Abstract
Bacillus sp. strains as attractive hosts for the production of heterologous secretory proteins usually play important roles in bio-industry. However, low transformation efficiency of exogenous plasmids limited the application of Bacillus species. Here, a novel plasmid interspecific transfer system, with high transformation efficiency, high positive rate, and convenient manipulation, has been successfully constructed. A high electrotransformation efficiency strain Bacillus subtilis F-168 containing the counter-selectable marker mazF was used as the plasmid donor strain in this transfer method. A shuttled plasmid pBE980 and its recombinant plasmids pBE980::pulA and pBE980::HSPA were successfully transferred into the recipient Bacillus strains (Bacillus amyloliquefaciens 66, Bacillus licheniformis 124 and Bacillus megaterium 258) by this method. After co-culturing the donor cells (OD600nm = 1.3–1.7) and the recipient cells (OD600nm = 0.5–0.9) for 24 h in 22 °C, more than 1.0 × 105 positive transformants were obtained and a interspecific transformation efficiency of 1.0 × 10−3. It would provide a new approach for genetic manipulation in Bacillus strains and accelerate the research progress of the wild Bacillus strains in bio-industry.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacillus is a large genus of bacteria which shows typical phenotypes including Gram-positive, spore-forming, strict aerobic or facultative anaerobic [5, 10, 15]. Most Bacillus species are non-pathogenic and are capable of secreting functional extracellular proteins directly to the culture medium. Moreover, the mature large scale fermentation conditions of many Bacillus sp. strains have now been acquired. Because of these superiorities, Bacillus species are usually supposed to attractive hosts for the production of heterologous secretory proteins and play important roles in bio-industrial production [3, 13, 24]. In the near past years, with the development of molecular biology and genetic engineering, the genetic manipulation of Bacillus sp. strains has become the research focus [16, 25, 30].
Bacillus subtilis 168 as a model strain for molecular manipulation was first discovered to be transformable in 1958 [17]. Hereafter, plasmids with antibiotic markers, such as pE194 and pUB110, were isolated from Staphylococcus aureus and used as vectors in B. subtilis and other Bacillus strains [11]. Nowadays, a series of genetically engineered Bacillus strains (such as Bacillus licheniformis and Bacillus megaterium) and the corresponding plasmids were constructed successfully [8, 29]. However, efficiently transformation of exogenous plasmids is a critical factor to the development of this field [21]. At present, foreign plasmids are transferred into Bacillus strains mostly by electroporation, protoplast method and biochemical transformation. These three transformation ways are well-developed, but they still have some shortcomings [27]. Biochemical transformation has limited universality and is usually used in B. subtilis 168. Electroporation and protoplast method are applicable to a great variety of Bacillus species, but the complex manipulation and the difficulty for cell recovering limited their application. On the other hand, low transformation rates are normally found in these two methods [1, 4, 28]. According to previous reports, there are another two means of gene transfer except transformation: conjugation and transduction [18]. Based on the transfer mechanism of these two ways, some transfer agents such as bacteriophage or transposon are necessary. However, an aggregation-mediated plasmid transfer was found between two Bacillus strains without any transfer agents neither in the donor nor the recipient strains [27]. The universality and mechanism of this aggregation-mediated plasmid transfer need to be further studied. Therefore, it is still necessary to make efforts to construct an applicable, simple and efficient transfer system in Bacillus species. Especially, among the wild Bacillus strains, there are no efficient plasmid transfer methods to be reported so far.
In this study, a convenient and efficient plasmid interspecific transformation procedure was developed for the wild Bacillus strains. A shuttle plasmid pBE980 was successfully transferred from B. subtilis F-168 to the wild Bacillus strains B. amyloliquefaciens 66, B. licheniformis 124 and B. megaterium 258, respectively. And the donor strain (B. subtilis F-168) was constructed by integrating an IPTG inducible toxin-gene mazF cassette to the chromosome of B. subtilis 168. So, an efficiency counter-selectable method was proved to work for the selection of the positive recipient Bacillus strains. Therefore, this reported transformation procedure is more applicative to the wild Bacillus recipients with high transfer efficiency and application prospects.
