Abstract
SARS-CoV-2 infection in the human body induces a severe storm of inflammatory factors. However, its specific mechanism is still not clear. Gas chromatography–mass spectrometry (GC–MS) technology is expected to explain the possible mechanism of the disease by detecting differential metabolites. 15 COVID-19 patients and healthy controls were included in this study. Immune indicators such as hs CRP and cytokines were detected to reflect the level of inflammation in patients with COVID-19. The distribution of lymphocytes and subpopulations in peripheral whole blood were detected using flow cytometry to assess the immune function of COVID-19 patients. The expression of differential metabolites in serum was analyzed using GC–MS non-targeted metabolomics. The results showed that hs CRP, IL-5/6/8/10 and IFN-α in the serum of COVID-19 patients increased to varying degrees, and CD3/4/8+ T lymphocytes decreased. Additionally, 53 metabolites in the serum of COVID-19 patients were up regulated, 18 metabolites were down regulated, and 8 metabolites remained unchanged. Increased Cholesterol, Lactic Acid and 1-Monopalmitin may be the mechanism that causes excessive inflammation in COVID-19 patients. The increase of D-Allose may be involved in the process of lymphocyte decrease. In conclusion, the significance of our study is to reveal the possible mechanism of inflammatory response in patients with COVID-19 from the perspective of metabolomics. This provided a new idea for the treatment of COVID-19.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
SARS-CoV-2 belongs to β-coronavirus. It is a non-segmented normal-stranded single-stranded RNA virus with a diameter of 60-140 nm. The reason it is named as coronavirus is that its membrane has protrusions around it, shaped like a corolla. The WHO named the disease caused by SARS-CoV-2 for coronavirus disease-2019 (COVID-19) based on the consultation and cooperation between the world organization for animal health (OIE) and the food and agriculture organization of the United Nations (FAO) on February 11, 2020. (http s://www.who.int/docs/default-source/coronaviruse/situation-reports/20200211-sitrep-22-ncov.pd f?sfvrsn = fb6d49b1_2).
The clinical manifestations of COVID-19 patients are mostly fever and cough, severe patients are manifested as respiratory distress, and occasionally patients have symptoms such as confusion, nausea and vomiting [1]. As of March 14, 2022, about 459 million of people worldwide had been infected with SARS-CoV-2, and approximately 6 million people have died from COVID-19. (COVID-19 Dashboard by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University (JHU), https://coronavirus.jhu.edu/).
At this stage, the clinical testing methods for COVID-19 patients are mainly nucleic acid testing and specific antibody testing. The disadvantage of nucleic acid testing is that it is difficult to accurately quantify and cannot reveal the severity and mechanism of the disease [2]. Specific antibody testing cannot distinguish SARS-CoV-2 infection from vaccination accurately [3]. Therefore, both are not ideal detection methods. However, the metabolomics technology can be used for the research and search of pathogenic mechanisms. Its basic principle is to detect all the small molecule metabolites in the body using chromatography-mass spectrometry technology and to obtain a large amount of data to find differential metabolic markers. These significantly different metabolic markers may play an important role in revealing mechanism of diseases.
This study included 15 cases of COVID-19 patients diagnosed between January and May 2020 in Mianyang, Sichuan, China, and selected samples collected from the health examination center during the same period as the control of non-COVID-19 patients. The controls were not diagnosed with COVID-19 and pneumonia caused by other pathogen infections. Informed signed consent was obtained from either the patient or from next of kin. We used flow cytometry to detect changes in cytokine levels during treatment of patients with COVID-19. In addition, we detected distribution of lymphocytes and subpopulations in peripheral whole blood to assess the immune function of COVID-19 patients. Gas chromatography–mass spectrometry technology was used to analyze the changes of metabolome in the serum of patients with COVID-19, to conduct an in-depth study on the inflammatory mechanism of SARS-CoV-2. The significance of our study is to reveal the possible mechanism of inflammatory response in patients with COVID-19 from the perspective of metabolomics. This provided a new idea for the treatment of COVID-19.
