Abstract
The fundamental Hennigian principle, grouping solely on synapomorphy, is seldom used in modern phylogenetics. In the submitted paper, we apply this principle in reanalyzing five datasets comprising 197 complete plastid genomes (plastomes). We focused on the latter because plastome-based DNA sequence data gained dramatic popularity in molecular systematics during the last decade. We show that pattern-cladistic analyses based on complete plastid genome sequences can successfully resolve affinities between plant taxa, simultaneously simplifying both the genomic and analytical frameworks of phylogenetic studies. We developed “Matrix to Newick” (M2N), a program to represent the standard molecular alignment of plastid genomes in the form of trees or relationships directly. Thus, massive plastome-based DNA sequence data can be successfully represented in a relational form rather than as a standard molecular alignment. Application of methods of median supertree construction (the Average Consensus method has been used as an example in this study) or Maximum Parsimony analysis to relational representations of plastome sequence data may help systematist to avoid the complicated assumption-based frameworks of Maximum Likelihood or Bayesian phylogenetics that are most used today in massive plastid sequence data analyses. We also found that significant amounts of pure genomic information that typically accommodate the majority of current plastid phylogenomic studies can be effectively dropped by systematists if they focus on the pattern-cladistics or relational analyses of plastome-based molecular data. The proposed pattern-cladistic approach is a powerful and straightforward heuristic alternative to modern plastome-based phylogenetics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
This paper demonstrates how to resolve affinities between plant taxa within the pattern-cladistic framework based on complete chloroplast genomes (plastomes). We focused on the latter because plastome-based DNA sequence data gained dramatic popularity in molecular systematics during the last decade. This Introduction must explain to the reader the pattern-cladistic framework and why resolving systematic relationships within this framework is beneficial for systematists. An articulation of the goals of this study follows this explanation.
1.1 What is the Pattern-Cladistic Framework?
Cladistics is a method of classification that arranges taxa hierarchically into nested sets based solely on the synapomorphies (or homologies) simultaneously representing such sets in the form of cladograms or, which is the same, branching diagrams that include no connotation of ancestry and has no implied time axis (Kitching et al. 1998; Williams and Ebach 2006, 2008, 2020). If homology is defined as a relationship between different organisms based on their parts (or homologues) (Kitching et al. 1998; Williams and Ebach 2006, 2008, 2020; Nelson 2011), then a pattern-cladistic (or Cladistics (written with a capital letter “C”)) (Williams and Ebach 2006, 2008, 2020) framework can be defined as a way of representing conventional character data in a matrix-free form or, which is the same, in the form of relationships, that is either in tree form (Fig. 1) (Williams and Ebach 2006, 2008, 2020; Mavrodiev et al. 2017, 2019) or, in a state of a standard three-taxon statement (3TS) matrix (reviewed in Kitching et al. 1998; Williams and Ebach 2008) but not in the form of a traditional (ordinary) data matrix (Williams and Ebach 2006, 2008; Mavrodiev et al. 2017, 2019). The other versions of pattern-cladism based on the semantic reinterpretation of conventional Maximum Parsimony (MP) analysis (Farris 1983) are also available (Brower 2000). Still, we prefer to focus on what we accept as the most comprehensive.
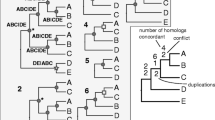
The central idea of this paper: the set of pre-selected informative characters from the plastome-based global alignment (a) can be represented as a forest (array) of the rooted trees (b) following the values of an a priori defined taxon (the operational outgroup). This forest becomes the subject of subsequent AC analysis (or the other methods of constructing median supertrees). In this way, process-related phylogenetic frameworks may be avoided by systematists. Relational representations of the genomic data (b) and the process-free semantic framework of pattern-cladistics (e.g., the AC analysis of the “maximal” relationships (Mavrodiev et al. 2017)) can be used instead
Standard molecular sequence alignments (e.g., of the complete plastid genomes or their subsets) are an excellent example of an ordinary data matrix (Williams and Ebach 2006). Thus, when analyzing plastome-based sequence data within the pattern-cladistic framework, researchers operate not with the standard molecular alignments but with relational representations of them. We call these representations 'relational' because they are establishing traditional informative DNA characters (the columns of the molecular alignment) either in the form of the maximal (reviewed in Williams and Ebach 2006, 2008; Mavrodiev et al. 2017) or minimal relationships (or 3TSs) (reviewed in Kitching et al. 1998; Williams and Ebach 2008; Mavrodiev et al. 2019) following the character-state values of the pre-selected outgroup taxon. Or if we repeat ourselves, either in a tree form (Fig. 1) or in the form of a 3TS matrix.
Because of this, pattern-cladistics is the one analytical approach that has today fully implemented the fundamental Hennigian principle: grouping solely on synapomorphy (Nelson 2011; Williams and Ebach 2008, 2020; Mavrodiev et al. 2017, 2019). Thus, analytical accuracy is the first and major benefit any researcher can get by working within the pattern-cladistic framework.
Analytical focus on the relational representation of the commonly used (ordinary) matrices is a new step in developing the pattern-cladistic methodology of pattern-cladistic methodology (Platnick 1979; Nelson 2011; Williams and Ebach 2008, 2020). Detailed reviews of the history of pattern-cladism and related topics have been given previously (Brower 2019; Williams and Ebach 2008, 2020), but here we would like to emphasize that the pattern-cladistic framework is based on the idea that patterns have priority over processes. Pattern-cladistics thus excludes all assumptions about hypothetical historical processes from the analytical component of the data analysis. In this sense, the pattern-cladistics framework is a model-free, static approach that focuses mainly on the comparative aspects of the data analysis.
1.2 Why Else is the Pattern-Cladistic Framework Beneficial for Plant Systematists Using Plastome-Based Data?
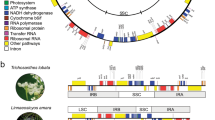
Complete plastid genomes are useful in molecular plant systematics, even if the discrimination power of plastomes is not always sufficient to resolve phylogenetic patterns at low taxonomic levels (e.g., Chen et al. 2022; Wagner et al. 2021). Plastomes are widely used in modern molecular plant systematics. However, current molecular systematic studies based on plastomes typically still resemble the early works within the field, where almost every paper on this topic includes detailed maps of selected gene regions (e.g., 26S and 18S loci), its statistics, architecture, discussion of its possible evolution, and the other details that had not been included into the actual phylogenetic analyses. More recent papers look at these issues in greater detail, but the fundamental approach and apparent interests are the same.
Today, almost every paper that uses plastome sequences for phylogenetic reconstructions typically contains a significant amount of genomic information that is interesting primarily to the experts in genetics and genomics but not to the plant systematists. Indeed, there tends to be a strong focus on issues that are not strictly necessary to the phylogenetic analysis itself, such as the detection of tandem repeats; detailed, frequently colored illustrations of the gene maps of the complete plastid genomes, including descriptions of its gene content based on the functional categories; data on nucleotide diversity of the entire plastomes; comparison of the borders of inverted repeat (and\or other selected regions of the genome) between the different taxa; detailed lists of genes in the analyzed plastomes; various statistics of protein-coding and non-coding regions; the estimations of the putative rates of positive selection of the particular genes (or sites) of the plastome; and so on, frequently accompanied by relatively complicated graphics and diagrams. The primary issue is that, in practice, only a tiny part of this genomic information can be helpful in phylogenetic analyses. We deliberately do not provide references to specific papers since we believe the outlined picture refers to most of the past and recent phylogenetic studies grounded in complete plastid genomes. Thus, we are not criticizing any specific work or research group but attempting to conduct a preliminary conceptual analysis of the situation within the given branch of modern molecular systematics.
