Abstract
Chloroplast-localized NAD kinase (NADK2) is responsible for the production of NADP+, which is an electron acceptor in the linear electron flow of photosynthesis. The Arabidopsis T-DNA-inserted mutant of NADK2 (nadk2) showed delayed growth and pale-green leaves under continuous light conditions. Under short-day conditions (8 h light / 16 h dark), the nadk2 mutant showed more severe growth inhibition.The genomic fragment containing the promoter and coding region of NADK2 complemented the phenotypes of nadk2 obtained under continuous light and short-day conditions. The nadk2 mutant produced higher amounts of H2O2 and O2−, which were reduced in the complementary line. Under short-day conditions, the nadk2 mutant accumulated more H2O2 than under continuous light conditions. The accumulation of ascorbate and up-regulation of the PDF1.2 and PR1 genes indicated that the nadk2 mutant is under ROS stress and responding to keep its living activities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
NAD(P)(H) are ubiquitous electron mediators that are involved in a variety of metabolic processes as they reversibly transfer electrons between the oxidized (NAD+, NADP+) and reduced (NADPH, NADH) forms in all organisms. NAD(H) is involved mainly in intracellular catabolic reactions, while NADP(H) is involved in anabolic reactions and as a defense against oxidative stress (Gakière et al. 2018; Pollak et al. 2007, Ziegler et al. 2000). NAD kinase (NADK) is the NADP+ biosynthetic enzyme that regulates the balance between NAD(H) and NADP(H) (Ohashi et al. 2011). NADK catalyzes the formation of NADP+ from NAD+ and ATP through phosphorylation (Hashida et al. 2009; Pétriacq et al. 2013, 2016).
Arabidopsis thaliana (L.) Heynh. (Arabidopsis) has three types of NADK (NADK1, NADK2 and NADK3) (Berrin et al. 2005; Turner et al. 2005). NADK1 is localized in the cytosol (Chai et al. 2006), NADK3 is reported to be localized in the peroxisomal matrix, and both NADK1 and NADK3 are involved in the oxidative stress response (Chai et al. 2006). NADK2 is a chloroplast-localized enzyme and is known to play a vital role in energy transduction through the photosynthetic electron transport chain (Chai et al. 2005; Takahashi et al. 2006; Turner et al. 2004). A novel NAD kinase, NADK C, was recently identified in Arabidopsis (Dell’ Aglio et al. 2019); NADK C is calmodulin/calcium dependent, is associated with the mitochondrial membrane and participates in oxidative bursts in response to attacks by pathogens.
In chloroplasts, the NADP+ produced by NADK2 accepts the electrons at the last step of photosystem I (PSI), and its reduced form NADPH, participates in carbon fixation in the Calvin cycle;
NADK2 is thus very important in photosynthesis. In a previous study, we demonstrated that alteration of the NADP+/NAD+ balance in rice and Arabidopsis affected the metabolism of these plants (Takahara et al. 2010; Takahashi et al. 2009). Specifically, rice overexpressing NADK2 showed an increased NADP+/NAD+ ratio, increased resistance to oxidative stress, and accumulation of certain Calvin cycle metabolites and amino acids (Kawai-Yamada et al. 2021; Onda et al. 2014; Takahara et al. 2010; Takahashi et al. 2009). Conversely, an NADK2 knockout mutant (nadk2) showed growth inhibition and produced smaller rosette leaves with a pale green color due to the reduced chlorophyll content (Chai et al. 2005). Our previous study demonstrated that photosynthetic activity in nadk2 was affected, as revealed by chlorophyll fluorescence analysis (Takahashi et al. 2006). Recently, Ji et al (2022) reported that photoinhibition susceptibility was increased in the nadk2 mutant. They demonstrated that a reduction in the translation activities of psaA and psaB caused a deficiency in the PSI complex in the nadk2 mutant.
As a photosynthetic organelle, the chloroplast has the potential to produce large amounts of reactive oxygen species (ROS), such as superoxide anions (O2−) and hydrogen peroxide (H2O2). The uncontrolled production of ROS can disrupt a wide variety of important molecular processes in plant cells (Apel and Hier 2004; Noctor et al. 2006). We hypothesized that ROS generated in the nadk2 mutant disrupted the photochemical system, and that this may be one of the factors affecting plant growth. In the present study, we first generated a complementary line of the nadk2 mutant to comprehensively evaluate the phenotype. We analyzed ROS generation in nadk2 to determine whether this mutant is under ROS stress.