Materials and methods
Bacterial strains, plasmids, oligonucleotides and materials
The bacterial strains and plasmids used in this study are listed in Table 1. B. subtilis 168 (ATCC23857) was provided by the American Tissue Culture Collection (ATCC). And the wild recipient strain B. amyloliquefaciens 66 was deposited in the China General Microbiological Culture Collection Center with the number CGMCC14363. The 16s rDNA sequences of Bacillus amyloliquefaciens 66, Bacillus licheniformis 124, and Bacillus megaterium 258 have been submitted to the GenBank database under accession No. MH150812, MH150817, and MH152576, respectively. The specific primers (Table S1) used for PCR amplification are synthesized by the Beijing Genomics Institute (BGI). The restriction endonucleases and DNA polymerase were commercially supplied by Thermo Fisher Scientific Co., Ltd. All other enzymes, chemicals and reagents were purchased from TaKaRa Biotechnology (Dalian, China) Co., Ltd.
Culture and growth conditions
All cells were routinely grown at 37 °C and 200 rpm in Luria–Bertani (LB) medium. LBG (LB containing 1% glucose) medium was used to prevent leaked expression of mazF gene in B. subtilis F-168. LB medium containing 0.8 mM IPTG was used to select transformants underlying the mazF counter-selectable system. Different antibiotics (100 μg/ml ampicillin, 25 μg/ml kanamycin, 150 μg/ml spectinomycin and 25 μg/ml chloramphenicol) were added in the medium for the relevant recombinant strains.
General molecular manipulations
The isolation and manipulation of recombinant DNA were performed using the methods as described by Sambrook and Russell [14]. The E. coli competent cells were obtained by regular way and transformed through chemical transformation method as described by Sambrook and Russell [14]. The electroporation transformation of B. subtilis 168 was carried out according to Tjalsma et al. [20].
Construction of the shuttle plasmid pBE980 and its recombinants
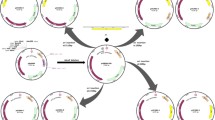
The ori gene from site 1694 to 2562 was amplified using primers ori-1/ori-2 (Table S1) from pUC19. Then the PCR products were inserted into pWB980 at the site between EcoRI and BglI which is on the upstream of P43 promoter (Fig. 1). The recombinant E. coli–B. subtilis shuttle expression vector was named as pBE980 in this work. Moreover, the recombinant vector pBE980::pulA and pBE980::HSPA were constructed by inserting a pullulanase encoding gene (pulA) and a protease encoding gene (HSPA) into the multiple cloning site (MCS) of pBE980, respectively. The PulA gene and HSPA gene were synthesized according to the sequences in GenBank database under HQ844266.1 and JX235368.1, respectively.
Relative plasmid copy number determination
The copy number of plasmids was determined by quantitative real-time PCR (qPCR) using iQ™ SYBR® Green Supermix (Bio-Rad). Primers were designed to have a predicted melting temperature of about 60 °C and to generate products of approximately 100 bp in length. Dilutions (10−1 to 10−4) of total cellular DNA (10 μg) isolated from B. subtilis F-168 (containing test plasmids) were analyzed by qPCR using primer pairs specific for the rep gene. The copy numbers were calculated as the mean threshold cycle (CT) values of the amplicons of the chromosomal genes (single-copy reference) compared to the amplicon of the plasmid-borne rep using the formula 2ΔCT. ΔCT is the difference between the threshold cycle number of the reference gene and that of the rep reaction [2].
Construction of B. subtilis F-168
Using E. coli chromosome DNA as the template, the lpp promoter and the mazE gene were amplified by PCR with primers P5/P6 and P7/P8 (Table S1), respectively. The two obtained fragments were then fused together by overlap PCR using primers P1/P2 (Table S1). After that, the PCR products were ligated to the sites between Xho I and Bgl II of pHT43 to form pHT43E. The mazF gene was amplified from the chromosome DNA of E. coli DH5α using the primers P9/P10 (Table S1), then inserted into the pHT43E to yield pHT43EF. With the primers P11/P12 and P13/P14 (Table S1), the upstream and downstream fragments of the aprx gene, aprxUP and aprxDOWN were amplified by PCR from B. subtilis 168 chromosome DNA and cloned into the corresponding site of pHT43EF resulting plasmid pHT43EUFD. When pHT43-EUFD was linearized by Xho I and then transformed into B. subtilis 168, a double crossover mutant strain was selected and named as B. subtilis F-168 (Fig. 2a).