However, if the goal of the systematic study is the estimation of taxonomic patterns among studied taxa, not the elucidation of genome sequences in terms of their structures and features, then the primary thing that a systematist must seek from plastome sequence data is the set of valuable relationships, regardless of their ‘nesting’ in the particular regions of the complete plastid genomes. Thus, massive chloroplast sequence data can be significantly simplified for systematic purposes. And the focus of molecular systematic studies can therefore be shifted from genomic phenomenology to relationships per se and, as the next step—to the taxonomy of the studied groups. Thus, simplifying the content of plastome-based data is the second benefit of the pattern-cladistic framework for the plant molecular systematist.
To make such simplification, here we would like to show that plant molecular systematists may handle plastome-based sequence data in the same way as a single locus sequence but with the final representation of all informative characters as a forest (array) of trees (Fig. 1). This means that much of the sequence data within complete plastid genomes may be effectively dropped by the researcher. Instead, the researcher may operate simply with the array of plastome-based relationships. The study’s first goal is to demonstrate the efficiency of such a simplification (Fig. 1).
To explain the study’s second goal, which also highlights the next benefit of the pattern-cladistic framework, we must address a common misunderstanding: that phylogenetic analyses reconstruct actual evolutionary processes. This is a mistake because the latter always remains unknown. However, current phylogenetics carries this as a basic assumption by including process-based models of character evolution (reviewed in Felsenstein 2004) implemented in analytical software for statistical phylogenetics (Felsenstein 2004). Most modern plastome-based phylogenetic analyses have been completed using complicated analytical tools from the arsenal of statistical or model-based phylogenetic analyses.
But for a systematist who compares the plastomes of the different plant taxa for systematic purposes, is it possible to analyze plastome-based sequence data more straightforwardly? E.g., within a framework free from either simple or complicated assumptions about any putative processes that might have taken place in the past? As we argue above, such a framework is the pattern-cladistic framework. Thus, the second goal of this study is to develop a program for pattern-cladistic analyses of plastome-based sequence data using a novel, simple, analytically designed approach. The simplification of the analytical context of the plastome-based molecular systematic studies is the third benefit of the pattern-cladistic framework.
2 Materials and Methods
Figures 1, S1 and S2 contain a graphical explanation of the main points of this section, but at the beginning, it is critical to clarify the taxonomic sampling of the study.
2.1 Taxonomic Sampling of the Study
For this study, we selected the following comprehensive plastome-based datasets:
-
1.
The set of the complete plastid genomes of the certain taxa of the recent subfamilies Agapanthoideae, Allioideae, Gilliesieae, Leucocoryneae, and Tulbaghieae (Amaryllidaceae) and its relatives (families Asparagaceae, Iridaceae, and Xanthorrhoeaceae) (32 plastomes) (Namgung et al. 2021);
-
2.
Of the tribe Gentianeae (Gentaniaceae) (33 plastomes) (Zhang et al. 2020);
-
3.
Of the subfamilies Borasseae, Calameae, Caryoteae, Chuniophoeniceae, Corypheae, Cryosophileae, Eugeissoneae, Lepidocaryeae, Phoeniceae, Sabaleae, and Trachycarpeae (Arecaceae) and genera Baxteria, Calectasia, Dasypogon, and Kingia (Dasypogonaceae) (42 plastomes) (Barrett et al. 2016);
-
4.
Of the tribe Tordylieae (Apiaceae) (39 plastoms) (Samigullin et al. 2022). To ensure the accuracy of the alignment, the newly reassembled genome of Angelica sinensis (Samigullin et al. 2022) has been excluded from future considerations. Because authors used their original nomenclatural solutions that are frequently different from the solutions proposed by GenBank, here we decided to provide a summary of discrepancies (Samigullin et al. 2022). Anethum foeniculum (binomial, assigned to taxon by the GenBank) = Foeniculum vulgare (binomioal in the tree from (Samigullin et al. 2022) (Fig. 5a)), Ligusticum hispidum = Ligusticopsis hispida, Ligusticum involucratum = Ligusticopsis involucrate, Ligusticum jeholense = Conioselinum smithii, Ligusticum pteridophyllum = Conioselinum pteridophyllum, Ligusticum sinense = Conioselinum anthriscoides, Ligusticum tenuissimum = Conioselinum tenuissimum, Ligusticum thomsonii = Seseli mucronatum, Tetrataenium candicans = Heracleum candicans, Tetrataenium yunnanense = Heracleum yunnanense (Samigullin et al. 2022);
-
5.
Of Diplazium (Athyriaceae) and related taxa (51 plastomes) (Wei and Zhang 2020). The summary of the nomenclature discrepancies is: Diplazium virescens (binomial, assigned to taxon by the GenBank) = D. conterminum (binomial in the tree from Wei and Zhang (2020) (Fig. 6a)), Athyrium opacum = Cornopteris opaca, Christella appendiculata = Cyclosorus procerus (Wei and Zhang 2020).
In summary, five plastome-based datasets that, in total, include 197 complete chloroplast genomes have been analyzed in this paper. Limniris sanguinea, Aucuba chlorascens (MT338539) (Huang et al. 2022), Typha latifolia, Carum carvi, and Macrothelypteris torresiana were selected accordingly as a single outgroup of each dataset.
2.2 Alignments
The first step (Fig. S2) in simplifying the plastome data is the construction of the standard global alignment of massive plastid DNA sequence data. The second step (Fig. S2) is the inclusion for future analyses of only molecular characters that give evidence for ‘grouping’ within the framework of phylogenetic systematics (parsimony-informative characters), but with the future exclusion of the poorly aligned (non-conserved, noisy, saturated, etc.) positions (Castresana 2000) (see below).
All genomic alignments were performed using MAFFT version 7.0 (Katoh and Standley 2013) as implemented in CIPRES (Miller et al. 2010) following the automatic selection of the appropriate strategy for the calculation. The algorithm's FFT-NS-2 strategy (Katoh and Standley 2013) was always selected by MAFFT version 7.0 as the best choice.
Informative characters were extracted from the matrices of the aligned plastomes by PAUP* version 4.0a (Swofford 2002) with the subsequent removal of the poorly aligned characters by the G-block (Castresana 2000) as implemented in SeaView version 4.0 (Gouyet al. 2010) following both “more” and “less” stringent strategies of the algorithm (Castresana 2000; Gouyet al. 2010).