Materials and methods
Plant materials and growth conditions
Arabidopsis ecotype Columbia was used as the wild type in this study. The nadk2 mutant was obtained from the T-DNA Express Collection at the Salk Institute Genomic Analysis Laboratory (http://signal.salk.edu) as described in Takahashi et al. (2006). Arabidopsis seeds were sown directly in Jiffy 7 (Jiffy Products International BV) and grown under CL (70 µmol m−2 s−1), SD conditions (8 h light/16 h dark, 70 µmol m−2 s−1), or long-day conditions (LD, 16 h light/8 h dark, 70 µmol m−2 s−1) at 23 °C. To measure the ROS in the nadk2 plants under SD conditions, the plants that were grown under CL for 2 weeks and transferred to SD conditions were also used. The gnadk line (nadk2 which includes a NADK2 genomic fragment containing 2 kb of the promoter region) was produced in this study. The genome fragment of NADK was amplified using the following primer set; forward, 5′-CCTGACAACGGCAAGGTAAACCATT-3′; reverse, 5′-AACCGTTTCTCGAGTTGTTTCTCTC-3′. The amplified fragment was cloned into the pHG plasmid containing a hygromycin resistant gene as a selection marker. The resultant plasmid (pHG-gnadk) was transformed to the nadk2 using an Agrobacterium-mediated method.
For the confirmation of WT, nadk2 and gnadk plants, genomic PCR and RT-PCR were performed using the following primers; AtNADK2-S2: 5′-GGCTTCTCTGCAGCCCCTATTGCTGTGCC-3′; AtNADK2-A2: 5′-GACTCGTTTGAGGTCTTGCCTGAAGTCCT-3′; LB: 5′-GCAAACCAGCGTGGACCGCTTGCTGCAAC-3′; Actin8 (forward, 5′-TGAGCCAGATCTTCATCGTC-3′, reverse, 5′-TCTCTTGCTCGTAGTCGACA-3′).
Measurement of chlorophyll contents
The chlorophyll (a + b) contents were measured spectrophotometrically. After elution of chlorophyll in N, N-dimethylformamide overnight, absorbance was measured at wavelengths of 647 nm and 664 nm using a UV–Vis spectrophotometer (Pharmacia Biotech Ultrospec® 3000 CT, USA). The pigment concentrations were calculated as described in Ceusters et al. (2019).
Assays of NAD kinase (NADK) activity and NAD(P)(H) contents
Leaves (20–30 mg) in 200 µL of protein extraction buffer (2.5 mM HEPES/KOH pH 8.0, 0.1 mM MgCl2, 0.001 mM CaCl2, 0.001 mM PMSF, 10% protease inhibitor [cOmplete™, Mini, Protease Inhibitor Cocktail, Roche]) were homogenized using a handy homogenizer (HOMOGENIZER S-203, IKEDA RIKA, Japan) and sonicated (5 s × 3 times) on ice. After centrifugation at 15,000 rpm for 10 min at 4 °C, the supernatant was transferred to new tubes, and the protein content was measured using a Bradford protein assay (Bio-Rad Laboratories, USA). NADK activity was measured using a discontinuous assay involving a cycling assay for NADP+, as described by Ishikawa et al. (2016; 2020).
For the NAD(P)(H) measurements, leaves (20–30 mg) were boiled in 200 µL of 0.2 N HCl (extraction of NAD+ and NADP+) and 0.2 N NaOH (extraction of NADH and NADPH) for 2 min. After homogenization with a handy homogenizer (IKEDA RIKA HOMOGENIZER S-203) and sonication (5 s × 3 times) on ice, samples were centrifuged at 15,000 rpm for 10 min at 4 °C. The supernatants were neutralized by adding 15 µL of 0.2 M NaH2PO4 (pH 5.6) and 120 µL of 0.2 N NaOH for NAD+ and NADP+, or adding 15 µL of 0.2 M HEPES/KOH pH8.0 and 120 µL of 0.2 N HCl for NADH and NADPH. The NAD(P)(H) contents were measured using a cycling assay, as described by Ishikawa et al. (2016; 2020).
Determination of H2O2 contents
Arabidopsis leaves (120–200 mg) were frozen in liquid nitrogen, crushed with a mortar and pestle, and extracted using 1 mL of 50 mM phosphate buffer (pH 7.0) supplemented with 10 mg of polyvinylpyrrolidone (PVP). The extract was centrifuged at 6,000 × g for 25 min at 4 °C, and the supernatant was collected, centrifuged at 6,000 × g for 25 min at 4 °C. The supernatant (300 μL) was added to 1 mL of titanium sulfate solution containing 1% titanium sulfate and 20% sulfuric acid (v/v). The mixture was centrifuged at 6,000 × g for 15 min at room temperature and hydrogen peroxide concentrations were determined by measuring the absorbance at 410 nm (Velikova et al. 2000). Calibration curves were obtained by adding several concentrations of H2O2 to the titanium sulfate solution.
