Abstract
Acidophilic microorganisms involved in uranium bioleaching are usually suppressed by dissolved fluoride ions, eventually leading to reduced leaching efficiency. However, little is known about the regulation mechanisms of microbial resistance to fluoride. In this study, the resistance of Acidithiobacillus ferrooxidans ATCC 23270 to fluoride was investigated by detecting bacterial growth fluctuations and ferrous or sulfur oxidation. To explore the regulation mechanism, a whole genome microarray was used to profile the genome-wide expression. The fluoride tolerance of A. ferrooxidans cultured in the presence of FeSO4 was better than that cultured with the S0 substrate. The differentially expressed gene categories closely related to fluoride tolerance included those involved in energy metabolism, cellular processes, protein synthesis, transport, the cell envelope, and binding proteins. This study highlights that the cellular ferrous oxidation ability was enhanced at the lower fluoride concentrations. An overview of the cellular regulation mechanisms of extremophiles to fluoride resistance is discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Uranium is a non-renewable resource used for the generation of nuclear power. To accommodate the demand for this limited resource, it is necessary to exploit low-grade and refractory uranium ores. The principal aim of uranium ore treatment is to obtain uranium concentrates with a U3O8 content of above 90 %. The fundamental step is to extract uranium from solid phase to solution phase. This extraction normally involves a process that employs strong acid as a reagent, which often creates environmental problems, requires large amounts of energy, and involves a complex operational plant. Biomining, the use of microorganisms to recover precious and base metals from low-grade mineral ores and concentrates, has developed into a successful and expanding area of biotechnology [38]. It has been widely used in the extraction of metals including copper, zinc, nickel, cobalt, gold, and uranium because of its economic and environmental benefits [35, 49]. Uranium bioleaching has been successfully applied in low-grade uranium ore in China, India, and Canada [29, 52].
In the uranium bioleaching process, iron- and sulfur-oxidizing microbes catalyze oxidation through indirect actions. They generate Fe(III) by oxidation of pyrite or additional soluble Fe(II). Fe(III) readily attacks minerals incorporating U(IV) and converts them to U(VI), which is soluble in dilute sulfuric acid generated by biological oxidation of pyrite or additional sulfur [3]. Acidithiobacillus ferrooxidans is the most widely used microorganism in such processes because of its stronger tolerance to heavy metal ions and ability to oxidize Fe- and S-compounds [21]. Along with other acidophilic Fe- or S-oxidizers, these bacteria commonly occur on the surfaces of exposed ores and in acid solutions in uranium mines [3]. The operating parameters of uranium bioleaching, including microbial species and community, pH, oxygen requirement, temperature, pyrite content, and additional energy substrate, have been previously described in detail [34, 37].
High-fluorine containing uranium ore accounts for a large proportion of the world’s uranium resources, with more than 60 % found in China. The fluorine concentration in the leachate of high fluorine uranium ore can reach 2–4 g/L. The deleterious fluoride ions released from fluorite (CaF2) and some other fluorine-bearing minerals strongly inhibited microbial growth, energy intake, enzyme activity, and metabolism, and in some cases, it killed the cells [22, 30, 31]. Research on fluoride inhibition of iron oxidation by Acidithiobacillus ferrooxidans illustrated that sodium fluoride at 0.4 mM caused 30 % inhibition of iron oxidation. Increasing the sodium fluoride concentration to 1.6 mM resulted in complete inhibition of microbial oxidation of ferrous iron [39]. It has also been demonstrated that the toxic fluoride concentrations at field sites can be significantly higher than the toxic levels reported in the laboratory, but still be inhibitory for the microorganisms [5]. Fluoride can negatively affect the process of bioleaching in a highly pH-dependent manner [30]. Therefore, microbial resistance to fluoride directly affects the growth activity and iron and sulfur oxidation rate, and in turn, the leaching efficiency [8, 30]. To obtain superior acid- and fluoride-tolerant stains, researchers have performed tests on acclimatizing and culturing the bacteria. After adaption to acid and fluoride, bacteria can grow in 0.8 M H2SO4 and 45 mM F−, respectively [50]. The level of microbial tolerance was not only related to the chemical factors of the growth environment, but also determined by the physical structure and genetic characteristics of the microorganisms [14, 43].
Microbes in extreme environments are normally equipped with complex metabolic pathways that allow them to adapt to certain stress situations. Regulation of these pathways is activated immediately after stimulation is introduced. Research into microbial resistance toward Ni2+, As3+, Hg+, and organic pollutants has been described in detail [4, 9, 17, 19, 55]. The tolerance of five typical bioleaching microorganisms to different concentrations of fluoride has been reported [26]. However, the regulation of microbial resistance to fluoride is still ambiguous at the genome level. The genome of A. ferrooxidans has been annotated and curated [47], making it possible to carry out genome-wide comprehensive studies through the construction and use of microarray technology [6]. Luo et al. have performed comparative genomic and phylogenetic analyses of different A. ferrooxidans strains based on microarray hybridization [25].
Herein, the growth responses of A. ferrooxidans ATCC 23270 to different concentrations of fluoride were compared in both FeSO4 and S0 substrate medium. Changes of the ferrous ion or sulfur oxidation rate were detected. A whole genome microarray was used to analyze the genome-wide expression profiling at different times after fluoride introduction in an attempt to identify the regulatory mechanism of microbial resistance to fluoride at the genome level.