The summary of the G-block methodology is already given in the literature (Talavera and Castresana 2007). The Gblocks algorithm removes essentially gapped as well as highly variable regions of an alignment according to several rules creating a unique set of “blocks” (or parts) ((Castresana 2000, p. 542) that are concatenated into a single “clean” alignment for use in further analyses (Castresana 2000). As pointed out by Talavera and Castresana (2007), “… Glocks selected for inclusion must be free from a large number of contiguous non conserved positions, must be flanked by highly conserved positions, and must have a minimum length, as controlled by the corresponding parameters. In addition, positions with gaps can be removed either always or only when more than half of the sequences contain gaps… The condition we call “stringent” does not allow any gap position. The relaxed condition allows gap positions if they are present in less than half of the sequences…” (Talavera and Castresana 2007, p. 567). In short, the method eliminates poorly aligned (i.e., essentially gapped and highly variable) positions of molecular alignment (Castresana 2000) and by this way “cleans” the latter from the over-divergent regions. From a cladistic perspective, the Gblock strategy can be viewed as an incomplete procedure to get an alignment rid of the mutually incongruent characters and, in this way, minimize the character conflicts. The Gblock method does not require any substitution matrix or the evolutionary model.
Finally, the sets of mutually compatible informative characters were extracted from the G-block outputs using PHYLIP version 3.69 (Baum 1989; Felsenstein 1993) and were the subjects of the 3TS permutations with fractional weighting in TAXODIUM version 1.2 (Mavrodiev and Madorsky 2012). Depending on the dataset, the values of the operational outgroups for the 3TS permutations (Williams and Siebert 2000) have been fixed as values of Limniris sanguinea (Iris sanguinea), Aucuba chlorascens, Typha latifolia, Carum carvi, and Macrothelypteris torresiana following the logic of Williams-Siebert (WS) representation of unordered multistate characters (Williams and Siebert 2000; Mavrodiev and Madorsky 2012).
2.3 Trees and Analyses
The analytical focus of the study includes the Average Consensus (AC) method (Lapointe and Cucumel 1997; Lapointe and Levasseur 2004; Swofford 2002; Creevey and McInerney 2009), as it applies to the forests of maximal relationships (Mavrodiev et al. 2017) as well as the three-taxon statement analysis (3TA) (Nelson and Platnick 1991, reviewed in Kitching et al. 1998; Williams and Ebach 2008, 2020), as conceptualized for unordered multistate characters (Williams and Siebert 2000; Mavrodiev and Madorsky 2012). Our current analytical focus on the AC method is non-mandatory (Mavrodiev et al. 2017, 2019) and, in theory, other methods of construction of median supertrees can be used to analyze the arrays of the maximal trees (Mavrodiev et al. 2017, 2019).
Again, following the values of the pre-selected outgroup taxa (all listed above), every G-block output with no mutually incompatible characters excluded was established as a forest of rooted trees using the newly designed program M2N (Fig. S1).
The M2N utility is written in portable C + + , and the source code compiles equally well with Microsoft Visual C + + and the GNU C + + compiler. M2N accepts input data files (including molecular alignments) in “relaxed” PHYLIP format. Output data files (tree files) are written in simplified NEWICK format (Fig. S1). The maximum taxa and characters in the input matrix must not exceed 5000 and 1,000,000, respectively. However, these limits can be modified within the source code if necessary. The pre-compiled versions of M2N for Windows, Linux, and MacOS are available for download here http://www.phys.ufl.edu/~madorsky/m2n
The M2N utility command line syntax is straightforward: m2n input_file_name, where input_file_name is the name of your input file in “relaxed” PHYLIP format (Figs. S1, S2).
A similar analytical approach to molecular data that requires its representation in tree form has been described previously (Mavrodiev et al. 2017). However, as suggested, we show how to obtain relationships directly from molecular alignments, avoiding their binary recoding (Mavrodiev et al. 2017). Still, M2N can successfully handle not only non-additive (unordered) multistate characters (i.e., up to 25 symbols representing IUB/IUPAC codes for DNA, RNA, and amino acids) but also binary characters.
MP analysis of the 3TS representations of molecular data has been conducted, as described in Mavrodiev and Madorsky (2012) and Mavrodiev et al. (2017). AC analysis of the M2N outputs has been performed as described in Mavrodiev et al. (2017), with the distance optimality criterion specified most simply as a “distance with non-weighted least squares” (Lapointe and Cucumel 1997; Swofford 2002; Creevey and McInerney 2009).
Manipulations of matrices and the tree files have been performed with Mesquite version 3.70. (Maddison and Maddison 2021), PAUP* version 4.0a (Swofford 2002), and FigTree version 1.4.4 (Rambaut and Drummond 2018).
Phylogenetic analyses summarized in Figs. 2a–6a (Namgung et al. 2021; Zhang et al. 2020; Barrett et al. 2016; Samigullin et al. 2022; Wei and Zhang 2020) have been conducted in PAUP* version 4.0a (Swofford 2002), TNT version 1.0 (Goloboff et al. 2008), MrBayes versions 1.0 and 3.2 (Huelsenbeck and Ronquist 2001; Ronquist et al. 2012), Tracer version 1.6 (Rambaut and Drummond 2009), RAxML versions 4.0 and 8.0 (Stamatakis 2006, 2014), IQ-TREE version 2.0 (Minh et al. 2020), and DAMBE5 (Xia 2013) with the selection (if any) of the best-fit models in jModelTest version 2.0 (Darriba et al. 2012), IQ-TREE version 2.0 (Minh et al. 2020) or ModelFinder (Kalyaanamoorthy et al. 2017). Coalescence analysis of Gentianaceae and outgroups (Zhang et al. 2020) was performed in ASTRAL-III version 5.6.2 (Zhang et al. 2018) and TREESPACE (Jombart et al. 2017) following previously published protocols (Zhang et al. 2018; Jombart et al. 2017; Gatesy et al. 2019; Goncalves et al. 2019).
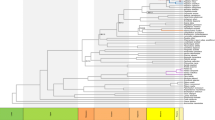
a BI and MP trees with the same topology inferred from 74 protein-coding regions of 32 complete chloroplast genomes (plastomes) of Allioideae (Amarylidaceae s.l.) and related taxa. This image is simplified from (Namgung et al. 2021). b The hierarchy of patterns with the most parsimonious fit of 2,258,281.6900 steps and RI equal to 1000, recovered from MP analysis (Wagner parsimony) of 3,638,687 3TS-WS representations of the 4136 bp modified 32 plastome-based alignment of Allioideae (Amarylidaceae s.l.) and related taxa, with only congruent characters included (in total, 9193 molecular characters are informative and simultaneously well conserved). c The hierarchy of patterns with the score 0.01927, recovered from AC analysis of 9193 rooted trees derived from the 9193 bp modified 32 plastome- based alignment of Allioideae (Amarylidaceae s.l.) and related taxa
3 Results
We developed “Matrix to Newick” (M2N), a program for the representation of standard molecular alignment in the form of trees or “maximal” relationships (Fig. 1) following the values of pre-selected outgroup taxon but requiring no binary recoding of the molecular outputs (Mavrodiev et al. 2017). This software can also deal with any extensive sequence alignment data set. M2N has a simple command line interface, briefly described in “Materials and Methods” and Fig. S1.
