Abstract
Filamentous fungi produce a variety of proteases with significant biotechnological potential and show diverse substrate specificities. Proteolytic analysis of the industrial enzyme producer Trichoderma reesei has been sparse. Therefore, we determined the substrate specificity of T. reesei secretome and its main protease Trichodermapepsin (TrAsP) up to P1 position using FRETS-25Xaa-libraries. The role of TrAsP was analyzed using T. reesei QM9414 and the deletant QM∆trasp in Avicel. We observed higher activities of CMCase, Avicelase, and Xylanase in QM∆trasp compared to that of QM9414. Saccharification rate of cellulosic biomass also increased when using secretome of QM∆trasp but the effect was not significant due to the absence of difference in BGL activity compared to QM9414. Higher TrAsP was produced when monosaccharides were used as a carbon source compared to cellulase inducers such as Avicel and α-sophorose. These results elucidate the relationship between TrAsP and cellulase production in T. reesei and suggest a physiological role for TrAsP.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Proteases are a complex group of hydrolytic enzymes with diverse physiological roles and potential use in biotechnology. These enzymes account for nearly 60% of the whole enzyme market and are extremely important in the medical, pharmaceutical, leather, food, and biotechnology industries [32]. Filamentous fungi have been utilized as a potential source of proteases due to their ability to secrete a broad range of enzymes using low-cost substrates and their susceptibility to genetic manipulation [4]. A large number of fungal strains belonging to the genera Aspergillus, Penicillium, Rhizopus, Mucor, Humicola, Thermoascus, Thermomyces, etc., with different chemical and physical characteristics have been used to produce industrial proteases [37]. In industrial applications, the catalytic site of the protease (serine, cysteine, aspartyl, threonine, glutamic or metal) is responsible for its substrate specificity. Catalysis is also related to a particular substrate that determines its enzyme specificity. Filamentous fungi produce proteases with diverse specificity and this has been investigated in detail [10].
Trichoderma reesei is one of the most effective industrial producers of cellulolytic enzymes for biotechnological applications and its enzymes have been extensively studied at both biochemical and genetic levels. Even though its ability to produce extracellular proteases in T. reesei has been investigated in the past, its proteolytic enzyme system has not been well studied. In comparison, considerable amount of work has focused on the identification and utilization of protease genes in the cellulytic enzyme producer Aspergillus genera [1, 4, 34]. To understand and regulate the proteases in the secretome, it is necessary to understand the specificity of each protease that determines the enzyme–substrate interaction and cleavage bonds. However, there is a paucity of information regarding T. reesei proteases and their substrate specificities. Therefore, this study aimed at developing a profile of enzyme–substrate interaction in T. reesei QM9414 secretome.
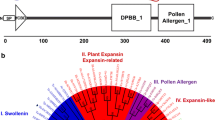
It has been reported that the interaction of cellulases and proteases in the T. reesei secretome results in proteolytic modifications of these enzymes [8]. The presence of multiple forms of cellobiohydrolases I and II in T. reesei was delineated using proteases [9] [16]. It has also been observed that proteases decrease heterologous protein expression in filamentous fungi [11, 29]. A large secretion of aspartic proteinases aspergillopepsin A and B has been recorded in A. awamori. The deletion of pep B genes results in an increase in thaumatin protein in this species [25]. The ultraviolet-mutant strain T. reesei A2221 that shows 94% reduction in acid protease activity also exhibited significant expression of the A. niger phytase gene [22]. However, the relationship between proteases and cellulases still remains unclear and poorly understood. This paper aims to develop a clearer view of how protease determines the fate of cellulases in the secretome in the model strain of T. reesei QM9414. To this end, we focused on the main protease in T. reesei genome the aspartic protease of the peptidase family A1 trichodermapepsin (TrAsP, MEROPS ID: A01.027) [28, 31]. The same protein was also called as pep1 in a previous report [17]. The precursor protein was recorded at a molecular weight of 40–42 kDa (JGI protein ID 74156) [8, 17] and the mature protein was recorded at 32 kDa (EC 3.4.23.18 and PDB accession code 3C9X) [6, 28]. The insights gained from this study will be helpful for optimizing cellulase production and for developing efficient strains. They will also provide an avenue for the study of other proteases in the T. reesei genome.
Materials and methods
Strain and media
T. reesei QM9414 strain and QM∆trasp were used in this study. Conidia from these strains were obtained from potato dextrose agar plate cultures. Submerged fermentation medium containing 0.14% (NH4)2SO4, 0.2% KH2PO4, 0.03% CaCl2·H2O, 0.03% MgSO4·7H2O, 0.05% yeast extract, 0.1% bacto polypeptone, 0.1% Tween 80, and 0.1% trace element solution (0.005 mg/L FeSO4, 0.002 mg/L MnSO4·H2O, 0.001 mg/L CuSO4·5H2O, 0.002 mg/L ZnSO4·7H2O, 0.001 mg/L (NH4)6Mo7O24·4H2O, 0.002 mg/L CoCl2·6H2O, and 0.001 mg/L NiCl2·6H2O) in a 50 mM Na-tartarate buffer (pH 4.0) was used. The medium was supplemented with 1% of galactose, mannose, xylose, arabinose, fructose, maltose, glucose, sucrose, galactomannan, xylo-oligosaccharide, β-1,3-glucan, birch wood xylan, glucomannan, pectin, beech wood xylan, cellobiose, oat-spelt xylan, glycerol, lactose, Avicel, starch, sorbose, CMC, chitin, lignin and xyloglucan as the carbon source. Spores (107 spores per 50 mL medium) were inoculated and incubated in a shaker at 28 °C and 220 rpm for 6 days. To analyze the response to changes in pH, T. reesei was grown on 1% Avicel medium adjusted to pH 3.0, 4.0, or 5.0. Spores (107 spores per 50 mL medium) were incubated at 28 °C and 220 rpm for 5 days. After cultivation, culture supernatants were obtained by centrifugation at 14,000 rpm. Samples of the culture supernatants were analyzed by 12.5% SDS-PAGE. The intensity of the bands in the gel was measured using ImageJ software (National Institutes of Health) [40]. Induction experiments were performed as previously described [19]. Carbon source inducers were 1% Avicel, 1% galactose, and 0.125% α-sophorose. Nitrogen source inducers were 1% bovine serum albumin (BSA), 1% bacto-peptone or 0.5% ammonium sulfate. Samples were taken at different time intervals. After induction, mycelia were harvested by filtration and rapidly frozen in liquid N2 for RNA extraction. For analyzing the saccharification rate, supernatants of 1% Avicel cultivation of QM9414 and QM∆trasp after 3 days were used.
