Abstract
Root nodules are highly organized root organs where nitrogen fixation takes place in, and its formation are the results of complicated interactions between legumes and rhizobia. Nodule number is tightly controlled by Legumes to ensure optimal growth without energy loses wasted by excessive number of nodules. One of major factors controlling of nodule number is nitrate availability in the soil. However, during symbiotic development, the details of Nod factor signaling associated with nitrate regulation of nodulation are unknown. NORK, the immediately downstream component of these Nod factor receptors is central to the Nod factor signaling. NIC1, a CLE peptide-encoding gene in soybean (Glycine max), play an important role in nitrate regulation of nodulation. In this study, specified RNAi construct of GmNORK was generated and transformed into soybean roots by agrobacterium rhizogenes-mediated hairy root transformation. We found that the nodule number decreased substantially in GmNORK knock-down soybean transgenic roots. The expression levels of GmNIC1 in the GmNORK RNAi soybean transgenic roots is substantially reduced after rhizobial inoculation compared with the control soybean transgenic roots. Our data suggest that nitrate regulation of nodulation was affected by Nod factor signaling during nodule development in soybean, providing valuable information toward understanding the functions of GmNORK and GmNIC1 in symbiotic signaling and nodule development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soybean (Glycine max) is the major crop legume in the world, which is mainly used as an important food and animal feed. Soybean has the capability to establish endosymbiotic relationships with nitrogen-fixing bacteria, called rhizobia (Gonzalez-Guerrero et al. 2014). The establishment of legume-rhizobium symbiosis is a multistage process including signal perception, signal transduction, and subsequent downstream developmental events that eventually give rise to a new organ, the nodule, in which the intracellular bacterial symbionts thrive (Broughton et al. 2000). In nodules, the bacteria are enclosed in a membrane of plant origin giving rise to vesicular-like compartments, termed symbiosomes to draw attention to their quasi-organellar like status (Emerich and Krishnan 2014). Within the symbiosomes, the rhizobia differentiate into nitrogen-fixing form called bacteroids (Udvardi and Poole 2013). Then bacteroids convert atmospheric nitrogen (N2) into ammonia (NH3) for plant use in exchange of photosynthate from the plant host (Gibson et al. 2008).
The formation of symbiotic nodules requires two parallel signaling pathways, one that promotes nodule organogenesis while the other allows bacterial infection (Madsen et al. 2010). These processes are coordinated in both a spatial and temporal manner to ensure successful symbiotic development (Oldroyd et al. 2011). Both processes require plant recognition of the Nod factors (NF). NF as signaling molecules secreted by rhizobia, are perceived by plant hosts through two plasma membrane-located receptor-like kinases (RLK) NFR1 and NFR5 (Arrighi et al. 2006; Indrasumunar et al. 2010; Limpens et al. 2005; Madsen et al. 2003), and activated a leucine rich repeat (LRR) serine/threonine kinase then triggers downstream signaling cascades. The leucine rich repeat (LRR) serine/threonine kinase is one of important genes involves nodule initiation. These receptor-like kinase were given different names depending on plant species, SYMbiosis Receptor-like Kinase (SymRK) in L. japonicus or Nodulation Receptor Kinase (NORK) in Medicago sativa, SYM19 in Pisum sativum, and DMI2 (Does not Make Infections 2) in Medicago truncatula, and GmNORKa/b in Glycine max (Indrasumunar et al. 2015; Capoen et al. 2005; Endre et al. 2002; Mitra et al. 2004; Stracke et al. 2002). Recently, SYMRK/NORK was shown to be a co-receptor interacting with NFR5 to regulate the symbiotic signaling cascade (Antolin-Llovera et al. 2012). Almost all the knowledge regarding the function of SymRK/NORK was obtained from model legumes, such as Lotus and Medicago, but not from soybean.