Construction of B. subtilis F-168 (a) and PCR identification of the recombinant clones (b). a Linearized plasmid pHT43-EUFD was inserted into the genome of B. subtilis 168 through double crossover events at the aprx locus. b PCR results using P7/P8 (mazE) and P9/P10 (mazF) as primers, respectively. M: DNA marker; 1: the negative clone; 2, 3, 4: the positive clones; +: using pHT43-EUFD as the template; −: using the genome of B. subtilis 168 as the template
The interspecific transformation
First, the target plasmids were transformed into B. subtilis F-168 using electroporation transformation method. The B. subtilis F-168 harboring exogenous plasmids was used as the plasmid donor. And the other Bacillus sp. strains were used as the plasmid recipients. The donor strain was cultured in LBG medium and the recipient strain was cultured in LB medium. They were separately cultured at 37 °C and 220 rpm for 10–12 h. And then 10% of each seed cultured broth was inoculated into the fresh LB medium for continuing cultivation. Until the donor cells grew to OD600nm = 1.3–1.8 and the recipient cells grew to OD600nm = 0.5–1.0, the cells were collected by centrifugating at 13,000g for 2 min. 10% of the supernatant was used to resuspend the cells. Immediately, the donor cells and recipient cells were mixed at a volume ratio of 3:7 into total 200 μl and co-incubated in 1.8 ml LB medium with 0.8 mM IPTG at 22 °C and 220 rpm for 24 h. As a control, 1 μg plasmid DNAs were mixed with the same volume of the recipient cells and co-incubated under the same condition. The positive recipient transformants were selected under the mazF counter-selectable system and confirmed by PCR or plasmid extraction.
The counter selection with mazF
After co-cultured of the donor cells and the recipient cells, the mixed cells (1 ml, OD600nm = 1.0) with ten times dilution was spread on a LB plate with 0.8 mM IPTG. Then the plate was cultured at 37 °C for 24 h. The positive recipient transformants could be found through colonial morphology and there remained minimal residual donor cells on the plate.
T1: Total amount of donor cells spread on the non-selectable plates
T2: Total amount of recipient cells spread on the non-antibiotic plates
R1: Residual donor cells on the selectable plates
R2: Recipient transformants on the selectable plates
Results and discussion
Construction of B. subtilis F-168 containing counter-selectable marker mazF
The mazF gene comes from E. coli and encodes an endoribonuclease MazF, which is lethal to bacteria by specifically cleaving free mRNAs at ACA sequences [7, 26]. In this work, a double crossover plasmid pHT43-mazE-aprxUP-mazF-cassette-aprxDOWN (pHT43-EUFD) was constructed by the procedure described in “Materials and methods” (Fig. 2a). To prevent the host death caused by MazF, an antitoxic gene mazE was introduced at the upstream of the mazF-cassette (Fig. 2a). After double crossover, the mazF-cassette containing mazF-lac operon-cat with the full length of 3131 bp were successfully inserted into the genome of B. subtilis 168 (Fig. 2a). The double crossover mutants were selected and identified by specific PCR. The positive transformants were obtained through PCR results which show the mazF gene band and without the mazE gene band (Fig. 2b). The positive transformant of B. subtilis 168 containing the mazF-cassette in its genome was named B. subtilis F-168 and will be used as the plasmid donor in the subsequent experiments. The expression of the toxic protein MazF was induced by IPTG for counter selecting the transformants in the plasmid transformation. After the plasmids transferring from the donor strains (B. subtilis F-168) to the recipient strains, the donor strains were death by IPTG inducing which facilitate the screening of the positive recipient strains.