Construction of QM∆trasp
For construction of the gene deletion plasmid, the fragments, upstream region (2.0 kbp), and downstream region (2.3 kbp) of trasp were amplified by polymerase chain reaction (PCR) using the genomic DNA of the standard strain QM9414 as a template and appropriate primers (Table S1). pUpyr4 plasmid was digested with ECORV for liberating pyr4 and pUC118 was digested with HindIII for generating a linear fragment. Deletion cassette containing plasmid was constructed using a pyr4 marker and pUC118 plasmid with Gibson Assembly Kit. trasp gene deleted cassette was prepared by HindIII enzyme excision of pUpyr4∆trasp. Prepared cassette was introduced into T. reesei QM∆pyr4 as described in [23] except that 10 mg/mL Yatalase (Takara Bio) was used for the preparation of protoplast. Transformants were selected by a uridine autotroph and purified through single conidia using MMRX and PDAX media. For the confirmation of QM∆trasp candidates, colony PCR was performed using the KOD FX Neo enzyme (TOYOBO). Homologous recombination and copy number of strains were confirmed by Southern hybridization. Independent transformants were analyzed after digestion of genomic DNA with KpnI and XbaI. After electrophoresis, the digested chromosomal DNA was transferred onto a Hybond N + membrane (GE Healthcare Bio Science, Waukesha, WI) and hybridized with an alkaline phosphatase-labeled probe of a 0.5 bp NcoI fragment of pyr4 gene.
Total RNA extraction and quantitative reverse transcription PCR (qRT-PCR)
Frozen samples were added to 750 μL TES buffer 10 mM Tris–HCl (pH 7.5), 10 mM EDTA and 0.5% SDS, and 750 μL acid-phenol preheated to 65 °C. Total RNA extraction was performed using a standard hot acid-phenol RNA extraction protocol. Samples were incubated at 65 °C for 15 min, followed by TRIzol LS (Invitrogen, USA) and chloroform extraction. Total RNA was purified by Illustra RNAspin (GE Health Care) according to the manufacturer’s instructions.
qRT-PCR was carried out in a Light Cycler 480 System (Roche Diagnostics). Amplification reactions were performed in a final volume of 20 μL using a Light Cycler 480 SYBR Green I Master kit (Roche Diagnostics) with 0.5 mM forward (Fw) primer, 0.5 mM reverse (Rv) primer and 2 μL of 100-fold diluted cDNA. The PCR primers used were cbh1 primers: Fw (5′-CTTGGCAACGAGTTCTCTT-3′) and Rv (5′-TGTTGGTGGGATACTTGCT-3′), egl1 primers: Fw (5′-CGGCTACAAAAGCTACTACG-3′) and Rv (5′-CTGGTACTTGCGGGTGAT-3′), xyn1 primers: Fw (5′-GGTTGGACGACTGGATCT-3′) and Rv (5′-GGTTGTCCTCCATGATGTAG-3′), xyn2 primers: Fw (5′-CCGTCAACTGGTCCAACT-3′) and Rv (5′-ACACGGAGAGGTAGCTGTT-3′), trasp primers: Fw (5′-GCTCAACCGCAACTCCATC-3′) and Rv (5′-GAAGACGACACCCTCCTGCT-3′), pac1 primers: Fw (5′-ACATCCGTGTTCACGTTC-3′), Rv (5′-GAGTCATCGGCATGAGTC-3′) and β-actin gene (act1) primers: Fw (5′-TCCATCATGAAGTGCGAC-3′) and Rv (5′-GTAGAAGGAGCAAGAGCAGTG-3′). Thermal cycling was conducted using the following conditions: 5 min at 95 °C followed by 45 cycles of 10 s at 95 °C, 10 s at 60 °C, and 10 s at 72 °C. All analyses were done independently in triplicate with a no-amplification control. The specificity of PCR amplification was documented by a melting curve analysis and act1 (Protein ID 44504) was used as a reference for quantification.
Mass spectrometry analysis
The band corresponding to a 40 kDa protein was digested in gel with trypsin. Peptides were recovered in the supernatant of the digest, dried by evaporation, reconstituted in 20 μL of H2O/acetonitrile 97:3 with 0.1% formic acid and analyzed by Matrix Assisted Laser Desorption/Ionization-Time of Flight (MALDI-TOF) mass spectrometry using Spectrometry Axima-CFR plus. From the raw files, peptide mass fingerprinting (PMF) analysis was searched using Mascot (Matrix Science, London, UK; version 2.3.0) against the JGI T. reesei QM6a genome database (http://genome.jgi.doe.gov/Trire2/) selected for fungi, trypsin digestion, and one missed cleavage. Mascot was searched with a fragmented ion mass tolerance of 0.50 Da and a parent ion tolerance of 10.0 ppm.
Purification of TrAsP
Extracellular TrAsP protease was purified from T. reesei QM9414 cultivated in 1% galactose media after 72 h of growth. Culture supernatants were collected by filtration on a mira cloth (Calbiochem). Supernatants were then precipitated with ammonium sulfate (80% saturation). Precipitates were collected by centrifugation at 14,000 rpm for 30 min and dissolved in 20 mM sodium acetate buffer (pH 4.0). Solutions were concentrated and buffer-exchanged using Vivaspin 20 (10 kDa) (GE Healthcare) to remove the ammonium sulfate, and then the solutions were applied to a HiPrep Q XL 16/10 (GE Healthcare) column, previously equilibrated with the same buffer. The fraction containing TrAsP was identified.