The NORK/SymRK gene is characterized with a signal peptide, three LRR motifs, a transmembrane domain and a Ser/Thr kinase domain. Mutations in SymRK/NORK not only affect nodulation, but also impair arbuscular mycorrhization (Endre et al. 2002; Stracke et al. 2002). The NORK gene is essential for Nod factor perception/transduction in Medicago sativa since it is involved in the common signaling pathway of symbiosis (Endre et al. 2002). Many researchers have suggested that NORK/SymRK is involved in a protein complex with the Nod factor receptor or/and transduces the signaling of Nod factor receptors to subsequent events (Hogg et al. 2006; Riely et al. 2004). However, there is little mechanistic information as to how NORK/SymRK actually does work. NORK/SymRK is a protein containing 919 amino acid, leucine rich repeat (LRR) receptor-like kinase, containing a 325 amino acid serine/threonine kinase domain (Yoshida and Parniske 2005). Previous research had shown that NORK/SymRK is an active kinase (Kevei et al. 2007).
NIC1, a CLE peptide-encoding genes in soybean (Glycine max), play an important role in nitrate regulation of nodulation. NIC1 strongly responds to nitrate treatment but not to Bradyrhizobium sp. Inoculation. NIC1 functions locally to inhibit nodulation depending on root GmNARK activity. A SDI-like nitrate induced inhibitor (NII) of nodulation produced by the interaction of NIC1 with NARK in the root lead to local nitrate inhibition of nodulation (Reid et al. 2011). GmNARK is a common component during autoregulation of nodulation (AON) and nitrate regulation of nodulation. Both systemic and local regulations of nodulation respond to nitrate (Jeudy et al. 2010).
In this study, we reported that rhizobial inoculation decreases the expression level of GmNIC1 in GmNORK RNAi soybean transgenic roots, suggesting that NORK directly regulates AON directed by GmNIC. Our results provide important information in understanding of the functions of GmNORK and GmNIC1 in symbiotic signaling and nodule development.
Materials and methods
Alignment and phylogenetic analysis
NCBI BLAST searches using GmNORKa and GmNORKb detected a number of highly similar peptide sequences. The alignments of the SymRK (NORK) genes are in terms of neighbor joining (NJ) tree in newick format.
Plasmid construction and transformation
A 188-bp DNA fragment from the 42 bp downstream of the GmNORKb stop codon to the 229-bp was amplified from William 82 using GmNORKb-RNAi F/R primer pairs (Table S1), and cloned into the binary vector pDONR222 to generate gateway entry plasmid using BP reaction. Then the 188-bp DNA fragment was cloned into pCAM-GWi(GWY RNAi) using LR reaction to generate ICNO1a-RNAi plasmid. Empty vector and CGT5200 (GUS RNAi) was used as the RNAi control vector, as described previously (Govindarajulu et al. 2009; Libault et al. 2010). This vector contains an RNAi construct directed against GUS.
Agrobacterium rhizogenes-mediated hairy root transformation
The constructs were electroporated into Agrobacterium rhizogenes strain K599 using a GenePulser apparatus with pulse controller (Bio-Rad Laboratories, Hercules, California) with settings at 25 μF, 200 Ω, 1.8 kV. After electroporation, 100 μL LB medium was added to the competent cells, which were allowed to recover at 30 °C with shaking at 180 rpm for at least 2 h, then plated onto LB agar supplemented with appropriate antibiotics. Plates were incubated at 28 °C for 2–3 days. Agrobacterium rhizogenes-mediated hairy root transformations were performed as described in our laboratory protocol.
Seedlings preparation: Seeds of Glycine max (L.) Merrill cultivar Williams 82 were surface sterilized by 10% bleach, rinsed several times with autoclaved distilled water (diH2O), once with 0.8% HCl for 10 min, and then several times with diH2O. Sterilized seeds were then sown onto 1% agar round plates (20 cm diameter) and incubated in chambers at 27 °C and 80% humidity for 3 days in darkness and 3 days at 22 °C with a light regime of 16 h light and 8 h darkness.