Materials and methods
Strains and culture media
The strain A. ferrooxidans ATCC 23270 was purchased from the American Type Culture Collection (ATCC, Manassas, VA). The culture medium was prepared using autoclave sterilized 9 K basal medium as described by Silverman and Lundgren [41] (initial pH 2.0) containing ultraviolet sterilized [23] FeSO4·7H2O (44.7 g/L) or S0 (10 g/L) as the energy substrate. The 9 K basal medium contained the following components: (NH4)2SO4, 3.0 g/L; MgSO4·7H2O, 0.5 g/L; K2HPO4, 0.5 g/L; KCl, 0.1 g/L, Ca(NO3)2, 0.01 g/L. The inoculum density of A. ferrooxidans was 5.6 × 105 cells/mL. It was cultivated aerobically at 30 °C in a rotary platform incubator at 170 rpm.
Fluoride stress treatments
The strains were triplicate inoculated and cultured in 1500-mL shake flasks with 600 mL 9 K basal medium added with energy substrate. According to an earlier experiment result, when the cells reached the mid-log phase (40 h), each 1500-mL shake flask was equally subpackaged into six 250-mL shake flasks by siphonage. They were treated by adding 476 mM sodium fluoride solution to the culture system, with a final fluoride concentration of 2.4, 4.8, 7.2, 9.6, and 12 mM. Both biotic and abiotic controls were set up without fluoride. Cell density was measured by hemacytometer and pH variation was measured by a digital pH meter (pHSJ-4A; Leici, Shanghai, China). Ferrous concentration was detected according to the 1,10-phenanthroline method [32]. The interference of fluoride on the reliability of 1,10-phenanthroline method was negligible as the analysis result of ferrous ion standard solution with and without the presence of F− presented high consistent. Values present in Fig. 1 were mean ± SD (Error bar) of triplicate samples analyzed individually.
Genome-wide expression profiling for fluoride-resistant regulation
Based on the results of the growth curves, pH and ferrous concentration variations, the appropriate concentration (4.8 mM) of the fluoride stimulus was selected for further research by microarray technique. At 10, 30, 60, 120, and 240 min after adding 4.8 mM NaF, 50 mL culture medium from the FeSO4 group was collected and quickly mixed with 100 mL RNAprotect bacteria regent (Qiagen, Valencia, CA). Then, they were harvested by centrifugation at 12,000×g for 20 min in a 5804R centrifuge (Eppendorf AG, Hamburg, Germany). Bacteria in the biotic control group without fluoride were also harvested at the same time points.
RNA extraction and reverse transcription
Total RNA was extracted using TRIzol (Invitrogen, Carlsbad, CA) and the RNeasy mini kit (Qiagen) and treated with RNase-free DNase I (Qiagen) to digest residual chromosomal DNA and subsequently purified using an RNeasy kit (Qiagen). RNA quality was assessed by Bioanalyzer (Agilent, Palo Alto, California) and spectrophotometer (Nanodrop Technologies, Wilmington, Delaware). Then, the pure RNA was converted to cDNA with random primers following the manufacturer’s protocol for the ImProm-II™ Reverse Transcription System (Promega Corporation, Madison, WI), and purified with the QIAquick PCR Purification Kit (Qiagen, Hilden, Germany). All experiments were done in triplicate.
Microarray hybridization and data visualization
The whole-genome microarrays of A. ferrooxidans were designed and synthesized on the basis of sequence information provided by the Institute for Genomic Research (TIGR). A whole-genome oligonucleotide array was developed based on the 3217 ORFs of the A. ferrooxidans ATCC 23270 genome, including the genes related to 20 categories [33]. The cDNA labeling, microarray hybridization, and data analysis were carried out sequentially [24]. The cDNA of samples with/without fluoride treatment were labeled using Cy5-/Cy3-dUTP fluorescent dyes. Hybridization was carried out on an HS4800 Hybridization Station (TECAN US, Durham, NC) in triplicate at 45 °C for 10 h. Microarray data were visualized by a GenePix Personal 4100A scanner (AXON Instruments, Burlingame, CA) and then was converted to digital signals using the Genepix Pro 6.1 software. Microarray raw and analyzed data have been deposited in Gene Expression Omnibus (GEO) under the accession code GSE76450 (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE76450). Statistical analysis of gene expression data was carried out using Microsoft Excel and the R (v. 3.1.2) package VennDiagram (www.r-project.org). After taking the logarithm of the data, cluster analysis based on the Euclidean distance was performed using the R package pheatmap.
Results
Growth in the presence of different levels of fluoride stress
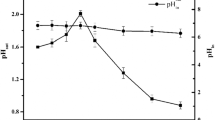
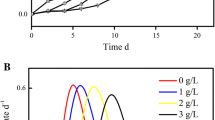
In the FeSO4 group, the growth of A. ferrooxidans was normal in the control group and inhibited in the groups containing 2.4, 4.8, 7.2, 9.6, and 12 mM fluoride (Fig. 1a). When the fluoride concentration was 2.4 mM, the maximum cell number was 7.3 × 107 cells/mL, which was 9.65 % lower than that grown in the non-fluoride condition. The maximum cell numbers decreased as the fluoride concentration increased. The critical fluoride concentration was about 7.2 mM, above which the growth curves trended down during the initial period after fluoride was introduced, but resumed to typical growth after a lag period. No significant difference was observed in the ferrous concentration among the experimental groups and control group when the fluoride concentration was less than 4.8 mM (Fig. 1b). In the group grown in the presence of 12 mM fluoride, the ferrous utilization was obviously decreased by high fluoride stress.