To further determine the lethal effects of the MazF in B. subtilis F-168, IPTG with a concentration of 1 mM was added into the culture medium at the beginning of cultivation. Different to the control (do not add IPTG), the B. subtilis F-168 begin to death at the end of logarithmic phase at 12–20 h due to the expression of toxic protein MazF (Fig. 3a). When B. subtilis F-168 was cultured at different inducing temperatures, the CFU of surviving stains showed obviously difference. At lower culture temperature (22 °C), the least CFU of B. subtilis F-168 was obtained with the highest lethality rate (Fig. 3b). Furthermore, the different lethality rates of B. subtilis F-168 were obtained according to various addition amounts of IPTG at 22 °C. After 24 h of cultivation, the highest lethality rate of 99.996% was reached at the IPTG concentration of 0.8 mM (Fig. 3c). When the additional amounts of IPTG were lower or higher than 0.8 mM, the lethality rate of B. subtilis F-168 began to decline (Fig. 3c).
These results demonstrate that the mazF gene from E. coli was a highly effective counter-selectable marker to eliminate the plasmid donor strain B. subtilis F-168 after plasmid interspecific transformation. So the positive recipient strains could easily be selected under the resistance of plasmid among the mix strains. The positive recipient recombinants also could be selected under double-resistance system. However, to implement double-resistance screening, it should prior import an appropriate resistant gene to the recipient strains. While most wild Bacillus strains do not have innate-resistant gene and hard to make molecular manipulate. Thus, the counter-selectable marker (mazF) makes this novel plasmid transfer method more suitable to transfer heterogeneous plasmids into the wild Bacillus strains which do not have any selectable markers or hard to manipulate in regular ways.
The high electrotransformation efficiency of pBE980 to B. subtilis F-168
B. subtilis 168 is a potential and attractive host for the transformation of heterologous plasmids. The high transformation efficiency of B. subtilis 168 was up to 1.0 × 104–1.0 × 106 transformants per microgramme plasmid DNA by electroporation method [9, 23]. Theoretically, the donor strain B. subtilis F-168, which derived from B. subtilis 168, could achieve the same level of transformation efficiencies of B. subtilis 168 using electroporation. However, it also determined by the qualified plasmid. pWB980 derived from pUB110 is commonly used for expression and secretion studies in B. subtilis [22]. Due to the smaller size, pWB980 could obtain relative higher electrotransformation efficiency than that of pUB110 (Table 2). While the connected products between pWB980 and the objective genes revealed much lower electrotransformation efficiency, even hard to obtain transformants. To overcome this defect, a novel shuttle plasmid pBE980 was constructed by inserting ori gene of pUC19 to the upstream of P43 promoter in pWB980 (Fig. 1). The electrotransformation efficiency and copy number of pBE980 to B. subtilis F-168 were 1.28 × 104 and 108, respectively (Table 2). Through plasmid enrichment in E. coli DH5α, the pBE980 and its recombinant plasmid pBE980::pulA and pBE980::HSPA could obtained the similar high electrotransformation efficiency with pWB980 to B. subtilis F-168 (Table 2). And the copy numbers of these plasmids are also high in the B. subtilis F-168 cells (Table 2) which ensure the required amount of the donor plasmids. All of these virtues provide the prerequisite for the plasmid interspecific transformation approach.
Plasmid interspecific transformation from B. subtilis F-168 to the wild strain B. amyloliquefaciens 66
B. amyloliquefaciens 66, a wild-type strain, with an attractive ability for secreting proteins was used as a plasmid recipient strain. And the plasmid pBE980 and its recombinant plasmid pBE980::pulA were successfully transferred from B. subtilis F-168 to B. amyloliquefaciens 66 according to the methods described in “Materials and methods”, respectively. The positive recombinant recipient clones were selected by their colonial morphologies (Fig. 4a, b). All the selected positive recipient strains could extract the target plasmids (Fig. S1c). To further identified the positive recipient stains, PCR for 16s DNA and desired gene (pulA) were used. The results showed that all the selected positive recipient strains were B. amyloliquefaciens 66 (data not shown). And the positive recipient strains B. amyloliquefaciens 66 (pBE980::pulA) have pulA gene, but do not have the mazF gene (Fig. S1a, b). Moreover, the amount of positive transformants harboring pBE980 was up to 3.4 × 103 which was a little higher than that (1.3 × 103) of transformants harboring pBE980::pulA (Fig. 4c). It may indicate that the efficiency of this plasmid interspecific transformation is closely related with the size of plasmid. The interspecific transformation efficiencies of plasmid pBE980 and pBE980::pulA were 1.1 × 10−5 and 2.8 × 10−5, respectively. In these results, plasmid pBE980 and its recombinant plasmid (pBE980::pulA) have proved to be feasibility of interspecific transformation from B. subtilis F-168 to the wild strain B. amyloliquefaciens 66. And those also illustrated that the interspecific transformation with counter-selectable marker mazF could screen the transformants without antibiotic selectable marker in recipient strains. That could provide convenient for wild Bacillus strains as the recipients. However, the transformation efficiency of the plasmid interspecific transformation still has room for improvement.