ROS staining
For 3,3′-diaminobenzidine (DAB) staining, the Arabidopsis plants were cut anad placed in 1 mg mL−1 3,3′-diaminobenzidine (DAB)-HCl, pH 3.8 (Sigma, MO, USA; # D-8001) (Thordal‐Christensen et al. 1997). Leaves were cleared by boiling in acetic acid/glycerol/ethanol (1:1:3[v/v/v]) solution for 5 min. Material was mounted on a glass slide in 60% glycerol for observation and H2O2 was detected as.
reddish-brown coloration.
NBT staining was performed to detect O2−,
Arabidopsis leaves were vacuum infiltrated with 10 mM NaN3 in 10 mM potassium phosphate buffer (pH 7.8) and incubated in 0.1% NBT (in 10 mM potassium phosphate buffer (pH 7.8) for 20 min at room temperature and cleared by boiling in acetic acid/glycerol/ethanol (1:1:3[v/v/v]) solution for 5 min. Material was mounted on a glass slide in 60% glycerol for observation.
To quantify the O2−, the blue formazan particles in the tissues were dissolved in 2 M potassium hydroxide and dimethylsulfoxide, and absorbance at 620 nm was measured using a UV–Vis spectrophotometer (Pharmacia Biotech Ultrospec® 3000 CT, USA) (Sim Choi et al. 2006).
Ascorbate contents
Leaves (50 mg-100 mg) harvested from 14 to day-old plants after germination, were rapidly frozen in liquid nitrogen and ground and homogenized in 0.3 mL / 100% (v/v) methanol containing 100 μM 1, 4-piperazine diethane sulfonic acid and 100 μM methionine sulfone as the internal standard. After centrifugation (15,000 rpm, 5 min at 4 °C), the supernatants were transferred to a 3 kDa Amicon ® Ultra Centrifugal Filter (Ultracel ®- 3 K, IRL). After centrifugation (12,000 rpm, 30 min at 4 °C), an aliquot of 13 μL of the resulting filtrate was used for metabolite analysis by capillary electrophoresis mass spectrometry (CE- MS/MS) (CE: G1600AX, MS: G1965B, Agilent Technologies, Waldbronn, Germany) according to the method of Miyagi et al. (2010).
Catalase activity
Catalase activity was determined using an extinction coefficient of 40 mM−1 cm−1 following the method of Abei (1984). The crude extracts of leaves were prepared using the same method as for the NADK activity assay. The reaction mixture contained 5 μL of 30% H2O2, 25 μL of 50 mM potassium phosphate buffer (pH 7.0) and 5 μL of enzyme extract dissolved in water to 500 μL, and the decrease in the absorbance at 240 nm was monitored. Enzyme activity was expressed as μmol min−1 g FW−1.
Quantitative real-time PCR (qRT-PCR)
Total RNA was isolated from Arabidopsis leaves using a RNeasy Plant Mini Kit (Qiagen, Venlo, The Netherlands), followed by DNase I treatment (Qiagen). The first strand cDNA was generated using reverse transcriptase and a random primer, according to the manufacturer’s instructions (Applied Biosystems™). qRT-PCR analysis was performed using the following primers: AT2G29350 (SAG13, forward, 5′-CAGCTTGCCCACCCATTGTTA-3′; reverse, 5′-GTCGTACGCACCGCTTCTTTC-3′), AT5G44420 (PDF1.2, forward, 5′-TAATCATCATGGCTAAGTTTGCTT-3′; reverse, 5′-ATACACACGATTTAGCACCAAAGA-3′), AT3G57260.1 (PR1, forward, 5′-AGTTTTGGGGACTGTTTCAT-3′; reverse, 5’-ATTTATGCTTGCAGCTTCAT-3′). qRT-PCR was then performed using Power SYBR Green PCR Master Mix (ABI Prism®) and the 7300 Real-Time PCR system (Applied Biosystems).