Both the cell density (Fig. 1c) and pH variations (Fig. 1d) were significantly suppressed by the presence of different concentrations of fluoride when A. ferrooxidans was grown in S0 substrate. Cell density was inhibited even in the presence of 2.4 mM fluoride as the maximum cell number was only 1.0 × 108 cells/mL, 50 % lower than that for the group grown without fluoride. When the fluoride concentration was higher than 4.8 mM, the growth immediately lagged and then the cell density exhibited a gradual decrease. Similarly, the pH variation was minimal in 2.4 or 4.8 mM fluoride, but increased when the medium contains over 7.2 mM fluoride.
Overview of differential gene expression profiles
Whole-genome microarrays were used to obtain a comprehensive description of the molecular response. The standardized data at 10 min are shown as an example (Fig. 2a). Experimental samples were Cy5 labeled and control samples were Cy3 labeled. The Cy5/Cy3 ratio represents the relative abundance of the 1278 up-regulated (ratio >2) and 465 down-regulated (ratio <0.5) gene signals.
Overview of differential gene expression profiles of A. ferrooxidans ATCC 23270 after 4.8 mM fluoride was added. a The scatter plots of hybridizing signals on gene chip at the point of 10 min (Numbers in plots indicate probes with a minimum change in expression of twofold (red, up-regulated; green, down-regulated). b Venn Diagram of differential expressed genes at different time points (Left, 1082 genes up-regulated within 240 min; Right, 430 genes down-regulated within 240 min). c Fold distribution of differential expressed genes at different time points
A total of 3217 unigenes were identified, of which 1354 genes were differently expressed. To access the temporal distribution of differentially expressed genes, overlapping and non-overlapping regions of differential gene expression in a subset of different time points are shown by Venn diagrams (Fig. 2b). There were 22 genes continuously up-regulated during the 240 min experimental process, while no genes underwent long-term down-regulation. The fold distribution of genes differentially expressed at a series of time points was also counted (Fig. 2c). Within the short-term stress period (0–120 min after fluoride was added), the gene expression was primarily up-regulated, while for the long-term stress time point (240 min), there were 211 down-regulated and 146 up-regulated genes differentially expressed (1.45-fold). Moreover, many genes were differently expressed less than 50-fold within 60 min, while genes differentially expressed over 50-fold were increased after 60 min.
According to the annotation results from the TIGR (www.tigr.org), the differentially expressed genes were divided into 19 categories by functional classification at each time point (Fig. S1). The distribution showed that genes expressed in response to fluoride stress were closely related to the cell envelope, cellular processes, energy metabolism, protein synthesis, transport, and binding proteins. There were also some related genes encoding conserved or non-conserved hypothetical proteins, and some genes with unknown functions that were differentially expressed.
Changes in gene expression related to energy metabolic systems
Cluster analysis showed that most genes associated with the iron/sulfur metabolic system of A. ferrooxidans were efficiently expressed from 10 to 120 min after 4.8 mM fluoride was added (Fig. 3). At the long-term stress time point of 240 min, gene expression returned to normal levels or even decreased below normal levels. The cyo operon coding for the cytochrome o ubiquinol oxidase and nuo operon coding for NADH-quinone oxidoreductase were typical examples that followed this expression pattern.
Energy metabolism-related genes in the carbon metabolic pathway were also differentially expressed after fluoride was introduced (Fig. 4). Genes involved in glycolysis/gluconeogenesis and the tricarboxylic acid (TCA) cycle were up-regulated within a short time and then returned to normal. For example, the genes pgi (AFE0185) and pgl (AFE0275) encoding the intermediate products of glucose-6-phosphate isomerase (G6PI) and 6-phosphogluconolactonase (6PGL) in the glycolysis/gluconeogenesis pathway were expressed in this manner.
Changes in gene expression related to carbon, nitrogen, and phosphate metabolic systems
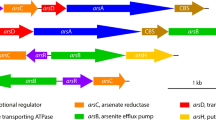
The results from the whole genome microarrays (Fig. 4) show that many genes of carbon fixation, including carboxysome encoding genes and those involved in the Calvin–Benson–Bassham cycle (CBB), were up-regulated most of the time. Most genes related to nitrogen metabolism were also up-regulated. For example, the genes encoding the domain protein, NifZ (AFE1548), and nitrogenase molybdenum-iron protein (AFE1563, nifD) appeared to be significantly up-regulated at multiple time points. The genes glnB-1 (AFE0640) and ntrC (AFE0208) encoding nitrogen regulatory proteins P-II and NR (I) were also up-regulated to some extent. By comparison, the genes involved in the phosphate transport system were partially up-regulated. A typical gene cluster named AFE1643-1648, including ppx (exopolyphosphatase), phoU (phosphate transport system protein PhoU), pstB (ATP-binding protein), pstA (permease protein PstA), pstC-2 (permease protein PstC), and pstS-2 (periplasmic phosphate-binding protein), was up-regulated significantly at several time points. Genes in the gene cluster AFE0811-0819, such as AFE0811 (transcriptional regulator), AFE0812 (PhnN protein), phnJ (phosphonate metabolism protein PhnJ), and phnH (phosphonate metabolism protein PhnH) were significantly up-regulated.