Optimization of plasmid interspecific transformation
Through previous study, some key factors such as the growth stage of the donor cells before co-culturing, the growth stage of the recipient cells before co-culturing, the mixed volume ratio and the co-incubation time could affect the interspecific transformation efficiency at different levels. To further improve the interspecific transformation efficiency, a series of transformation conditions were optimized.
Effects of the growth stage of the donor cells before co-culturing
The donor strain B. subtilis F-168 (pBE980::pulA) were cultured previously to different OD600nm (from 0.3 to 2.0). And then were mixed with the recipient strain B. amyloliquefaciens 66 in its OD600nm at 0.85 and the volume ratio was 5:5. After co-incubating at 22 °C and 220 rpm for 12 h, the mixed cells were properly diluted and coated on the selected plates. In the results, when the OD600nm of donor cells were between 1.3 and 1.7, which was at the later of exponential stage of the cells, the positive transformants were higher than 1.5 × 104 (Fig. 5a). The highest transformant amount was up to 2.38 × 104 at OD600nm = 1.387, while the interspecific transformation efficiency was up to 1.98 × 10−4 correspondingly (Fig. 5a). It probably because the plasmids were more facile to be released into the extracellular at the later of logarithmic phase (Fig. S2a). When the cells in the later of the logarithmic phase were inoculated into fresh medium, more plasmids were free in the solution resulting in high interspecific transformation efficiency to the recipient cells.
Optimization of plasmid interspecific transformation from B. subtilis F-168 to B. amyloliquefaciens 66. a Effects on interspecific transformation efficiency by different OD600nm of the donor cells. b Effects on interspecific transformation efficiency by different OD600nm of the recipient cells. c Effects on interspecific transformation efficiency by different volume ratios of the donor to the recipient in the mixture. d Effects on interspecific transformation efficiency by different co-incubation times of the donor and recipient mixture
Effects of the growth stage of the recipient cells before co-culturing
When the donor cells were cultured to OD600nm = 1.4, various growth stages (OD600nm = 0.3–1.5) of the recipient cells were mixed with those at volume ratio of 5:5. The mixed strains were then co-cultured at 22 °C and 220 rpm for another 12 h. The positive transformants were at least 1.5 × 104 when the OD600nm of recipient cells before co-culturing was 0.5–0.9 (Fig. 5b). And the optimal OD600nm of recipient cells was 0.85 which lead to the highest positive transformants amount of 2.22 × 104 and the highest interspecific transformation efficiency of 1.85 × 10−4 (Fig. 5b). In this experiment, we conclude that the recipient cells were inoculated at the early stage of the logarithmic phase (OD600nm = 0.5–0.9) could facilitate absorption of exogenous plasmids due to their active growth (Fig. S2b).
Effects of the volume ratio of the donor to the recipient in the mixture
Before the two kinds of cells were mixed, the donor B. subtilis F-168 (pBE980:: pulA) and the recipient B. amyloliquefaciens 66 were cultured to OD600nm= 1.4 and OD600nm = 0.85, respectively. And then they were mixed at different volume ratios (1:9; 2:8; 3:7; 4:6; 5:5; 6:4; 7:3; 8:2; 9:1) to the total volume of 200 μl. For co-incubating, the 200 μl mixed cells were incubated into 1.8 ml fresh LB medium with 0.8 mM IPTG. The culture condition was also at 22 °C and 220 rpm for 12 h. The most positive transformants (6.83 × 104) were obtained at a volume ratio of 3:7 (the donor to the recipient, v/v). And at that ratio, the highest interspecific transformation efficiency was up to 1.85 × 10−4 (Fig. 5c). Though the volume ratio of 8:2 and 9:1 also showed relative high interspecific transformation efficiency (Fig. 5c), the residue donor cells simultaneously occupied a high proportion in the mixed cells which would hinder the screen of positive recombinant. Thus, the optimum mix ratio was not only lie on the high-positive transformants but also need to consider the convenient for positive recipient cells selection.