Materials and Methods
Detection of Cytokines in the Plasma of COVID-19 Patients
According to the sample collection time, we divided all the samples collected from 15 patients into 3 groups, which were first day of infection, advanced stage of infection and recovered stage of infection.
The cytokine detection kit was purchased from Jiangxi Saiji Biotechnology Co., Ltd. (Nanchang, China), all operations were carried out in accordance with the instruction. The standard was serial diluted by 10 concentration gradients. Capture microsphere mixture (25 μL), sample (25 μL), and PE (Phycoerythrin) labeled antibody (25 μL) were added to each sample tube. After vortexing and mixing the reaction system, each tube was incubated in the dark at room temperature for 2.5 h. PBS (Phosphate Buffer Saline, 1 mL) was added to each tube, centrifuged at 200 × g for 5 min, and the supernatant was discarded carefully. PBS (100μL) was added to each sample and perform fluorescence detection on a flow cytometer (Navios, BECKMAN COULTER, USA).
Analysis of T Lymphocyte Subpopulations in Peripheral Blood
The purple tube containing EDTA anticoagulant was used to collect the peripheral blood sample. The four-color labeled monoclonal antibody (10 μL) CD3-PC5/CD4-PE/CD8-ECD/CD45-FITC (BECKMAN COULTER, USA) and 100 μL of anticoagulated whole blood were added into the test tube. After shaking and mixing, the tube was placed in the dark for 15 min. Then hemolysin (500 μL), sheath fluid (500 μL) and flow count (100 μL) were added into tube, Beckman Coulter Fc500 flow cytometer was used to detect T lymphocyte subsets. Specific operations were referred to the reagent instructions.
In the analysis of T lymphocyte subsets, 15 COVID patients were divided into two groups. Common cases refer to patients with upper respiratory tract infection symptoms such as moderate to low grade fever, dry throat, sore throat, nasal congestion, runny nose, and imaging findings of pneumonia. An adult confirmed case would be diagnosed as severe if one of the following occurred: (1). Shortness of breath, RR (Respiratory Rate) ≥ 30 beats min−1; (2). In resting state, oxygen saturation ≤ 93%; (3). Arterial blood Partial pressure of oxygen (PaO2)/FiO2 ≤ 300 mmHg; (4). Lung imaging showed that the lesion progressed > 50% within 24 h. According to the above criteria, among the 15 COVID-19 patients, 11 were common cases and 4 were sever cases.
Detection of Hypersensitive C Reactive Protein
We tested the concentration of hs CRP (hypersensitive C Reactive Protein) in the serum samples collected from acute phase of COVID-19 patients. Serum samples from healthy medical examiners during the same period were collected as a control group. The concentration of hs CRP was detected using immunochemiluminescence apparatus (i4000, ABBOTT, USA).
Pretreatment of Serum Metabolism Analysis
The controls in this part were collected from the health examination center. They were not diagnosed with COVID-19 and pneumonia caused by other pathogen infections.
The serum (100 µL) was mixed with ice methanol (400 µL), and 3-phenylbutyric acid (80 µL, (Kangbo Rui Biotechnology Co., Ltd., Chengdu, China)) were added into the reaction system to vortex. After centrifugation at 16 000 rpm at 4 °C for 15 min, 400 µL of supernatant was taken for freeze-vacuum drying. Methoxyamine pyridine solution (50 µL) were added into the lyophilized product, and the oximation reaction was carried out in a water bath at 70 ℃ for 1 h. Then, methoxyamine hydrochloride-(trimethylsilyl)-trifluoroacetamide (Shanghai Xinyu Biological Technology Co., Ltd., Shanghai, China) derivatization reagent (containing 1.0% trimethylchlorosilane, Shanghai Xinyu Biological Technology Co., Ltd., Shanghai, China) was added to finish derivatization reaction in a 50 ℃ water bath. The prepared samples were placed at room temperature for 2 h and used for gas chromatography-mass spectrometer (GC–MS) analysis.
