobia, Planctomycetes, Chloroflexi and the candidatus phyla WPS-2 and AD3), with percentages of OTUs above 1%, represented more than 90% of the total OTUs. Among the most abundant phyla, six (Proteobacteria, Acidobacteria, Actinobacteria, Firmicutes, Verrucomicrobia and Planctomycetes) were shared by the four Cerrado areas.
Unclassified OTUs accounted for 5.42% of the total dataset. Twenty-nine phyla (Bacteroidetes, Chlamydiae, AD3, Cyanobacteria, TM6, Elusimicrobia, Gammatimonadetes, Armatimonadetes, Tenericutes, Chlorobi, TM7, OD1, GAL15, OP3, FCPU426, Spirochaetes, BHI80-139, Nitrospirae, Synergistetes, Deferribacteres, SR1, Caldithrix, SBR1093, OP11, WS3, Fibrobacteres, BRC1, GN04, and Lentisphaerae) were in low abundance, with percentages below 1% (Fig. S1).
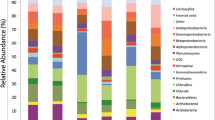
The relative abundance of dominant phyla varied through the different sites (Fig. 2). Proteobacteria were dominant in ‘Cerrado stricto sensu’ and ‘Cerradao’, Acidobacteria were dominant in ‘Campo Graminoide’ and Proteobacteria and Actinobacteria were more abundant in ‘Floresta decidual’.
Relative abundance (greater than 1%) of bacterial phyla from each soil library derived from Cerrado areas. a ‘Campo graminoide’; b ‘Cerrado stricto sensu’; c ‘Cerradao’; d ‘Floresta decidual’. Bars standard error. Different letters indicate statistically significant differences between groups (p < 0.01)
The relative abundance of the fifteen most abundant (above 1%) bacterial classes is presented in Fig. 3. Eleven classes (Alphaproteobacteria, Acidobacteria, Thermoleophilia, Solibacteres, Bacilli, DA052, Planctomycetia, Actinobacteria, Deltaproteobacteria, Gammaproteobacteria and Candidatus Spartobacteria) were found in all the four Cerrado areas. On the other hand, Ktedonobacteria (Phylum Chloroflexi) were not detected in ‘Floresta decidual’, Acidimicrobiia (Phylum Actinobacteria) were not detected in ‘Cerrado stricto sensu’, and Betaproteobacteria were only found in ‘Cerrado strico sensu’. Alphaproteobacteria was the dominant class in ‘Cerrado stricto sensu’ and ‘Cerradao’. Alphaproteobacteria, Acidobacteria and Thermoleophilia were dominant in Campo graminoide, while Alphaproteobacteria and Thermoleophilia were dominant in ‘Floresta decidual’. Considering low abundance classes (below 1%), Betaproteobacteria and Chlamydia were the most abundant across the gradient of Cerrado (Fig. S2). Rhodoplanes stood out as the most abundant bacterial genus, followed by Candidatus Xiphinematobacter, Bacillus, DA101, Candidatus Koribacter, Mycobacterium, Alicyclobacillus, Conexibacter, Candidatus Solibacter, Paenibacillus Pedosphaera, Cohnella, and Burkholderia (Fig. 4).
The rarefaction curves revealed higher number of OTUs in ‘Cerradao’ (1253 OTUs) followed by ‘Campo graminoide’ (1179 OTUs), ‘Floresta decidual’ (1149 OTUs), and ‘Cerrado stricto sensu’ (1140 OTUs), but the full extent of the bacterial diversity has not been sampled, since the asymptote was not reached (Fig. S3). The alpha diversity estimators also indicated ‘Cerradao’ as the richest habitat, however no differences (p > 0.05) were found between the Cerrado areas for any of the diversity estimators (Table 3). The analysis of bacterial beta diversity showed that ‘Cerrado stricto sensu’ and ‘Cerradao’ are more closely related, whereas ‘Floresta decidual’ and ‘Campo graminoide’ were different from each other, and form two distinct groups (Fig. 5). The RDA of OTUs from tropical Cerrado included the variables TOC (p = 0.027), N (p = 0.016) and P (p = 0.003) and showed significant marginal effects and VIF < 10. The selected RDA model for these variables was significant (F = 6.26, df = 3, p < 0.001) with an explanatory power of 29.9% (R2 = 0.356; adjusted R2 = 0.299) (Fig. 6).