Changes in gene expression related to the cell membrane and detoxification systems
Investigation of global gene expression with a focus on the cell membrane (Fig. 5) showed that the synthetase-related genes of the surface polysaccharide, lipopolysaccharide (LPS), cytoplasm, and peptidoglycan were efficiently expressed under conditions of fluoride stress. These included genes such as rfaD (AFE1125) encoding ADP-L-glycero-D-mannoheptose-6-epimerase, mdoG (AFE2087) encoding periplasmic glucan biosynthesis protein mdoG, AFE2752 encoding dolichyl phosphate-mannose:protein O-mannosyltransferases (PMT/POMTs), AFE0135 encoding capsule polysaccharide exports protein from the BexD/CtrA/VexA family, AFE0179/AFE0025 encoding membrane proteins, and AFE1235/AFE0134 encoding a glycosyl transferase. However, several genes such as AFE0875 encoding penicillin-binding protein 1A, glmM (AFE0471) encoding phosphoglucosamine mutase, and nagZ (AFE0283) encoding beta-hexosaminidase were found to be down-regulated under conditions of long-term fluoride stress.
Differentially expressed genes related to the detoxification system (Fig. 5) included genes associated with toxin-resistant transporters and ion channel proteins, which were highly up-regulated at 240 min under fluorine stress. Examples included genes such as AFE2124, AFE0913, and AFE1004. On the contrary, AFE2737 and AFE2849, encoding glutathione S-transferase family proteins, were down-regulated.
Gene expression of co-expressed genes and expression regulators
Gene expression patterns based on nonnegative matrix factorization (NMF) were further analyzed. The 22 up-regulated genes in all 240 min were clustered into three fine modules based on their patterns of co-expression (Fig. S2). Gene 9 to gene 22 (AFE0913, AFE0677, AFE0427, AFE0597, AFE0645, AFE0868, AFE2124, AFE0847, AFE0926, AFE1041, AFE0929, AFE0821, AFE2209, and AFE0375) in Cluster III were significantly correlated, in particular from gene 14 to gene 22. These genes encoding proteins in detoxification (AFE2124, AFE0847), transportation (AFE0926, AFE0929, AFE0821), and energy metabolism (AFE1041, AFE0375) were simultaneously significantly induced by fluoride stress.
The genes contained in co-expression clusters at each time point (Fig. 6) were differentially expressed over 20-fold in 120 min short time stress, and differentially expressed only twofold at the long-term stress time point of 240 min, except genes in Cluster I (AFE0202, AFE0381, AFE0903, AFE1091, AFE0126, and AFE1867). Genes in Cluster I were differentially expressed over 20-fold throughout the time of the experiment for those whose functions were closely related to basal growth and metabolism of the cells, such as conserved protein synthesis (AFE0202, AFE0381, AFE1091, AFE0126, and AFE1867), DNA replication, recombination, and repair (AFE0903).
The fold changes of 22 up-regulated co-expressed genes at different time point. The radial plot displays serial number of genes on the circumference and the expression fold changes of genes on the radius. Genes: 1 transglutaminase-like domain protein (AFE0202); 2 conserved hypothetical protein (AFE0381); 3 ATP-dependent DNA helicase RecQ (AFE0903); 4 conserved hypothetical protein (AFE1091); 5 PIN domain protein (AFE0126); 6 hypothetical protein (AFE1867); 7 transcriptional regulator, GntR family (AFE0811); 8 hypothetical protein (AFE1300); 9 drug resistance transporter, EmrB/QacA family (AFE0913); 10 hypothetical protein (AFE0677); 11 glycogen synthase (AFE0427); 12 transposon, transposition protein B, putative (AFE0597); 13 amino acid permease family protein (AFE0645); 14 hypothetical protein (AFE0868); 15 drug resistance transporter, EmrB/QacA family (AFE2124); 16 PQQ enzyme repeat domain protein (AFE0847); 17 GTP-binding protein (AFE0926); 18 xylulose-5-phosphate/fructose-6-phosphate phosphoketolase (AFE1041); 19 outer membrane toxin secretion efflux protein, putative (AFE0929); 20 major facilitator family transporter (AFE0821); 21 hypothetical protein (AFE2209); 22 ubiquinol-cytochrome c reductase, cytochrome b subunit (AFE0375)
Discussion
Each type of extremophile has different survival strategies to help it adapt to the environment [36]. The conversion of F− to HF spontaneously occurs in the acidic leaching environment as shown in Fig. 7. As a highly permeable solute, the permeability of HF through the phospholipid bilayer is about seven orders of magnitude higher than that of F− [15, 46]. When HF is transferred into cells, it decomposes into H+ and F−. Concomitantly, the cytoplasm is acidified by H+ while F− combines with certain biosynthetic endoenzymes, inhibiting microbial growth or even killing the cells [27, 48]. The Fe3+ could react with F− to form soluble charged compounds and reduce the fluoride toxicity [40]. As a result, cells in the group with FeSO4 presented higher tolerance to different fluoride concentrations than those with the S0. Cells grown with the S0 as the substrate were strongly suppressed under minimum fluoride stimulation, leaving no opportunity for the microbes to adapt to the stimulation. The group with FeSO4 as the substrate was selected to explore the regulation mechanism of microbial resistance to the stimulation of fluorine.
The result of gene expression analysis based on the whole genome microarrays showed that many genes related to several metabolic pathways were continually up-regulated in response to the stressful environment. Genes differentially expressed at only one time point outnumbered those at two or more time points, indicating that many genes respond to the fluoride environment rapidly and efficiently as a result of the inherent cellular defense mechanisms [7] that function to adapt to environmental changes during periods of short-term stress. However, under conditions of long-term stress, the expression levels of many genes returned to normal or were even down-regulated. The number of genes differentially expressed over 50-fold was increased after 60 min. The gene expression levels were far away from the normal baselines for the long-term fluoride stress condition. From these results, we concluded that long-term fluoride stress at a certain level resulted in minor effects on the expression of a majority of genes [53]. This suggests that the cells have adapted to fluoride stress after a long-term exposure to fluoride. However, certain genes were strongly inhibited, leading to cases where the expression level significantly deviated from the normal level.
