Abstract
Bitter taste perception enables the detection of potentially toxic molecules and thus evokes avoidance behavior in vertebrates. It is mediated by bitter taste receptors, TAS2Rs. One of the best-studied TAS2R is TAS2R38. Phenylthiocarbamide (PTC) perception and TAS2R38 receptors vary across primate species, and this variation may be related to variation in dietary preferences. In particular, we previously found that the low sensitivity of TAS2R38s in Asian colobines likely evolved as an adaptation to their leaf-eating behavior. However, it remains unclear whether this low PTC sensitivity is a general characteristic of the subfamily Colobinae, a primate group that feeds predominantly on leaves. We performed genetic analyses, functional assays with mutant proteins, and behavioral analyses to evaluate the general characteristics of TAS2R38 in colobines. We found that PTC sensitivity is lower in TAS2R38s of African colobines than in TAS2R38s of omnivorous macaques. Furthermore, two amino acids shared between Asian and African colobines were responsible for low sensitivity to PTC, suggesting that the last common ancestor of extant colobines had this phenotype. We also detected amino acid differences between TAS2R38s in Asian and African colobines, indicating that they evolved independently after the separation of these groups.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bitter taste perception enables the detection of potentially toxic molecules and thus evokes avoidance behavior in vertebrates. It is mediated by bitter taste receptors, TAS2Rs (Meyerhof 2005). A well-studied TAS2R, TAS2R38, recognizes thioamides, including both natural (sinigrin, allyl isothiocyanate, and goitrin) and synthetic ligands (phenylthiocarbamide [PTC] and propylthiouracil [PROP]) (Meyerhof et al. 2010). Previous studies have shown that variation exists in PTC perception and TAS2R38 receptors among primate species and have suggested that PTC perception reflects dietary preferences (Chiarelli 1963; Wooding et al. 2006; Hayakawa et al. 2012, 2014; Suzuki-Hashido et al. 2015). For example, most species of primates that belong to the subfamily Cercopithecinae, which includes omnivores and frugivores, are generally characterized as PTC-tasters (Chiarelli 1963; Suzuki-Hashido et al. 2015; Widayati et al. 2019).
Colobines (subfamily Colobinae), which include reciprocally monophyletic Asian and African groups (Ting et al. 2008; Roos et al. 2011; Wang et al. 2012), are unique among primates because their diet is composed largely of plant material, predominantly leaves (Tsuji et al. 2019; Oates 1978). The colobines possess large, specialized stomachs (Lambert 1998), and this may be an adaptive trait to increase the efficiency of processing fibrous foods such as foliage (Caton 1999). Bitter taste perception in colobines is particularly interesting, because plants use chemical defenses to deter herbivores (Ames et al. 1990), and these chemicals could be bitter or toxic (Wooding et al. 2010).
In cellular and behavioral studies, we found that Asian colobines have low sensitivity to the bitter compound PTC (Purba et al. 2017). However, it remains unclear whether low PTC sensitivity is a general characteristic of the subfamily Colobinae. In this study, we performed genetic analyses, functional assays with mutant proteins, and behavioral assays to evaluate the general characteristics of TAS2R38 of extant species and the last common ancestor of extant colobines with respect to PTC sensitivity. We here show that TAS2R38 in African colobines and the predicted last common ancestor of colobines as well as Asian colobines have lower PTC sensitivity than that of macaque TAS2R38. This result supports our previous hypothesis that the low sensitivity to bitter flavors in the colobine lineage is an adaptation to their leaf-eating behavior. The molecular mechanisms underlying variation in sensitivity are also discussed based on point mutations in TAS2R38 in macaques and colobines.
Materials and methods
Genotyping of TAS2R38 in African colobines
Genomic DNA of African colobines (Colobus guereza [Cg] and C. angolensis [Ca]; Fig. 1) was extracted from either tissue or fecal samples. Tissue samples were collected from dead individual monkeys provided by the Japan Monkey Centre. Fecal samples were collected from live animals at the Japan Monkey Centre (Table S1). Genomic DNA was extracted using a DNeasy Blood & Tissue Kit (Qiagen GmbH, Hilden, Germany) for tissue samples or QIAamp DNA Stool Mini Kit (Qiagen GmbH) for fecal samples.
The entire coding region of TAS2R38 from each DNA sample was amplified and sequenced using primers described in a previous study (Suzuki-Hashido et al. 2015). TAS2R38 amplification was performed by PCR using ExTaq DNA Polymerase (Takara Bio Inc., Shiga, Japan) under the following conditions: initial denaturation at 94 °C for 10 min; 40–45 cycles of denaturation at 94 °C for 10 s, annealing at 56 °C for 30 s, and extension at 72 °C for 1 min, followed by a final extension at 72 °C for 10 min. PCR products were sequenced using BigDye Terminator v3.1 (Applied Biosystems, Carlsbad, CA), and the sequencing products were separated by capillary electrophoresis using a 3130xl Genetic Analyzer (Applied Biosystems). Five samples of C. guereza (three tissue samples and two fecal samples) and two samples from two individuals of C. angolensis (one tissue sample and one fecal sample) were sequenced. The sequences were deposited in the DNA Data Bank of Japan (DDBJ) under accession numbers LC500224 (C. angolensis-A TAS2R38), LC500225 (C. angolensis-B TAS2R38), and LC500226 (C. guereza TAS2R38).