Transformation: Agrobacterium rhizogenes K599 carrying the respective constructs was inoculated into liquid LB medium with appropriate antibiotics and incubated overnight at 30 °C with shaking at 200 rpm. Approximately 500 µl of the overnight bacterial culture were plated onto LB plates with antibiotics and bacteria were grown overnight in a 30 °C chamber. The 6-day old seedlings were cut at the base of the hypocotyl and the shoot was dipped into the bacterial confluent lawn that had developed overnight on the plates. The seedlings were placed onto square Petri dishes with Fahräeus medium (Fahraeus 1957) supplemented with 1 mM CaCl2 and 1 mM KNO3. Plantlets were incubated in the growth chamber at 22 °C for 2 days in darkness. Plantlets were transferred into plastic growth pouches (Mega International, Minneapolis, MN) containing 50 ml solid Fahraeus medium supplemented with CaCl2 and KNO3 and grown under a 16-h photoperiod for 5–7 days at 22 °C. After 5–7 days, adventitious roots were removed and the composite plants transferred into growth pouches with liquid Fahraeus medium supplemented with CaCl2 and KNO3. Approximately 3 weeks after transformation, formation of transgenic roots was checked by GFP florescence and non-transgenic roots were removed. If the roots were long enough (at least ca. 3 cm), the seedlings were planted into a vermiculite and perlite mixture (3:1), grown in greenhouse conditions and inoculated with the soybean rhizobial symbiont, Bradyrhizobium japonicum.
Nodulation assay
For soybean inoculation with rhizobia, B. japonicum wild-type strain USDA110 (Halverson and Stacey 1986), or a derivative strain constitutively expressing β-glucuronidase (GUS) (Loh et al. 2001)were grown at 30 °C for 3 days prior to inoculation in liquid HM medium (Cole and Elkan 1973) supplemented with appropriate antibiotics (50 μg/ml tetracycline, 100 μg/ml spectinomycin) Cells were grown to an OD600 between 0.5 and 1.0. Cells were then pelleted by centrifugation at 4000 rpm at 20 °C for 10 min and resuspended in sterilized MilliQ water to a final OD600 of 0.05. Composite plants growing in a vermiculite and perlite mixture were inoculated 2 days later with 2 mL of the respective B. japonicum strain. Plants were left in the laboratory for 2 days for acclimatization and then transferred into the greenhouse.
The nodulation phenotypes of soybean roots inoculated with B. japonicum were analyzed 4 weeks post-inoculation (wpi) using a Leica M205 FA stereo microscope with a high resolution Leica DFC295 color camera and Leica Application Suite Software. Only the nodules formed on transgenic roots, as determined by expression of the GFP marker, were analyzed.
ß-glucuronidase assay
For histochemical ß-glucuronidase staining, transgenic roots of composite plants were cut and were placed into 15-ml Falcon tubes containing 10 ml of GUS staining solution as previously described (Toth et al. 2012). Vacuum was applied three times for 3 min, and then the roots were incubated at 37 °C in the dark for 2 days overnight. Infected roots and nodules were checked for blue staining under a stereomicroscope. Evaluation and documentation of nodules and primordia were performed using stereomicroscope (Olympus SZX12; Olympus, Tokyo, Japan) fitted with an Olympus DP10 camera.