Cellular oxidation ability
The pH variation of the S0 group was minimal at the lower fluoride concentration and significantly increased at the higher fluoride concentration, primarily because of the weakly alkaline aqueous solutions resulting from the partial hydrolysis of NaF. However, in the FeSO4 groups grown in the presence of the lower fluoride concentration, the same ferrous oxidation rate was maintained at a lower cell density indicating that the oxidation ability of each cell was enhanced by fluoride stress. A previous study also showed that an increase of fluoride concentration resulted in growth delays of S. thermosulfidooxidans, but the oxidation rate was not inhibited [48]. The expression data for genes involved in energy metabolism were highly consistent with the ferrous oxidation rate, which was increased without an equivalent increase in cell density. This resulted because the increased energy intake was used for synthesizing the electron transport proteins, which acted as primary proton pumps to export protons and generated energy for other homeostatic mechanisms [51].
A general view of the cellular regulation mechanisms responsible for the fluoride resistance
When A. ferrooxidans was challenged by fluoride, the gene regulatory networks continued to adjust the metabolic systems to cope with the fluorine stress. The pattern of up-regulated genes coding for cell envelope, transport, binding, and regulatory proteins is consistent with the findings of previous studies on the effects of heat, cold, salt, and alkali stresses, however, genes related to energy metabolism were also up-regulated, a finding that was not in accordance with previous studies [10–12, 16, 42, 44, 45].
Genes involved in energy metabolism were efficiently expressed 10–120 min after fluoride addition, but at the long-term stress time point of 240 min, the gene expression levels returned to normal or were even down-regulated. This gene expression pattern observed for A. ferrooxidans may be the result of the differences in the amounts of energy expended by autotrophic acidophiles compared with heterotrophic acidophiles. A previous study reported that A. ferrooxidans lacks the key enzymes of phosphofructokinase in the Embden–Meyerhof–Parnas pathway, 6-phosphogluconate dehydratase and 2-keto-3-deoxy-6-phosphogluconate aldolase in the Entner–Doudoroff pathway, and α-ketoglutarate dehydrogenase in the TCA cycle [28]. This makes it impossible for A. ferrooxidans to obtain energy by substrate-level phosphorylation, leaving ferrous oxidation as the only energy source. However, the conversion of the Fe(II)/Fe(III) couple is a low-energy yielding reaction and, therefore, A. ferrooxidans needs to oxidize more substrate to maintain its own metabolism. Alternatively, different ferrous and sulfur metabolism pathways may be activated depending on the concentration of fluoride present. As discussed above, the ferrous oxidation ability of each cell was enhanced by a reduction in fluoride stress. Thus, the genes were up-regulated in 120 min to satisfy the cells’ requirement for more energy before they entered a period of long-term stress and were down-regulated.
The cluster of genes involved in carbon dioxide, nitrogen fixation, and phosphorus metabolism were up-regulated to obtain cellular carbon, nitrogen, and phosphorus sources for new protein biosynthesis [47]. This agrees well with the up-regulated genes found in the functional ‘protein synthesis’ group in Fig. S1 (Column k). The genes related to carbon metabolism displayed similar expression patterns as those involved in energy metabolism. A. ferrooxidans expanded its carbon sources by up-regulating those in the CBB pathway under fluoride stress. The genes involved in the synthesis of organic carbon were enhanced, especially those responsible for the production of fatty acids, phospholipids, and glycogen. These substances provided important components for the synthesis of the cell membrane, and played a crucial role in maintaining the membrane integrity and biological activity [18]. The highly expressed genes involved in glycolysis and the pentose phosphate pathway were relevant to DNA mismatch repair and NADPH synthesis. Additionally, genes involved in the expression of the pentose derivatives of NAD+, FAD2+, and CoA, important coenzymes in the electron transport chain, were up-regulated. As a result, carbon metabolism-related pathways, especially CBB, glycolysis, and the pentose phosphate pathway, played key roles in maintaining cell growth and metabolism under fluoride stress.
Most genes related to nitrogen and phosphorus metabolism were up-regulated to maintain the physiological balance under the fluoride stress, similar to those genes of sediment microbes involved in nitrogen and phosphorus cycling processes after nitrate injection [54]. It has been reported that the gene ppx was relevant to the stress resistance of A. ferrooxidans [2]. In this study, it was up-regulated in a gene cluster typically involved in resisting fluoride. Nitrogen and phosphorus metabolism are closely related to protein folding and stability, DNA recombination, and repair [1, 13]. Several associated pathways involved in the nitrogen and phosphorus metabolism of A. ferrooxidans were enhanced, presumably to reduce or repair the protein and nucleotide damage caused by fluoride stress. Additionally, the synthesis of lipid molecules was intensified to maintain the integrity, permeability, and fluidity of the cell membrane.