Amino acid differences were compared between TAS2R38 in African colobines and humans (Homo sapiens; AY258597.1, Kim et al. 2003), chimpanzees (JQ272199.1, Pan troglodytes, Wooding et al. 2006), Japanese macaques (Macaca fuscata, AB623012.1, Suzuki et al. 2010), and Asian colobines (Trachypithecus auratus LC167094.1, T. cristatus LC167094.1 and LC167093.1, Presbytis femoralis LC167095.1, and Nasalis larvatus LC167091.1, Purba et al. 2017) (Table S2).
Expression vector
The coding regions of Asian (Purba et al. 2017) and African colobine TAS2R38 were appended to the last eight amino acids of bovine rhodopsin (ID4) at the C-terminal end. Subsequently, the tagged TAS2R38 fragments and 45 amino acids of rat somatostatin receptor type 3 (ssr3) were inserted into the mammalian expression vector pEAK10 (Edge BioSystems, Inc., Gaithersburg, MD) using the In-Fusion HD Cloning Kit (Takara Bio Inc.). Japanese macaque (Mf) TAS2R38 was prepared as described previously (Suzuki-Hashido et al. 2015). The vectors were transfected to cells for functional assays.
Site-directed mutagenesis and cell-based functional assay
Site-directed mutagenesis was performed using the QuikChange Mutagenesis Kit (Agilent Technologies, Santa Clara, CA) following the manufacturer’s protocol. Site-directed mutations were introduced by annealing mutagenic primers to the Macaca fuscata and/or colobine vectors. After extension, the products were digested with DpnI and transformed into XL-Gold ultracompetent cells. All clones were sequenced to verify mutations and to exclude amplification errors.
The TAS2R38 expression vector was transfected along with Gα16gust44 into HEK 293T cells using Lipofectamine 2000 (Invitrogen, Carlsbad, CA) (Ueda et al. 2003). For the functional assay, the fluorescence dye Calcium 4 (Molecular Devices, Inc., Eugene, OR) was used as an intracellular Ca2+ indicator. Fluorescence was measured at 525 nm following excitation at 485 nm using the FlexStation 3 Multi-Mode Microplate Reader (Molecular Devices Japan, Inc.). The calcium response amplitudes (ΔF/F) were defined as the ratio of the ligand-dependent increase in fluorescence to the fluorescence before ligand addition. The response of cells that were transfected with the empty pEAK10 vector and Gα16gust44 was defined as the mock response. The average ΔF/F values from at least three independent experiments were plotted, and a nonlinear regression function was fitted to the data as follows: f(x) = Imin + (Imax−Imin)/(1 + (x/EC50)h), where x is the ligand concentration and h is the Hill coefficient used to calculate the EC50 (half maximal effective concentration) for the ligand–receptor interaction between PTC and TAS2R38. Curve fitting and parameter estimation were performed using the drc package (Ritz et al. 2015) in R v3.4.3 (https://www.r-project.org/). The maximum activity (maximum responses) to various concentrations of PTC were calculated as the normalized peak response (F) relative to background fluorescence (F0), ΔF/F [= (F − F0)/F0]. Original functional data are available from the Dryad Digital Repository ( https://doi.org/10.5061/dryad.908jf3r).
Behavioral assay
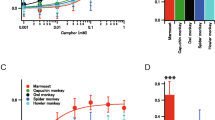
A behavioral experiment was performed using a single C. angolensis (adult male, born at the Japan Monkey Centre) and two C. guereza (juvenile females, born at the Japan Monkey Centre) (Table S1). Monkeys were given slices of apple that had been soaked overnight in the test solution (2 mM PTC in water) (Purba et al. 2017; Suzuki et al. 2010). A trial began when a monkey put an apple slice in its mouth. Whether they ate (accepted) or spat out (rejected) the apple was recorded. The rate (%) of acceptance was calculated for PTC and baseline (0 mM PTC). To confirm that the PTC-treated apples contained PTC, the two types of apple slices were provided to a single Japanese macaque (M. fuscata) (adult male, kept at the Primate Research Institute, Kyoto University). Differences in the frequency of acceptance between PTC and the control were assessed by proportion test (Newcombe 2016) using the stats package in R.
Results
Asian colobines and the predicted last common ancestor of Colobinae
TAS2R38 in Asian colobines differed at four amino acid residues (V44I/Q93E/I148F/R330K) from TAS2R38 in humans, chimpanzees, and macaques, which are conserved. Single-site mutations of macaque TAS2R38 at these amino acids to mimic colobine TAS2R38 result in remarkably decreased responses to PTC (Purba et al. 2017).