Measurement of proteolytic activity
Total protease activity was measured by Azocasein assay. A solution of Azocasein was prepared at 5 mg/mL in a 50 mM sodium acetate buffer (pH 5). Azocasein (400 μL) was then mixed with 100 μL of the samples and incubated at 50 °C for 90 min. The reaction was then stopped by the addition of 150 μL of 12% trichloroacetic acid (TCA) and incubated at room temperature for 30 min. These samples were then centrifuged at 8000 × g for 3 min. Subsequently, 150 μL of supernatant was taken and mixed with 150 μL of 1 M NaOH and the absorbance of the solution was measured as 436 nm. One unit of protease activity at OD 436 is equal to 1.0 per mL. Activity of the purified protease was measured by hydrolysis of the fluorogenic peptide MOCAc-Ala-Pro-Ala-Lys-Phe-Phe-Arg-Leu-Lys(Dnp)-NH2. The optimal pH for protease activity was determined by incubating the purified enzyme at 50 °C for 30 min in 0.5 M citrate and 0.5 M NaOH buffer (pH 2–6). To determine the optimal temperature, the enzyme was incubated in 0.5 M citrate and 0.5 M NaOH buffer (pH 5) for 30 min from 20–80 °C. Fluorescence was measured in a microplate reader Powerscan HT (DS Pharma Biomedical) at Ex = 328 nm and Em = 393 nm. Substrate specificity and kinetic parameters of purified TrAsP were measured using the Fluorescence-Quenching Substrate Library FRETS-25 Series, Peptide Institute, Inc [39]. Kinetic parameters were determined by fitting the experimental data to the Michaelis–Menten equation using Excel Solver (Microsoft) by nonlinear least-square fitting with different concentrations (0.391–25.0 μM) of FRETS-25 as substrates. The enzyme reaction was performed in 50 mM sodium acetate buffer pH 5.0 at 298 K for 5 min and standard deviations were calculated from three independent experiments. The florescence intensity of the released 2-(N-methylamino) benzoyl (Nma) group was measured with excitation at 340 nm and emission at 440 using Infinite 200 PRO microplate reader (Tecan).
Biomass saccharification
Cellulosic biomass (Erianthus) was subjected to alkaline treatment as previously reported [14]. Saccharification by the cellulases was performed in a 20 ml plastic bottle containing 5% dry biomass (w/v) in 100 mM sodium acetate buffer (pH 5.0), with enzyme loading at 3 mg of protein per gram of dry biomass. The reaction was performed at 50 °C with shaking at 150 rpm for 72 h. The supernatants were boiled for 5 min to inactivate the enzymes, and the produced sugar was measured by the 3, 5-dinitrosalicylic acid (DNS) method. The biomass saccharification ratio (%) was calculated as the ratio of the sugar content to cellulose and hemicellulose in the dry mass.
Results
Specificity of proteases secreted by of T. reesei
To determine the activity of proteases in T. reesei secretome, supernatants of T. reesei in 4- and 6-day cultivations were used (Fig. 1a). Azocasein activity, which represents total proteases in the culture was shown to be 1.1 ± 0.03 U/mL in 4 days and 1.4 ± 0.01 U/mL in 6 days. Proteinase A activity in the culture measured using the specific synthetic substrate, MOCAc-Ala-Pro-Ala-Lys-Phe-Phe-Arg-Leu-Lys(Dnp)-NH2, was 0. 76 ± 0.21 U/mL in 4 days which increased to 1.16 ± 0.07 U/mL in 6 days. Substrate specificity of protease secretion by T. reesei was measured by FRESTS-25Xaa substrate containing a highly fluorescent 2-(N-methylamino) benzoyl (Nma) group linked to the side chain of the amino-terminal D-2,3-diamino propionic acid (D-A2Pr) residue, which is efficiently quenched by 2-4-dinitrophenyl (Dnp) group linked to the amino acid Lys (Fig. 1b). Xaa represents the fixed position for any of the 20 natural amino acids excluding Cys. Yaa and Zaa position represents a mixture of five amino acid residues (P, Y, K, I, and D) and (F, A, V, E, and R), respectively. FRETS-25Xaa-libraries were prepared by 25 combinations of each Xaa series and provided a total of 475 peptide substrates in 19 separate sections. Primary screening of selection of favored Xaa (P1 position) was carried out using T. reesei 4-day culture supernatants. The results indicated the total proteases in the secretome with preferred residues Phe, Ala, Tyr, Val, Leu, Ile, Met and Glu in P1 position in descending order.
Substrate specificity of the secretome of T. reesei.a Azocasein activity and proteinase A activity using the fluorogenic peptide MOCAc-Ala-Pro-Ala-Lys-Phe-Phe-Arg-Leu-Lys (Dnp)-NH2 in 1% Avicel cultivation of T. reesei QM9414 after 4 and 6 days. b FRETS-25Xaa substrate with its subsites of P1, P2, P3 P1′, P2′, P3′. c Identification of substrate specificity in T. reesei secretome by FRETS-Series. Each bar shows the mean of three independent experiments with SD indicated by a vertical line
To determine the effect of the main protease on substrate specificity, a deletion strain of trasp was constructed (Fig. S1) and its protease activity was measured compared to wild type using T. reesei QM∆trasp cultivation supernatant. Compared to that of QM9414 strain, Azocasein activity was reduced to 0.96 ± 0.02 U/mL and MOCA peptidase activity was reduced to 0.56 ± 0.23 U/mL in 4 days (Fig. 2a). The secretome preferred the residues Ala, Leu, Val, Glu, Met, Pro, Lys, and Ile in the P1 position (Fig. 2b). It looked like the preference to substrate shifted from aromatic (Phe) to aliphatic (Ala) in the deletion strain compared to QM9414. However, it was difficult to get a clear understanding of TrAsP substrate specificity by increasing or decreasing some of the other proteases in trasp deletant stain. Therefore, we decided to use purified TrAsP for analyzing preference of P1 position by measuring specific activity (Fig. 3). Results indicated that TrAsP had a higher preference for positively charged (Arg). Polar uncharged amino acids such as Asn, Gln, Ser, and Thr and negatively charged Asp were indicated a lower preference. TrAsP showed significant low preference for the smallest amino acid Gly and the largest amino acid Trp and a special amino acid Pro with imino acid group (dihedral angle of − 60°), suggesting the protein was restricted by size. Further kinetic studies were carried out using purified TrAsP (Table S3). With the agreement of Fig. 3, the highest kcat value was obtained in Arg (1.22 ± 0.27 s−1). According to kcat/Km of the protein, higher values were shown for positively charged amino acids Lys, Arg, and His as 145 ± 12 mM−1 s−1, 130 ± 12 mM−1 s−1 and 108 ± 11 mM−1 s−1, respectively. Based on the specific activity and kinetic analysis, we concluded that positively charged amino acids may be more preferable substrates for TrAsP.