A summary of the potential cellular regulation processes involved in fluoride resistance is shown in Fig. 7. Previous research has also indicated that cells in an arsenic environment maintained the fluidity of the membrane by regulating LPS and lipoprotein composition, or by self-tuning the polyunsaturated/saturated fatty acid ratio to reduce toxic incursion [20]. The analysis of the expression of genes related to the cell membrane suggested that A. ferrooxidans subjected to fluoride stress maintained the osmotic balance and membrane fluidity by regulating the biosynthesis of diverse membrane components such as the dermatoplasm, peptidoglycan, polysaccharide, LPS, and lipoprotein, or by self-tuning the polyunsaturated/saturated fatty acid ratio, as well as down-regulating some unnecessary pathways to improve the tolerance to fluoride. The genes associated with the expression of the resistance transporter and regulators involved in the detoxification system were highly expressed. The mercuric and arsenic resistance genes were also up-regulated, indicating that there is similarity in the ability of microbes to resist the toxicity of metal and nonmetallic ions. In this way, cells can resist external environmental stresses and maintain metabolic balance and physiological processes.
A total of 22 genes in three co-expression clusters with different functions were continuously up-regulated in 240 min. The genes in Cluster I were closely related to cell growth and metabolism as a result of the basal physiological need to cope with the exposure to the fluoride environment. The genes in Cluster III were related to detoxification, transportation, and energy metabolism, and were up-regulated because of their role in resisting fluoride stress.
References
Acosta M, Beard S, Ponce JVG, Vera M, Mobarec JC, Jerez CA (2005) Identification of putative sulfurtransferase genes in the extremophilic Acidithiobacillus ferrooxidans ATCC 23270 genome: structural and functional characterization of the proteins. OMICS 9:13
Alvarez S, Jerez CA (2004) Copper ions stimulate polyphosphate degradation and phosphate efflux in Acidithiobacillus ferrooxidans. Appl Environ Microbiol 70:5177–5182
Bhatti TM, Antti V, Martti L, Tuovinen OH (1998) Dissolution of uraninite in acid solutions. J Chem Technol Biotechnol 73:259–263
Borole AP, Hamilton CY (2011) Using acidithiobacillus, leptospirillium and/or sulfolobus as bioreactor for removing mercury from fossil fuels; bioremediation; pollution control; biodegradation; biooxidation: US 7998724 B2
Brierley J, Kuhn M (2010) Fluoride toxicity in a chalcocite bioleach heap process. Hydrometallurgy 104:410–413
Brigham CJ, Speth DR, Rha C, Sinskey AJ (2012) Whole-genome microarray and gene deletion studies reveal regulation of the polyhydroxyalkanoate production cycle by the stringent response in Ralstonia eutropha H16. Appl Environ Microbiol 78:8033–8044
Chakraborty S, Mukherjee A, Khuda-Bukhsh AR, Das TK (2014) Cadmium-induced oxidative stress tolerance in cadmium resistant Aspergillus foetidus: its possible role in cadmium bioremediation. Ecotoxicol Environ Saf 106:46–53
Dopson M, Halinen AK, Rahunen N, Boström D, Sundkvist JE, Riekkola-Vanhanen M, Kaksonen AH, Puhakka JA (2008) Silicate mineral dissolution during heap bioleaching. Biotechnol Bioeng 99:811–820
Dopson M, Holmes DS (2014) Metal resistance in acidophilic microorganisms and its significance for biotechnologies. Appl Microbiol Biotechnol 98:8133–8144
Fisher MA, Plikaytis BB, Shinnick TM (2002) Microarray analysis of the Mycobacterium tuberculosis transcriptional response to the acidic conditions found in phagosomes. J Bacteriol 184:4025–4032
Gao H, Wang Y, Liu X, Yan T, Wu L, Alm E, Arkin A, Thompson DK, Zhou J (2004) Global transcriptome analysis of the heat shock response of Shewanella oneidensis. J Bacteriol 186:7796–7803
Gao H, Yang ZK, Wu L, Thompson DK, Zhou J (2006) Global transcriptome analysis of the cold shock response of Shewanella oneidensis MR-1 and mutational analysis of its classical cold shock proteins. J Bacteriol 188:4560–4569
Gehrke T, Hallmann R, Kinzler K, Sand W (2001) The EPS of Acidithiobacillus ferrooxidans—a model for structure-function relationships of attached bacteria and their physiology. Water Sci Technol 43:159–167
Gholami RM, Borghei SM, Mousavi SM (2011) Bacterial leaching of a spent Mo–Co–Ni refinery catalyst using Acidithiobacillus ferrooxidans and Acidithiobacillus thiooxidans. Hydrometallurgy 106:26–31
Gutknecht J, Walter A (1981) Hydrofluoric and nitric acid transport through lipid bilayer membranes. BBA Bioenergetics 644:153–156
Helmann JD, Wu MFW, Kobel PA, Gamo F-J, Wilson M, Morshedi MM, Navre M, Paddon C (2001) Global transcriptional response of Bacillus subtilis to heat shock. J Bacteriol 183:7318–7328
Jin D, Kong X, Li Y, Bai Z, Zhuang G, Zhuang X, Deng Y (2015) Biodegradation of di-n-butyl phthalate by Achromobacter sp. isolated from rural domestic wastewater. Int J Env Res Public Health 12:13510–13522