Results
Complementation of nadk2 mutant using a NADK2 genome fragment
Previous studies have reported that the nadk2 mutant shows delayed growth, but it has not yet been determined whether a genomic fragment complements this phenotype. Therefore, we first generated a complementary line of the nadk2 mutant. The results revealed that the nadk2 mutant has lost its function due to a T-DNA insertion in the NADK motif. which is required for enzyme activity (Fig. 1a). A complementary line (gnadk) was created in which the genomic fragment of NADK2 containing a promoter (~ 2 kb) and a coding region was introduced into the nadk2 mutant (Fig. 1b, c). Under continuous light (CL) conditions, the phenotype of the nadk2 mutant was characterized by having pale green leaves and slow growth; however, the mutant was capable of matureing and producing flowers and seeds, as described previously (Chai et al. 2005; Takahashi et al. 2006). However, under short-day (SD) conditions (8 h light/16 h dark), the nadk2 mutant could germinate, but could not develop any further. The complementary line (gnadk) showed similar growth to wild-type plants (Fig. 1d). To investigate whether the gnadk line completely recovered the nadk2 phenotype, we measured the fresh weight (Fig. 2a, b) and chlorophyll content of the plants (Fig. 2c, d). No data could be collected for the nadk2 mutants grown under SD conditions because the nadk2 mutant cannot develop further grow under the growth conditions employed after germination (Fig. 1d). As shown in Fig. 2a-d, under conditions of CL and SD, the gnadk and WT plants had comparable fresh weights and chlorophyll contents. In addition, the gnadk showed the same levels of NADK activity as the WT plants (Fig. 2e, f). The NAD(P)(H) measurements demonstrated that the nadk2 mutant showed decreased NADP+ levels, and increased NAD+ and NADPH levels compared to WT plants. As a result, the phosphorylation ratio (NADP+/NAD+) and redox reactions (NADH/NAD+) in the nadk2 mutant were decreased, but NADPH/NADP+ was increased compared to that of WT plants (Fig. 3). In contrast, the gnadk plants demonstrated recovery to the WT (Fig. 3). Under long-day conditions (16 h light/8 h dark), the nadk2 mutant showed better growth than under SD conditions, but it did not grow further and was unable to reach the flowering stage (Fig. S1).
Arabidopsis nadk2 mutant and the complementary line (gnadk). (a) Diagrammatic illustration of the Arabidopsis NADK2 gene showing the site of the T-DNA insertion. Gray boxes indicate the NADK-motif. PCR primers (LB, S2 and A2) used in (b) and (c) are shown as arrows. (b) Genomic PCR analysis of wild-type NADK2 (S2-A2 primers) and T-DNA inserted NADK2 (LB-A2) in wild-type (WT), nadk2 and a complementary line (gnadk). (c) RT-PCR analysis of NADK2 in WT, nadk2 and gnadk plants. The Actin8 gene was used as a control. (d) Plant phenotype of WT, nadk2, and gnadk grown for 28 days under continuous light (CL) and short-day (SD) conditions
Comparison of WT, nadk2 mutant, and gnadk plants. Fresh weight (a, b), chlorophyll contents (c, d) and NADK activity (e, f) were measured in plants grown under continuous light (CL) (a, c, e) or under short-day (SD) (b, d, f) conditions for 28 days. Leaves were harvested during the light period. n = 3, *p < 0.05, **p < 0.01, compared with WT (t-test), n.d. = not determined, Error bars indicate SD
Comparison of NAD(P)(H) contents. Leaves obtained from plants grown for 28 days under continuous light (CL) were used for NAD(P)(H) measurement. The phosphorylation ratio (b, c) and redox (d, e) reaction were calculated from (a). n = 3 *p < 0.05, **p < 0.01, compared with WT (t test). Error bars indicate SD
These results indicate that the phenotypes of the nadk2 plants were complemented by the introduction of a genomic fragment of NADK2. In addition, the 2 kb promoter used in this study was shown to be suitable for the correct functioning of NADK2.
nadk2 produced more H2 O 2 and O 2 − than WT plants
Since NADK2 is responsible for producing NADP+, which acts as an electron acceptor for photosynthetic electron transfer in the chloroplast, we hypothesized that the lack of electron acceptors in the nadk2 mutant would result in a surplus of electrons and the generation of ROS, which could inhibit the growth of the nadk2 mutant. Therefore, we examined H2O2 and O2− accumulation in WT, nadk2, and gnadk plants grown under CL conditions. As shown Fig. 4a, H2O2 was highly accumulated in the nadk2, and 3,3′-diaminobenzidine (DAB) staining also showed strong signal in the nadk2 mutant. In addition, we performed Nitro blue tetrazolium chloride (NBT) staining to detect O2−. The results showed strong staining in nadk2 mutants grown under CL conditions (Fig. 4b). Furthermore, the antioxidant ascorbate was also accumulated in the nadk2 plant, suggesting that nadk2 is under ROS stress (Fig. 4c).