Substrate specificity of the secretome of T. reesei QM∆trasp.a Azocasein activity and proteinase A activity using fluorogenic peptide MOCAc-Ala-Pro-Ala-Lys-Phe-Phe-Arg-Leu-Lys (Dnp)-NH2 in 1% Avicel cultivation of T. reesei QM∆trasp after 4 and 6 days. b Identification of substrate specificity in T. reesei secretome by FRETS-Series. Each bar shows the mean of three independent experiments with SD indicated by a vertical line
Role of TrAsP in T. reesei culture and saccharification condition
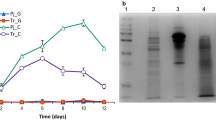
T. reesei secretome is naturally an enzyme cocktail with synergetic activity for biomass degradation. To determine the role of trasp in cellulase inducing media, QM9414 and QM∆trasp were cultivated in 1% Avicel. Supernatants were collected after 2, 4, and 6 days and cellulase activity was measured (Fig. 4a–d). A marginal increase was observed in CMCase activity of QM∆trasp compared to that of QM9414 and activity increased gradually from 2 to 6 days of cultivation (Fig. 4a). There was a significant increase in QM∆trasp in Avicelase activity compared to that of QM9414, but there was no significant difference between 4 and 6 days (Fig. 4b). In xylanase activity analysis, higher xylanase activity was observed in 4 days but no significant change was seen between the strains in 6 days of cultivation (Fig. 4c). It seemed, therefore, that there was no effect in cellobiase activity in cultivation due to deletion of trasp gene (Fig. 4d).
Analysis of cellulase activity in T. reesei QM9414 and QM∆trasp after cultivation in 1% Avicel. Samples were collected after 2, 4, and 6 days a CMCase activity b Avicelase activity c Xylanase activity d Cellobiase activity. Each bar shows the mean of three independent experiments with SD indicated by a vertical line
NaOH–pretreated Erianthus was used for analyzing the saccharifying activity of cellulases of QM∆trasp due to its higher cellulose and hemicellulose composition in pretreated biomass. The cellulose and hemicellulose composition after pretreatment were 50.1% and 25.2%, respectively, and was similar to that reported previously [14]. Glucose and xylose yields were slightly higher for QM∆trasp compared to that of QM9414 (Table 1). We did not observe a significant increase in saccharification rate, perhaps because the deletion of trasp did not have an effect on the cellobiase activity. BGL activity is a critical point for achieving a higher saccharification rate. However, our results show that there is a marginal effect of trasp in altering the saccharification rate. It also predicts that if the strain is genetically modified to increase cellobiase activity, the effect of trasp may be significantly higher.
TrAsP production condition in T. reesei
In the enzyme production or purification process, it is vital to know the production condition of each of the enzymes in the secretome. In addition, T. reesei has an environmental factor-responsive enzyme secretion mechanism [2] and to develop an optimal enzymatic cocktail, it is important to understand the natural regulation of the enzyme profile as a response to environmental factors. Therefore, TrAsP production condition was analyzed based on carbon, nitrogen, and pH values. The extracellular protein expression pattern of T. reesei is regulated by the carbon source in the cultivation medium. To analyze the extracellular protein profile of T. reesei, SDS-PAGE was carried out using supernatants of monosaccharides, disaccharides, oligosaccharides, polysaccharides, and other carbon sources after submerged cultivation (Fig. 5a). The different protein expression patterns were compared with Avicel. The expression patterns during growth on lactose, xylo-oligosaccharide, β-1,3-glucan and xyloglucan were similar to that of Avicel. A protein that resolved at approximately 40 kDa and appeared in almost all the samples was identified as an unknown protein and chosen for further analysis. To investigate the 40 kDa protein further, we digested the band with trypsin, and analyzed the extracted peptides by MALDI-TOF and PMF analysis. The analysis indicated that the peptides were derived from a 40 kDa aspartic protease known as TrAsP [28]. The intensity of this 40 kDa protein band was determined by imageJ analysis (Fig. 5b). TrAsP was highly expressed in the T. reesei supernatants from cultures grown on monosaccharides including arabinose, glucose, mannose, galactose, fructose, and xylose. Interestingly, expression was lower during growth on well-known inducers of cellulase including Avicel and L-sorbose.