In the present study, multiple M. fuscata TAS2R38 mutants were created to evaluate the effects of multiple amino acid changes on receptor sensitivity. In particular, we performed site-directed mutagenesis of macaque TAS2R38 at positions 44, 93, 148, and 330 to mimic TAS2R38 in Asian colobines. We also performed site-directed mutagenesis of TAS2R38 in three Asian colobine species (T. auratus, P. femoralis, and N. larvatus) at amino acid positions 44, 93, 148, and 330 to mimic TAS2R38 in macaques. The macaque TAS2R38 mimicking TAS2R38 in Asian colobines showed a significantly weaker response to PTC (M. fuscata-V44I/Q93E/I148F/R330K, EC50 = 28.78 ± 4.00 µM, pairwise t test, p < 0.05) than that of wild-type macaque TAS2R38 (M. fuscata-WT, EC50 = 1.56 ± 0.11 µM) (Fig. 2a). The colobine TAS2R38s of T. cristatus and P. femoralis mimicking macaque TAS2R38 showed increased sensitivity to PTC (T. cristatus-I44V/E93Q/F148I/K330R, EC50 = 3.72 ± 0.14 µM; P. femoralis-I44V/E93Q/F148I/K330R, EC50 = 1.29 ± 0.14 µM) (Fig. 2b, c, Table 1). These results implied that amino acid positions 44, 93, 148, and 330 are important determinants of PTC sensitivity, supporting our previous hypothesis (Purba et al. 2017). N. larvatus TAS2R38 mimicking macaque TAS2R38 (N. larvatus-I44V/E93Q/F148I/K330R) showed no difference from wild-type TAS2R38 (Fig. 2d). We found four amino acid mutations at positions 184, 199, 216, and 243 specific to TAS2R38 of N. larvatus. These four mutations might be responsible for the low sensitivity of N. larvatus TAS2R38. We constructed a mutant N. larvatus TAS2R38 with eight amino acid substitutions (N. larvatus-I44V/E93Q/F148I/R184W/C199Y/M216L/M243I/K330R) to mimic TAS2R38 of macaques. The mutant showed a similar response to that of WT M. fuscata, used as a positive control (EC50 = 3.45 ± 0.47 µM, p = 0.43) (Fig. 2d). These results showed that the four additional amino acid residues were responsible for the low sensitivity of N. larvatus TAS2R38 to PTC. We also induced two types of TAS2R38 mutations at seven sites (N. larvatus-I44V/E93Q/F148I/R184W/C199C/M216L/M243I/K330R; N. larvatus-I44V/E93Q/F148I/R184W/C199Y/M216M/M243I/K330R). The N. larvatus-I44V/E93Q/F148I/R184W/C199C/M216L/M243I/K330R showed no response to PTC, while N. larvatus-I44V/E93Q/F148I/R184W/C199Y/M216M/M243I/K330R showed intermediate response to PTC (EC50 = 7.2 ± 1.96 µM) (Fig. S1). These results showed that an amino acid Y199 partly contributes to the response to PTC.
Dose–response curves for mutant macaque TAS2R38 mimicking Asian colobine TAS2R38 (a) and reverse mutants of Asian colobine TAS2R38 mimicking macaque TAS2R38 (b–d). Squares indicate wild-type TAS2R38 of the species (filled box), mutant TAS2R38 (blank box), and negative control. Values are derived from at least three independent experiments. The EC50 value for each variant of TAS2R38 was compared with that of M. fuscata wild type as a positive control (p < 0.05, pairwise t test)
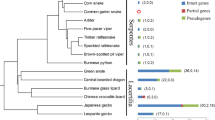
Based on the phylogenetic relationships among TAS2R38 proteins, we predicted that two amino acid substitutions (I148F and R330K) occurred before the separation of the Asian and African colobine lineages, while V44I and Q93E occurred after the separation of these lineages (Fig. 3). We hypothesized that the last common ancestor of colobines possessed F148 and K330. Thus, to study the evolution of TAS2R38 in Colobinae, we evaluated the effects of combinations of these four amino acid mutations on sensitivity in the Asian lineage.
Phylogenetic relationships among primates based on amino acid sequences of TAS2R38. Numbers at the nodes represent bootstrap values. The relationships are consistent with the species phylogeny obtained using other molecular data (Roos et al. 2011)
We engineered several macaque TAS2R38 variants mimicking colobine TAS2R38 (Fig. 4, Table 1). The two-site mutations of macaque TAS2R38 mimicking colobine TAS2R38 at amino acid positions 148 and 330 caused a significantly weaker response to PTC (M. fuscata-I148F/R330K, EC50 = 7.91 ± 1.39 µM) than that of wild-type macaque TAS2R38 (pairwise t test, p < 0.05). These results suggest that the predicted TAS2R38 of the last common ancestor of colobines showed an intermediate response to PTC compared with those of macaques and extant colobines.