Kates M (1996) Structural analysis of phospholipids and glycolipids in extremely halophilic archaebacteria. J Microbiol Methods 25:113–128. doi:10.1016/0167-7012(96)00010-3
Ko M-S, Park H-S, Kim K-W, Lee J-U (2013) The role of Acidithiobacillus ferrooxidans and Acidithiobacillus thiooxidans in arsenic bioleaching from soil. Environ Geochem Health 35:727–733
Kruger MC, Bertin PN, Heipieper HJ, Arsène-Ploetze F (2013) Bacterial metabolism of environmental arsenic—mechanisms and biotechnological applications. Appl Microbiol Biotechnol 97:3827–3841
Lee J-U, Kim S-M, Kim K-W, Kim IS (2005) Microbial removal of uranium in uranium-bearing black shale. Chemosphere 59:147–154. doi:10.1016/j.chemosphere.2004.10.006
Li Q, Ding D, Sun J, Wang Q, Hu E, Shi W, Ma L, Guo X, Liu X (2014) Community dynamics and function variation of a defined mixed bioleaching acidophilic bacterial consortium in the presence of fluoride. Ann Microbiol 65:121–128. doi:10.1007/s13213-014-0843-x
Li Q, Ren Y, Qiu G, Li N, Liu H, Dai Z, Fu X, Shen L, Liang Y, Yin H (2011) Insights into the pH up-shift responsive mechanism of Acidithiobacillus ferrooxidans by microarray transcriptome profiling. Folia Microbiol 56:439–451
Liang Y, Van Nostrand JD, Wang J, Zhang X, Zhou J, Li G (2009) Microarray-based functional gene analysis of soil microbial communities during ozonation and biodegradation of crude oil. Chemosphere 75:193–199
Luo H, Shen L, Yin H, Li Q, Chen Q, Luo Y, Liao L, Qiu G, Liu X (2009) Comparative genomic analysis of Acidithiobacillus ferrooxidans strains using the A. ferrooxidans ATCC 23270 whole-genome oligonucleotide microarray. Can J Microbiol 55:587–598. doi:10.1139/W08-158
Ma LY, Li Q, Xiao YH, Wang QL, Yin HQ, Liang YL, Qiu GZ, Liu XD (2013) Comparative study of fluoride-tolerance of five typical bioleaching microorganisms. Adv Mat Res 825:214–218
Marquis RE, Clock SA, Mota-Meira M (2003) Fluoride and organic weak acids as modulators of microbial physiology. FEMS Microbiol Rev 26:493–510
Matin A, Rittenberg SC (1971) Enzymes of carbohydrate metabolism in Thiobacillus species. J Bacteriol 107:179–186
Pandey B (2013) Microbial processing of apatite rich low grade Indian uranium ore in bioreactor. Bioresour Technol 128:619–623
Parsonage D, Singh P, Nikoloski AN (2014) Adverse effects of fuoride on hydrometallurgical operations. Miner Process Extr Metall Rev 35:44–65
Peng ZJ, Yu RL, Qiu GZ, Qin WQ, Gu GH, Wang QL, Li Q, Liu XD (2013) Really active form of fluorine toxicity affecting Acidithiobacillus ferrooxidans activity in bioleaching uranium. T Nonferr Metal Soc 23:812–817. doi:10.1016/s1003-6326(13)62533-9
Pham ALT, Lee C, Doyle FM, Sedlak DL (2009) A silica-supported iron oxide catalyst capable of activating hydrogen peroxide at neutral pH values. Environ Sci Technol 43:8930–8935
Qihou L, Nuo L, Xueduan L, Zhijun Z, Qian L, Yun F, Xiangru F, Xian F, Yi L, Huaqun Y (2012) Characterization of the acid stress response of Acidithiobacillus ferrooxidans ATCC 23270 based on the method of microarray. J Biol Res 17:3–15
Qiu G, Li Q, Yu R, Sun Z, Liu Y, Chen M, Yin H, Zhang Y, Liang Y, Xu L (2011) Column bioleaching of uranium embedded in granite porphyry by a mesophilic acidophilic consortium. Bioresour Technol 102:4697–4702
Quatrini R, Jedlicki E, Holmes DS (2005) Genomic insights into the iron uptake mechanisms of the biomining microorganism Acidithiobacillus ferrooxidans. J Ind Microbiol Biotechnol 32:606–614
Raddadi N, Cherif A, Daffonchio D, Neifar M, Fava F (2015) Biotechnological applications of extremophiles, extremozymes and extremolytes. Appl Microbiol Biotechnol 99:7907–7913
Rashidi A, Roosta-Azad R, Safdari S (2014) Optimization of operating parameters and rate of uranium bioleaching from a low-grade ore. J Radioanal Nucl Chem 301:341–350
Rawlings DE, Johnson DB (2007) The microbiology of biomining: development and optimization of mineral-oxidizing microbial consortia. Microbiology 153:315–324
Razzell W, Trussell P (1963) Isolation and properties of an iron-oxidizing Thiobacillus. J Bacteriol 85:595–603
Rodrigues MLM (2015) Biolixiviação de cobre com micro-organismos mesófilos e termófilos moderados: sulfetos secundários contendo flúor e placas de circuito impresso. Universidade Federal de Ouro Preto, Ouro Preto
Silverman MP, Lundgren DG (1959) Studies on the chemoautotrophic iron bacterium Ferrobacillus ferrooxidans: I. An improved medium and a harvesting procedure for securing high cell yields. J Bacteriol 77:642
Smoot LM, Smoot JC, Graham MR, Somerville GA, Sturdevant DE, Migliaccio CAL, Sylva GL, Musser JM (2001) Global differential gene expression in response to growth temperature alteration in group A Streptococcus. Proc Natl Acad Sci USA 98:10416–10421
Stepanauskas R, Glenn TC, Jagoe CH, Tuckfield RC, Lindell AH, McArthur J (2005) Elevated microbial tolerance to metals and antibiotics in metal-contaminated industrial environments. Environ Sci Technol 39:3671–3678