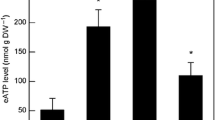
Reactive oxygen species (ROS) detection in WT, nadk2, and gnadk plants. a H2O2 measurement and 3, 3-Diaminobenzidine (DAB) staining were performed in plants grown under continuous light (CL) conditions for 28 days. Bars = 20 μm. b Nitro blue tetrazolium chloride (NBT) staining was performed in plants grown under CL conditions for 28 days. Stainability was quantified as described in the Materials and methods. Bars = 20 μm. c Ascorbate contents were measured in plants grown under CL conditions for 28 days. n = 3, *p < 0.05, **p < 0.01, compared with WT (t test). Error bars indicate SD
However, since nadk2 mutants do not grow at all under SD conditions as shown in Fig. 1d, it is not possible to measure ROS levels in these mutants. Therefore, we transferred plants grown for 2 weeks under CL conditions to SD conditions, and collected samples at the Light End (LE), Dark Initial (DI), Dark End (DE), and Light Initial (LI) to quantify H2O2. The results showed that nadk2 consistently accumulated large amounts of H2O2, especially under LE time point (Fig. 5a). H2O2 levels decreased during the dark period (DI and DE), but never reached the same levels as in WT plants. In addition, the catalase activity that decomposes H2O2 to water and oxygen showed a tendency to be increased in the nadk2 mutant (Fig. 5b). Comparison of H2O2 accumulation between CL and SD (LI) showed that H2O2 accumulated more under the SD conditions (Fig. S2). The amount of H2O2 detected under SD conditions was always higher than that detected under CL conditions, implying that this was one of the reasons for the significant inhibition in the growth of nadk2 mutants under SD conditions.
H2O2 contents a and catalase activity b under dark/light conditions. Plants grown under continuous light for 14 days were transferred to short-day conditions. After 6 days, H2O2 contents and catalase activity were measured at LE (Light End), DI (Dark Initial), DE (Dark End), and LI (Light Initial) time points. n = 3, *p < 0.05, **p < 0.01, compared with WT (t test). Error bars indicate SD
To determine whether the nadk2 mutant is under ROS stress, expression levels of several ROS-related genes were analyzed by qRT-PCR. Among the examined genes, plant defensin 1.2 (PDF1.2) and pathogenesis-related 1 (PR1) showed up-regulation in the nadk2 mutant, but senescence-associated gene 13 (SAG13) and others (WRKY70, DND1, and SGT1b) did not (Fig. 6, Fig. S3). These results suggest that the PR1 and PDF1.2-related H2O2 stress responses are activated in nadk2, and that these responses are not related to senescence or cell death response involving other genes (SAG13, WRKY70, DND1 and SGT1b).
qRT-PCR analysis of the plant defensin 1.2 (PDF1.2), pathogenesis-related 1 (PR1) gene and senescence-associated gene 13 (SAG13) in leaves obtained from plants grown under continuous light conditions for 28 days. The transcript levels of PDF1.2, PR1 and SAG13 were compared among the WT, nadk2 mutant and gnadk plants. The expression levels were normalized to Actin8. n = 3 *p < 0.05, **p < 0.01
Discussion
NADK2 has been demonstrated to play important roles in the stress response and in controlling cellular metabolism in Arabidopsis (Hashida et al. 2009; Takahashi et al. 2009). The NADP+ produced mediates photosynthetic energy transfer as the final electron acceptor and promotes anabolism to support plant growth (Hashida et al. 2009). In addition to being used for carbon and nitrogen assimilation, and lipid and chlorophyll metabolism, NADPH provides the reducing power, and functions to maintain redox homeostasis by regulating the scavenging of ROS in plant cells (Noctor 2006). The nadk2 mutant has been demonstrated to have a broad range of phenotypes, including photosynthetic defects, leaf color variations, metabolic changes and growth retardation (Chai et al. 2005; Ji et al. 2022; Kawai-Yamada et al. 2021; Takahashi et al. 2006, 2009).
For the first time, a complementary experiment using a genomic fragment was conducted in this study. In a recent study, Ji et al. (2022) reported that the expression of NADK2 cDNA under the control of CaMV 35S promoter was capable of mitigating growth defects in nadk2. Our results indicate that the nadk2 phenotype was complemented by the introduction of a genomic fragment of NADK2. In addition, the 2 kb promoter used in this study was shown to be effective for the correct functioning of NADK2.
In the nadk2 mutant, a decrease in ΦII (a parameter indicating the efficiency of photosynthetic electron transport) has been reported (Takahashi et al. 2006). Ji et al. (2022) suggested that PSI was reduced in the nadk2 mutant in response to disrupted NAD(P)(H) balance. The present study also showed that a decrease in NADP+ and increase in NAD+ and NADPH contents in the nadk2 mutant affect phosphorylation ratios (NADP+/NAD+) and redox reactions (NADH/NAD+, NADPH/NADP+), and disrupt the photosynthetic electron transport chain.