SDS-PAGE analysis of culture supernatants of T. reesei. T. reesei QM9414 was cultivated in medium containing various carbon sources for 6 days. a Supernatant (15 µl) was subjected to 12.5% SDS-PAGE. Arrow indicates an approximate 40 kDa band. b Bands on SDS-PAGE were quantified using ImageJ software
To analyze trasp gene expression as a function of T. reesei growth on different carbon sources, T. reesei was induced in α-sophorose, Avicel, galactose and the mixed media of α-sophorose and galactose. The expression of trasp was analyzed compared to that of cellulase and hemicellulase genes and to the particular carbon source. To avoid the effects caused by differential germination on different substrates, the experiments were conducted with glucose-grown mycelia inoculated in the culture. T. reesei produces cellulase when grown on cellulose, α-sophorose, L-sorbose, and lactose [15, 20, 21, 38]. In agreement with the previous reports, expression of cbh1 and egl1 was found to be significantly higher in T. reesei induced on crystalline cellulose Avicel or α-sophorose than on galactose (Fig. 6). Minimal expression of xyn1 was detected with induction on Avicel or α-sophorose. However, compared to α-sophorose, Avicel showed significant trasp expression during late induction (Fig. 6b). During induction on galactose, trasp was highly expressed after 8 and 16 h, while the expression levels of glycoside hydrolase-encoding genes were quite low. The expression of trasp from T. reesei cultivated on galactose (at 16 h) was 106-fold higher than on Avicel and 516-fold higher than on α-sophorose (Fig. 6c). In addition, the expression of trasp was very low in samples taken at an early induction time during T. reesei induction on Avicel or α-sophorose. With the addition of 0.125% α-sophorose to 1% galactose induction medium, trasp expression was significantly reduced and showed lower cbh1 expression (Fig. 6d). It suggested that there is a different regulatory system for both cellulases and proteases when T. reesei is induced by a mixed carbon source.
Transcriptional analysis of carbon induction of genes encoding TrAsP and glycoside hydrolases, cbh1, egl1, and xyn1. T. reesei QM9414 was precultured on glucose and transferred to a medium containing a 0.125% (w/v) α-sophorose b 1% (w/v) Avicel c 1% (w/v) galactose d and mix of 0.125% (w/v) α-sophorose and 1% (w/v) galactose. All values were normalized to β-actin expression under the same conditions. Each bar shows the mean of three independent experiments with SD indicated by a vertical line
In addition to carbon sources, nitrogen sources play an important role in regulating proteases. Therefore, it is vital to understand trasp expression patterns when using different nitrogen sources. To accomplish this, T. reesei was induced with 1% BSA, 1% peptone, or 0.5% ammonium sulfate. cbh1, egl1, and xyn2 were found to be expressed at extremely low levels under these conditions (data not shown). In contrast, T. reesei trasp expression was induced by BSA and peptone. While the expression level of trasp on BSA was higher than on peptone, no induction was observed when T. reesei was induced with ammonium sulfate (Fig. 7a). The expression of trasp was also measured in T. reesei grown on a combination of 1% galactose and all the individual nitrogen sources tested (Fig. 7b). We found that the expression of trasp was much lower upon addition of 1% galactose as compared to expression with the individual nitrogen sources alone (Fig. 7b).
Transcriptional analysis of nitrogen-induction genes encoding TrAsP. aT. reesei QM9414 was precultured on glucose and transferred to a medium containing 1% (w/v) peptone, 1% (w/v) BSA, 0.5% (w/v) ammonium sulfate and incubated for 2, 4, 8, and 16 h. b T. reesei QM9414 was precultured on glucose and transferred to a medium containing 1% (w/v) peptone + 1% (w/v) galactose, 1% (w/v) BSA + 1% (w/v) galactose, 0.5% (w/v) ammonium sulfate + 1% (w/v) galactose and incubated for 2, 4, 8, and 16 h. All values were normalized to β-actin expression under same conditions. Each bar shows the mean of three independent experiments with SD indicated by a vertical line
When determining T. reesei expression of trasp following galactose induction, the effect of pH was needed to be analyzed. To this end, RNA extracted from T. reesei grown on galactose medium adjusted to pH 3, 4, or 5 was analyzed by qRT-PCR (Fig. S2). These pH values were selected because TrAsP is an acidic protease. The relative expression levels of trasp were high at pH 3 and 4, with a 16.5-fold and 21-fold higher expression, respectively, as compared to the very low trasp expression at pH 5 (after 12 h). This result indicates that trasp expression is more favorable in acidic conditions in a galactose medium.
In conclusion, monosaccharides especially galactose and peptone, and acidic pH conditions are preferred for optimal TrAsP expression. However, TrAsP production was low in mixed culture condition. Therefore, we suggest that TrAsP has its own regulatory mechanism and it needs to be elucidated using T. reesei as a model organism.
Discussion
The analysis of fungal proteases can provide information on various aspects, such as biochemical characteristics, specificity, and subsite cooperativity. This information is required to determine the optimal substrate and reaction conditions that will greatly improve the biotechnological applications of these proteases. While specificity analysis of some proteases secreted by filamentous fungi has been carried out [10], there is little information on the protease of Trichoderma reesei. In this study, first we measured the substrate preference of T. reesei secretome, which contains a mixture of proteases. It helped to predict the possibility of protein degradation in the medium and to understand how it affected specificity when genes were deleted. The substrate specificity of P1 position of T. reesei secretome was determined as Phe, followed by Ala and Tyr in the cellulase-inducing condition (Fig. 1c). We observed some differences in substrate preference in secretome of QM∆trasp (Fig. 2b) compared to that of QM9414. This result could be due to deletion of gene affecting the composition and by the other proteases in secretion. Therefore, we used the purified protein of TrAsP to measure specific activity and kinetic parameters. We could determine that basic amino acids, Lysine and Arginine were more preferable substrates of the main protease TrAsP (Fig. 3 and Table S3). Similar results were reported for an aspartic peptidase from filamentous fungi Rhizomucor miehei [36] and Phanerochaete chrysosporiumn [35]. Further, it was recorded that the Lys-containing substrates were hydrolysed because of the conserved aspartic acid residue in the flap of crystal structure (Asp 77 in TrAsP) [12, 28] and this fact also explained the preference of basic amino acids in the P1 position. Additionally, we found that P1 position was restricted by size. For supporting our results, da Silva and co-authors [35] were reported little or no hydrolysis when nonpolar residues Gly, Pro and Trp were at not only P1 position but also other subsites. As this study is only the primary screening, we need to elaborate this further to find other subsites. However, this is the first study that focused on elucidating amino acid substrate specificity of a protease in T. reesei. Recent enzyme specificity studies have focused on developing a computational design of remodeling [18] and on modifying biochemical properties using recombinant DNA techniques [1]. These advances can be extended to studying the proteases of filamentous fungi considering their biotechnological potential and the need for improvement in productivity.