Quantitative real-time RT-PCR analysis
Transgenic root were collected at 28 d after inoculation (DAI), All samples were immediately frozed in liquid nitrogen and stored at − 80 °C untile use. Total RNA from transgenic root tissues used in this study were isolated by TRizol (Ambion, Austin, TX, USA), followed by a DNase treatment (Turbo DNase, Ambion), and 1 micrograms root total RNA were used for complementary DNA synthesis using M-MLV Reverse Transcriptase (Promega, Madison, WI, USA) according to the manufacturer’s instructions.
qPCR experiments were conducted with gene-specific primers (Table S1.) in the reaction system of SYbGreen mix (Bio-Rad) on the 7500 System (Applied Biosystems) according to the manufacturer’s instructions. The thermal profile of the qRT-PCR reactions was 50 °C for 2 min, 95 °C for 10 min, 40 cycles of 95 °C for 15 s, and 60 °C for 1 min. The geometric mean of cons4 and cons6, encoding an ATP-binding cassette transporter and an F-box protein (Govindarajulu et al. 2009), were used as reference genes to normalize the expression levels. The relative expression levels of the gene between the two samples is calculated using 2 (−ΔΔCt) method. The average of the expression levels between three different replicates was calculated.
Recombinant protein purification and kinase assay
The plasmid pGENORK was transformed into BL21 (DE3). Protein production was induced by adding 0.3 mM isopropyl b-D-thiogalactopyranoside (IPTG) when the culture reached an OD600 of 0.6. The protein was extracted after inducing the cultures for 24 h at 16 °C. Recombinant protein purification was done according to the following procedure. Bacterial cells from 200 ml LB medium were pelleted by centrifugation at 4 °C at 8000 rpm for 10 min and resuspended in 10 ml 1 × PBS solution supplemented with 1 × EDTA-free protease inhibitor (Roche, Indianaplois, IN), 0.5% Triton X-100, 1 mg/ml lysozyme and put on ice for 30 min. The cells were lysed by sonication before centrifuging at 13,000 rpm for 10 min at 4 °C. Supernatants were used for protein affinity purification using Glutathione Sepharose 4B (GE Healthcare, Milwaukee, WI). The column was washed with at least 20 bed volumes of 1 × PBS solution. The eluted proteins, including GST and GSTNORKKD, were dialyzed with buffer [50 mM Tris (PH 7.5), 50 mM KCl, 2 mM DTT, and 10% glycerol].
The in vitro kinase assays used 1 mg purified protein in a buffer containing 50 mM Tris (PH 7.5), 50 mM KCl, 2 mM DTT, 5 mM MnCl2, 5 mM MgCl2, 10% glycerol, 10 mM ATP and 5mMCi[γ-32P]-ATP. The assay mix was incubated at 28 °C for 30 min and the reaction was stopped by adding 1 × SDS loading buffer. The samples were separated on 10% SDS-PAGE gel and the gel imaged by autoradiography using phosphor screens and a phosphor imager (Emerich and Krishnan 2014).
Statistical analysis
Data for nodule numbers were statistically analyzed using a one-way analysis of variance (ANOVA). The expression levels of genes were analyzed using Student’s t test.
Results
GmNORK structure
As a diploidized ancient tetraploid species, soybean has twocopies of NORK genes in its genome, referred to as GmNORKa and GmNORKb. Through analysis of their genomic features, both GmNORKa and GmNORKb have 15 Exons and 14 introns. Both the predicted proteins have signal peptides (SP) at N-terminal, leucine-rich-repeat domain (LRR), trans-membrane domain (TM) and Serine-Threonine/tyrosine kinase domain (PK) (Fig. 1). Both GmNORKa and GmNORKb both contain 919 a.a and share 94.9% identity (Fig. 2).
NORK1a and NORK1b gene structure. Genomic structure of NORK1a and NORK1b with indicated predicted protein domains. Exons are indicated as boxes, introns as black line. SP, predicted signal peptide; EC, extracellular domain; LRR, Leucine-rich repeat motifs; TM, transmembrane domain; PK, protein kinase domain
Molecular phylogenetic analysis of GmNORKa and GmNORKb
To investigate the relationship of GmNORKa and GmNORKb proteins with other homologous proteins in soybean and other plant species, a phylogenetic tree based on amino acid sequences was generated (Fig. 3). GmNORKa and GmNORKb share 94.9% identity, and they are the closest homologs. They likely derive from a common ancestor and have close homologs in Phaseolus vulgaris (ADQ74920.1) and Sesbania rostratate (AAV88623.1), both are legumes that support a rhizobium symbiosis. It shows that the SymRK (NORK) orthologs are divided into three groups, Rhizobium (Fabaceae, legume), actinorhizal (non-legume) nodule-forming plants and non-nodule-forming plants. G.max NORK and P. vulgaris NORK are highly similar. Within the legume orthologs, A. hypogaea is the most distant ancestor since the SymRK of A. hypogaea shows the lowest similarity to GmNORK (Fig. 3).