When plants are exposed to environmental stress, ROS are produced as metabolic byproducts. Abiotic stresses, such as cold, heat, salt, and drought, as well as biotic stresses generate ROS (Edreva 2005; Foyer and Noctor 2005; Neill et al. 2002). Photosynthesis is a major source of ROS, such as H2O2, O2−, singlet oxygen (1O2) and hydroxyl radicals (-OH), which generate oxygen molecules and the reducing power through the decomposition of water by light energy (Maruta et al. 2016). As shown in Fig. 4, the nadk2 mutant accumulated considerable quantities of H2O2 and O2− under CL conditions. Since nadk2 cannot grow under SD conditions, we developed a system to transfer plants grown under CL conditions for 2 weeks to SD conditions. Under these dark/light growth conditions, high concentrations of H2O2 were consistently generated in nadk2 mutants (Fig. 5), suggesting that the nadk2 mutant is under ROS stress. In plant cells, NADP+ is increased under light conditions, and decreased under dark conditions. Hashida et al. (2018) demonstrated that the NADP+ content of the nadk2 mutant is always low, and that there are no quantitative fluctuations over time. The inability to quantitatively regulate NADP+ under light and dark conditions may be related to the fact that more ROS are produced under dark/light conditions and nadk2 causes more severe growth inhibition.
Since plant cells are continuously exposed to ROS, ROS scavenging mechanisms in different organelles play key roles in cell survival. The chloroplast, one of the major sources of ROS, has evolved numerous antioxidative enzymes, including catalases, monodehydroascorbate reductase, dehydroascorbate reductase, glutathione S-transferase, glutathione reductase, superoxide dismutase, ascorbate peroxidase and NADPH thioredoxin reductase. All of these enzymes act synergistically in ROS scavenging in plants, and most utilize NADPH as a reducing energy source, either directly or indirectly (Mittler 2002). As shown in Fig. 3, the amount of NADPH in whole cells is not reduced compared to WT plants, but the NADP(H) pool size (NADP+ + NADPH) was reduced and the NAD(P)(H) balance was disrupted. Since there are isoforms (NADK1, NADK3 and NADK C) with different intracellular localizations in the cells, it is possible that they are activated in different cellular locations in cases of NADK2 deficiency. It is also possible that the ROS elimination system does not work well due to disruption of the NAD(P)(H) balance at the organelle level.
Furthermore, we investigated expression levels of certain ROS-related genes in nadk2 mutants. The PDF1.2 gene, which is related to the defense response to biotrophic and hemi biotrophic microorganisms, has been reported to be upregulated by ROS, and to play a central role in the induction of cellular death and pathogen confinement (Alvarez et al. 1998; Mukherjee et al. 2010). The defense machinery of plants has been forced to evolve continuously to combat a wide range of abiotic and biotic stress factors. These responses typically involve a series of events including the production of ROS and the synthesis of antimicrobial molecules and PR proteins. PR proteins, which induce programmed cell death, in order to inhibit the spread of infection, also contribute to systemic acquired resistance (Chassot et al. 2007; Van Baarlen et al. 2007). Increased expression of PDF1.2 and PR1 indicate that nadk2 is under ROS stress, and that it is responding to ROS-related signals. However, although the leaves of nadk2 plants are pale green in color, they continue to grow under CL conditions and cell death does not occur. In addition, the senescence marker gene SAG13, ROS-related transcription factor (WRKY70), cell death regulator (DND1), and hormone regulator (SGT1b) were not upregulated in nadk2 plants (Fig. 6, Fig. S3). Based on these results, it is suggested that the physiological phenomena occurring in nadk2 differ from typical senescence or cell death. Chai et al. (2005) reported that the nadk2 mutants accumulate considerably more pchlide and Mg-protoporphyrin IX than WT plants. Because the conversion of pchlide to chlide is dependent upon NADPH as a reducing agent, the lack of NADK2 may affect chlorophyll biosynthesis and be responsible for the pale green color of the leaves in these plants. The growth inhibition observed in the nadk2 mutant may therefore be the result of a combination of multiple defects in photosynthetic electron transport, ROS scavenging systems, and chlorophyll biosynthesis.
Conclusion
In conclusion, the growth retardation of nadk2 was restored in a gnadk line in which the genomic fragment of NADK2 was introduced. These findings indicate that the phenotypes observed in nadk2 plants grown under CL or SD conditions were caused by the loss of NADK2. The ROS analysis showed that the accumulation of H2O2 under CL and SD conditions may be responsible for the growth inhibition of nadk2 mutants. Furthermore, catalase activity tended to be increased at all stages, and ascorbate increased significantly in nadk2. These findings show that nadk2 is under ROS stress and is responding and trying to adapt.
Data availability
No new datasets were generated or analyzed in this study.
