Abstract
A rod shaped, Gram-stain positive, non-motile, facultative anaerobic and gelatin hydrolysing bacterium, strain PG1T, was isolated from reclaimed land soil in Kyehwa-do, Republic of Korea. Strain PG1T showed highest 16S rRNA gene sequence similarity (97.4 and 96.5 %, respectively) to Paenibacillus shenyangensis A9T and Paenibacillus hunanensis FeL05T, and clustered closely with the members of the family Paenibacillaceae. DNA–DNA hybridization studies revealed a genomic relatedness of 47 ± 9 % with P. shenyangensis A9T. The predominant fatty acids of strain PG1T were identified to be anteiso-C15:0 (46.7 %) and C16:0 (22.7 %). Diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine and an unidentified phospholipid were found to be the major polar lipids. The genomic DNA G+C content was found to be 47.7 mol%. This polyphasic characterisation of the newly isolated strain PG1T justifies its description as representative of a novel species in the genus Paenibacillus, for which the name Paenibacillus gelatinilyticus sp. nov., (type strain = PG1T = KCTC 33642T = JCM 30624T) is proposed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Paenibacillus was described by Ash et al. (1993) based on the analysis of 16S rRNA gene sequences of group 3 bacilli. Members of the genus Paenibacillus are facultatively anaerobic or strictly aerobic, Gram-stain positive or Gram-variable endospore forming rods with oval endospores and swollen sporangia. The major fatty acid is anteiso-C15:0 and meso-diaminopimelic acid (meso-DAP) is commonly present in the peptidoglycan (Priest 2009). They are widely distributed in different environments and are of interest because of their ability to promote plant growth (Jin et al. 2011), degrade complex organic molecules (Wang et al. 2012) and act as bio-control agents (Kim et al. 2004). In this communication we describe the isolation and characterisation of a rod shaped, psychrotolerant bacterium, strain PG1T, and propose it as the type strain of a novel species belonging to the genus Paenibacillus based on polyphasic taxonomy.
Materials and methods
Strains and culture conditions
Strain PG1T was isolated from reclaimed land soil in Kyehwa-do, Republic of Korea. Purification of the isolate was achieved by repeated streaking on tryptone soy agar plates at 30 °C. The strain was maintained under refrigeration at 4 °C and also preserved as glycerol stocks at −80 °C. For comparative taxonomic studies, the type strain of Paenibacillus shenyangensis A9T (=JCM19307T) was obtained from Dr. Xiaomin Hu (Jiang et al. 2015) and also procured from Japan Collection of Microorganisms. Comparative polyphasic characterisation of strain PG1T with P. shenyangensis A9T was performed in this study following the recommended standards proposed by Tindall et al. (2010) and Logan et al. (2009).
Phenotypic characterisation
Morphological properties (cell shape and cell size) of strain PG1T were observed by phase contrast light microscopy (Zeiss Primo Star) after 24 h incubation in tryptone soy broth. Motility was examined using the same microscopy after 24 and 48 h of incubation in nutrient agar at 25 °C. Gram reaction was performed by the non-staining method according to Buck (1982). Spore formation was observed after growing strain PG1T in tryptone soy broth supplemented with MnCl2 (50 μM), CaCl2 (700 μM) and MgCl2 (1 mM) (Priest 2009). Growth at different concentrations of NaCl was tested in 1 % tryptone soy broth supplemented with varying amounts of NaCl (0-10 % w/v). Ability to grow at different pH was tested in tryptone soy broth adjusted to varying pH using phosphate buffer and glycine-NaOH buffer. Similarly growth temperature range was tested in tryptone soy broth with incubation at different temperatures (5–50 °C). Utilisation of various substrates as sole carbon source and growth in the presence of inhibitory compounds was determined using GEN III MicroPlate test panel (BIOLOG) following the manufacturer’s instructions. In addition, production of acid from carbohydrates, enzyme activities and other biochemical tests were performed using API 50CHB and API 20E strips (bioMérieux).