Effects of co-incubation time of the donor and recipient mixture
Based on the determined transformation condition above, different co-culturing times from 2 to 36 h were test for the higher interspecific transformation efficiency. The results indicated that the transformation amount reached to 1.0 × 105 after co-cultured for 24–28 h (Fig. 5d). The highest positive transformants amount and the highest interspecific transformation efficiency were 1.10 × 105 and 9.19 × 10−4, respectively, when they co-cultured for 24 h (Fig. 5d). Moreover, the interspecific transformation efficiency was increasing gradually in the co-cultured time from 12 to 24 h. It indicates that the plasmid interspecific transformation in this study needs a relatively long period of incubation time. The process of plasmid transfer was during the growth of the mix strains.
The plasmid interspecific transformation system for various wild Bacillus species
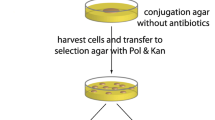
Based on the optimized transformation conditions above, B. subtilis F-168 was further used as the donor to introduce plasmids pBE980, pBE980::pulA and pBE980::HSPA into B. licheniformis 124 and B. megaterium 258 through the interspecific transformation, respectively (Table 3). The total flowchart of the plasmid interspecific transfer method was shown in Fig. 6. After co-cultured for 24 h at 22 °C in LB medium containing 0.8 mM IPTG, the mixture was plated on selective plates (kan25 + 0.8 mM IPTG) with an appropriate dilution (Fig. 6). The positive transformants were confirmed by plasmid extraction and PCR identification using primers 980S/980A. Comparing the recombinant plasmids pBE980::pulA and pBE980::HSPA, pBE980 showed the highest interspecific transformation efficiency to the three recipient Bacillus species, respectively (Table 3). In a word, the plasmid interspecific transformation method was confirmed to be valid at least to the three wild-type recipient Bacillus strains in this study. It can be largely solved the difficulty in transferring plasmids into the three wild-type strains. Moreover, many Bacillus species are of considerable interest to industry and agriculture [30]. Both gene function research and strain improvement require effective plasmid transfer method for genetic manipulation in these species. Despite the fact that a number of plasmid-borne genes have been cloned and expressed in Bacillus species, genetic studies on wild Bacillus species have been seriously hampered by the lack of efficient methods for the plasmid transformation. Various studies have reported the transfer of plasmid DNAs into Bacillus species by biochemical transformation, electroporation, and protoplast method. While these methods suffer from serious shortcomings such as cumbersome and very low transformation efficiencies [9]. In this work, a simpler whole cell operate procedure has been achieved, which do not need a complicated process of making competent cells (Fig. 6). However, the detailed transfer mechanism remains to be further research.
The plasmid pBE980 was an E. coli/B. subtilis shuttle plasmid compounding from pWB980 and pUC19. And the B. subtilis plasmid pWB980 was derived from pUB110 with destroyed mob region [22]. Thus, the plasmid pBE980 has no autogenous transfer ability in this study. And it have no mediated plasmids been found neither in the donor strains nor in the recipient strains. The result of control experiment, which mixed the plasmid DNAs and the recipient cells under the same procedure, showed that the plasmids could not transfer directly to the recipient strains. It indicated that the novel plasmid interspecific transformation method reported here mainly based on natural transfer system of the donor and the recipient strains. The plasmids pWB980 and pHT43 were also tried to perform interspecific transformation under the optimum co-culture conditions, respectively. As a result, pWB980 showed little higher transformation efficiencies to all of the three recipient strains than that of pBE980 (Table 3). However, pHT43 could not be obtained any positive recipient transformant (Table 3). The necessary optimization of the transformation conditions has also been performed and resulted in no improvement (data not shown). But when B. subtilis WB600 was used as the recipient cells, a very low transformation efficiency (1 × 10−6) of pHT43 was obtained using this method. So the main reason may be that pHT43 cannot replicate in those different species strains. Besides, the lower copy number (Table 2) of pHT43 and the different replicate mode (θ-type replicate mode) [12, 19] with pWB980 were possibly resulted in the low transformation efficiency. Fortunately, the plasmid pBE980 used in this work showed well compatibility in all these selected strains. And these three strains have typical evolutionary relationship with the most of the type or industrially important Bacillus strains (Fig. S3). Furthermore, the derivative plasmids of pUB110 are still the main current Bacillus research tools. So the pBE980 with the replicon derived from pUB110 used in this work has certain representativeness. As to whether other plasmids (pUB110 derivative or not pUB110 derivative) could be transformed using this method, the replication ability in different Bacillus species strains is prerequisite and needs to be further studied. Moreover, according to a previous report, an aggregation phenotype of Bacillus thuringiensis subsp. israelensis has supposed to be correlated with conjugation-like plasmid transfer [6]. In this study, the aggregation phenotype and the growth periods of both donor and recipient strains also showed important effect on the natural plasmid transfer phenomenon.