Abbreviations
- DAB 3:
-
3-Diaminobenzidine
- H2O2 :
-
Hydrogen peroxide
- NADP:
-
Nicotinamide adenine dinucleotide phosphate
- NAD+ :
-
Nicotinamide adenine dinucleotide
- NBT:
-
Nitro blue tetrazolium chloride
- ROS:
-
Reactive oxygen species
- WT:
-
Wild type
References
Aebi H (1984) Catalase in vitro. Methods in Enzymology, vol. 105. Elsevier, pp 121–126
Alvarez MaE, Pennell RI, Meijer P-J, Ishikawa A, Dixon RA, Lamb C (1998) Reactive oxygen intermediates mediate a systemic signal network in the establishment of plant immunity. Cell 92:773–784
Apel K, Hier H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Berrin J-G, Pierrugues O, Brutesco C, Alonso B, Montillet J-L, Roby D, Kazmaier M (2005) Stress induces the expression of AtNADK-1, a gene encoding a NAD (H) kinase in Arabidopsis thaliana. Mol Genet Genom 273:10–19
Ceusters N, Valcke R, Frans M, Claes JE, Van den Ende W, Ceusters J (2019) Performance index and PSII connectivity under drought and contrasting light regimes in the CAM orchid Phalaenopsis. Front Plant Sci 10:1012
Chai M-F, Chen Q-J, An R, Chen Y-M, Chen J, Wang X-C (2005) NADK2, an Arabidopsis chloroplastic NAD kinase, plays a vital role in both chlorophyll synthesis and chloroplast protection. Plant Mol Biol 59:553–564
Chai MF, Wei PC, Chen QJ, An R, Chen J, Yang S, Wang XC (2006) NADK3, a novel cytoplasmic source of NADPH, is required under conditions of oxidative stress and modulates abscisic acid responses in Arabidopsis. Plant J 47:665–674
Chassot C, Nawrath C, Métraux JP (2007) Cuticular defects lead to full immunity to a major plant pathogen. Plant J 49:972–980
Dell’Aglio E, Giustini C, Kraut A, Couté Y, Costa A, Decros G (2019) Identification of the Arabidopsis calmodulin-dependent NAD+ kinase that sustains the elicitor-induced oxidative burst. Plant Physiol 181:1449–1458
Edreva A (2005) Generation and scavenging of reactive oxygen species in chloroplasts: a submolecular approach. Agric Ecosyst Environ 06:119–133
Foyer CH, Noctor G (2005) Redox homeostasis and antioxidant signaling: a metabolic interface between stress perception and physiological responses. Plant Cell 17:1866–1875
Gakière B, Fernie AR, Pétriacq P (2018) More to NAD+ than meets the eye: a regulator of metabolic pools and gene expression in Arabidopsis. Free Radic Biol Med 122:86–95
Hashida S-N, Takahashi H, Uchimiya H (2009) The role of NAD biosynthesis in plant development and stress responses. Ann Bot 103:819–824
Hashida S-N, Miyagi A, Nishiyama M, Yoshida K, Hisabori T, Kawai-Yamada M (2018) Ferredoxin/thioredoxin system plays an important role in the chloroplastic NADP status of Arabidopsis. Plant J 95:947–960
Ishikawa Y, Kawai-Yamada M, Hashida S-N (2020) Measurement of chloroplastic NAD kinase activity and whole tissue NAD kinase assay. Bio-Protoc 10:e3480–e3480
Ishikawa Y, Miyagi A, Haishima Y, Ishikawa T, Nagano M, Yamaguchi M, Hihara Y, Kawai-Yamada M (2016) Metabolomic analysis of NAD kinase-deficient mutants of the cyanobacterium Synechocystis sp. PCC 6803. J Plant Physiol 205:105–112
Ji D, Li Q, Guo Y, An W, Manavski N, Meurer J, Chi W (2022) NADP+ supply adjusts the synthesis of photosystem I in Arabidopsis chloroplasts. Plant Physiol 189:2128–2143
Kawai-Yamada M, Miyagi A, Sato Y, Hosoi Y, Hashida S-N, Ishikawa T, Yamaguchi M (2021) Altered metabolism of chloroplastic NAD kinase-overexpressing Arabidopsis in response to magnesium sulfate supplementation. Plant Signal Behav 16:1844509
Maruta T, Sawa Y, Shigeoka S, Ishikawa T (2016) Diversity and evolution of ascorbate peroxidase functions in chloroplasts: more than just a classical antioxidant enzyme? Plant Cell Physiol 57:1377–1386
Mittler R (2002) Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci 7:405–410
Miyagi A, Takahashi H, Takahara K, Hirabayashi T, Nishimura Y, Tezuka T, Kawai-Yamada M, Uchimiya H (2010) Principal component and hierarchical clustering analysis of metabolites in destructive weeds; polygonaceous plants. Metabolomics 6:146–155
Mukherjee M, Larrimore KE, Ahmed NJ, Bedick TS, Barghouthi NT, Traw MB, Barth C (2010) Ascorbic acid deficiency in Arabidopsis induces constitutive priming that is dependent on hydrogen peroxide, salicylic acid, and the NPR1 gene. Mol Plant Microbe Interact 23:340–351