Stintzi A (2003) Gene expression profile of Campylobacter jejuni in response to growth temperature variation. J Bacteriol 185:2009–2016
Sun J, Feng X, Liang D, Duan Y, Lei H (2011) Down-regulation of energy metabolism in Alzheimer’s disease is a protective response of neurons to the microenvironment. J Alzheimer’s Dis 28:389–402
Suzuki I, Lee D, Mackay B, Harahuc L, Oh JK (1999) Effect of various ions, pH, and osmotic pressure on oxidation of elemental sulfur by Thiobacillus thiooxidans. Appl Environ Microbiol 65:5163–5168
Valdes J, Pedroso I, Quatrini R, Dodson RJ, Tettelin H, Blake RC, Eisen JA, Holmes DS (2008) Acidithiobacillus ferrooxidans metabolism: from genome sequence to industrial applications. BMC Genom 9:597
Veloso TC, Sicupira LC, Rodrigues IC, Silva LA, Leão VA (2012) The effects of fluoride and aluminum ions on ferrous-iron oxidation and copper sulfide bioleaching with Sulfobacillus thermosulfidooxidans. Biochem Eng J 62:48–55
Vera M, Schippers A, Sand W (2013) Progress in bioleaching: fundamentals and mechanisms of bacterial metal sulfide oxidation—part A. Appl Microbiol Biotechnol 97:7529–7541
Wang Q, Qiu G (2011) Study on bacteria domestication and application of heap leaching in uranium mine. In: Remote sensing, environment and transportation engineering (RSETE), International Conference on, 2011 IEEE, pp 8522–8525
Watkin E, Zammit C (2016) Adaption to extreme acidity and osmotic stress. In: Quatrini R, Johnson D (eds) Acidophiles: life in extremely acidic environments. Caister Academic Press, Norfolk
Wen JK, Chen BW, Shang H, Zhang GC (2016) Research progress in biohydrometallurgy of rare metals and heavy nonferrous metals with an emphasis on China. Rare Met 35:433–442. doi:10.1007/s12598-016-0739-y
Xiong J, He Z, Van Nostrand JD, Luo G, Tu S, Zhou J, Wang G (2012) Assessing the microbial community and functional genes in a vertical soil profile with long-term arsenic contamination. PLoS One 7:e50507
Xu M, Zhang Q, Xia C, Zhong Y, Sun G, Guo J, Yuan T, Zhou J, He Z (2014) Elevated nitrate enriches microbial functional genes for potential bioremediation of complexly contaminated sediments. ISME J 8:1932–1944
Xu Y, Yin H, Jiang H, Liang Y, Guo X, Ma L, Xiao Y, Liu X (2013) Comparative study of nickel resistance of pure culture and co-culture of Acidithiobacillus thiooxidans and Leptospirillum ferriphilum. Arch Microbiol 195:637–646
Acknowledgment
This work was supported by the National Natural Science Foundation of China (NSFC 31570113) and Fundamental Research Funds for the Central Universities of Central South University (2016zzts110).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10295_2016_1827_MOESM1_ESM.tif
Fig. S1 Differentially expressed genes grouped by functional classification according to the TIGR A. ferrooxidans ATCC 23270 genome database. Columns: a biosynthesis of cofactors, prosthetic groups, and carriers; b central intermediary metabolism; c mobile and extrachromosomal element functions; d amino acid biosynthesis; e cell envelope; f cellular processes; g DNA metabolism; h energy metabolism; i fatty acid and phospholipid metabolism; j protein fate; k protein synthesis; l purines, pyrimidines, nucleosides, and nucleotides; m regulatory functions; n signal transduction; o transcription; p transport and binding proteins; q hypothetical Proteins-Conserved; r hypothetical Proteins; s unknown functions (TIFF 335 kb)
10295_2016_1827_MOESM2_ESM.tif
Fig. S2 The heatmap exhibits the four co-expression clusters of 22 genes. Red and blue represent co-expression values as 1 and −1, respectively. Genes: 1 transglutaminase-like domain protein (AFE0202); 2 conserved hypothetical protein (AFE0381); 3 ATP-dependent DNA helicase RecQ (AFE0903); 4 conserved hypothetical protein (AFE1091); 5 PIN domain protein (AFE0126); 6 hypothetical protein (AFE1867); 7 transcriptional regulator, GntR family (AFE0811); 8 hypothetical protein (AFE1300); 9 drug resistance transporter, EmrB/QacA family (AFE0913); 10 hypothetical protein (AFE0677); 11 glycogen synthase (AFE0427); 12 transposon, transposition protein B, putative (AFE0597); 13 amino acid permease family protein (AFE0645); 14 hypothetical protein (AFE0868); 15 drug resistance transporter, EmrB/QacA family (AFE2124); 16 PQQ enzyme repeat domain protein (AFE0847); 17 GTP-binding protein (AFE0926); 18 xylulose-5-phosphate/fructose-6-phosphate phosphoketolase (AFE1041); 19 outer membrane toxin secretion efflux protein, putative (AFE0929); 20 major facilitator family transporter (AFE0821); 21 hypothetical protein (AFE2209); 22 ubiquinol-cytochrome c reductase, cytochrome b subunit (AFE0375) (TIFF 1147 kb)
Rights and permissions
About this article
Cite this article
Ma, L., Li, Q., Shen, L. et al. Insights into the fluoride-resistant regulation mechanism of Acidithiobacillus ferrooxidans ATCC 23270 based on whole genome microarrays. J Ind Microbiol Biotechnol 43, 1441–1453 (2016). https://doi.org/10.1007/s10295-016-1827-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-016-1827-6