Serum GC–MS Analysis
The samples (1.0 µL) were analyzed in random sequence, using Shimadzu GC–MS (TQ8050, Shimadzu Corporation, Tokyo, Japan). Shimadzu HP-5 capillary column (30 m × 250 µm × 0.25 µm) was used for the separation of GC compounds. High-purity helium was used as the carrier gas with a carrier gas flow rate of 1.0 mL min−1. The initial temperature of the column was set to 70 ℃, and maintained at this temperature for 4 min, and then increased to 300℃ at a rate of 8℃ min−1. The ionization mode was electron bombardment, the electron energy was 70 eV, and the temperature was 200 ℃. A mass range of 35 ~ 800 m z−1 was used for the mass spectra.
Statistical Analysis
GC–MS Solution software (version 2.1) were used to perform mass spectrometry deconvolution, peak alignment and peak identification on the collected mass spectra. The mass spectrum was matched with the National Institute of Standards and Technology (NIST) mass spectrum library for qualitative analysis. The data were analyzed with SIMCA-P (version 13.0), including principal component analysis (PCA), partial least squares discriminant analysis (PLS-DA) and orthogonal partial least squares discriminant analysis (OPLS-DA). At last, SPSS 17.0 was used to perform t test on the data, and P value < 0.05 was considered to be statistically significant.
Results
Changes in Cytokine Concentration
Figures 1 shows the concentration of 11 cytokines in the plasma samples of COVID-19patients. The red dotted line in the figure indicates the reference range. Compared with the first day of infection, the concentration of IL-4, IL-2,IL-10,TNF-α,IL-1 and IL-12P70 increased in the advanced stage. Subsequently,
T Lymphocyte Subpopulations in Peripheral Blood
The distribution of peripheral blood lymphocytes and T lymphocyte subsets of COVID-19 patients are shown in Fig. 2. Compared with the healthy control group, the absolute lymphocyte value, CD3+, CD4+, CD8+ T cells in peripheral blood of COVID-19 patients were significantly decreased, P˂0.01. This alteration was more obvious in severe cases of COVID-19, that was, lymphocytes and T cell subsets decreased as the patient’s condition worsened.
Concentration of hs CRP in Serum
As shown in Fig. 3, the hs CRP level of some patients on the first day of infection was higher than the reference upper limit. Compared with the first day of infection, the concentration of hs CRP increased significantly in the advanced stage, P < 0.05. The hs CRP levels in advanced stage were all above the upper reference range, 20 mgL−1. And compared with the advanced stage of infection, the concentration of hs CRP were significantly decreasd in recovered stage, P < 0.05.
Serum Metabolomics Analysis
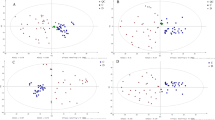
As shown in Fig. 4, we selected the acute phase serum of COVID-19 patients for metabonomic analysis. Compared to the control group, 53 metabolites were up regulated, 18 were down regulated, and 8 were unchanged.
Using PCA to analyze the two groups of samples, the results showed that there were significant differences between the serum of control group and COVID-19 group. The PCA score scatter plot in Fig. 5 showed that the contribution rates of the two principal components were 58.1% and 10.6%, respectively, and it could explain 95.0% of the data difference. Subsequently, based on the successful verification of the OPLS-DA and PLS-DA models, the metabolites with VIP value > 1, correlation coefficient value > 0.5 in the (V + S) scatter plot and t test P value < 0.05 were considered to be the differential metabolites with the most contribution. Seven different metabolites were obtained through the above-mentioned multivariate statistical analysis, which were: Lactic Acid, D-Glucose, D-Allose, 1-Monopalmitin, Glycerol monostearate, Cholesterol, Urea.
Metabolite Pathway Analysis
MetaboAnalyst 5.0 was used for enrichment analysis of pathways related to differential metabolites and to investigate alterations in their abundance. As shown in Fig. 6A, the differential metabolites in the serum of COVID-19 patients covered several biochemical pathways: glycolysis/gluconeogenesis, arginine biosynthesis, pyruvate metabolism, steroid biosynthesis, primary bile acid biosynthesis, purine metabolism, steroid hormone biosynthesis. It's expected to reveal the metabolic mechanism of body’s response to infection through analysis of regulatory metabolic pathways. The metabolite sets enrichment overview shown in Fig. 6B demonstrated that glycolysis/gluconeogenesis metabolism, warburg effect and lactose degradation were the metabolite concentrated set most altered by the SARS-CoV-2, confirming that glycometabolism was a key factor in the process of virus infection.