Discussion
This study examined the bacterial communities in soils under influence of the preserved Cerrado in Sete Cidades National Park, Northeast Brazil, which can be distinguished from other Brazilian savannas mostly by its high diversity of plant species (Castro et al. 1998, 1999).
Our dataset encompasses a more comprehensive survey of bacteria communities from Brazilian Cerrado soils, comprising 36 libraries of 16S rRNA gene amplicons, with 9 replicates per sample. We found more than 4000 OTUs distributed in 37 phyla, while Araujo et al. (2012) retrieved 17 phyla from similar Cerrado samples from Central Brazil region. On a higher taxonomic level, soils exhibit a remarkably stable community structure, sharing five of the most abundant phyla (Proteobacteria, Acidobacteria, Actinobacteria, Verrucomicrobia, and Planctomycetes) found in the Brazilian Central Cerrado (Araujo et al. 2012). These results suggest that, despite the expected differences due to different next-generation sequencing platforms used in each study, both surveys have retrieved most of the abundant phyla found in soils worldwide (Janssen 2006).
Taking into account the significant differences in plant richness, diversity and density and soil properties in Cerrado areas from Sete Cidade National Park, the rarefaction curves showed small variation in OTUs bacterial richness at species level. ‘Cerradao’ appears to be more diverse than the other sites, but the indices of alpha diversity showed no significant difference (p > 0.05) among them. Therefore, local bacterial species richness was not significantly different among the habitats, but community composition was significantly distinct. According to Cassman et al. (2016), different plant litter and root exudates provide different carbon resources to soil microbes and can select a subset of the bacterial community.
The data from UniFrac phylogenetic distances demonstrated that the bacterial communities across the vegetation gradient have significant differences. ‘Cerrado stricto sensu’ and ‘Cerradao’ that share more similarities in their soil physicochemical properties and vegetation also present more similar bacterial communities. ‘Floresta decidual’ and ‘Campo graminoide’ that show the largest differences in soil physicochemical properties and vegetation composition, house more distinct bacterial communities. Similar results were found by Araujo et al. (2012) from soils in Central Cerrado, except that they found ‘Mata de Galeria’ as the most distinct and richness habitat.
Weighted PCoA explained 68.21% of beta diversity variation, demonstrating that overall differences between bacterial communities within the samples were more related to the abundance of specific taxon OTUs compared to their presence or absence, which explained only 24.95% of variation (Unweighted PCoA).
The phylum Proteobacteria was dominant (26% of the total OTUs) and presented a constant abundance in ‘Cerrado stricto sensu’, ‘Cerradao’ and ‘Floresta decidual’, and it was less abundant in ‘Campo graminoide’. Among the classes of Proteobacteria, the dominance of Alphaproteobacteria is notable in the Cerrado soils. Representatives of this class were dominant in ‘Cerrado stricto sensu’ and ‘Cerradao’, and shared with Thermoleophilia (Phylum Actinobacteria) the dominance in ‘Floresta decidual’, and with Thermoleophilia and Acidobacteriia the dominance in ‘Campo graminoide’. Previous studies also reported the dominance of Alphaproteobacteria in natural Brazilian Cerrado from Central region (Quirino et al. 2009; Araujo et al. 2012). Representatives of this class include important bacteria that are capable of nitrogen fixation in symbiosis with plants (Zhang and Xu 2008). The genus Bradyrhizobium (Order Rhizobiales) was commonly found in ‘Cerrado stricto sensu’ and ‘Floresta de Galeria’ in Brazilian Central Cerrado (Araujo et al. 2012), while in our study, the genus Rhodoplanes (Order Rhizobiales) appears as the most abundant along the four Cerrado areas. The genus Rhodoplanes comprises a group of phototrophic, purple non-sulfur bacteria (Okamura et al. 2009) recognized as good producers of hopanoids (Lodha et al. 2015), a class of lipids associated with N-fixing bacteria to protect nitrogenase from oxygen destruction (Berry et al. 1993). Interestingly, hopanoids are involved in allowing bacteria to adapt to changes in soil and, in particular, confer adaptation advantages to bacteria living under tropical conditions (Kannenberg et al. 1996). Therefore, our results suggest that the different environmental factors found in Cerrado from Sete Cidades National Park could select groups of bacteria that are more adapted to semiarid region. In addition, this may be the first report on the dominance of Rhodoplanes in Brazilian Cerrado soils.