Sensitivities of macaque TAS2R38 and mutants mimicking colobine TAS2R38 to PTC. Each point represents the mean ± standard error of the mean [SEM] determined from at least three independent measurements. The EC50 value for each variant of TAS2R38 was compared with that of M. fuscata wild type as a positive control (p < 0.05, pairwise t test)
The macaque TAS2R38 variant with three mutations at amino acid positions 44, 148, and 330 showed lower sensitivity to PTC than that of wild-type macaque TAS2R38 (M. fuscata-V44I/I148F/R330K, EC50 = 6.75 ± 0.02 µM, pairwise t test, p < 0.05) but showed similar sensitivity to that of the M. fuscata-I148F/R330K variant (pairwise t test, p = 0.73). We also found that another macaque TAS2R38 variant with three mutations at positions 93, 148, and 330 showed a significantly weaker response to PTC than those of wild-type TAS2R38 in macaques and the ancestral TAS2R38 of colobines (M. fuscata-I148F/R330K) (M. fuscata-Q93E/I148F/R330K, EC50 = 15.64 ± 4.81 µM, pairwise t test, p < 0.05). These results confirmed that the mutations at I148F/R330K reduce sensitivity and clarified the molecular basis of the low PTC sensitivity in Asian colobines and predicted last common ancestor of colobines. The amino acid change from Q to E at position 93 is crucial for the reduced sensitivity to PTC in Asian colobines.
Variation in TAS2R38 genotypes and phenotypes of African colobines
Since Asian colobines showed lower responses to PTC than those of macaques, we expanded our analysis to African colobines to evaluate whether this trait is a general characteristic of present-day colobines. We sequenced the coding regions of TAS2R38 in two species of African colobines, C. angolensis and C. guereza. Orthologous TAS2R38 sequences from each species were intact (1002 bp).
African colobine TAS2R38 differed at four amino acid positions, 34, 148, 167, and 330 (A34G/H167P/I148F/R330K or A34G/H167P/I148F/R330M), from human, chimpanzee, and macaque TAS2R38 (Table S2). Two amino acid mutations (I148F/R330K) were shared among all Asian and African colobines. Glycine at position 34 was unique to TAS2R38 of the African colobine lineage. An amino acid substitution at site 167 of TAS2R38 was shared among all African colobines and some Asian colobines in the genus Trachypithecus (T. cristatus and T. auratus) (Table S2), which had proline while other species of Asian colobines (P. femoralis and N. larvatus) had histidine at this position, consistent with human, chimpanzee, and macaque TAS2R38. There were three TAS2R38 variants in African colobines (Colobus angolensis-A, C. angolensis-B, and C. guereza). C. angolensis-A and C. angolensis-B differed only at amino acid position 125 (Table S2).
We performed an in vitro analysis of African colobine TAS2R38 functions in response to PTC. TAS2R38 in C. angolensis-A and C. guereza showed dose-dependent responses to PTC, and these responses were weaker than that of TAS2R38 in macaques (C. angolensis-A, EC50 = 13.34 ± 1.09 µM; C. guereza, EC50 = 5.99 ± 1.10 µM, Fig. 5, Table 1) but slightly higher than those of Asian colobine TAS2R38. Additionally, TAS2R38 in C. angolensis-B did not respond to PTC, even at the highest concentration (100 µM PTC), as in Asian colobines (Fig. 5). The responses of African colobine TAS2R38s were significantly lower than those of Japanese macaque TAS2R38 (pairwise t test, p < 0.05). C. guereza showed a significantly higher response to PTC than that of C. angolensis-A (pairwise t test, p < 0.05). These results indicated that the responses of African colobine TAS2R38 receptors to PTC varied from no response to an intermediate response.
In vitro responses of TAS2R38 of African colobines against PTC compared with those of wild-type TAS2R38 of M. fuscata. Each point represents the mean ± standard error of the mean [SEM] determined from three independent measurements. a Responses of TAS2R38 receptors of wild-type African colobines and their mutants. b Responses of TAS2R38 receptors of mutant TAS2R38 of macaques mimicking those of African colobines. The EC50 value for each variant of TAS2R38 was compared with that of M. fuscata wild type as a positive control (p < 0.05, pairwise t test)
To evaluate the effects of the three amino acids on PTC sensitivity, we performed site-directed mutagenesis at positions 34, 148, and 330 of macaque TAS2R38 to mimic the TAS2R38 in African colobines. The mutant macaque TAS2R38 mimicking African colobine TAS2R38 showed a significantly weaker response to PTC (M. fuscata-A34G/I148F/R330K, EC50 = 6.47 ± 1.30 µM; M. fuscata-A34G/I148F/R330M, EC50 = 7.44 ± 1.14 µM; M. fuscata-H167P/I148F/R330K, EC50 = 10.47 ± 1.69 µM; M. fuscata-A34G/H167P/I148F/R330K, EC50 = 13.34 ± 3.40 µM, pairwise t test, p < 0.05) than that of wild-type macaque TAS2R38 (Fig. 5b, Table 1). The three-site macaque TAS2R38 mutants A34G/I148F/R330M and A34G/I148F/R330K had similar EC50 values as those of the predicted last common ancestor of colobines, I148K/R330K (p = 0.93 and 0.60, respectively). In addition, we performed site-directed mutagenesis and functional assays to evaluate the effects of the mutation at amino acid residue 167. We engineered the three-site mutant of macaque TAS2R38 M. fuscata-H167P/I148F/R330K to mimic colobine TAS2R38. This mutant showed significantly lower PTC sensitivity than that of wild-type macaque TAS2R38 (M. fuscata-H167P/I148F/R330K, EC50 = 10.47 ± 1.69 µM, pairwise t test, p < 0.05), but had similar EC50 values as those of the predicted last common ancestor of colobines, I148K/R330M (p = 0.30). Mutations from alanine to glycine at site 34 (A34G) and from histidine to proline at site 167 (H167P) did not significantly reduce the sensitivity to PTC in the African colobine lineage.