Differential metabolites in serum of COVID-19 patients were mainly enriched in the two pathways of glycolysis/gluconeogenesis and arginine metabolism. The up regulated D-glucose in the serum of COVID-19 patients activated the glycolysis pathway, accelerated the occurrence of anaerobic oxidation, and increased the content of lactic acid. Increased lactic acid could further participate in the gluconeogenesis metabolic pathway. Another enriched metabolic pathway was arginine metabolism. Active arginine metabolism promoted the occurrence of ornithine cycle and induced an increase in urea synthesis.
Discussion
In this study, we tested the serum cytokine levels of COVID-19 patients at different stages of the disease course. As shown in Fig. 1, IL-6 and IL-8 in some patients exceeded the upper limit of the reference range on the first day of infection. The reason for it may be that the patients were already febrile at the time of presentation. Many studies have shown that the levels of pro-inflammatory cytokines (IL-6, IL-1β, IL-8 and TNF-α) and chemokines (GCSF, IP10 and MCPl) in the plasma of COVID-19 patients in the intensive care unit are high within normal reference range [1, 4]. Our research results are similar to these, the cytokines (IL-6, IL-8, IL-10, IL-5 and TNF-α) in the serum of patients with COVID-19 all increased to varying degrees.
Through the analysis of the peripheral blood T lymphocyte subsets of COVID-19 patients, we found that compared with healthy controls, both CD4+ and CD8+ T cells in the peripheral blood of COVID-19 patients were reduced. And in severe cases, the absolute value of lymphocytes and the counts of each subsets decreased the most, suggesting that the decrease of CD3+, CD4+, CD8+ counts in peripheral blood may be an important factor leading to progression of COVID-19, and the degree of their decline could be used as one of the indicators to judge the severity of COVID-19. The results of this study were consistent with the results of a retrospective study [5]: Compared with healthy controls, the total number of lymphocytes, CD4+ and CD8+ T cells in patients with COVID-19 were reduced. And another study also showed that the level of lymphocytes in severe cases of COVID-19 was lower than that of common cases [6].
At the same time, we also detected an increase in hs CRP in the serum of patients in the different stages of COVID-19. Figure 3 depicted that the concentration of hs CRP increased gradually with the aggravation of disease course. Whereas it decreased at the recovered stage. CRP, a widely used acute phase protein, is an objective biomarker of infection, inflammation and tissue damage [7]. The increase in hs CRP indicates a severe inflammatory response in COVID-19 patients. And the increase of IL-6 and IL-8 to different degrees may be one of the mechanisms that cause acute inflammation and fever in COVID-19 patients.
GC–MS is an important method to study metabolomics. The metabolomics research on COVID-19 has been carried out rapidly, bringing hope to the discovery of reliable biomarkers to assess disease progression and the development of therapeutic drugs [8]. Many intermediate products in the body’s metabolism have been shown to be key signaling molecules that regulate the occurrence, progress and outcome of immunity [9]. The main purpose of this study is to detect metabolic markers related to COVID-19 to better understand the pathogenesis of COVID-19.
In the present study, we detected 53 metabolites were up regulated, 18 metabolites were down regulated, and 8 metabolites were unchanged in the serum of COVID-19 patients using Gas chromatography–mass spectrometry technology (Fig. 4). As shown in Table 1, seven different metabolites were obtained through multivariate statistical analysis, which were Lactic Acid, D-Glucose, D-Allose, 1-Monopalmitin, Glycerol monostearate, Cholesterol, Urea. And compared with the control group, the contents of 7 different metabolites were all up regulated. These differential metabolites were significantly related to human immune system function and clinical features.