Other dominant phyla in our dataset were Acidobacteria (21.5% of OTUs) and Actinobacteria (21.2% of total OTUs). The relative abundance of representatives of these phyla varied along the Cerrado areas, which suggests influence of different soil conditions and plant coverage. Representatives of Acidobacteria were only dominant in ‘Campo graminoide’, less abundant in ‘Floresta decidual’, and similarly abundant in ‘Cerrado stricto sensu’ and ‘Cerradao’. These results confirm the tolerance of this group to acid pH, low moisture and TOC (Quirino et al. 2009; Araujo et al. 2012), but contrast to Acidobacteria as dominant phylum in all Cerrado areas (Araujo et al. 2012). Also, our results disagree with those from Rampelotto et al. (2013) who found Proteobacteria (30.2 ± 2.3%) and Acidobacteria (30.3 ± 9.0%) as dominant phyla in Brazilian Central Cerrado.
Actinobacteria were also detected across the gradient of Cerrado vegetation, but with higher abundance in ‘Floresta decidual’. Representatives of this group are among the most important organic matter decomposers in soil, and rich actinobacterial communities can be expected in sites with high quantities of organic matter (Kopecky et al. 2011). The most abundant class of Actinobacteria was Thermoleophilia, and this finding disagrees with a previous study (Araujo et al. 2012) that reported the class Rubrobacter as most abundant in Brazilian Central Cerrado. Thermoleophilia are involved in P cycling (Steenbergh et al. 2015), and the high abundance of these organisms may be associated with the permanent presence of organic residues from plants as expected in ‘Floresta decidual’. According to Pitombo et al. (2015), the presence of organic straw influences the abundance of Thermoleophilia in soils. Representatives of the genus Conexibacter, belonging to the class Thermoleophilia, and found among the most abundant genera (greater than 1%) in our dataset, were previously detected in forest soils in Italy and Japan (Monciardini et al. 2003).
Firmicutes were found in the four Cerrado areas with highest relative abundance in ‘Campo graminoide’ and ‘Cerrado stricto sensu’, confirming the high resistance of these bacteria to unfavorable conditions (Hartmann et al. 2014). This group contains important genera, such as Bacillus and Paenibacillus, which are involved in the promotion of plant growth (Pindi et al. 2014). Our results agree with previous studies that have shown that Bacillus species are ubiquitous in soils from several biomes (Janssen 2006; Bergmann et al. 2011), including Cerrado (Rampelotto et al. 2013).
Candidatus Xiphinematobacter, which belongs to Verrucomicrobia, was the second most abundant genus in our dataset after Rhodoplanes. This genus comprises obligate endosymbionts of nematodes, which could explain its widespread distribution in the four Cerrado areas. It may be considered a bioindicator of soil quality (Vandekerckhove et al. 2000).
Regarding the low-abundant phyla (<1% of OTUs), which can act in important soil functions, our study identified Chlamydiae and Bacteroidetes among the most abundant. Chlamydiae were also previously included in the rare group in Brazilian Cerrado (Araujo et al. 2012). Representatives of this phylum are obligate intracellular pathogens (Wyrick 2000) that are present in different soils. Bacteroidetes are composed of Gram-negative bacteria that are widely distributed in soil and sediments (Gupta 2004), ranging from 0 to 18% (5% in average) of soil bacterial communities (Janssen 2006). Although Bacteroidetes are among the dominant phyla in the Brazilian Central Cerrado (Araujo et al. 2012), our results showed this phylum in very low relative abundance in the Cerrado areas, except in the ‘Floresta decidual’ which showed abundance of Bacteroidetes higher than 1%.
From the total OTUs, 5% were unclassified, suggesting that a significant amount of bacterial phyla could not be classified by the chosen experimental approach or are still unknown, and indicating that further studies should assess this important source for new microbes.
Previous studies have either demonstrated shifts in soil bacterial communities structure and diversity as consequence of land use of native Cerrado (Peixoto et al. 2006; Mota et al. 2008; Bresolin et al. 2010; Rachid et al. 2013; Rampelotto et al. 2013) or retrieved the bacterial communities in native Cerrado gradient under the same edaphoclimatic conditions (Araujo et al. 2012; Castro et al. 2016).
This study, therefore, represents the first study on soil bacterial communities along Cerrado gradient from a different edaphoclimatic Brazil region. The differences observed in soil bacterial communities in Cerrado biome from different regions may be related with the biogeographic pattern, since the bacterial communities are both affected by environmental factors and distance (Xiong et al. 2012, Zhang et al. 2016). Previous studies have shown that the dissimilarity of the bacteria communities increased according with the distance (Rysanek et al. 2015; Martiny et al. 2011). It may mean that the environmental factors and vegetation may have been affected by the geographic distances in Brazilian Cerrado, and these factors significantly influenced the bacterial communities, since the evaluated sites by Araujo et al. (2012) and Castro et al. (2016) presented different vegetation pattern and soil properties.