We conducted behavioral assays using two African colobine species to evaluate PTC sensitivity at the behavioral level. An individual with the C. angolensis-B haplotype homozygously ate both control and PTC-treated apples (prop.test, p = 0.2841, Table 2), suggesting that it could not detect PTC. These results were consistent with those of the functional assay for TAS2R38 in C. angolensis-B, which also showed no response to PTC. However, two individuals of C. guereza rejected 2 mM PTC-soaked apple pieces, as observed in Japanese macaques (prop.test, p < 0.05, Table 2). This observation indicated that C. guereza is able to detect PTC, consistent with the intermediate response of TAS2R38 in African colobines in the functional assay.
Discussion
Our results revealed that TAS2R38 in African colobines and the predicted TAS2R38 of the last common ancestor in colobines showed weaker responses to PTC than that of macaque TAS2R38 but a greater response than that of Asian colobine TAS2R38. Based on these results, low sensitivity to PTC is a general characteristic of colobines, but the magnitude of the reduction in sensitivity varies, especially in African colobines. This result strengthens our previous hypothesis that low sensitivity to bitter flavors in the colobine lineage is an adaptation to their leaf-eating behavior (Purba et al. 2017).
There is a clear difference between plant species consumed by C. angolensis and C. guereza (Anderson 2005; Dunham 2017; Oates 1977; Fashing et al. 2007). Even in two colobus species found in sympatry in Ituri Forest, Democratic Republic of Congo, only 28.5% of the plant species in their diet is shared (Bocian 1997). C. angolensis populations consume the leaves of Trichilia emetica (family Meliaceae), which are considered toxic and are avoided by many animals (Dunham 2011; Komane et al. 2011). C. guereza do not eat these leaves, even though they are available in the habitat in Kakamega Forest, Kenya (Fashing et al. 2004, 2007). Based on phytochemical studies, plant species belonging to the family Meliaceae contain limonoids (Roy and Saraf 2005), which are secondary metabolites in the triterpenoid group. One limonoid, limonin, is an agonist of TAS2R38 (Meyerhof et al. 2010). Thus, we predict that the reduced sensitivity of TAS2R38 enables C. angolensis to eat the leaves of T. emetica with high limonoid content.
Two amino acids responsible for low sensitivity to PTC at positions 148 and 330 are shared between Asian and African colobines. Based on a phylogenetic analysis, we predicted that these amino acids were present in the last common ancestor of these colobines. There were amino acid differences between Asian and African colobines, indicating that TAS2R38 in the two lineages evolved independently after their separation around 10 million years ago (Stewart and Disotell 1998). Based on molecular markers, the most parsimonious explanation for the origin of colobines is that they first evolved in Africa and subsequently dispersed to Asia (Zhang and Ryder 1998). We predicted that after their separation, the TAS2R38 in Asian and African lineages evolved independently in the context of their local environments. Such independent adaptation has also been observed for their digestive RNases (Zhang 2006).
The TAS2R38 of the predicted common ancestor of colobines showed a weaker response to PTC than that of macaque TAS2R38 and greater sensitivity than that of Asian colobine TAS2R38. Moreover, TAS2R38 in African colobines showed variation in responses to PTC. Some variants showed intermediate responses to PTC (C. angolensis-A and C. guereza) but C. angolensis-B showed no response to PTC. The similarity in the response to PTC between African and ancestral colobines may be explained by the relatively long history of shared environmental and climatic patterns. Intermediate responses to PTC have also been observed in human TAS2R38, typically in populations in Africa (Campbell et al. 2012). We speculate that the similarity in environmental conditions mediated the convergent evolution of TAS2R38 functions in Africa. A similar pattern has been found for the dentition of colobines; the relationships between teeth and body size of African colobines are more similar to those of cercopithecines than to those of Asian colobines (Pan 2006).