The details of the full results are given in Figs. 2, 3, 4, 5, 6 and their legends. The general result of the study is that the outputs of pattern-cladistic analyses yield tree topologies similar to those obtained previously using model-based phylogenetics (or even the same [Fig. 5a,b)]. However, in most cases (Figs. 2, 3, 4, and 6, see also Fig. 5c), remarkable differences have also been obtained. The latter appears to be either subtle, as in Figs. 1 and 3, where distinctions in the tree shapes can be described as small smooth changing the topological placement of a few taxa, such as Allium obliquum, A. senescens, and A. sativum (Fig. 1) or Gentiana siphonantha, G. macrophylla, and G. officinalis (Fig. 3). Or, on the contrary, be “global”, as on the Figs. 5 or 6, where different analyses lead to the different relationships between the subgenera of Diplazium (Fig. 6) or the parts of non-monophyletic tribe Tordyliinae (Apiaceae) (Fig. 5). From the traditional biogeographic and taxonomic standpoints (i.e., intuitively), however, such differences, whether subtle or “global” (e.g., the sisterhoods of the mostly Asian Allium obliquum and A. koreanum (Fig. 2c) or the American Diplazium subg. Diplazium and the Afro-Eurasian Diplazium subg. Pseudallantodia, all obtained in this paper [Fig. 5c)], are easily understood as new strong hypotheses of the relationship, rather than an artifact of the suggested method.
a Tree constructed from 76 protein-coding regions of 32 plastomes of Gentianaceae and related taxa by the coalescent method in ASTRAL-III (Zhang et al. 2020). 76 unrooted individual gene trees were inferred separately in RAxML and were inputted into ASTRAL-III to estimate the presented species tree (Zhang et al. 2020). The ML tree of Gentianaceae and relatives from the partitioned concatenated matrix of 76 plastid protein-encoding genes generated using RAxML (not shown here), looks almost the same in shape (Zhang et al. 2020). The image is simplified from (Zhang et al. 2020). b The hierarchy of patterns with the most parsimonious fit of 3,929,531.8770 steps and RI equal to 1000, recovered from MP analysis (Wagner parsimony) of 5961,440 3TS-WS representation of the 7350 bp modified 33 plastome-based alignment of Gentianaceae and outgroups (incl. Aucuba chlorascens (Garryaceae)), with only congruent characters included (in total, 11,579 molecular characters are informative and simultaneously well conserved). c The hierarchy of patterns with the score 0.01075, recovered from AC analysis of 11,579 rooted trees derived from the 11,579 bp modified 33 plastome-based alignment of Gentianaceae and outgroups
a ML Tree of Arecales, Dasypogonales and Typha latifolia L. (Poales, outgroup) based analysis of 42 complete plastomes minus one copy of the Inverted Repeat (Barrett et al. 2016). Image is simplified from (Barrett et al. 2016). b The hierarchy of patterns with the most parsimonious fit of 4,291,739.8001 steps and RI equal to 1000, recovered from MP analysis (Wagner parsimony) of 7,635,676 3TS-WS representation of the 7450 bp modified 42 plastome-based alignment of Arecales, Dasypogonales and Typha latifolia L. (Poales, outgroup), with only congruent characters included (in total, 14,081 molecular characters are informative and simultaneously well conserved). c The hierarchy of patterns with the score 0.01170, recovered from AC analysis of 14,081 rooted trees derived from the 14,081 bp modified plastome-based alignment of Arecales, Dasypogonales and Typha latifolia L. (Poales) (Barrett et al. 2016).
a Maximum a posteriori probability tree (PP = 1) inferred with BI of complete plastome sequences from 40 representatives of the apioid ‘superclade’ (Tordylieae and related taxa) (Apiaceae, Apioideae) (Samigullin et al. 2022). The image is simplified from (Samigullin et al. 2022). The position of Angelica sinensis is not shown because we excluded this taxon from future analyses. b The hierarchy of patterns with the most parsimonious fit of 862,711.3312 steps and RI equal to 1000, recovered from MP analysis (Wagner parsimony) of 1,410,397 3TS WS representation of the 2195 bp modified 39 plastome-based alignment of Tordylieae and related taxa (Apiaceae, Apioideae), with only congruent characters included (in total, 3744 molecular characters are informative and simultaneously well conserved). c The hierarchy of patterns with the score 0.01027, recovered from AC analysis of 3744 rooted trees derived from the 3744 bp modified 39 plastome-based alignment of Tordylieae and related taxa (Apiaceae, Apioideae)
a The ML phylogram of Diplazium (Athyriaceae) and related taxa based on the plastome sequence data (Wei and Zhang 2020). b The hierarchy of patterns with the most parsimonious fit of 6,374,449.8548 steps and RI equal to 1000, recovered from MP analysis (Wagner parsimony) of 8,595,914 3TS-WS representation of the 3905 bp modified 51 plastome-based alignment of Diplazium (Athyriaceae) and related taxa, with only congruent characters included (in total, 23,180 molecular characters are informative and simultaneously well conserved). c The hierarchy of patterns with the score 0.01547, recovered from AC analysis of 23,180 rooted trees derived from the 23,180 bp modified 51 plastome-based alignment of Diplazium (Athyriaceae) and related taxa
Therefore, our strict analytical focus on the informative characters instead of the detailed phenomenology of the genomic architecture of plastomes, as well as on the representations of such informative characters in the form of the 3TS matrices or the arrays of the trees, is heuristic (Figs. 2,3, 4, 5, 6).
4 Discussion
Many aspects of genomic annotation that typically frame plastome-based systematic studies today are unnecessary. In that sense, the approach to molecular systematics presented here is independent of all aspects of genomics above the simple sequences of bases. Plastome-based genomic information may be helpful for other purposes, but it must be published separately to avoid the distraction of the taxonomic focus in systematic studies. For example, all the original studies used here as data sources of the reanalyzed sequences of the plastomes (Namgung et al. 2021; Zhang et al. 2020; Barrett et al. 2016; Samigullin et al. 2022; Wei and Zhang 2020) partitioned the sequence data according to complicated genomic annotations. Without using the latter, we could recover cladograms with a similar shape to the phylogenetic trees resulting from the original studies (Namgung et al. 2021; Zhang et al. 2020; Barrett et al. 2016; Samigullin et al. 2022; Wei and Zhang 2020) (Figs. 1, 2, 3, 4, 5, 6).
Someone may note that the assembling of the plastomes from the raw sequence data (e.g., from the reads of the Illumina platform (https://www.illumina.com)), it already implies the essential genomic information. We agree with this point but still argue that even if such information is implied, it is not necessary to explicate it in a plant systematic paper, especially considering that analytical usage of the essential part of such information is doubtful.
The alignment of entire plastomes may be a flawed approach because plastid genomes may contain structural rearrangements (e.g., large or small inversions). Again, referring to the results of our Cladistic analyses (Figs. 1, 2, 3, 4, 5, 6), here we explicitly demonstrate that the standard G-block method (Castresana 2000), if applied to the set of the informative characters, extracted from the global plastome-based alignment (see Materials and Methods), in combination with the matrix-free representation of these data (Fig. 1, Fig. S1), is analytically sufficient to compensate the possible structural issues within the aligned plastomes.