Under aerobic conditions, glucose generates pyruvate through glycolysis, then pyruvate is converted into carbon dioxide for oxidative phosphorylation in mitochondria; under anaerobic conditions, pyruvate is converted into lactic acid in the cytoplasm, and the importance is to rebuild NAD+ (Nicotinamide Adenine Dinucleotide) required for glycolysis to maintain the synthesis of ATP (Adenosine triphosphate) [10]. Therefore, the increase in lactic acid in serum of COVID-19 patients that we have detected reflects hypoxia of cells and tissues. Moreover, studies have found that lactic acid can promote the proliferation of endothelial cells and the maturation of new blood vessels by increasing the production of IL-8 [11]. It is speculated that lactic acid may be a possible cause of increase of IL-8 in serum of COVID-19 patients and occurrence of inflammation.
A study showed that lactic acid and cholesterol in the serum of COVID-19 patients were enriched compared with healthy controls [12]. Our research results are consistent with it. SREBPs (Sterol Regulatory Element-Binding Proteins) are major transcription factors that regulate cholesterol biosynthesis [12]. A recent study reported that it is highly expressed in severe cases of COVID-19 and can be used as an indicator for severity diagnosis [13]. It indicates that lipid and cholesterol synthesis pathways play an important role in the pathogenesis of COVID-19. The research of Sohrabi, Yahya et al. have also shown that increased cholesterol synthesis combined with fructose metabolic pathway mediates excessive inflammation in COVID-19 patients [14].
Not only that, as an important lipid molecule that maintains the fluidity of cell membranes, the biosynthesis and regulation pathways of cholesterol are ubiquitous in various cells, including immune cells [15]. Cholesterol in the tumor microenvironment induces CD8+ T cell exhaustion in an ER-stress-XBP1 (Endoplasmic Reticulum-stress-transcription factor X-box binding protein1) dependent manner [16]. Whether the decrease of CD8+ T cells in the peripheral blood of COVID-19 patients is related to cholesterol remains unclear. Nevertheless, the importance of cholesterol in regulating immune cell function and inflammatory response suggests that patients with COVID-19 should maintain healthy eating habits and avoid high fructose and high cholesterol intake during treatment.
We detected that D-Allose in the serum of COVID-19 patients was more abundant than the control group. D-Allose is a rare 6-carbon monosaccharide in nature, which can down-regulate the endocytic activity and subsequent antigen presentation to T cells of dendritic cells, thereby triggering immunosuppression [17]. Experiments in vitro also verified that D-Allose inhibits cell proliferation by inducing cell G2/M phase arrest [18]. A study showed that D-Allose were depleted in the feces of COVID-19 patients [19]. We cannot yet explain the difference in the distribution of D-Allose in the serum and feces of COVID-19 patients. However, it can be speculated that the increase of D-Allose in the serum of COVID19 patients may be one of the reasons for the decreased cellular immune function.
In addition, D-glucose was increased in the serum of COVID-19 patients compared with the control group, MetaboAnalyst 5.0 analysis also demonstrated that glycometabolism was the main factor in the process of SARS-CoV-2 infection (Fig. 6). Glycometabolism has been shown to have a close relationship with immune system function. Hyperglycemia results in an increase in peripheral blood mononuclear cells and the production of TNF-α (Tumor Necrosis Factor) and IL-1β [20], induction of oxidative stress [21], and the activation of leukocyte [22]. Hyperglycemia leads to reduction of glucose to sorbitol due to the increase of aldose reductase [23]. NADPH is consumed in the above process, and the antioxidant function of cells is reduced. Therefore, inhibition of glycometabolism pathways has gradually become a potential target for the treatment of inflammation [24]. In summary, disorder of glycometabolism may be a key event for COVID-19 patients.