Chemotaxonomic characterisation
Polar lipids were extracted from 1 g freeze-dried cells using methanol/chloroform/saline (2:1:0.8, v/v/v), as described by Kates (1986). Separation of the lipids was achieved by two-dimensional chromatography on silica gel TLC plates (Kieselgel 60 F254, Merck) using chloroform:methanol:water (65:25:4, v/v/v) in the first dimension and chloroform:methanol:acetic acid:water (80:12:15:4, v/v/v/v) in the second dimension as mobile phase (Tindall et al. 2007). Total polar lipids were detected using 5 % ethanolic molybdophosphoric acid and specific functional groups were detected using spray reagents, i.e. ninhydrin (specific for amino groups), molybdenum blue (specific for phosphates) or α-naphthol (specific for sugars) (Kates 1972; Oren et al. 1996). Quinones were extracted with a chloroform–methanol (2:1 v/v) mixture, purified by TLC and analysed by HPLC as described by Tamaoka et al. (1983). For the detection of meso-DAP in the cell wall, whole cell hydrolysates were prepared (6 N HCl, 100 °C, 18 h) and examined by TLC on cellulose plates (DC Cellulose F, Merck) using n-butanol:water:acetic acid (50:25:25) as the solvent system (Komagata and Suzuki 1987; Rhuland et al. 1955). For determination of cellular fatty acid composition, strain PG1T was grown on tryptone soy agar (Difco) at 30 °C for 2 days. Fatty acid methyl esters were prepared, separated and identified according to the instructions for the Microbial Identification System (Microbial ID; MIDI 6.0 version) (Sasser 2001).
Molecular and phylogenetic analysis
For 16S rRNA gene sequencing, biomass of strain PG1T was taken from 1.5 ml of well-grown liquid cultures. DNA was extracted and purified by using the GeneAll Exgene™ Cell SV kit. PCR amplification of 16S rRNA gene and sequencing were performed as described previously (Imhoff et al. 1998). Recombinant Taq polymerase and universal primers 27F (5′-AGAGTTTGATCATGGCTCAG-3) and 1522R (5′-AAGGAGGTGATCCAGCCGAC-3′) were used for PCR. Sequencing were performed by a chain termination reaction (Sanger et al. 1977) using the BigDye® Terminator v3.1 Cycle Sequencing Kit and an ABI 3730xl DNA Analyzer (Applied Biosystems). The 16S rRNA gene sequences obtained were compiled using SeqMan software (DNASTAR) and the almost complete sequence (1,470 bp) was compared with available 16S rRNA gene sequences of cultured species by NCBI-BLAST search (Altschul et al. 1990) and the EzTaxon-e server (http://eztaxon-e.ezbiocloud.net/; Kim et al. 2012). For phylogenetic analysis, 16S rRNA gene sequences of the most closely related type strains of the species belonging to the genus Paenibacillus and of strain PG1T were aligned using the CLUSTAL_W algorithm and MEGA6 software (Tamura et al. 2013) was used for phylogenetic analyses. Evolutionary distances were calculated by using the Kimura 2-parameter correction (Kimura 1983) in a pair-wise deletion procedure. Neighbour-joining (Saitou and Nei 1987) and the maximum-parsimony method (Fitch 1972) were used to construct phylogenetic trees. Percentage support values were obtained using a bootstrap procedure with 1000 replications.
Genomic DNA was extracted and purified according to the method of Marmur (1961) and the mol% G+C of the DNA was determined following the fluorimetric procedure described by Gonzalez and Saiz-Jimenez (2002). The degree of DNA–DNA relatedness between strain PG1T and P. shenyangensis A9T was determined by the fluorimetric method described by Gonzalez and Saiz-Jimenez (2005). This method measures the divergence between the thermal denaturation midpoint of homoduplex DNA and heteroduplex DNA (ΔT m) using a real-time PCR thermocycler that obtains fluorescence determinations. Prior to analysis, isolated DNA was renatured at the optimum temperature for renaturation (T or) which was approximated according to the method of De Ley et al. (1970) using the equation T or = 0.51(% G+C) + 47.0. Renaturation conditions consisted of a denaturation step of 99 °C for 10 min, followed by an annealing period of 8 h at T or and by progressive 60 min steps, each at 10 °C below the previous one, until room temperature was reached. To the renatured DNA, SYBR Green I (LONZA) dye was added at a final concentration of 1:100000. The thermal denaturation of the labeled mixture and measurement of fluorescence during denaturation was carried out using a real-time PCR machine (CFX Connect™ Real-Time System, Bio-Rad). The thermal profile consisted of a 15 min hold at 25 °C followed by a 25–99 °C ramp in 0.2 °C steps with a 12 s hold. The mean of three replicates was used to determine the T m (melting temperature). Using the associated software, Bio-Rad CFX Manager, T m values of homologous and hybrid DNAs were calculated as the temperatures corresponding to a 50 % decrease in fluorescence. Percentage similarity was estimated based on the ΔT m values (Gonzalez and Saiz-Jimenez 2005).