In filamentous fungi, the enzymes in the secretion exert a synergetic activity contributing to their function and survival. The stability of cellulases and the amount of secreted cellulases are crucial in industrial cellulase producers. In this study, we were able to determine that there is an impact of TrAsP on cellulases in the medium under industrial saccharification conditions (Table 1). We also observed a stability of cellulases in SDS-PAGE after the enzymes were produced by QM∆trasp in Avicel incubated at 50 °C (Fig. S3). The results of this study also suggest that deletion of trasp also supports the stability of cellulases under cellulase-inducing conditions (Fig. 4) and this will be helpful for commercial applications. The absence of an effect of TrAsP on BGL activity may be due to the low increase in saccharification rate. Alternatively, when the main protease was deleted, other proteases may have compensated for the loss as its function is critical for the survival of T. reesei. We believe that our results are significant and provide a basis for improving the saccharification rate and for increasing cellulase and xylanase stability in these genetically improved strains.
In this work, we analyzed the analysis of TrAsP production mechanism on carbon, nitrogen, and pH conditions. These results are important for predicting cellulase stability under similar conditions. TrAsP was expressed when T. reesei was grown on various carbon sources (Fig. 5). In particular, trasp was highly expressed on a monosaccharide-containing medium except sorbose and trasp expression was low when cellulase inducers were present. Under these conditions, we were able to identify galactose as the carbon source that best induced trasp expression. In contrast, T. reesei induced on Avicel medium led to minimal trasp expression. With respect to protease activity, cultivation on Avicel produced proteinase A activity, which may indicate that many other proteinase A isoforms exist in the culture medium. When analyzing the T. reesei genome, we found 12 extracellular peptidase A pepsins besides TrAsP, identified as JGI protein ID: 53961, 68662, 81004, 121306, 79807, 105781, 111818, 76887, 77579, 119876, 122076, and 108686. The expression of proteases in response to cellulosic substrates including Avicel has been demonstrated in T. reesei RUT C-30 [30]. Peptidase A proteins were found in late cultivations on Avicel, similar to those observed in our results.
This is the first study highlighting the effect of carbon sources on the production of TrAsP, as other reports have mainly focused on the effects of nitrogen and protein sources. It is worth noting that trasp expression was only induced by galactose after a lengthy period of induction (Fig. 6). In T. reesei, galactose is reported to induce cellulases gene expression at low growth rates [13]. This fact also supports the delay in TrAsP expression when T. reesei was grown on galactose medium, as TrAsP is not well expressed when cellulase inducers are present in the medium. On the other hand, we speculated that the true inducer of trasp expression might be a galactose metabolite. A recent study on the optimization of protease production of some filamentous fungi has recorded that lactose elicited the highest protease production among the carbon sources tested [27]. Several studies have been performed to solve the mechanism of cellulase expression in lactose medium that has been extracellularly hydrolyzed by ß-galactosidase to glucose and galactose, and found that neither of those led to cellulase expression [24, 33]. In the metabolism pathway, galactose can either be phosphorylated and enter into the Leloir pathway or be reduced to galactitol. Further investigations will be necessary to determine the true inducer of trasp expression following galactose induction in T. reesei.
Many fungi produce proteases in the presence of a nitrogen source. In T. reesei, TrAsP expression was also induced by organic nitrogen but not by ammonium sulfate (Fig. 7). This result concurs with the previous studies showing that peptone strongly stimulated protease formation in T. reesei [8, 30]. However, trasp expression in response to the combination of galactose and organic nitrogen was lower than to galactose or organic nitrogen alone. These results suggest that the mechanism of induction differs when inducers are mixed, and they may compete given that they act via different regulatory pathways. Interestingly, sequence analysis of the promoter region of TrAsP found the existence of putative binding sites (GATA) of the nitrogen metabolite regulator AreA (data not shown), indicating that trasp expression influences AreA. Under nitrogen-limiting conditions, AreA activates the transcription of many structural gene encoding enzymes for nitrogen source catabolism. However, there are very few reports on the involvement of AreA in both cellulase and protease production in filamentous fungi.
A number of proteases are induced in different fungal species in response to pH changes [5, 7]. In A. niger, pH and the availability of organic nitrogen influence protease production [3, 23]. Indeed, the induction of T. reesei mycelia at pH 3 or 4 resulted in higher trasp expression. T. reesei also produces an acidic protease at pH 3 or 4 during late growth stages [8, 23]. In contrast, trasp expression was lower at pH 5. These results indicate that trasp expression is regulated by pH of the galactose medium. Interestingly, we found that purified TrAsP has higher activity at pH 5 (Fig. S4). A similar phenomenon was observed in T. harzanium with its aspartic protease, papA gene (accession number: AJ276388) (85.9% identity with TrAsP) showing a maximal expression at pH 3.7 and maximal activity at pH 4.5 [26]. The Pac1 transcription factor responds to extracellular pH and activates genes that are expressed at an alkaline pH and represses genes that are expressed under acidic growth conditions. Its putative binding sequence was found in the promoter region of TrAsP revealing that Pac1 may regulate trasp. Pac1 expression at pH 3 and 4 was extremely low, however, it was much higher at pH 5 (Fig. S2), indicating that Pac1 could repress the expression of trasp.