Conclusion
This study constructed a novel, efficient and convenient plasmid interspecific transformation method to solve the puzzle for plasmid transformation to the wild-type Bacillus strains. This approach expands the alternatives of the traditional transformation methods and makes the more successful plasmid transformation into the wild-type Bacillus strains become possible. Compared with traditional transformation methods, this new method has three superiorities. First, it revealed high transformation success rate to the wild-type Bacillus strains, while the traditional transformation generally difficult to achieve. Second, the manipulation is simple and convenient. It needs no preparation of competent cells. It can be performed under normal cultivating process. Third, it showed wider applicable range in recipient Bacillus species such as B. amyloliquefaciens, B. licheniformis and B. megaterium. These features make it an attractive technique to facilitate genetic studies and industrial application of these Bacillus species. And this technique may prove to be applicable to other members of the genus Bacillus.
References
Akamatsu T, Taguchi H (2012) Plasmid transformation of competent Bacillus subtilis by lysed protoplast DNA. J Biosci Bioeng 114:138–143
Chen Y-YM, Shieh H-R, Lin C-T et al (2011) Properties and construction of plasmid pFW213, a shuttle vector with the oral Streptococcus origin of replication. Appl Environ Microb 77:3967–3974
Chowdhury SP, Hartmann A, Gao X et al (2015) Biocontrol mechanism by root-associated Bacillus amyloliquefaciens FZB42—a review. Front Microbiol 6:780
Gao C, Xue Y, Ma Y (2011) Protoplast transformation of recalcitrant alkaliphilic Bacillus sp. with methylated plasmid DNA and a developed hard agar regeneration medium. PLoS One 6:e28148
Hmani H, Daoud L, Jlidi M et al (2017) A Bacillus subtilis strain as probiotic in poultry: selection based on in vitro functional properties and enzymatic potentialities. J Ind Microbiol Biot 44:1157–1166
Jensen GB, Wilcks A, Petersen SS et al (1995) The genetic basis of the aggregation system in Bacillus thuringiensis subsp. israelensis is located on the large conjugative plasmid pXO16. J Bacteriol 177:2914–2917
Kolodkin-Gal I, Hazan R, Gaathon A et al (2007) A linear pentapeptide is a quorum-sensing factor required for mazEF-mediated cell death in Escherichia coli. Science 318:652–655
Li Y, Gu Z, Zhang L et al (2017) Inducible expression of trehalose synthase in Bacillus licheniformis. Protein Expr Purif 130:115–122
Liu L, Liu Y, Shin HD et al (2013) Developing Bacillus spp. as a cell factory for production of microbial enzymes and industrially important biochemicals in the context of systems and synthetic biology. Appl Microbiol Biot 97:6113–6127
Melo AL, Soccol VT, Soccol CR (2016) Bacillus thuringiensis: mechanism of action, resistance, and new applications: a review. Crit Rev Biotechnol 36:317–326
Naglich JG, Andrews RE Jr (1988) Tn916-dependent conjugal transfer of PC194 and PUB110 from Bacillus subtilis into Bacillus thuringiensis subsp. israelensis. Plasmid 20:113–126
Nguyen HD, Nguyen QA, Ferreira RC et al (2005) Construction of plasmid-based expression vectors for Bacillus subtilis exhibiting full structural stability. Plasmid 54:241–248
Philibert T, Rao Z, Yang T et al (2016) Heterologous expression and characterization of a new heme-catalase in Bacillus subtilis 168. J Ind Microbiol Biot 43:729–740
Sambrock J, Russel D (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York