In summary, our findings show the bacterial community structure was clearly different among the sites, however, we neither detect higher frequency of Acidobacteria (40–47%) in all sites nor significant species richness among the sites, as previously reported in other Brazilian savannas (Araujo et al. 2012). However, we detected that Brazilian savannas share a bacterial phylum core comprising Proteobacteria, Acidobacteria, Actinobacteria, Verrucomicrobia and Planctomycetes.
Conclusion
This study provides the first in depth assessment of soil bacterial composition and community structure along the Cerrado gradient from Northeast Brazil, revealing 37 phyla and a high number of 29 rare phyla. Moreover, unclassified OTUs accounted for 5.42% of the total dataset, highlighting the potential of the preserved Cerrado from Northeast Brazil for further studies regarding with microbial ecology in tropical soils, ecological services and evolution, as well as a potential source of new bacterial species for biotechnological purposes and industrial applications.
References
Araujo JF, Castro AP, Costa MMC, Togawa RC, Pappas Júnior GJ, Quirino BF, Bustamante MMC, Williamson L, Handelsman J, Krüger RH (2012) Characterization of soil bacterial assemblies in Brazilian savanna-like vegetation reveals acidobacteria dominance. Microb Ecol 64:760–770
Bardgett RD, Shine A (1999) Linkages between litter diversity, soil microbial biomass and ecosystem function in temperate grasslands. Soil Biol Biochem 31:317–321
Bergmann GT, Bates ST, Eilers KG, Lauber CL, Caporaso JG, Walters WA, Knight R, Fierer N (2011) The under-recognized dominance of verrucomicrobia in soil bacterial communities. Soil Biol Biochem 43:1450–1455
Berry AM, Harriott OT, Moreau RA, Osman SF, Benson DR, Jones AD (1993) Hopanoid lipids compose the Frankia vesicle envelope, presumptive barrier of oxygen diffusion to nitrogenase. Proc Natl Acad Sci USA 90:6091–6094
Bresolin JD, Bustamante MMC, Krüger RH, Silva MRSS, Perez KS (2010) Structure and composition of bacterial and fungal community in soil under soybean monoculture in the Brazilian Cerrado. Braz J Microbiol 41:391–403
Bru D, Ramette A, Saby NP, Dequiedt S, Ranjard L, Jolivet C, Arrouays D, Philippot L (2011) Determinants of the distribution of nitrogen-cycling microbial communities at the landscape scale. ISME J 5:532–542
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010a) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010b) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 108:4516–4522
Cassman NA, Leite MFA, Pan Y, de Hollander M, van Veen JA, Kuramae EE (2016) Plant and soil fungal but not soil bacterial communities are linked in long-term fertilized grassland. Sci Rep 6:23680
Castro AAJF, Martins FR, Fernandes AG (1998) The woody flora of Cerrado vegetation in the state of Piauí, Northeastern Brazil. Edinb J Botany 55:455–472
Castro AAJF, Martins FR, Tamashiro JY, Shepherd GJ (1999) How rich is the flora of Brazilian Cerrados? Ann Missouri Bot Gard 86:192–224
Castro AP, Silva MRSS, Quirino BF, Bustamante MMC, Krüger RH (2016) Microbial diversity in Cerrado biome (Neotropical Savanna) soils. PLoS ONE 11:e0148785
Coutinho LM (1978) O conceito de cerrado. Rev Bras Bot 1:17–23
DeMandal S, Zothansanga AKP, Bisht SS, Kumar NS (2015) First report of bacterial community from a bat guano using Illumina next-generation sequencing. Gen Data 4:99–101
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. App Environ Microbiol 72:5069–5072
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200
Forzza RC, Leitman PM, Costa A, Carvalho AA, Peixoto AL, Walter BMT, Bicudo C, Zappi D, Costa DP, Lleras E, Martinelli G, Lima HC, Prado J, Stehmann JR, Baumgratz JFA, Pirani JR, Sylvestre LS, Maia LC, Lohmann LG, Paganucci L, Silveira M, Nadruz M, Mamede MCH, Bastos MNC, Morim MP, Barbosa MR, Menezes M, Hopkins M, Secco R, Cavalcanti T, Souza VC (2010) Catalogo de plantas e fungos do Brasil. Instituto de Pesquisas Jardim Botanico, Rio de Janeiro