Process-based phylogenetics is today’s analytical standard for massive molecular alignments (including plastome-based alignments). The quintessence of process-based phylogenetics is statistical or parametric phylogenetics, in particular: Maximum Likelihood (ML) (reviewed in Felsenstein 2004) and Bayesian (BI) (Rannala and Yang 1996) analyses. All characters are analytically important, not solely informative, within these frameworks (Felsenstein 2004; Lewis 2001). This leads to unexpected consequences within the genomic context. In fact, the strict link with ML or BI methodologies now encourages most researchers frequently to save constant and autapomorphic characters within the final dataset (e.g., within the molecular alignment of the plastomes or the parts of it) (Namgung et al. 2021; Zhang et al. 2020; Barrett et al. 2016; Samigullin et al. 2022; Wei and Zhang 2020).
The theoretical issues of this practice have already been discussed: From the standpoint of pattern-cladism, the model-based phylogenetics that treats all characters as informative undoubtfully belongs to the phenetic stream (Williams et al. 2010). Thus, Hennigian principle, grouping solely on synapomorphy, is seldom used in modern, primarily model-based phylogenetics.
The applied aspect of this practice is that the size of plastome alignment that contains constant and autapomorphic characters is much bigger if compared with the size of the alignment with all non-informative characters excluded before analyses.
The principal limitation of the taxonomic sampling is just the next step from the same observation regarding the larger size of the plastome-based alignments with all non-informative characters included. However, it was known a while ago that limited taxon sampling in analyzed data could lead to artifactual results (Graybeal 1998; Soltis and Soltis 2004), even if the number of the included characters in the matrix is significant (Soltis and Soltis 2004).
Someone may note that many statistical phylogeneticists, for example, who would prefer the ML approach, however, would agree that synapomorphies give the most analytical information. Also, considering all positions of the alignment as potentially informative does not imply that all these positions are equally influential to the results, as in the mainstream phenetic works. Therefore, there is no direct link between the ML method and phenetic.
Indeed, “As originally formulated, numerical phenetics based its estimates of similarity on many characters with equal a priori weights” (Sokal 1986, p. 426, see also Michener and Sokal 1957, pp. 130–131). However, “Strictly considered, equal weighting of characters is impossible in an actual study.” (Sokal 1986, p. 426). As Michener and Sokal (1957, p. 131) pointed out, “If only a few characters are used, studies of relationships require weighting of certain conservative characters in the mental processes of the systematist.” Later, exact phenetic methods that operate with non-equally weighted characters were designed (for example, Hartigan 1975, reviewed in De Soete et al. 1985; DeSarbo et al. 1984 reviewed in De Soete et al. 1985; De Soete et al. 1985, see also Sokal 1986, p. 434). Therefore, the phenetic classification does not necessarily have to treat all characters as equally influential on the results. Nevertheless, all the characters must be included in the phenetic analysis. The same is true in ML method. Even if, within statistical phylogenetics, most information is given by synapomorphies (Townsend 2007; Townsend and Lopez-Giraldez 2010; Assis 2015), all characters can influence the shape of the final phylogenetic tree.
What makes phenetic and ML philosophy different is the meaning of the term “character.” Again, as was stressed by the founders of phenetics:
“The word ”character “has been used in two rather different ways by taxonomists. In its commonest usage, any feature of one kind of organism that differentiates it from another kind is a character. Thus, the red abdomen of one bee is a character distinguishing it from another bee with the abdomen black. In this paper we use the word in a second connotation only; that is, as a feature which varies from one kind of organism to another. Now, to use the above example, abdominal color is the character; it occurs in two “states” or alternatives, red and black.” (Michener and Sokal 1957, p. 137, italics is ours).
Therefore, from the beginning of the phenetic stream, researchers who advocate the “overall similarity” as a significant principle in resolving relationships between studied taxa still treated characters traditionally, i.e., did not consider constant (or invariant) characters.
Hennig (1966) and cladists did the same, even detaching with phenetics regarding the vision of what character is. For example, for Platnick (1979, p. 542):
‘a character consists of two or more different attributes (character states) found in two or more specimens that, despite their differences, can be considered alternate forms of the same thing (the character)’
Massive analytical usage of the constant characters happened only during the molecular age (Lewis 2001). However, only within statistical phylogenetics (not within the standard maximum parsimony or pattern-cladism) the constant characters are analytically important, up to the formal "correction" of branch lengths of the trees obtained in the analysis of matrices that contain no constant characters (Lewis 2001). The purpose of such a posterior “correction” is formal compensation for the lack of invariant characters within analyzed matrices because the absence of constant columns in the matrix can essentially overestimate the branch lengths of the obtained tree. (Lewis 2001). But “… because branch lengths play an important role in determining the overall likelihood for a particular tree topology, such overestimation, if not corrected, would lead to bias in tree topology inferences (Lewis 2001, p. 917, italics is ours). Thus, within ML method, constant characters can influence not only the branch lengths, but the shape of the tree i.e., relationships between clades.
From this unusual perspective, the ML analysis, even if it treats all charters of the analyzed matrix as analytically important, cannot be considered a direct successor of phenetic ideology and, as it looks to us, represent a unique phenomenon in the history of logic. From the formal logical point of view, the invariant characters of one group can help distinguish this group from the other ones but cannot do anything to distinguish the parts of this group from each other. The rounded shape can quickly help distinguish balls from prisms and cubes. However, if all balls are round, it cannot help to determine red balls from black and white in any way (unless within the semantic frames of the ML analysis).
Thus, here we are arguing that despite theoretical requirements from the background of statistical phylogenetic methods, molecular characters that provide no evidence for the between-taxa relationship (e.g., the constant characters) must be excluded from the plastome-based global alignment. There is no “problem of constant characters” (Lewis 2001), as this “problem” was outlined by the ML Method advocates (Tuffley and Steel 1997; Lewis 2001). Ignoring constant characters, systematists (either conventional or molecular) never create any “problem” (Lewis 2001); they simply follow common sense and the long-term practice of traditional systematic research.
Someone may note incomplete lineage sorting is a significant challenge for synapomorphy-based inference. Even if this phenomenon is not an issue for plastomes-based phylogenetics, this problem may affect the other implications of the pattern-cladistic analyses of the different data sources. Moreover, similarly to ML method, the recognition of incomplete lineage sorting may also challenge the commonsense reasoning of systematists because the most common gene tree seems to be a reliable indicator of the species tree. From the pattern-cladistic point of view, it is first critical to talk about the incongruence of the trees based on different sources of evidence (Nelson 1979). The phrase “incomplete lineage sorting” explains this incongruence within some hypothetical process-based context. From that, a species tree must be viewed as a consensus of the gene trees that are not necessarily congruent to each other and the resulting consensus. Perhaps, the earliest discussion of the topic is given in the papers of Baum (1992) and Ragan (1992a, b), where they described the method that is known as “Matrix Representation with Parsimony” (or “MRP” (Ragan 1992b, p. 47)) approach and explicitly showed how to combine different trees based on the standard synapomorphies. For the cases of the massive incongruence of the source trees, a softly positive assessment of the accuracy of MRP was given by Bininda-Emonds and Sanderson (2001). Williams (1994) discussed Baum (1992) and Ragan’s (1992a, b) method and suggested a way of combining trees using 3TA instead of the standard parsimony (Williams 1996, 2004). Later literature on the topic mainly focused on the statistic framework is hard to observe (Degnan et al. 2009, Mossel and Roch 2010, among many others). From that, it is perhaps clear that the hypothetical explanation, such as incomplete lineage sorting, is not moving incongruence of gene and species trees from the "common-sense" context. The situation with the ML approach is different because, within the framework of the latter, the constant characters in some way gain the power to discriminate between alternatives (see above). Lewis (2001, p. 917), for example, is talking about “the systematic mission” of constant characters. Referring to the link of the likelihood of mutation probabilities with the number of constant characters in the matrix that makes such gaining technically possible (Tuffley and Steel 1997, p. 602) is just the rational reduction of the logic of the ML method to absurd.