RNA silencing GmNORK resulted in reduced nodule number
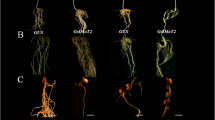
To study the function of GmNORK involved in root nodule symbiosis in soybean, specified RNAi contruct of GmNORK was generated and transformed into soybean roots by agrobacterium rhizogenes-mediated hairy root transformation. Transgenic roots were identified with the green fluorescent protein (GFP) marker expressed in the binary vector (Fig. 4a). Nodule morphology was examined by using GUS staining. The nodule morphology from the RNAi roots showed no obvious structural changes relative to the controls (Fig. 4b, c). The number of nodules formed on the GmNORK RNAi was approximately 6.4% that of the controls (with P < 0.01) (Fig. 4d). The results demonstrated that GmNORKa and GmNORKb are involved in nodule formation in soybean plants after inoculated with B. japonicum.
RNA silencing of GmNORK resultes in reduced nodule number. a Representative transgenic root and nodules expressing GUS RNAi, EV RNAi and GmNORK RNAi constructs. Scale bars represent 1 mm. b Stained micrographs of nodule derived from RNAi transgenic roots at 21 days after inoculate with B. japonicum. Scale bars represent 1 mm. c Stained micrographs of nodule sections derived from RNAi transgenic roots at 21 days after inoculate with B. japonicum. The nodule morphology was similar among the nodules formed on roots expressing the GUS RNAi, Empty vector (EV), and GmNORK RNAi constructs. Scale bars represent 1 mm. EV RNAi indicates Empty vector RNAi. d Nodulation was measured as nodule number per transgenic root. The nodule numbers per root in GmNORK RNAi transgenic lines decreased significantly compared to which in GUS RNAi and Empty vector transgenic lines. ** denotes significantly different at P < 0.01. Bars indicate standard error of the mean from 30 plants
The expression of GmNORKa, GmNORKb, and GmNIC1 in GmNORK RNAi lines
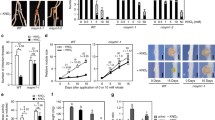
Due to the lack of homozygous knockout mutants for GmNORKa and GmNORKb in soybeans, RNA silencing was applied to investigate their functions in nodulation. Soybean roots were transformed with GmNORK RNAi, an empty vector and a vector expressing an RNAi construct specifically targeted to β-glucuronidase (GUS) (GUS RNAi) as controls. GmNORKa and GmNORKb expression levels in GmNORK RNAi are approximately 37% and 24% of that in the control transgenic roots, respectively (Fig. 5a, b). The results suggest that GmNORK plays a critical role in mediating root nodule symbiosis (RNS). GmNIC1 play a critical role in AON system, which tightly control the soybean nodule numbers. In order to study whether GmNORK involves the GmNIC1 associated nodulation, we analyzed the GmNIC1 expression in GmNORK RNAi lines. GmNIC1 expression levels in GmNORK RNAi are approximately 21% of that in the control transgenic roots (Fig. 5c).