In conclusion, we found that trasp was induced by monosaccharides, especially, galactose, and organic nitrogen, but not by the typical inducers of cellulase. Expression of trasp was regulated by pH and increased during the late growth stage. These results indicate a novel form of regulation unrelated to known mechanisms governing cellulase gene expression. This knowledge improved our understanding of T. reesei behavior in cellulase-producing media. Therefore, considering the physiological role of TrAsP, we propose an enzyme secretion regulatory model for T. reesei QM9414 with enzymes helping in the uptake of nutrients and for survival in a cellulase-inducing culture medium. Previous studies have indicated beyond doubt that T. reesei QM9414 produces large number of cellulases in cellulase-inducing media and we have also detected TrAsP expression in very small amounts under that condition. However, in later stages of cultivation, degradable products of cellulase-inducing substrates are formed due to secreted cellulases, and these monosaccharides induce TrAsP expression. We surmise that cellulases are modified or degraded by the secreted TrAsP in the media and allow T. reesei to take up nutrients essential for survival in later stages of cultivation thus exerting a physiological role. However, for this mechanism to work, QM∆trasp strain should not show growth in later stages of cultivation, but we did not observe such a lack of growth. It is possible that some other proteases take over the role of TrAsP when nutrients are depleted in the culture medium to enhance chances of survival. Furthermore, many aspects of TrAsP regulation in different carbon sources are still unknown. With respect to trasp expression, a detailed analysis of transcription factors possessing binding sites in the promoter region and transporters involved in the nitrogen assimilation pathway and sugar uptake, need to be elucidated to gain a clear view of protease regulation in comparison to cellulase regulation.
References
Ahmed ME (2018) Extraction and purification of protease from Aspergillus niger isolation. Pharm Pharmacol Int J 6:96–99. https://doi.org/10.15406/ppij.2018.06.00162
Amore A, Giacobbe S, Faraco V (2013) Regulation of cellulase and hemicellulase gene expression in fungi. Curr Genom 14:230–249. https://doi.org/10.2174/1389202911314040002
Braaksma M, Smilde AK, van der Werf MJ, Punt PJ (2009) The effect of environmental conditions on extracellular protease activity in controlled fermentations of Aspergillus niger. Microbiology 155:3430–3439. https://doi.org/10.1099/mic.0.031062-0
de Castro RJS, Sato HH (2014) Protease from Aspergillus oryzae: biochemical characterization and application as a potential biocatalyst for production of protein hydrolysates with antioxidant activities. J Food Process 2014:1–11. https://doi.org/10.1155/2014/372352
Delgado-Jarana J, Rincón AM, Benítez T (2002) Aspartyl protease from Trichoderma harzianum CECT 2413: cloning and characterization. Microbiology 148:1305–1315. https://doi.org/10.1099/00221287-148-5-1305
Eneyskaya EV, Kulminskaya AA, Shabalin KA et al (1999) Acid protease from Trichoderma reesei: limited proteolysis of fungal carbohydrases. Appl Microbiol Biotechnol 52:226–231. https://doi.org/10.1007/s002530051513
Gente S, Billon-Grand G, Poussereau N, Févre M (2001) Ambient alkaline pH prevents maturation but not synthesis of ASPA, the aspartyl protease from Penicillium roqueforti. Curr Genet 38:323–328
Haab D, Hagspiel K, Szakmary K, Kubicek CP (1990) Formation of the extracellular proteases from Trichoderma reesei QM 9414 involved in cellulase degradation. J Biotechnol 16:187–198. https://doi.org/10.1016/0168-1656(90)90035-A
Hagspiel K, Haab D, Kubicek C (1989) Protease activity and proteolytic modification of cellulases from a Trichoderma reesei QM 9414 selectant. Appl Microbiol Biotechnol 32:61–67. https://doi.org/10.1007/BF00164824
Hamin Neto YAA, da Rosa Garzon NG, Pedezzi R, Cabral H (2017) Specificity of peptidases secreted by filamentous fungi. Bioengineered 5979:1–8. https://doi.org/10.1080/21655979.2017.1373531
Van Den Hombergh JPTW, Van De Vondervoort PJI, Fraissinet-Tachet L, Visser J (1997) Aspergillus as a host for heterologous protein production: the problem of proteases. Trends Biotechnol 15:256–263. https://doi.org/10.1016/S0167-7799(97)01020-2
Kamitori S, Ohtaki A, Ino H, Takeuchi M (2003) Crystal structures of Aspergillus oryzae aspartic proteinase and its complex with an inhibitor pepstatin at 1.9 Å resolution. J Mol Biol 326:1503–1511. https://doi.org/10.1016/S0022-2836(03)00078-0
Karaffa L, Fekete E, Gamauf C et al (2006) D-Galactose induces cellulase gene expression in Hypocrea jecorina at low growth rates. Microbiology 152:1507–1514. https://doi.org/10.1099/mic.0.28719-0
Kawai T, Nakazawa H, Ida N et al (2012) Analysis of the saccharification capability of high-functional cellulase JN11 for various pretreated biomasses through a comparison with commercially available counterparts. J Ind Microbiol Biotechnol 39:1741–1749. https://doi.org/10.1007/s10295-012-1195-9
Kawamori M, Morikawa Y, Shinsha Y et al (1985) Preparation of mutants resistant to catabolite repression of Trichoderma reesei. Agric Biol Chem 49:2875–2879. https://doi.org/10.1080/00021369.1985.10867203
Kubicek-Pranz EM, Gsur A, Hayn M, Kubicek CP (1991) Characterization of commercial Trichoderma reesei cellulase preparations by denaturing electrophoresis (SDS-PAGE) and immunostaining using monoclonal antibodies. Biotechnol Appl Biochem 14:317–323
Landowski CP, Huuskonen A, Wahl R et al (2015) Enabling low cost biopharmaceuticals: a systematic approach to delete proteases from a well-known protein production host Trichoderma reesei. PLoS One 10:1–28. https://doi.org/10.1371/journal.pone.0134723