Results and discussion
Phenotypic, physiological and biochemical characteristics
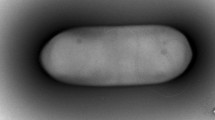
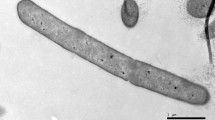
Strain PG1T was observed to be a rod shaped, aerobic or facultatively anaerobic, Gram-positive and non-motile bacterium measuring 5.0–6.0 µm long and 0.5–0.7 µm wide (Fig. S1), with terminal endospores and swollen sporangia (Fig. S2). Aerobically grown colonies of strain PG1T on tryptone soy agar were observed to be orange pink in colour, elastic, circular, convex with entire margins (Fig. S3). Strain PG1T does not require NaCl for its growth but can tolerate NaCl concentrations up to 8 % (w/v). Growth was observed at a pH range of 5.0–9.5 (optimum at 7.0) and temperature of 10–45 °C (optimum at 25–30 °C). Comparison of different morphological, physiological and biochemical characteristics of strain PG1T and the closely related strains P. shenyangensis A9T and P. hunanensis FeL05T, is shown in Table 1 and Supplementary Table S1. It was reported in the species description by Jiang et al. (2015) that P. shenyangensis A9T could hydrolyse gelatin and also utilise citrate as sole carbon source. When the tests were repeated in the present study using API 20E strips (bioMérieux) and following the manufacturer’s instructions, it was observed that P. shenyangensis A9T showed negative reactions for gelatin hydrolysis and citrate utilisation.
Genetic characteristics
Phylogenetic analysis based on the 16S rRNA gene sequence of strain PG1T demonstrated its relationship with members of the genus Paenibacillus and the highest 16S rRNA gene sequence similarity with P. shenyangensis A9T (97.4 %) and P. hunanensis FeL05T (96.5 %) supported its affiliation to the genus. The neighbor-joining phylogenetic tree showed that strain PG1T clustered with P. shenyangensis A9T with high bootstrap support (Fig. 1) and a similar topology was also observed in the maximum parsimony tree (Fig. S4). The G+C content of strain PG1T was 47.7 mol% as determined by fluorimetric method. The ΔT m between genomic DNA of strain PG1T and P. shenyangensis A9T and their hybrid DNA mixture was 8.6 ± 1.6 °C, equivalent to DNA–DNA relatedness similarity of about 47 ± 9 % (Gonzalez and Saiz-Jimenez 2005), indicating a relatedness below the 70 % cut-off point for the circumscription of bacterial species according to Wayne et al. (1987).
Phylogenetic analysis based on 16S rRNA gene sequences showing the relationship of strain PG1T to closely related species. Accession numbers are given in parentheses. Multiple alignment, distance calculations (distance options according to the Kimura 2-parameter model) and clustering with the neighbour-joining method were performed by using the software MEGA version 6. Bootstrap values based on 1,000 replications are listed as percentages at the branching points. Bar 0.01 nucleotide substitutions per nucleotide position
Chemotaxonomic characteristics
The major polar lipids of strain PG1T were identified as diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine and an unidentified phospholipid (PL1). The presence of three unidentified phospholipids (PL2, 3, and 4), two amino phospholipids (APL1 and 2) and three unidentified lipids (UL1, 2 and 3) in minor amounts was also observed (Fig. S5). The presence of a phospholipid of unknown structure containing glucosamine (NPG) in P. shenyangensis A9T as reported by Jiang et al. (2015) could not be confirmed in the present study because the spot was stained only with phosphomolybdic acid and molybdenum blue reagents but not with α-naphthol reagent. Hence the spot is labeled as an unknown phospholipid (PL1) in Supplementary Fig. S5. Also minor quantities of three unidentified lipids (UL1, 2 and 3), three unknown phospholipids (PL2, 3, and 4) and two amino phospholipids (APL1 and APL2) were detected in the present study in P. shenyangensis A9T which were not reported by Jiang et al. (2015). Two glycolipids (GL1 and GL2) which were reported earlier in P. shenyangensis A9T was not detected in the present study (Fig. S5).
MK-7 was detected as the predominant menaquinone in strain PG1T as is in accordance with the description of the genus Paenibacillus. Whole-cell fatty acid analysis revealed that anteiso-C15:0 and C16:0 are the predominant fatty acids (Table 2). The cell wall peptidoglycan of strain PG1T was found to contain meso-DAP.