The expression of GmNORK1a, GmNORK1b and GmNIC1 in soybean roots was significantly reduced in GmNORK RNAi lines. aGmNORK1a relative expression levels in roots of soybean transformed with negative control vector, GmNORK RNAi plasmids. bGmNORK1b relative expression levels in roots of soybean transformed with negative control vector, GmNORK RNAi plasmids. cGmNIC1 relative expression levels in roots of soybean transformed with negative control vector, GmNORK RNAi plasmids. ** denotes significantly different at P < 0.01. Bars indicate standard error of the mean from 3 plants. Control indicates Control indicates geometric mean of GUS RNAi and Empty vector RNAi
Recombinant protein purification and kinase assay
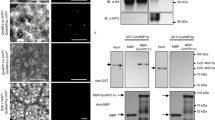
In order to confirm GmNORK phosphorylating activation, we performed in vitro phosphorylation assays. GmNORK kinase domain (538 a.a. to 919 a.a.) was expressed by fusing with the glutathione sulfotransferase domain. As shown in Fig. 6, GmNORK was shown to exhibit autophosphorylation in vitro, consistent with previous reports (Samaddar et al. 2013; Yoshida and Parniske 2005).
Discussion
GmNIC1 functions locally to inhibit nodulation depend on root GmNARK activity and strongly responds to nitrate treatment but not to Bradyrhizobium sp. Inoculation (Reid et al. 2011). The interaction of NIC1 with NARK in the root probably produces a SDI-like nitrate induced inhibitor (NII) of nodulation leading to local nitrate inhibition of nodulation. Thus, GmNARK is a common component during AON and nitrate regulation of nodulation (Reid et al. 2011). Both systemic and local regulations of nodulation respond to nitrate (Jeudy et al. 2010).
SYMRK/NORK, a component immediately downstream of these Nod factor receptors, is central to the Nod factor signaling cascade (Indrasumunar et al. 2010; Stracke et al. 2002). Several interacting NFRs and SYMRK/NORK proteins have been identified, including SINA4 (Seven In Absentia 4), HMGR1 (3-hydroxy-3-methylglutaryl-CoA reductase 1), PUB1 (Plant U-box protein 1) and SYMREM1 (Symbiotic Remorin 1). How the signal is transferred from the plasma membrane to the nuclear envelop remains unclear. Three nucleoporin (NUP85, NUP133, and NENA) and two cation channel proteins (CASTOR and POLLUX) are invoved in symbiotic calcium oscillations generation. The nucleus-localized Ca2+/calmodulin dependent protein kinase and CYCLOPS are used in decoding these calcium oscillations, and activate the transcription factor (TF) for symbiosis-associated gene expression (Mitra et al. 2004). However, the mechanism via which Nod factor signaling affects nodule development, nuclear size and the chromatin structure remains unknown.
Soybean nodule numbers are tightly controlled by AON. GmNIC1 play a critical role in AON system (Reid et al. 2011). GmNORK plays a critical role in mediating root nodule symbiosis (RNS). GmNORK probably involves the GmNIC1 associated nodulation. As seen in Fig. 4, RNA silencing GmNORK resulted in reduced nodule number, consistent with previous reports (Indrasumunar et al. 2015). Our studies show that the GmNIC1 expression level is decreased in GmNORK RNAi transgenic soybean roots. These data suggest that GmNORK affect nodule development and nitrate regulation of nodulation by decreasing the GmNIC1 expression level. The details of the mechanism behind the GmNORK affect GmNIC expression level require further elucidation.