Li Q, Yi L, Marek P, Iverson BL (2013) Commercial proteases: present and future. FEBS Lett 587:1155–1163. https://doi.org/10.1016/j.febslet.2012.12.019
Mach-Aigner AR, Pucher ME, Steiger MG et al (2008) Transcriptional regulation of xyr1, encoding the main regulator of the xylanolytic and cellulolytic enzyme system in Hypocrea jecorina. Appl Env Microbiol 74:6554–6562. https://doi.org/10.1128/AEM.01143-08
Mandels M, Reese ET (1957) Induction of cellulase in Trichoderma viride as influenced by carbon sources and metals. J Bacteriol 73:269–278. https://doi.org/10.1002/path.1700730133
Mandels M, Parrish FW, Reese ET (1962) Sophorose as an inducer of cellulase in Trichoderma viride. J Bacteriol 83:400–408
Mäntylä M, Paloheimo MSP (1998) Industrial mutants and recombinant strains of Trichoderma reesei. In: Christian P, Harman GE, Ondik KL, Kubicek CP (eds) Trichoderma and Gliocladium. Taylor and Francis, London, pp 291–309
Margolles-Clark E, Hayes CK, Harman GE, Penttilä M (1996) Improved production of Trichoderma harzianum endochitinase by expression in Trichoderma reesei. Appl Env Microbiol 62:2145–2151
Mojzita D, Herold S, Metz B et al (2012) l- xylo -3-Hexulose reductase is the missing link in the oxidoreductive pathway for d-galactose catabolism in filamentous fungi. J Biol Chem 287:26010–26018. https://doi.org/10.1074/jbc.M112.372755
Moralejo FJ, Cardoza RE, Gutierrez S et al (2002) Silencing of the aspergillopepsin B (pepB) gene of Aspergillus awamori by antisense RNA expression or protease removal by gene disruption results in a large increase in thaumatin production. Appl Environ Microbiol 68:3550–3559. https://doi.org/10.1128/AEM.68.7.3550-3559.2002
Moreno-Mateos MA, Delgado-Jarana J, Codón AC, Benítez T (2007) pH and Pac1 control development and antifungal activity in Trichoderma harzianum. Fungal Genet Biol 44:1355–1367. https://doi.org/10.1016/j.fgb.2007.07.012
Muthukrishnan S, Mukilarasi K (2016) Industrial important protease screening and optimization from micro-fungal isolates of Ayyanar Falls Forest Samples, Rajapalalyam. World Appl Sci J 34:343–347. https://doi.org/10.5829/idosi.wasj.2016.34.3.23113
Nascimento AS, Krauchenco S, Golubev AM et al (2008) Statistical coupling analysis of aspartic proteinases based on crystal structures of the Trichoderma reesei enzyme and its complex with pepstatin A. J Mol Biol 382:763–778. https://doi.org/10.1016/j.jmb.2008.07.043
Nevalainen KMH, Te’o VSJ, Bergquist PL (2005) Heterologous protein expression in filamentous fungi. Trends Biotechnol 23:468–474. https://doi.org/10.1016/j.tibtech.2005.06.002
Peciulyte A, Anasontzis GE, Karlström K et al (2014) Morphology and enzyme production of Trichoderma reesei Rut C-30 are affected by the physical and structural characteristics of cellulosic substrates. Fungal Genet Biol 72:64–72. https://doi.org/10.1016/j.fgb.2014.07.011
Pitts JE, Crawford MD, Nugent PG et al (1995) The three-dimensional X-ray crystal structure of the aspartic proteinase native to Trichoderma reesei complexed with a renin inhibitor CP-80794. Springer, Boston, pp 543–547
Rao MB, Tanksale AM, Ghatge MS, Deshpande VV (1998) Molecular and biotechnological aspects of microbial proteases. Microbiol Mol Biol Rev 62:597–635
Rauscher R, Wurleitner E, Wacenovsky C et al (2006) Transcriptional regulation of xyn1, encoding xylanase I, in Hypocrea jecorina. Eukaryot Cell 5:447–456. https://doi.org/10.1128/EC.5.3.447-456.2006
Siala R, Sellami-Kamoun A, Hajji M et al (2009) Extracellular acid protease from Aspergillus niger I1: purification and characterization. Afr J Biotechnol 8:4582–4589. https://doi.org/10.5897/AJB09.063
da Silva RR, de Oliveira LCG, Juliano MA et al (2017) Biochemical and milk-clotting properties and mapping of catalytic subsites of an extracellular aspartic peptidase from basidiomycete fungus Phanerochaete chrysosporium. Food Chem 225:45–54. https://doi.org/10.1016/j.foodchem.2017.01.009
da Silva RR, Souto TB, de Oliveira TB et al (2016) Evaluation of the catalytic specificity, biochemical properties, and milk clotting abilities of an aspartic peptidase from Rhizomucor miehei. J Ind Microbiol Biotechnol 43:1059–1069. https://doi.org/10.1007/s10295-016-1780-4
de Souza PM, de Assis Bittencourt ML, Caprara CC et al (2015) A biotechnology perspective of fungal proteases. Braz J Microbiol 46:337–346. https://doi.org/10.1590/S1517-838246220140359
Sternberg D, Mandels GR (1979) Induction of cellulolytic enzymes in Trichoderma reesei by sophorose. J Bacteriol 139:761–769
Tanskul S, Oda K, Oyama H et al (2003) Substrate specificity of alkaline serine proteinase isolated from photosynthetic bacterium, Rubrivivax gelatinosus KDDS1. Biochem Biophys Res Commun 309:547–551
Yu L-X, Gray BN, Rutzke CJ et al (2007) Expression of thermostable microbial cellulases in the chloroplasts of nicotine-free tobacco. J Biotechnol 131:362–369. https://doi.org/10.1016/J.JBIOTEC.2007.07.942
Acknowledgements
This work was partially supported by a grant from the New Energy and Industrial Technology Development Organization (NEDO) Project (P16009). Furthermore, our sincere thanks go to Nobuyuki Homma; Nagaoka University of Technology, for contributing the substrate specificity and kinetic analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Daranagama, N.D., Shioya, K., Yuki, M. et al. Proteolytic analysis of Trichoderma reesei in celluase-inducing condition reveals a role for trichodermapepsin (TrAsP) in cellulase production. J Ind Microbiol Biotechnol 46, 831–842 (2019). https://doi.org/10.1007/s10295-019-02155-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-019-02155-9