Our results showed that three amino acid mutations at positions 93, 148, and 330 contribute to the reduced sensitivity to PTC of TAS2R38 in Asian colobines. Amino acid residue 330 is located in the intracellular region of the receptor (Fig. S2), which may mediate interactions with G proteins. Amino acid residue 93 is located in the extracellular loop (Fig. S2), which may mediate extracellular ligand interactions. In more detail, according to previous modeling studies, it is located close to TM3 in extracellular loop 1 (ECL1), which is not close to the PTC binding site. It may indirectly affect ligand binding or the distance between the ligand and the binding pocket (Biarnes et al. 2010; Marchiori et al. 2013). Amino acid residue 148 is located in the middle of transmembrane (TM) 4 (Fig. S2). Based on a homology modeling study, TM4 does not directly affect the binding site (Biarnes et al. 2010; Marchiori et al. 2013). The substitution at position 148 may have an indirect effect to binding pocket or effect for receptor folding. Based on a functional assay, we confirmed that an amino acid substitution from non-polar glutamine (Q) to electric-charge glutamic acid (E) at position 93 had the largest effect on the sensitivity of TAS2R38 in Asian colobines. The extracellular regions of G protein-coupled receptors are important for ligand recognition (Vaidehi et al. 2002). Thus, the amino acid mutation at position 93 may alter the three-dimensional structure of Asian colobine TAS2R38, which could alter binding to PTC. African colobine showed a lineage-specific amino acid at position 34, in TM1. However, this amino acid mutation did not significantly alter the sensitivity to PTC in comparison with that of the predicted last common ancestor of colobines. Among variants of TAS2R38 in African colobines, C. angolensis-B had no response to PTC. This variant possessed one amino acid mutation at position 125, from serine to phenylalanine, different from C. angolensis-A, which responds to PTC. The amino acid mutation was located in intracellular loop (IL) 2. In many G protein-coupled receptors, IL2 is an important region for G protein coupling (Wong 2003). Thus, the mutation might alter proper coupling between TAS2R38 and G proteins.
In conclusion, two amino acids at positions 148 and 330 shared between Asian and African colobines were responsible for low sensitivity to PTC. We predicted that these amino acids were already present in the last common ancestor of Asian and African colobines. The amino acid differences between Asian and African colobines indicated that TAS2R38 evolved independently in these lineages after their separation. The amino acid mutation at position 93 might substantially reduce sensitivity in nearly all Asian colobines, while African colobines still retain the ancestral sequence or distinct alterations. Our findings improve our understanding of the evolution of bitter taste perception in colobine lineages.
References
Anderson J (2005) Habitat fragmentation and metapopulation dynamics of the Angola black-and-white colobus (Colobus angolensis palliatus) in coastal Kenya. In: Dissertation, University College London.
Ames BN, Profet M, Gold LS (1990) Nature’s chemicals and synthetic chemicals: comparative toxicology. Proc Natl Acad Sci 87:7782–7786
Biarnes X, Marchiori A, Giorgetti A, Lanzara C, Gasparini P, Carloni P, Born S, Brockhoff A, Behrens M, Meyerhof W (2010) Insights into the binding of phenyltiocarbamide (PTC) agonist to its target human TAS2R38 bitter receptor. PLoS One 5(8):e12394. https://doi.org/10.1371/journal.pone.0012394
Bocian (1997) Niche separation of black-and-white colobus (Colobus angolensis and C. guereza) in the Ituri forest. In: Dissertation, City University of New York
Campbell MC, Ranciaro A, Froment A, Hirbo J, Omar S, Bodo JM, Nyambo T, Lema G, Zinshteyn D, Drayna D, Breslin PAS, Tishkoff SA (2012) Evolution of functionally diverse alleles associated with PTC bitter taste sensitivity in Africa. Mol Biol Evol 29:1141–1153. https://doi.org/10.1093/molbev/msr293
Caton JM (1999) Digestive strategy of the Asian colobine genus Trachypithecus. Primates 40:311–325
Chiarelli B (1963) Sensitivity to PTC (phenyl-thio-carbamide) in primates. Folia Primatol 1:88–94
Dunham NT (2017) Feeding ecology and dietary flexibility of Colobus angolensis palliatus in relation to habitat disturbance. Int J Primatol 38(3):553–557
Fashing PJ, Dierenfeld ES, Mowry CB (2007) Influence of plant and soil chemistry on food selection, ranging patterns, and biomass of Colobus guereza in Kakamega Forest, Kenya. Int J Primatol 28(3):673–703. https://doi.org/10.1007/s10764-006-9096-2
Fashing PJ, Forrestel A, Scully C, Cords M (2004) Long-term tree population dynamics and their implications for the conservation of the Kakamega Forest, Kenya. Biodivers Conserv 13:753–771