References
Antolin-Llovera M, Ried MK, Binder A, Parniske M (2012) Receptor kinase signaling pathways in plant–microbe interactions. Annu Rev Phytopathol 50:451–473
Arrighi JF, Barre A, Ben Amor B, Bersoult A, Soriano LC, Mirabella R, de Carvalho-Niebel F, Journet EP, Gherardi M, Huguet T, Geurts R, Dénarié J, Rougé P, Gough C (2006) The Medicago truncatula lysin [corrected] motif-receptor-like kinase gene family includes NFP and new nodule-expressed genes. Plant Physiol 142:265–279
Broughton WJ, Jabbouri S, Perret X (2000) Keys to symbiotic harmony. J Bacteriol 182:5641–5652
Capoen W, Goormachtig S, De Rycke R, Schroeyers K, Holsters M (2005) SrSymRK, a plant receptor essential for symbiosome formation. Proc Natl Acad Sci USA 102:10369–10374
Cole MA, Elkan GH (1973) Transmissible resistance to penicillin G, neomycin, and chloramphenicol in Rhizobium japonicum. Antimicrob Agents Chemother 4:248–253
Emerich DW, Krishnan HB (2014) Symbiosomes: temporary moonlighting organelles. Biochem J 460:1–11
Endre G, Kereszt A, Kevei Z, Mihacea S, Kalo P, Kiss GB (2002) A receptor kinase gene regulating symbiotic nodule development. Nature 417:962–966
Fahraeus G (1957) The infection of clover root hairs by nodule bacteria studied by a simple glass slide technique. J Gen Microbiol 16:374–381
Gibson KE, Kobayashi H, Walker GC (2008) Molecular determinants of a symbiotic chronic infection. Annu Rev Genet 42:413–441
Gonzalez-Guerrero M, Matthiadis A, Saez AN, Long TA (2014) Fixating on metals: new insights into the role of metals in nodulation and symbiotic nitrogen fixation. Front Plant Sci 5:45
Govindarajulu M, Kim SY, Libault M, Berg RH, Tanaka K, Stacey G, Taylor CG (2009) GS52 ecto-apyrase plays a critical role during soybean nodulation. Plant Physiol 149:994–1004
Halverson LJ, Stacey G (1986) Effect of lectin on nodulation by wild-type Bradyrhizobium japonicum and a nodulation-defective mutant. Appl Environ Microbiol 51:753–760
Hogg BV, Cullimore JV, Ranjeva R, Bono JJ (2006) The DMI1 and DMI2 early symbiotic genes of medicago truncatula are required for a high-affinity nodulation factor-binding site associated to a particulate fraction of roots. Plant Physiol 140:365–373
Indrasumunar A, Kereszt A, Searle I, Miyagi M, Li D, Nguyen CD, Men A, Carroll BJ, Gresshoff PM (2010) Inactivation of duplicated nod factor receptor 5 (NFR5) genes in recessive loss-of-function non-nodulation mutants of allotetraploid soybean (Glycine max L. Merr.). Plant Cell Physiol 51:201–214
Indrasumunar A, Wilde J, Hayashi S, Li D, Gresshoff PM (2015) Functional analysis of duplicated Symbiosis Receptor Kinase (SymRK) genes during nodulation and mycorrhizal infection in soybean (Glycine max). J Plant Physiol 176:157–168
Jeudy C, Ruffel S, Freixes S, Tillard P, Santoni AL, Morel S, Journet EP, Duc G, Gojon A, Lepetit M, Salon C (2010) Adaptation of Medicago truncatulato nitrogen limitation is modulated via local and systemic nodule developmental responses. New Phytol 185:817–828
Kevei Z, Lougnon G, Mergaert P, Horvath GV, Kereszt A, Jayaraman D, Zaman N, Marcel F, Regulski K, Kiss GB, Kondorosi A, Endre G, Kondorosi E, Ane JM (2007) 3-Hydroxy-3-methylglutaryl coenzyme a reductase 1 interacts with NORK and is crucial for nodulation in Medicago truncatula. Plant Cell 19:3974–3989
Libault M, Zhang XC, Govindarajulu M, Qiu J, Ong YT, Brechenmacher L, Berg RH, Hurley-Sommer A, Taylor CG, Stacey G (2010) A member of the highly conserved FWL (tomato FW2.2-like) gene family is essential for soybean nodule organogenesis. Plant J 62:852–864