Gupta RS (2004) The phylogeny and signature sequences characteristics of fibrobacteres, chlorobi, and bacteroidetes. Critical Rev Microbiol 30:123–140
Hartmann M, Niklaus PA, Zimmermann S (2014) Resistance and resilience of the forest soil microbiome to logging-associated compaction. ISME J 8:226–244
IBDF (1979) Plano de manejo: parque nacional de sete cidades. Fundação Brasileira para a Conservação da Natureza, Brasília
Illumina (2013) 16S metagenomic sequencing library preparation. Illumina Guides http://support.illumina.com/downloads/16s_metagenomic_sequencing_library_preparation.html. Accessed 6 Mar 2014
Janssen PH (2006) Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes. Appl Environ Microbiol 73:1719–1728
Kannenberg EL, Perzi M, Muller P, Hartner T, Poralla K (1996) Hopanoid lipids in Bradyrhizobium and other plant-associated bacteria and cloning of the Bradyrhizobium squalene-hopene cyclase. Pl Soil 186:107–112
Kopecky J, Kyselkova M, Omelka M, Cermak L, Novotna J, Grundmann GL, Moenne-Loccoz Y, Sagova-Mareckova M (2011) Actinobacterial community dominated by a distinct clade in acidic soil of a waterlogged deciduous forest. FEMS Microbiol Ecol 78:386–394
Lodha TD, Srinivas A, Sasikala C, Ramana CV (2015) Hopanoid inventory of Rhodoplanes spp. Arch Microbiol 197:861–867
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71:8228–8235
Martiny JB, Eisen JA, Penn K, Allison SD, Horner-Devine MC (2011) Drivers of bacterial beta-diversity depend on spatial scale. Proc Natl Acad Sci 108:7850–7854
Mishra S, Lee WA, Hooijer A, Reuben S, Sudiana IM, Idris A, Swarup S (2014) Microbial and metabolic profiling reveal strong influence of water table and land-use patterns on classification of degraded tropical peatlands. Biogeosciences 11:1727–1741
Monciardini P, Cavaletti L, Schumann P, Rohde M, Donadio S (2003) Conexibacter woesei gen. nov., sp. nov., a novel representative of a deep evolutionary line of descent within the class Actinobacteria. Int J Syst Evol Microbiol 53:569–576
Mota FF, Gomes EA, Marriel IE, Paiva E, Seldin L (2008) Tolerant and aluminum-sensitive maize (Zea mays L.) lines cultivated in unlimed and limed Cerrado soil. Microbiol Biotechnol 18:805–814
Myers N, Mittermeier RA, Mittermeier CG, Fonseca GAB, Kent J (2000) Biodiversity. hotspots for conservation priorities. Nature 403:853–858
Okamura K, Kanbe T, Hiraishi A (2009) Rhodoplanes serenus sp. nov., a purple non-sulfur bacterium isolated from pond water. Int J Syst Evol Microbiol 59:531–535
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Wagner H (2015) Vegan: community ecology package. R package version 2.2–1. http://cran.r-project.org/package=vegan
Oliveira MEA, Martins FR, Castro AAJF, Santos JR (2007) Classes de cobertura vegetal do parque nacional de sete cidades (transição campo-floresta) utilizando imagens TM/Landsat, NE do Brasil. Proceedings XIII Simpósio Brasileiro de Sensoriamento Remoto Anais 13:1775–1783
Peixoto RS, Coutinho HLC, Madari B, Machado PLOA, Rumjanek NG, Van Elsas JD, Seldin L, Rosado AS (2006) Soil aggregation and bacterial community structure as affected by tillage and cover cropping in the Brazilian Cerrados. Soil Till Res 90:16–28
Pindi PK, Sultana T, Vootla PK (2014) Plant growth regulation of Bt-cotton through Bacillus species. 3 Biotech 4:305–315
Pitombo LM, Carmo JB, Hollander M, Rossetto R, López MV, Cantarella H, Kuramae EE (2015) Exploring soil microbial 16S rRNA sequence data to increase carbon yield and nitrogen efficiency of a bioenergy crop. GCB Bioenergy. doi:10.1111/gcbb.12284