The object of any analysis is not the data per se (appearances) but its representation. Tree-based and matrix-based representations of the systematic data may be viewed as technical equivalents but as semantic alternatives (Williams and Ebach 2006, 2008). Similarly, the matrix-based representation of the array of the minimal relationships (3TSs) is also a semantic alternative to the standard matrix-based representation of the data (Williams and Ebach 2008, 2020), for example, a plastome-based molecular alignment. Why is it worth stressing this difference? It is worth doing so because many of the standard phylogenetic procedures, such as reconstructions of character-states, identification of ancestral areas, molecular dating of the phylogenetic trees, calculations of the evolutionary rates within given branches of the tree, etc., are strictly resulting from the form of the data representation. Such procedures are irrelevant if systematists operate not with the conventional molecular alignment but with the relationships, which are the arrays of the single-character trees or the 3TS matrices (Williams and Ebach 2006, 2008, 2020; Mavrodiev et al. 2017, 2019).
Someone may note that no phylogenetic reconstruction method is free of assumptions about historical processes; therefore, an evolutionary model is always implied. Suppose the object of analysis is not an ordinary data matrix but an array of relationships. In that case, the assumption about the historical process is a way to explain the obtained patterns, not to get the patterns. No assumptions about historical processes are necessary to acquire the hierarchy of patterns within the pattern-cladistic framework. The latter is true because relationships are not physical objects (Nelson 2011) but are concepts (hypotheses, propositions, etc. (Platnik 1993; Kitching et al. 1998; Williams and Ebach 2006, 2008, 2020; Mavrodiev 2016)). Thus, the lack of a standard evolutionary model may imply a posteriori assumption about the process(es), simultaneously making no such assumptions a priori. In other words, the pattern-cladistic framework we are defending here is flexible regarding any historical and process-based explanations of the obtained results. Simultaneously it does not require any analytical parallels for hypothetical processes (e.g., the formal process of the optimization of the character-states) that might (or might not) have taken place in the past.
Someone may note that algorithms of multiple alignments are actually model-based: due of the probability scoring of the alternative alignments, such algorithms are making assumptions about molecular evolutionary processes. Therefore, nucleotide homology generated by multiple sequence alignment procedures is not purely structural, but rather the process-based.
Concepts of molecular homology were indeed developed entirely within an evolutionary context, which differs them from those of morphology (Patterson 1982, p. 619; Fitch 1966; Morrison et al. 2015). However, as it was shown by Patterson (1982, p. 622), "most aspects of molecular homology do have exact parallels or counterparts in morphology." The simplest way to establish such a link is the evident ability to create multiple molecular alignments just manually. Perhaps Watson and Kendrew (1961) were the first authors who aligned the sequences of the proteins of different taxa (in particular, the amino-acid sequences of sperm whale myoglobin and human hemoglobin). They did it by eye, with no usage of any evolutionary models. In their paper, Watson and Kendrew (1961) talk only about the models of molecules, i.e., about purely structural entities.
As it was summarized by Morrison et al. (2015, p. 49), “in molecular biology the word homology has long been used as a synonym for similarity.” The first paper that suggested a general method of aligning the amino acid sequences of two proteins was published in 1970 by Needleman and Wunsch, who talked solely about the similarity of the proteins as evidence for their homology and, therefore, possible models of protein evolution. Remarkably, Needleman and Wunsch (1970) didn’t refer to the paper of Jukes and Cantor (1969), who published the first stochastic substitution model of molecular evolution. Manual correction of the automatically generated multiple alignments is typical today (as a random example, see the paper Samigullin et al. 2022 we already cited above). A well-known result of such efforts is the BAliBASE, a database of manually refined multiple sequence alignments established as a benchmark alignment collection for evaluating numerous alignment programs (Thompson et al. 1999, p. 87). Therefore, the relation of the nucleotides or amino acids as homologous can be established simply by visual comparing the primary structures of the biological molecules, such as DNA or proteins. From that, the probability scoring of the molecular alignments by different softwareis just a procedure that can increase the efficiency of the whole multiple alignment process. This procedure remains optional (Edgar 2004, 2010).
It is perhaps clear from above that this paper considers alignment columns as a pure structural phenomenon, even if the evolutionary (or process-based) vision of such alignment as well as of molecular homology itself gained popularity from the beginning of the molecular age (e.g., Sankoff et al. 1973; Fitch 1966; Morrison et al. 2015).
For either traditional or molecular systematists, the most important phenomena are the potential homologies or the relations between different organisms based on their “parts” (or homologues) (Williams and Ebach 2006, 2008, 2020; Nelson 2011). Today the best way to formalize such relations is a Newick’s notation, e.g., the language that includes only the names of the studied taxa (or instead the symbols, such as A, B, C, etc.), parentheses, commas, and semicolons (Williams and Ebach 2006, 2008) (Fig. 1, Fig. S1). In fact, the same is also true for molecular systematics as well, where the relations between different organisms may also be easily written in a tree form (Fig. 1, Fig. S1), not in the form of the standard molecular alignment, and thus be the subject of future consensus analyses. This paper shows that such analyses are successful (Figs. 2b, c–6b, c). Therefore, the tree-based or relational representation of systematic data may effectively replace the standard “processes-based” approach with a purely comparative view of the problem (Mavrodiev et al. 2017). Instead of the standard ML or BI analytical frameworks and conventional molecular alignments of the complete plastid genomes (either partitioned or global alignments), systematists can now use a pattern-cladistic framework as well as the alignment-free representation of the same data (Fig. 1).
In theory, strict focus on the relational (tree-based or standard 3TS) representation of molecular data, and particularly molecular alignments of the plastomes, may almost automatically encourage botanists to center attention on the systematics context of the data itself (e.g., on the taxonomic relations of the studied genera, tribes, families and other taxa), instead of speculative model-based reconstructions of the hypothetical evolution of the studied group or the details of the genomic architecture of the plastomes (and multiple related summaries and statistics).