Limpens E, Mirabella R, Fedorova E, Franken C, Franssen H, Bisseling T, Geurts R (2005) Formation of organelle-like N2-fixing symbiosomes in legume root nodules is controlled by DMI2. Proc Natl Acad Sci USA 102:10375–10380
Loh JT, Yuen-Tsai JP, Stacey MG, Lohar D, Welborn A, Stacey G (2001) Population density-dependent regulation of the Bradyrhizobium japonicum nodulation genes. Mol Microbiol 42:37–46
Madsen EB, Madsen LH, Radutoiu S, Olbryt M, Rakwalska M, Szczyglowski K, Sato S, Kaneko T, Tabata S, Sandal N, Stougaard J (2003) A receptor kinase gene of the LysM type is involved in legume perception of rhizobial signals. Nature 425:637–640
Madsen LH, Tirichine L, Jurkiewicz A, Sullivan JT, Heckmann AB, Bek AS, Ronson CW, James EK, Stougaard J (2010) The molecular network governing nodule organogenesis and infection in the model legume Lotus japonicus. Nat Commun 1:10
Mitra RM, Gleason CA, Edwards A, Hadfield J, Downie JA, Oldroyd GE, Long SR (2004) A Ca2+/calmodulin-dependent protein kinase required for symbiotic nodule development: gene identification by transcript-based cloning. Proc Natl Acad Sci USA 101:4701–4705
Oldroyd GE, Murray JD, Poole PS, Downie JA (2011) The rules of engagement in the legume-rhizobial symbiosis. Annu Rev Genet 45:119–144
Reid DE, Ferguson BJ, Gresshoff PM (2011) Inoculation- and nitrate-induced CLE peptides of soybean control NARK-dependent nodule formation. Mol Plant Microbe Interact 24:606–618
Riely BK, Ane JM, Penmetsa RV, Cook DR (2004) Genetic and genomic analysis in model legumes bring Nod-factor signaling to center stage. Curr Opin Plant Biol 7:408–413
Samaddar S, Dutta A, Sinharoy S, Paul A, Bhattacharya A, Saha S, Chien KY, Goshe MB, DasGupta M (2013) Autophosphorylation of gatekeeper tyrosine by symbiosis receptor kinase. FEBS Lett 587:2972–2979
Stracke S, Kistner C, Yoshida S, Mulder L, Sato S, Kaneko T, Tabata S, Sandal N, Stougaard J, Szczyglowski K, Parniske M (2002) A plant receptor-like kinase required for both bacterial and fungal symbiosis. Nature 417:959–962
Toth K, Stratil TF, Madsen EB, Ye J, Popp C, Antolin-Llovera M, Grossmann C, Jensen ON, Schussler A, Parniske M, Ott T (2012) Functional domain analysis of the Remorin protein LjSYMREM1 in Lotus japonicus. PLoS ONE 7:e30817
Udvardi M, Poole PS (2013) Transport and metabolism in legume-rhizobia symbioses. Annu Rev Plant Biol 64:781–805
Yoshida S, Parniske M (2005) Regulation of plant symbiosis receptor kinase through serine and threonine phosphorylation. J Biol Chem 280:9203–9209
Acknowledgements
This work was supported by the PhD Start-up Fund of Natural Science Foundation under Grant No. 801100010121; Open fund project of State Key laboratory of Crop Genetics and Germplasm Enhancement under Grant No. ZW201706; Open fund project of oil crops biology and genetics and breeding key laboratory of Ministry of Agriculture under Grant No. 2016006. Open funds of State Key Laboratory of Agricultural Microbiology under Grant No. AMLKF201608.
Author information
Authors and Affiliations
Contributions
LW, YW, YJ, YC, LD, and XB contributed equally to this work.
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, L., Deng, L., Bai, X. et al. Regulation of nodule number by GmNORK is dependent on expression of GmNIC in soybean. Agroforest Syst 94, 221–230 (2020). https://doi.org/10.1007/s10457-019-00382-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10457-019-00382-8