Pylro VS, Roesch LFW, Ortega JM, Amaral AM, Tótola MR, Hirsch PR, Rosado AS, Góes-Neto A, Silva ALC, Rosa CA, Morais DK, Andreote FD, Duarte GF, Melo IS, Seldin L, Lambais MR, Hungria M, Peixoto RS, Kruger RH, Tsai SM, Azevedo V (2014) Brazilian microbiome project: revealing the unexplored microbial diversity–challenges and prospects. Microb Ecol 67:237–241
Pylro VS, Morais DK, Oliveira FS, Santos FG, Lemos LN, Oliveira G, Roesch LFW (2016) BMPOS: a flexible and user-friendly tool sets for microbiome studies. Microb Ecol 73:443–447
Quirino BF, Pappas GJ, Tagliaferro AC, Collevatti RG, Neto EL, Silva MR, Bustamante MM, Kruger RH (2009) Molecular phylogenetic diversity of bacteria associated with soil of savanna-like Cerrado vegetation. Microbiol Res 164:59–70
Rachid CTCC, Santos AL, Piccolo MC, Balieiro FC, Coutinho HLC, Peixoto RS (2013) Effect of sugarcane burning or green harvest methods on the Brazilian Cerrado soil bacterial community structure. PLoS ONE 8:e59342
Rampelotto PH, Ferreira AS, Barboza AD, Roesch LF (2013) Changes in diversity, abundance, and structure of soil bacterial communities in Brazilian Savanna under different land use systems. Microb Ecol 66:593–607
Rodrigues JLM, Pellizari VH, Mueller R, Baek K, Jesus EC, Paula FS, Mirza B, Hamaoui GS, Tsai SM, Feigl B, Tiedje JM, Bohannan JM, Nüsslein K (2013) Conversion of the Amazon rainforest to agriculture results in biotic homogenization of soil bacterial communities. Proc Natl Acad Sci USA 110:988–993
Rysanek D, Hrckova K, Skaloud P (2015) Global ubiquity and local endemism of free-living terrestrial protists: phylogeographic assessment of the streptophyte alga Klebsormidium. Environ Microbiol 17:689–698
Steenbergh AK, Bodelier PLE, Hoogveld HL, Slomp CP, Laanbroek HJ (2015) Phylogenetic characterization of phosphatase-expressing bacterial communities in baltic sea sediments. Microbes Environ 30:192–195
Tedesco MJ, Gianello C, Bissani CA, Bohnen H, Volkweiss SJ (1995) Análise de solo, plantas e outros materiais. Universidade Federal do Rio Grande do Sul, Porto Alegre, p 147
Vandekerckhove TTM, Willems A, Gillis M, Coomans A (2000) Occurrence of novel verrucomicrobial species, endosymbiotic and associated with parthenogenesis in Xiphinema americanum-group species (Nematoda, Longidoridae). Int J Syst Evol Microbiol 50:2197–2205
Wyrick P (2000) Intracellular survival by Chlamydia. Cell Microbiol 2:275–282
Xiong J, Liu Y, Lin X, Zhang H, Zeng J, Hou J, Yang Y, Yao T, Knight R, Chu H (2012) Geographic distance and pH drive bacterial distribution in alkaline lake sediments across Tibetan Plateau. Environ Microbiol 14:2457–2466
Yeomans JC, Bremner JM (1988) A rapid and precise method for routine determination of organic carbon in soil. Comm Soil Sci Plant Anal 19:1467–1476
Zhang L, Xu Z (2008) Assessing bacterial diversity in soil. J Soils Sedim 8:379–388
Zhang B, Wu X, Zhang G, Li S, Zhang W, Chen X, Sun L, Zhang B, Liu G, Chen T (2016) The diversity and biogeography of the communities of Actinobacteria in the forelands of glaciers at a continental scale. Environ Res Lett 11:054012
Acknowledgements
The authors thank “Fundação de Amparo a Pesquisa no Estado do Piauí” (FAPEPI) and “Conselho Nacional de Desenvolvimento Científico e Tecnológico” (CNPq) for financial support to this project through of PRONEX (FAPEPI/CNPq 004/2012). A.S.F. Araujo, W.M. Bezerra, A.C.A. Lopes, and V.M.M. Melo are supported by grants from CNPq (Research’s Productivity). V.M Santos is supported by grants from FAPEPI (Regional Scientific Development).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
de Araujo, A.S.F., Bezerra, W.M., dos Santos, V.M. et al. Distinct bacterial communities across a gradient of vegetation from a preserved Brazilian Cerrado. Antonie van Leeuwenhoek 110, 457–469 (2017). https://doi.org/10.1007/s10482-016-0815-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-016-0815-1