Sella SR, Vandenberghe LP, Soccol CR (2015) Bacillus atrophaeus: main characteristics and biotechnological applications—a review. Crit Rev Biotechnol 35:533–545
Song P, Liu S, Guo X et al (2015) Scarless gene deletion in methylotrophic Hansenula polymorpha by using mazF as counter-selectable marker. Anal Biochem 468:66–74
Spizizen J (1958) Transformation of biochemically deficient strains of Bacillus subtilis by deoxyribonucleate. P Natl Acad Sci USA 44:1072–1078
Strohmaier H, Noiges R, Kotschan S et al (1998) Signal transduction and bacterial conjugation: characterization of the role of ArcA in regulating conjugative transfer of the resistance plasmid R1. J Mol Biol 277:309–316
Titok MA, Chapuis J, Selezneva YV et al (2003) Bacillus subtilis soil isolates: plasmid replicon analysis and construction of a new theta-replicating vector. Plasmid 49:53–62
Tjalsma H, Koetje EJ, Kiewiet R et al (2004) Engineering of quorum-sensing systems for improved production of alkaline protease by Bacillus subtilis. J Appl Microbiol 96:569–578
Tominaga Y, Ohshiro T, Suzuki H (2016) Conjugative plasmid transfer from Escherichia coli is a versatile approach for genetic transformation of thermophilic Bacillus and Geobacillus species. Extremophiles 20:375–381
Wu SC, Wong SL (1999) Development of improved pUB110-based vectors for expression and secretion studies in Bacillus subtilis. J Biotechnol 72:185–195
Xue G-P, Johnson JS, Dalrymple BP (1999) High osmolarity improves the electro-transformation efficiency of the gram-positive bacteria Bacillus subtilis and Bacillus licheniformis. J Microbiol Methods 34:183–191
Yzturk S, Yalik P, Yzdamar TH (2016) Fed-batch biomolecule production by Bacillus subtilis: a state of the art review. Trends Biotechnol 34:329–345
Zhang K, Duan X, Wu J (2016) Multigene disruption in undomesticated Bacillus subtilis ATCC 6051a using the CRISPR/Cas9 system. Sci Rep UK 6:27943
Zhang XZ, Yan X, Cui ZL et al (2006) mazF, a novel counter-selectable marker for unmarked chromosomal manipulation in Bacillus subtilis. Nucleic Acids Res 34:e71
Zhang XZ, You C, Zhang YH (2014) Transformation of Bacillus subtilis. In: Sun L, Shou W (eds) Engineering and analyzing multicellular systems. Methods in molecular biology (methods and protocols), vol 1151. Humana Press, New York
Zhang Z, Ding ZT, Shu D et al (2015) Development of an efficient electroporation method for iturin A-producing Bacillus subtilis ZK. Int J Mol Sci 16:7334–7351
Zheng H, Liu Y, Liu X et al (2012) Overexpression of a Paenibacillus campinasensis xylanase in Bacillus megaterium and its applications to biobleaching of cotton stalk pulp and saccharification of recycled paper sludge. Bioresour Technol 125:182–187
Zhou C, Shi L, Ye B et al (2017) pheS (*), an effective host-genotype-independent counter-selectable marker for marker-free chromosome deletion in Bacillus amyloliquefaciens. Appl Microbiol Biot 101:217–227
Acknowledgements
This work was supported by the National Natural Science Fund of China (Grant 31701534), the Key Deployment Project in Chinese Academy of Sciences (Grant KFJ-STS-ZDTP-016-1, KFZD-SW-211-2), and the Tianjin Science & Technology Planning Project (Grant 16YFZCSY00790,16YFXTSY00530 and 15YFYssy00040).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, X., Xu, J., Tan, M. et al. Construction of a plasmid interspecific transfer system in Bacillus species with the counter-selectable marker mazF. J Ind Microbiol Biotechnol 45, 417–428 (2018). https://doi.org/10.1007/s10295-018-2038-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-018-2038-0