On the basis of differentiating physiological, biochemical, and genotypic characteristics (Table 1), we conclude that strain PG1T represents a new species belonging to the genus Paenibacillus, for which the name Paenibacillus gelatinilyticus sp. nov. is proposed. During the present study it was also observed that biochemical characteristics and polar lipid profile of P. shenyangensis A9T are not in accordance with the species description by Jiang et al. (2015). Hence, we also propose an emended description of the species P. shenyangensis.
Description of Paenibacillus gelatinilyticus sp. nov.
Paenibacillus gelatinilyticus (ge.la.ti.ni.ly’ti.cus. N.L. neut n. gelatinum, gelatin; N.L. adj. lyticus (from Gr. adj. lytikos), able to loosen, able to dissolve; N.L. masc. adj. gelatinilyticus, gelatin-dissolving).
Cells are rod shaped, 5.0–6.0 µm long and 0.5–0.7 µm wide with terminal endospores and swollen sporangia, aerobic or facultatively anaerobic, Gram-positive, non-motile, alkali tolerant (pH range 5.0–9.5; optimum 7.0) and psychrotolerant (temperature range 10–45 °C; optimum 25–30 °C). On tryptone soy agar, aerobically grown colonies are orange pink in colour, elastic, circular, convex with entire margins. NaCl is not required for growth but can tolerate NaCl concentrations up to 8 % (w/v). Positive for catalase, β-galactosidase, MR, VP test, hydrolysis of gelatin, starch, esculin and Tween 40, but negative for arginine dihydrolase, lysine decarboxylase, ornithine decarboxylase, citrate utilisation, urea hydrolysis, nitrate reduction, H2S production, indole production, and oxidase. Assimilation of the following carbon sources is positive: glycerol, l-arabinose, ribose, d-xylose, galactose, glucose, fructose, mannose, mannitol, N-acetyl-glucosamine, amygdalin, arbutin, salicin, d-cellobiose, d-maltose, lactose, melibiose, sucrose, trehalose, d-raffinose, starch, glycogen, 3-gentiobiose, dextrin, d-turanose, stachyose, β-methyl-d-glucoside, N-acetyl-β-d-mannosamine, 3-methyl glucose, l-rhamnose, inosine, d-sorbitol, l-alanine, pectin, l-galactonic acid lactone, d-gluconic acid, methyl pyruvate, l-lactic acid, acetoacetic acid, acetic acid and formic acid. Can tolerate and grow in the presence of 1 % sodium lactate, guanidine HCl, nalidixic acid, lithium chloride, potassium tellurite and aztreonam. The cell wall peptidoglycan contains meso-DAP. Predominant fatty acids include anteiso-C15:0 and C16:0. MK7 is the major quinone. Major polar lipids are diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine and an unidentified phospholipid, along with three unidentified phospholipids (PL2, 3, 4), two amino phospholipids (APL1, 2) and three unidentified lipids (UL1, 2, 3) in minor quantities. The DNA G + C content of the type strain is 47.7 mol%.
The type strain PG1T (=KCTC 33642T = JCM 30624T) was isolated from a reclaimed land soil in Kyehwa-do, Republic of Korea. The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain PG1T is KP231650.