1-Monopalmitin is a MAGs (monoacylglycerols) that is implicated in the pathogenesis of inflammation via being hydrolyzed by ABHD6 (α/β Hydrolase Domain Containing 6) [25]. Accumulation of 1-Monopalmitin depend on suppression of ABHD6 directly binds to PPARs (Peroxisome Proliferators Activated Receptors) to participate in the inflammatory process [26]. In addition, 1-Monopalmitin was up regulated in GVHD (Graft Versus Host Disease) and could be used as a pro-inflammatory metabolic biomarker [27]. The urea cycle is a central metabolic pathway in the liver, which converts toxic ammonia into non-toxic urea, its disorders can cause hyperammonemia and irreversible damage to the brain [28]. Increases in blood urea are commonly seen in nephropathy, dehydration, prostatic hypertrophy and chronic urinary tract obstruction. However, a recent study showed that virus infection can inhibit T cell function due to alteration of the urea cycle [29]. Additionally, urea-to-creatinine ratio were independent predictors of COVID-19 patients in the intensive care unit [30]. Hence, we speculated that the increase in 1-Monopalmitin may be one of the mechanisms of decreased cell function in COVID-19 patients, and urea may reflect the severity of the disease. Studies of metabolomics in COVID-19 have attracted more and more attention. Infected patients have widespread imbalances in nitrogen metabolism, and disorders in amino acid and fatty acid metabolism [31]. The appearance of differential metabolites such as leukotrienes and purines also participates in the inflammation and hypoxia response of COVID-19 patients [32]. This study screened the differential metabolites in the serum of patients with COVID-19 through non-targeted metabolomics and explained the possible mechanism of the decrease in peripheral blood lymphocytes and the storm of inflammatory factors. Nonetheless, the conclusions may be limited due to the low patient number Next, we will quantitatively analyze the differential metabolites in the serum of COVID-19 patients and evaluate their diagnostic value in COVID-19.
Conclusion
There was a decline in cellular immune function and a severe storm of inflammatory factors in COVID-19 patients. 53 metabolites were up regulated, 18 were down regulated and 8 were unchanged in the serum of COVID-19 patients. These differential metabolites affected the body’s immune system through different mechanisms. The increase of D-Allose, cholesterol, lactic acid and 1-monopalmitin may be one of the mechanisms of severe inflammation in COVID-19 patients.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article.
References
Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, Qiu Y, Wang J, Liu Y, Wei Y, Xia J, Yu T, Zhang X, Zhang L (2020) Lancet 395:507–513
Udugama B, Kadhiresan P, Kozlowski HN, Malekjahani A, Osborne M, Li V, Chen H, Mubareka S, Gubbay J, Chan W (2020) ACS Nano 14(4):3822–3835
Li D, Li J (2021) J Clin Microbiol 59(5):e02160-e2120
Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M, Xiao Y, Gao H, Guo L, Xie J, Wang G, Jiang R, Gao Z, Jin Q, Wang J, Cao B (2020) Lancet 395:497–506
Sun DW, Zhang D, Tian RH, Li Y, Wang YS, Cao J, Tang Y, Zhang N, Zan T, Gao L, Huang YZ, Cui CL, Wang DX, Zheng Y, Lv GY (2020) Clin Chim Acta 508:122–129
Wang F, Nie J, Wang H, Zhao Q, Xiong Y, Deng L, Song S, Ma Z, Mo P, Zhang Y (2020) J Infect Dis 221:1762–1769
Pepys MB (1982) Nature 296:12
Ji D, Zhang M, Qin E, Zhang L, Xu J, Wang Y, Cheng G, Wang F, Lau G (2021) Metabolism 115:154437
Jung J, Zeng H, Horng T (2019) Nat Cell Biol 21:85–93
Choi SY, Collins CC, Gout PW, Wang Y (2013) J Pathol 230:350–355