Neill S, Desikan R, Hancock J (2002) Hydrogen peroxide signalling. Curr Opin Plant Biol 5:388–395
Noctor G (2006) Metabolic signalling in defence and stress: the central roles of soluble redox couples. Plant Cell Environ 29:409–425
Ohashi K, Kawai S, Koshimizu M, Murata K (2011) NADPH regulates human NAD kinase, a NADP+-biosynthetic enzyme. Mol Cell Biochem 355:57–64
Onda Y, Miyagi A, Takahara K, Uchimiya H, Kawai-Yamada M (2014) Effects of NAD kinase 2 overexpression on primary metabolite profiles in rice leaves under elevated carbon dioxide. Plant Biol 16:819–824
Penninckx I, Eggermont K, Terras FR, Thomma B, De Samblanx GW, Buchala A, Métraux J-P, Manners JM, Broekaert WF (1996) Pathogen-induced systemic activation of a plant defensin gene in Arabidopsis follows a salicylic acid-independent pathway. Plant Cell 8:2309–2323
Pétriacq P, de Bont L, Tcherkez G, Gakière B (2013) NAD: not just a pawn on the board of plant-pathogen interactions. Plant Signal Behav 8:e22477
Pétriacq P, Ton J, Patrit O, Tcherkez G, Gakière B (2016) NAD acts as an integral regulator of multiple defense layers. Plant Physiol 172:1465–1479
Pollak N, Dölle C, Ziegler M (2007) The power to reduce: pyridine nucleotides–small molecules with a multitude of functions. Biochem J 402:205–218
Sim Choi H, Woo Kim J, Cha YN, Kim C (2006) A quantitative nitroblue tetrazolium assay for determining intracellular superoxide anion production in phagocytic cells. J Immunoassay Immunochem 27:31–44
Takahara K, Kasajima I, Takahashi H, Hashida S-n, Itami T, Onodera H, Toki S, Yanagisawa S, Kawai-Yamada M, Uchimiya H (2010) Metabolome and photochemical analysis of rice plants overexpressing Arabidopsis NAD kinase gene. Plant Physiol 152:1863–1873
Takahashi H, Takahara K, Hashida S-N, Hirabayashi T, Fujimori T, Kawai-Yamada M, Yamaya T, Yanagisawa S, Uchimiya H (2009) Pleiotropic modulation of carbon and nitrogen metabolism in Arabidopsis plants overexpressing the NAD kinase2 gene. Plant Physiol 151:100–113
Takahashi H, Watanabe A, Tanaka A, Hashida S-n, Kawai-Yamada M, Sonoike K, Uchimiya H (2006) Chloroplast NAD kinase is essential for energy transduction through the xanthophyll cycle in photosynthesis. Plant Cell Physiol 47:1678–1682
Thordal-Christensen H, Zhang ZG, Wei YD, Collinge DB (2002) Subcellular localization of H2O2 in plants. H2O2 accumulation in papillae and hypersensitive response during the barley-powdery mildew interaction. Plant J 11:1187–1194
Turner WL, Waller JC, Snedden WA (2005) Identification, molecular cloning and functional characterization of a novel NADH kinase from Arabidopsis thaliana (thale cress). Biochem J 385:217–223
Turner WL, Waller JC, Vanderbeld B, Snedden WA (2004) Cloning and characterization of two NAD kinases from Arabidopsis. Identification of a calmodulin binding isoform. Plant Physiol 135:1243–1255
Van Baarlen P, Van Belkum A, Summerbell RC, Crous PW, Thomma BP (2007) Molecular mechanisms of pathogenicity: how do pathogenic microorganisms develop cross-kingdom host jumps? FEMS Microbiol Rev 31:239–277
Velikova V, Yordanov I, Edreva A (2000) Oxidative stress and some antioxidant systems in acid rain-treated bean plants: protective role of exogenous polyamines. Plant Sci 151:59–66
Ziegler M (2000) New functions of a long-known molecule: emerging roles of NAD in cellular signaling. Chem Eur J 267:1550–1564
Funding
This work was supported by KAKENHI Grant Numbers 19H04715 and 21H05647 to M. K-Y.
Author information
Authors and Affiliations
Contributions
MKY. and C. designed the study. C., YZ., AM., SNH, and MKY. performed the experiments. C., AM., TI., MY. and MKY. interpreted the data. C. and MKY. wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
No conflicts of interest declared.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chaomurilege, Zu, Y., Miyagi, A. et al. Loss of chloroplast-localized NAD kinase causes ROS stress in Arabidopsis thaliana. J Plant Res 136, 97–106 (2023). https://doi.org/10.1007/s10265-022-01420-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-022-01420-w