Emended description of Paenibacillus shenyangensis
The description of the species is as given previously (Jiang et al. 2015) except that the type strain is not able to hydrolyse gelatin and cannot utilise citrate as sole carbon source. The polar lipid profile is composed of diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine and an unidentified phospholipid (PL1) as major components. Minor quantities of three unidentified lipids (UL1, 2 and 3), three unknown phospholipids (PL2, 3 and 4), two amino phospholipid (APL1 and 2) and a glycolipid (GL1) are present.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Ash C, Priest FG, Collins MD (1993) Molecular identification of rRNA group 3 bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test. Proposal for the creation of a new genus Paenibacillus. Antonie Van Leeuwenhoek 64:253–260
Buck JD (1982) Nonstaining (KOH) method for determination of Gram reactions of marine bacteria. Appl Environ Microbiol 44:992–993
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Fitch WM (1972) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Gonzalez JM, Saiz-Jimenez C (2002) A fluorimetric method for the estimation of G + C mol% content in microorganisms by thermal denaturation temperature. Environ Microbiol 4:770–773
Gonzalez JM, Saiz-Jimenez C (2005) A simple fluorimetric method for the estimation of DNA–DNA relatedness between closely related microorganisms by thermal denaturation temperatures. Extremophiles 9:75–79
Imhoff JF, Süling J, Petri R (1998) Phylogenetic relationships among the Chromatiaceae, their taxonomic reclassification and description of the new genera Allochromatium, Halochromatium, Isochromatium, Marichromatium, Thiococcus, Thiohalocapsa and Thermochromatium. Int J Syst Bacteriol 48:1129–1143
Jiang B, Zhao X, Liu J, Fu L, Yang C, Hu X (2015) Paenibacillus shenyangensis sp. nov., a bioflocculant-producing species isolated from soil under a peach tree. Int J Syst Evol Microbiol 65:220–224
Jin HJ, Lv J, Chen SF (2011) Paenibacillus sophorae sp. nov., a novel nitrogen-fixing species isolated from the rhizosphere of Sophora japonica. Int J Syst Evol Microbiol 61:767–771
Kates M (1972) Techniques of lipidology. Elsevier, New York
Kates M (1986) Techniques of lipidology: isolation, analysis, and identification of lipids. Elsevier, Amsterdam
Kim DS, Bae CY, Jeon JJ, Chun SJ, Oh HW, Hong SG, Baek KS, Moon EY, Bae KS (2004) Paenibacillus elgii sp. nov., with broad antimicrobial activity. Int J Syst Evol Microbiol 54:2031–2035
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge
Komagata K, Suzuki K (1987) Lipids and cell-wall analysis in bacterial systematics. Methods Microbiol 19:161–207
Liu Y, Liu L, Qiu F, Schumann P, Shi Y, Zou Y, Zhang X, Song W (2010) Paenibacillus hunanensis sp. nov., isolated from rice seeds. Int J Syst Evol Microbiol 60:1266–1270
Logan NA, Berge O, Bishop AH, Busse H-J, De Vos P, Fritze D, Heyndrickx M, Kämpfer P, Rabinovitch L, Salkinoja-Salonen MS, Seldin L, Ventosa A (2009) Proposed minimal standards for describing new taxa of aerobic, endospore-forming bacteria. Int J Syst Evol Microbiol 59:2114–2121
Marmur J (1961) A procedure for the isolation of deoxyribonucleic acid from microorganisms. J Mol Biol 3:208–218
Oren A, Duker S, Ritter S (1996) The polar lipid composition of Walsby’s square bacterium. FEMS Microbiol Lett 138:135–140
Priest FG (2009) Genus I. Paenibacillus. In: De Vos P, Garrity G, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman WB (eds) The firmicutes, Bergey’s manual of systematic bacteriology, vol 3, 2nd edn. Springer, New York, pp 269–295
Rhuland LE, Work E, Denman RF, Hoare DS (1955) The behavior of the isomers of α, ε-diaminopimelic acid on paper chromatograms. J Am Chem Soc 77:4844–4846
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Sasser M (2001) Identification of bacteria by gas chromatography of cellular fatty acids. Technical Note 101. MIDI Inc, Newark
Tamaoka J, Fujimura Y-K, Kuraishi H (1983) Analysis of bacterial menaquinone mixtures by high performance liquid chromatography. J Appl Microbiol 54:31–36
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tindall BJ, Sikorski J, Smibert RM, Kreig NR (2007) Phenotypic characterization and the principles of comparative systematics. In: Reddy CA, Beveridge TJ, Breznak JA, Marzluf G, Schmidt TM, Snyder LR (eds) Methods for general and molecular microbiology, 3rd edn. Snyder ASM Press, Washington DC, pp 330–393
Tindall BJ, Rosselló-Móra R, Busse H-J, Ludwig W, Kämpfer P (2010) Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 60:249–266
Wang L, Baek SH, Cui Y, Lee HG, Lee ST (2012) Paenibacillus sediminis sp. nov., a xylanolytic bacterium isolated from a tidal flat. Int J Syst Evol Microbiol 62:1284–1288
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Trüper HG (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Acknowledgments
We thank Prof. Aharon Oren for his expert suggestion for correct species epithet and Latin etymology. This work was carried out with the support of “Cooperative Research Program for Agriculture Science & Technology Development (Project No. PJ009801)” Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Additional information
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain PG1T is KP231650.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Padakandla, S.R., Lee, GW. & Chae, JC. Paenibacillus gelatinilyticus sp. nov. a psychrotolerant bacterium isolated from a reclaimed soil and amended description of Paenibacillus shenyangensis . Antonie van Leeuwenhoek 108, 1197–1203 (2015). https://doi.org/10.1007/s10482-015-0574-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-015-0574-4