Polet F, Feron O (2013) J Intern Med 273:156–165
Shi D, Yan R, Lv L, Jiang H, Lu Y, Sheng J, Xie J, Wu W, Xia J, Xu K, Gu S, Chen Y, Huang C, Guo J, Du Y, Li L (2021) Metabolism 118:154739
Lee W, Ahn JH, Park HH, Kim HN, Kim H, Yoo Y, Shin H, Hong KS, Jang JG, Park CG, Choi EY, Bae JS, Seo YK (2020) Signal Transduct Target Ther 5:186
Sohrabi Y, Reinecke H, Godfrey R (2021) Trends Endocrinol Metab 32:132–134
Aguilar-Ballester M, Herrero-Cervera A, Vinue A, Martinez-Hervas S, Gonzalez-Navarro H (2020) Nutrients 12(7):1–19
Ma X, Bi E, Lu Y, Su P, Huang C, Liu L, Wang Q, Yang M, Kalady MF, Qian J, Zhang A, Gupte AA, Hamilton DJ, Zheng C, Yi Q (2019) Cell Metab 30:143-156.e145
Tanaka S, Sakamoto H (2011) Cell Immunol 271:141–146
Kanaji N, Kamitori K, Hossain A, Noguchi C, Katagi A, Kadowaki N, Tokuda M (2018) Oncol Rep 39:1292–1298
Lv L, Jiang H, Chen Y, Gu S, Xia J, Zhang H, Lu Y, Yan R, Li L (2021) Anal Chim Acta 1152:338267
Deopurkar R, Ghanim H, Friedman J, Abuaysheh S, Sia CL, Mohanty P, Viswanathan P, Chaudhuri A, Dandona P (2010) Diabetes Care 33:991–997
Wang L, Guo L, Zhang L, Zhou Y, He Q, Zhang Z, Wang M (2013) J Diabetes Res 2013:849295
Sampson MJ, Davies IR, Brown JC, Ivory K, Hughes DA (2002) Arterioscler Thromb Vasc Biol 22:1187–1193
Brownlee M (2001) Nature 414:813–820
Kim YH, Nakayama T, Nayak J (2018) Allergy Asthma Immunol Res 10:6–11
Alhouayek M, Masquelier J, Cani PD, Lambert DM, Muccioli GG (2013) Proc Natl Acad Sci U S A 110:17558–17563
Poursharifi P, Madiraju SRM, Prentki M (2017) Diabetes Obes Metab 19(Suppl 1):76–89
Contaifer D, Roberts CH, Kumar NG, Natarajan R, Fisher BJ, Leslie K, Reed J, Toor AA, Wijesinghe DS (2019) Cancers 11(8):1051
Matsumoto S, Hberle J, Kido J, Mitsubuchi H, Endo F, Nakamura K (2019) J Hum Genet 64:833–847
Lercher A, Bhattacharya A, Popa AM, Caldera M, Schlapansky MF, Baazim H, Agerer B, Gurtl B, Kosack L, Majek P, Brunner JS, Vitko D, Pinter T, Genger JW, Orlova A, Pikor N, Reil D, Ozsvar-Kozma M, Kalinke U, Ludewig B, Moriggl R, Bennett KL, Menche J, Cheng PN, Schabbauer G, Trauner M, Klavins K, Bergthaler A (2019) Immunity 51:1074-1087.e1079
Solimando AG, Susca N, Borrelli P, Prete M, Lauletta G, Pappagallo F, Buono R, Inglese G, Forina BM, Bochicchio D, Capobianco M, Carrieri V, Cicco S, Leone P, Silvestris N, Saracino A, Ria R, Procacci V, Migliore G, Vacca A, Racanelli V (2020) Front Med (Lausanne) 7:625176
Thomas T, Stefanoni D, Reisz JA, Nemkov T, Bertolone L, Francis RO, Hudson KE, Zimring JC, Hansen KC, Hod EA, Spitalnik SL, D’Alessandro A (2020) JCI Insight. 5(14):e140327
Doan HO, Senol O, Bolat S, Yldz N, Büyüktuna SA, Sarıismailoğlu R, Doğan K, Hasbek M, Hekim SN (2021) J Med Virol 93:2340–2349
Acknowledgements
Thanks to the Emergency Research Projects of COVID-19 in Mianyang (2020YJKY012) for supporting this study. Thank you to everyone who has contributed to the fight against COVID-19 .
Funding
This study was supported by the Emergency Research Projects of COVID-19 in Mianyang (2020YJKY012).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors (Xiangwa Chen, Xiujuan Gu, Jie Yang, Zhengfang Jiang and Jianjun Deng) declare that they have no conflict of interest. This article does not contain any studies with human or animal subjects.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, X., Gu, X., Yang, J. et al. Gas Chromatography–Mass Spectrometry Technology: Application in the Study of Inflammatory Mechanism in COVID-19 Patients. Chromatographia 86, 175–183 (2023). https://doi.org/10.1007/s10337-022-04222-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10337-022-04222-3