Hayakawa T, Sugawara T, Go Y, Udono T, Hirai H, Imai H (2012) Eco-geographical diversification of bitter taste receptor genes (TAS2Rs) among subspecies of chimpanzees (Pan troglodytes). PLoS One 7:e43277. https://doi.org/10.1371/journal.pone.0043277
Hayakawa T, Suzuki-Hashido N, Matsui A, Go Y (2014) Frequent expansions of the bitter taste receptor gene repertoire during evolution of mammals in the Euarchontoglires clade. Mol Biol Evol 31:2018–2031
Kim UK, Jorgenson E, Coon H, Leppert M, Risch N, Drayna D (2003) Positional cloning of the human quantitative trait locus underlying taste sensitivity to phenylthiocarbamide. Science 299:1221–1225. https://doi.org/10.1126/science.1080190
Komane BM, Olivier EI, Viljoen AM (2011) Trichilia emetica (Meliaceae)—a review of traditional uses, biological activities and phytochemistry. Phytochem Lett 4:1–9
Lambert JE (1998) Primate digestion: interactions among anatomy, physiology, and feeding ecology. Evol Anthropol 7:8–20
Marchiori A, Capece L, Giorgetti A, Gasparini P, Behrens M, Carloni P, Meyerhof W (2013) coarse-grained/molecular mechanics of the TAS2R38 bitter taste receptor: experimentally-validated detailed structural prediction of agonist binding. PLoS One 8(5):e64675. https://doi.org/10.1371/journal.pone.0064675
Meyerhof W (2005) Elucidation of mammalian bitter taste. Rev Physiol Biochem Pharmacol 154:37–72. https://doi.org/10.1007/s10254-005-0041-0
Meyerhof W, Batram C, Kuhn C, Brockhoff A, Chudoba E, Bufe B, Appendino G, Behrens M (2010) the molecular receptive ranges of human TAS2R bitter taste receptors. Chem Senses 35:157–170. https://doi.org/10.1093/chemse/bjp092
Newcombe RG (2016) Interval estimation for the difference between independent proportions: comparison of eleven methods. Statist Med 17:873–890
Oates JF (1977) The guereza and its food. In: Clutton-Brock TH (ed) Primate ecology. Academic Press, New York, pp 275–321
Oates JF (1978) Water-plant and soil consumption by guereza monkeys (Colobus guereza): a relationship with minerals and toxins in the diet. Biotropica 1978:241–253
Pan R (2006) Dental morphometric variation between African and Asian colobines, with special reference to the other Old World monkeys. J Morphol 267:1087–1098. https://doi.org/10.1002/jmor.10463
Purba LHPS, Widayati KA, Tsutsui K, Suzuki-Hashido N, Hayakawa T, Nila S, Suryobroto B, Imai H (2017) Functional characterization of the TAS2R38 bitter taste receptor for phenylthiocarbamide in colobine monkeys. Biol Lett 13:20160834. https://doi.org/10.1098/rsbl.2016.0834
Ritz C, Baty F, Streibig JC, Gerhard D (2015) Dose-response analysis using R. PLoS One 10:e0146021. https://doi.org/10.1371/journal.pone.0146021
Roos C, Zinner D, Kubatko L, Schwarz C, Yang M, Meyer D, Nash SD, Xing J, Batzer MA, Brameier M, Leenderzt FH, Ziegler T, Perwitasari-Farajallah D, Nadler T, Walter L, Osterholz M (2011) Nuclear versus mitochondrial DNA: evidence for hybridization in colobine monkeys. BMC Evol Biol 11:77
Roy A, Saraf S (2005) Limonoids: overview of significant bioactive triterpenes distributed in plants kingdom. Biol Pharm Bull 29(2):191–201
Stewart CB, Disotell TR (1998) Primate evolution: in and out of Africa. Curr Biol 8:582–588
Suzuki N, Sugawara T, Matsui A, Go Y, Hirai H, Imai H (2010) Identification of non-taster Japanese macaques for a specific bitter taste. Primates 51:285–289. https://doi.org/10.1007/s10329-010-0209-3
Suzuki-Hashido N, Hayakawa T, Matsui A, Go Y, Ishimaru Y, Misaka T, Abe K, Hirai H, Satta Y, Imai H (2015) Rapid expansion of phenylthiocarbamide non-tasters among Japanese macaques. PLoS One 10:e0132016. https://doi.org/10.1371/journal.pone.0132016
Ting N, Tosi AJ, Li Y, Zhang Y-P, Disotell TR (2008) Phylogenetic incongruence between nuclear and mitochondrial markers in the Asian colobines and the evolution of the langurs and leaf monkeys. Mol Phylogenet Evol 46(2):466–474. https://doi.org/10.1016/j.ympev.2007.11.008
Tsuji Y, Mitani M, Widayati KA, Suryobroto B, Watanabe K (2019) Dietary habits of wild Javan lutungs (Trachypithecus auratus) in a secondary-plantation mixed forest: effects of vegetation composition and phenology. Mamm Biol 98:80–90. https://doi.org/10.1016/j.mambio.2019.08.001
Ueda T, Ugawa S, Yamamura H, Imaizumi Y, Shimada S (2003) Functional interaction between T2R taste receptors and G-protein α subunits expressed in taste receptor cells. J Neurosci 23:7376–7380
Vaidehi N, Floriano WB, Trabanino R, Hall SE, Freddolino P, Choi EJ, Zamanakos G, Goddard WA (2002) Prediction of structure and function of G protein-coupled receptors. Proc Natl Acad Sci 99:12622–12627. https://doi.org/10.1073/pnas.122357199
Wang XP, Yu L, Roos C, Ting N, Chen CP, Wang J, Zhang YP (2012) Phylogenetic relationships among the colobine monkeys revisited: new insights from analyses of complete mt genomes and 44 nuclear non-coding markers. PLoS One 7(4):e36274. https://doi.org/10.1371/journal.pone.0036274