Despite intensive past discussions (e.g., Hull 1967; O'Rourke 2004), the metaphysical foundations of the evolutionary theory itself still need to be thoroughly thought out (Tremblay 2013, 2020). Remaining completely agnostic about any historical events, pattern-cladistic thinking focuses only on the sets of given phenomena and tries to estimate the taxonomic relationships based solely on the latter, ignoring any assumptions on how characters or taxa may evolve in the past.
References
Assis LCS (2015) Homology assessment in parsimony and model–based analyses: two sides of the same coin. Cladistics 31:315–320
Barrett CF, Baker WJ, Comer JR, Conran JG, Lahmeyer SC, Leebens-Mack JH, Li J, Lim GS, Mayfield-Jones DR, Perez L (2016) Plastid genomes reveal support for deep phylogenetic relationships and extensive rate variation among palms and other commelinid monocots. New Phytol 209:855–870
Baum BR (1989) PHYLIP: phylogeny inference package Version 3.2. Q Rev Biol 64:539–541
Baum BR (1992) Combining trees as a way of combining data sets for phylogenetic inference and the desirability of combining gene trees. Taxon 41:3–10
Bininda-Emonds ORP, Sanderson MJ (2001) Assessment of the accuracy of matrix representation with parsimony analysis supertree construction. Syst Biol 50:565–579
Brower AVZ (2000) Evolution is not a necessary assumption of cladistics. Cladistics 16:143–154
Brower AVZ (2019) Background knowledge: the assumptions of pattern cladistics. Cladistics 35:717–731
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Chen YP, Zhao F, Paton AJ, Sunojkumar P, Gao L-M, Xiang C-L (2022) Plastome sequences fail to resolve shallow level relationships within the rapidly radiated genus Isodon (Lamiaceae). Front Plant Sci 13:985488
Creevey CJ, McInerney JO (2009) Trees from trees: construction of phylogenetic supertrees using Clann. In: Posada D (ed) Springer protocols: methods in molecular biology Bioinformatics for DNA Sequence Analysis, vol 537. Humana Press Totowa, Totowa, pp 139–161
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models new heuristics and parallel computing. Nat Methods 9:772–772
De Soete G, DeSarbo WS, Carroll JD (1985) Optimal variable weighting for hierarchical clustering—an alternating least-squares algorithm. J Classif 2:173–192
Degnan JH, DeGiorgio M, Bryant D, Rosenberg NA (2009) Properties of consensus methods for inferring species trees from gene trees. Syst Biol 58:35–54
DeSarbo WS, Carroll JD, Clark LA, Green PE (1984) Synthesized clustering—a method for amalgamating alternative clustering bases with differential weighting of variables. Psychometrika 49:57–78
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Edgar RC (2010) Quality measures for protein alignment benchmarks. Nucleic Acids Res 38:2145–2153
Farris JS (1983) The logical basis of phylogenetic analysis. In: Platnick NI, Funk V (eds) Advances in cladistics, 2. Proceedings of the 2nd meeting of the Willi Hennig society; Ann Arbor, Mich., USA, Oct. 1–4; 1981. Columbia University Press, New York, pp. 7–36
Felsenstein J (1993) PHYLIP (Phylogeny Inference Package) Version 3.5c Distributed by the author Department of Genetics University of Washington Seattle USA https://csbf.stanford.edu/phylip/
Felsenstein J (2004) Inferring phylogenies. Sinauer Associates Inc Sunderland, Sunderland
Fitch WM (1966) An improved method of testing for evolutionary homology. J Mol Biol 16:9–16
Gatesy J, Sloan DB, Warren JM, Baker RH, Simmons MP, Springer MS (2019) Partitioned coalescence support reveals biases in species-tree methods and detects gene trees that determine phylogenomic conflicts. Mol Phylogenet Evol 139:106539
Goloboff PA, Farris JS, Nixon KC (2008) TNT, a free program for phylogenetic analysis. Cladistics 24:774–786
Goncalves DJ, Simpson BB, Ortiz EM, Shimizu GH, Jansen RK (2019) Incongruence between gene trees and species trees and phylogenetic signal variation in plastid genes. Mol Phylogenet Evol 138:219–232
Gouy M, Guindon S, Gascuel O (2010) SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27:221–224
Graybeal A (1998) Is it better to add taxa or characters to a difficult phylogenetic problem? Syst Biol 47:9–17
Hartigan JA (1975) Clustering Algorithms. John Wiley and Sons, New York
Hennig W (1966) Phylogenetic systematics. University of Illinois Press, Urbana
Huang Y, Fan L, Huang J, Zhou G, Chen X, Chen J (2022) Plastome phylogenomics of Aucuba (Garryaceae). Front Genet 13:753719
Huelsenbeck JP, Ronquist F (2001) MrBayes: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Hull DL (1967) The metaphysics of evolution. Br J Hist Sci 3:309–337
Jombart T, Kendall M, Almagro-Garcia J, Colijn C (2017) treespace: statistical exploration of landscapes of phylogenetic trees. Mol Ecol Resour 17:1385–1392
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic Press, New York, pp 21–132
Kalyaanamoorthy S, Bui Quang M, Wong TKF, von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Kitching IJ, Forey PL, Humphries CJ, Williams DM (1998) Cladistics: the theory and practice of parsimony analysis, 2nd ed. Systematics Association publications (Book 11). Oxford University Press, Oxford, UK
Lapointe FJ, Cucumel G (1997) The average consensus procedure: a combination of weighted trees containing identical or overlapping sets of taxa. Syst Biol 46:306–312
Lapointe FJ, Levasseur C (2004) Everything you always wanted to know about the average consensus, and more. In: Bininda-Emonds RP (ed) Phylogenetic supertrees: combining information to reveal the Tree of Life. Computational biology series, vol 4. Kluwer Academic Publishers, Dordrecht, pp 87–105
Lewis PO (2001) A likelihood approach to estimating phylogeny from discrete morphological character data. Syst Biol 50:913–925
Maddison WP, Maddison DR (2021) Mesquite: a modular system for evolutionary analysis Version 3.70 http://mes.quite.project.org
Mavrodiev EV (2016) Dealing with propositions not with the characters: the ability of three-taxon statement analysis to recognize groups based solely on ‘reversals’ under the maximum-likelihood criteria. Aust Syst Bot 29:119–125
Mavrodiev EV, Madorsky A (2012) TAXODIUM Version 10: a simple way to generate uniform and fractionally weighted three-item matrices from various kinds of biological data. PloS one 7:e48813
Mavrodiev EV, Dell C, Schroder L (2017) A laid-back trip through the hennigian forests. Peer J 5:e3578
Mavrodiev EV, Williams DM, Ebach MC (2019) On the typology of relations. Evol Biol 46:71–89
Michener CD, Sokal RR (1957) A quantitative approach to a problem in classification. Evolution 11:130–162
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic tres In: Pirece M (ed) Proceedings of the gateway computing environments workshop (GCE) 14 Nov 2010 New Orleans pp 1–8
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, Von Haeseler A, Lanfear R (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol 37:1530–1534