Widayati KA, Yan X, Suzuki-Hashido N, Itoigawa A, Purba LHPS, Fahri F, Terai Y, Suryobroto B, Imai H (2019) Functional divergence of the bitter receptor TAS2R38 in Sulawesi macaques. Ecol Evol 00:1–7. https://doi.org/10.1002/ece3.5557
Wong SKF (2003) G protein selectivity is regulated by multiple intracellular regions of GPCR. Neurosignals 12:1–12. https://doi.org/10.1159/000068914
Wooding S, Bufe B, Grassi C, Howard MT, Stone AC, Vazquez M, Dunn DM, Meyerhof W, Weiss RB, Bamshad MJ (2006) Independent evolution of bitter-taste sensitivity in humans and chimpanzees. Nature 440:930–934. https://doi.org/10.1038/nature04655
Wooding S, Gunn H, Ramos P, Thalmann S, Xing C, Meyerhof W (2010) Genetics and bitter taste responses to goitrin, a plant toxin found in vegetables. Chem Senses 35(8):685–692
Zhang J (2006) Parallel adaptive origins of digestive RNases in Asian and African leaf monkeys. Nat Genet 38:819–823. https://doi.org/10.1038/ng1812
Zhang Y, Ryder OA (1998) Mitochondrial cytochrome b gene sequences of Old World Monkeys: with special reference on evolution of Asian Colobines. Primates 39:39–49. https://doi.org/10.1007/BF02557742
Acknowledgements
The authors would like to thank Dr. Takashi Ueda, Dr. Yoshiro Ishimaru, Dr. Takumi Misaka, and Dr. Keiko Abe for providing the Ga16/gust44 and pEAK10 vectors, as well as Dr. Hiroaki Matsunami for providing HEK293T cells. We also wish to express our gratitude to Yumi Tsujiuchi, Taiki Okumura, and other keepers and veterinarians at the Japan Monkey Centre for taking care of the colobines, supporting behavioral assays, and collecting genetic samples, as well as Atsushi Yamanaka and the Center for Human Evolution Modeling Research at the Primate Research Institute, Kyoto University for taking care of the Japanese macaques. This study was funded by JSPS KAKENHI (#16H01338, #15H02421, #18H0400, and #19K21586 to HI; #16K18630 to TH; and #17K15203 to NSH); research grants from the Kobayashi International Scholarship Foundation (to HI); PMDSU nos. 1050/IT3.11/LT/2017 and 1486/IT3.11/PN/2018 and nos. 219.6/D3/PG/2019, 121/SP2H/LT/DRPM/2019 from the Ministry of Research, Technology and Higher Education to BS; Cooperative Research Programme of Kyoto University (Kyodo Riyo Theme nos. 2017-B-51, 2018-B-35, 2019-B-55 and Sandwich-like PMDSU No: 1056/D3.2/PG/2018 to LHPSP). This work was also supported by JSPS and DG-RSTHE under the Japan-Indonesia Research Cooperative Program and Core-to-Core Program.
Author information
Authors and Affiliations
Contributions
LHPSP conducted the experiments, wrote the first draft, and analyzed and interpreted the data. KAW conducted the experiments, analyzed and interpreted data, and revised the manuscript. NSH and TH conducted the experiments, analyzed the data, and revised the draft. AI analyzed the data and revised the draft. SN collected samples. BJ analyzed the data and revised the draft. BS designed the experiments, and revised the manuscript. HI designed the experiments, and wrote the final draft. All authors agree to be held accountable for the content in the manuscript and approve the final version.
Corresponding authors
Ethics declarations
Ethical approval
Colobine behavioral assays were conducted at the Japan Monkey Centre, Inuyama, Japan. The study was approved by the Research Ethics Committee of the Japan Monkey Centre in accordance with the Ethical Guidelines for Research of the institution (April 1, 2016) as Collaborative Research with the Japan Monkey Centre (#2018015). The Japanese macaque assay was conducted at the Primate Research Institute, Kyoto University. Both experiments were approved by the Animal Welfare and Animal Care Committee of Primate Research Institute, Kyoto University (#2018–201) and were conducted in compliance with the Guidelines for Care and Use of Nonhuman Primates of the Primate Research Institute, Kyoto University (Version 3, issued in 2010). These guidelines were prepared based on the provisions of the Guidelines for Proper Conduct of Animal Experiments (June 1, 2006; Science Council of Japan), Basic Policies for the Conduct of Animal Experiments in Research Institutions under the Jurisdiction of the Ministry of Health, Labour and Welfare (effective on June 1, 2006; Ministry of Health, Labour and Welfare), Fundamental Guidelines for Proper Conduct of Animal Experiment and Related Activities in Academic Research Institutions (Notice No. 71 of the Ministry of Education, Culture, Sports, Science and Technology dated June 1, 2006), and Standards Relating to the Care and Management of Laboratory Animals and Relief of Pain (Notice No. 88 of the Ministry of the Environment dated April 28, 2006).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
About this article
Cite this article
Purba, L.H.P.S., Widayati, K.A., Suzuki-Hashido, N. et al. Evolution of the bitter taste receptor TAS2R38 in colobines. Primates 61, 485–494 (2020). https://doi.org/10.1007/s10329-020-00799-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10329-020-00799-1