Morrison DA, Morgan MJ, Kelchner SA (2015) Molecular homology and multiple-sequence alignment: an analysis of concepts and practice. Aust Syst Bot 28:46–62
Mossel E, Roch S (2010) Incomplete lineage sorting: consistent phylogeny estimation from multiple loci. IEEE/ACM Trans Comput Biol Bioinform 7:166–171
Namgung J, Do HDK, Kim C, Choi HJ, Kim JH (2021) Complete chloroplast genomes shed light on phylogenetic relationships divergence time and biogeography of Allioideae (Amaryllidaceae). Sci Rep 11:1–13
Needleman SB, Wunsch CD (1970) A general method applicable to the search for similarities in the amino acid sequence of two proteins. J Mol Biol 48:443–453
Nelson G (1979) Cladistic analysis and synthesis - principles and definitions, with a historical note on Adanson’s Familles des Plantes (1763–1764). Syst Zool 28:1–21
Nelson G (2011) Resemblance as Evidence of Ancestry. Zootaxa 2946:137–141
Nelson G, Platnick NI (1991) Three-taxon statements—a more precise use of parsimony? Cladistics 7:351–366
O’Rourke F (2004) Aristotle and the Metaphyics of Evolution. Rev Metaphys 58:3–59
Patterson C (1982) Homology in classical and molecular biology. Mol Biol Evol 5:603–625
Platnick NI (1979) Philosophy and the transformation of cladistics. Syst Zool 28:537–546
Platnick NI (1993) Character optimization and weighting - differences between the standard and three-taxon approaches to phylogenetic inference. Cladistics 9:267–272
Ragan MA (1992a) Phylogenetic inference based on matrix representation of trees. Mol Biol Evol 1:53–58
Ragan MA (1992b) Matrix representation in reconstructing phylogenetic relationships among the eukaryotes. BioSystems 28:47–55
Rambaut A, Drummond AJ (2009) Tracer v. 16. http://beast.bio.ed.acuk/
Rambaut A, Drummond AJ (2018) FigTree v. 1.4. Molecular evolution, phylogenetics and epidemiology. University of Edinburgh, Edinburgh. http://tree.bio.ed.ac.uk/software/figtree/
Rannala B, Yang ZH (1996) Probability distribution of molecular evolutionary trees: a new method of phylogenetic inference. J Mol Evol 43:304–311
Ronquist F, Teslenko M, Van Der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 32: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Samigullin T, Logacheva M, Terentieva E, Degtjareva G, Pimenov M, Valiejo-Roman C (2022) Plastid Phylogenomic analysis of tordylieae tribe (Apiaceae Apioideae). Plants 11:709
Sankoff D, Morel C, Cedergren RJ (1973) Evolution of 5S RNA and non-randomness of base replacement. Nature New Biol 245:232–234
Sokal RR (1986) Phenetic taxonomy—theory and methods. Annu Rev Ecol Evol Syst 17:423–442
Soltis DE, Soltis PS (2004) Amborella not a “basal angiosperm”? not so fast. Am J Bot 91:997–1001
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Swofford DL (2002) PAUP*. Phylogenetic analysis using parsimony (*and other methods). Sinauer Associates Inc, Sunderland
Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56:564–577
Thompson JD, Plewniak F, Poch O (1999) BAliBASE: a benchmark alignment database for the evaluation of multiple alignment programs. Bioinformatics 15:87–88
Townsend JP (2007) Profiling phylogenetic informativeness. Syst Biol 56(2):222–231
Townsend JP, Lopez-Giraldez F (2010) Optimal selection of gene and ingroup taxon sampling for resolving phylogenetic relationships. Syst Biol 59:446–457
Tremblay F (2013) Nicolai Hartmann and the metaphysical foundation of phylogenetic systematics. Biol Theory 7:56–68
Tremblay F (2020) Nikolai Lossky’s evolutionary metaphysics of reincarnation. Sophia 59:733–753
Tuffley C, Steel M (1997) Links between maximum likelihood and maximum parsimony under a simple model of site substitution. Bull Math Biol 59:581–607
Wagner ND, Volf M, Hörandl E (2021) Highly diverse shrub willows (Salix L.) share highly similar plastomes. Front Plant Sci 12:662715
Watson HC, Kendrew JC (1961) Comparison between the amino-acid sequences of sperm whale myoglobin and of human hemoglobin. Nature 190:670–672
Wei R, Zhang X-C (2020) Phylogeny of Diplazium (Athyriaceae) revisited: resolving the backbone relationships based on plastid genomes and phylogenetic tree space analysis. Mol Phylogenet Evol 143:106699
Williams DM (1994) Combining trees and combining data. Taxon 43:449–453
Williams DM (1996) Characters and cladograms. Taxon 45:275–283
Williams DM (2004) Supertrees, components and three-item data. In: Bininda-Emonds ORP (ed) Phylogenetic supertrees: Combining information to reveal the Tree of Life. Springer-Kluwer Academic Publisher, Dordrecht, The Netherlands, pp 389–408
Williams DM, Ebach MC (2006) The data matrix. Geodiversitas 28:409–420
Williams DM, Ebach MC (2008) Foundations of systematics and biogeography. Springer, New York
Williams DM, Siebert DJ (2000) Characters, homology and three-item analysis. In: Scotland RW, Pennington TR (eds) Homology and systematics: coding characters for phylogenetic analysis Systematics Association Special Volume (Book 58). Taylor and Francis, London, pp 183–208
Williams DM, Ebach MC, Wheeler QD (2010) Beyond belief: The steady resurrection of phenetics. In: Williams DM, Knapp S (eds) Beyond cladistics: The branching of a paradigm. University of California Press, Berkley, California, USA, pp 169–197
Williams DM, Ebach MC (2020) Cladistics: A guide to biological classification, 3rd ed, Systematics Association Special Volume (Book 88). Cambridge University Press, Cambridge, UK
Xia X (2013) DAMBE5: a comprehensive software package for data analysis in molecular biology and evolution. Mol Biol Evol 30:1720–1728
Zhang C, Rabiee M, Sayyari E, Mirarab S (2018) ASTRAL–III: polynomial time species tree reconstruction from partially resolved gene trees. BMC Bioinform 19:15–30
Zhang X, Sun Y, Landis JB, Lv Z, Shen J, Zhang H, Lin N, Li L, Sun J, Deng T (2020) Plastome phylogenomic study of Gentianeae (Gentianaceae): widespread gene tree discordance and its association with evolutionary rate heterogeneity of plastid genes. BMC Plant Biol 20:1–15
Acknowledgements
The authors thank Prof. Richard Buggs (School of Biological and Chemical Sciences, Queen Mary University of London, UK) for his helpful notes and comments on an early version of the article. We acknowledge Dr. David M. Williams (the Natural History Museum, London, UK) and Prof. Malte C. Ebach (University of New South Wales & the Sydney’s Australian Museum, AU) for their helpful discussion. We also thank Dr. Williams for bringing the study of Nelson (1979) to our attention. Two anonymous Reviewers (especially Reviewer 2) are highly acknowledged for their elegant comments, that helped to improve the article's content.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mavrodiev, E.V., Madorsky, A. On Pattern-Cladistic Analyses Based on Complete Plastid Genome Sequences. Acta Biotheor 71, 22 (2023). https://doi.org/10.1007/s10441-023-09475-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10441-023-09475-5