Abstract
The centromere and telomere are universal heterochromatic domains; however, the proper positioning of those domains in nuclear space during the mitotic interphase differs among eukaryotes. Consequently, the question arises how and why this difference occurs. Studies over the past 2 decades have identified several nuclear membrane proteins, nucleolar proteins, and the structural maintenance of a chromosome complex as factors involved in the positional control of centromeres and/or telomeres during the mitotic interphase in yeasts, animals, and plants. In this review, with a primary focus on plants, the roles of those factors are summarized, and the biological significance of proper centromere and telomere positionings during the mitotic interphase is discussed in an effort to provide guidance for this question.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
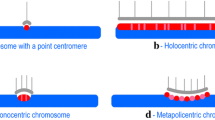
In eukaryotic genomes, chromatin is organized into functionally and structurally distinct domains, which are directly associated with genome functions, such as replication, repair, and transcription. Among the chromatin domains, the centromere and telomere are universal to all eukaryotic chromosomes. With regard to plants, as with other eukaryotes, some species show characteristic distributions of centromeres and telomeres in the mitotic interphase, in which the centromeres show a clustered distribution at one side of the nucleus whereas telomeres show a clustered distribution at the opposite side of the nucleus. Such a structure is termed the Rabl configuration (Dong and Jiang et al. 1998; Rabl 1885) (Fig. 1a). This chromosome configuration is seen in both monocots and dicots, such as Triticum aestivum, Hordeum vulgare, Avena sativa, and Vicia faba, whereas other species, such as Sorghum bicolar and Zea mays adopt a non-Rabl configuration with centromeres and telomeres more dispersed in the nucleus (Dong and Jiang et al. 1998; Santos and Shaw 2004) (Fig. 1c). A model plant, Arabidopsis thaliana, does not exhibit the Rabl configuration; instead, it shows a dispersion of centromeres at the entire nuclear periphery and a localization of telomeres at the nucleolar periphery (Fang and Spector 2005; Roberts et al. 2009) (Fig. 1d). Several studies, thus far, have indicated that whether the chromosomes adopt a non-Rabl configuration is not dependent on the genome size in plants (Table 1) (Matsunaga et al. 2013; Schubert and Shaw 2011). However, the type of configuration is considered to be likely dependent on the amount and genome organization of the repetitive regions, such as transposable elements, satellite DNAs, and ribosomal DNAs, that form heterochromatin (Santos and Shaw 2004; Schubert and Shaw 2011). For example, in A. thaliana, which shows a non-Rabl configuration, most of the heterochromatin is found around the centromere, whereas in T. aestivum, which exhibits a Rabl configuration, the genome predominantly consists of heterochromatin distributed throughout the chromosome arms (Santos and Shaw 2004). However, Z. mays, which shows a non-Rabl configuration, has a genome organization of heterochromatin similar to T. aestivum (Choulet et al. 2010). In addition, in Oryza sativa, most cells show a non-Rabl configuration (Dong and Jiang et al. 1998), but premeiotic anther cells and xylem-vessel precursor cells show a Rabl configuration (Santos and Shaw 2004). Therefore, the genome organization of heterochromatin is unlikely to be a definitive factor that determines the type of chromosome configuration. Thus, how the chromosomes acquire a non-Rabl configuration is still an open question for plants as well as other eukaryotes.
Distribution patterns of centromeres and telomeres in the Rabl (a, b) and the non-Rabl (c, d) configurations. a Clustered distributions of centromeres and telomeres seen in plants such as T. aestivum. b Clustering of all centromeres beneath the SPB seen in S. pombe and S. cerevisiae. c Dispersed distributions of centromeres and telomeres seen in plants such as S. bicolar. It should be noted that the peripheral distributions of centromeres and telomeres in plants which do not adopt a Rabl configuration are not clearly confirmed yet. d Dispersed centromere distribution around nuclear periphery and nucleolar peripheral distribution of telomeres in A. thaliana
Given that the Rabl configuration is the default chromosome configuration, as it retains the anaphase chromosome orientation (Schubert and Shaw 2011), the acquisition of certain systems that alter and fix the distribution of centromeres and telomeres is one potential explanation for the establishment of a non-Rabl configuration. To date, whether there is any biological significance to adopting a non-Rabl instead of a Rabl configuration is unknown. The identification and characterization of such systems will help us to clarify the answer to this question. The distributions of centromeres and telomeres are representative indices that distinguish the chromosome configuration, as Rabl or non-Rabl type. Therefore, we focus on the factors that affect the distribution of either centromeres or telomeres during the interphase, such as nuclear membrane proteins, nucleolar proteins, and the structural maintenance of the chromosome complex (Table 2). In addition, we compile the roles of those factors in intranuclear events, such as gene expression and chromosome organization. Through a comparison with the findings for other eukaryotes, the mechanisms of positional control of centromeres and telomeres and the biological significance of the proper distribution of those domains in plants are discussed.
Positional control of centromeres during the interphase
Although the distribution pattern of centromeres differs from species to species, they tend to be localized at the nuclear periphery in plants as well as in many other eukaryotes (Muller et al. 2019; Vanrobay et al. 2013), suggesting that the factors around the nuclear periphery may participate in the positional control of centromeres. Indeed, recent studies of the interphase nuclei in Schizosaccharomyces pombe where chromosomes show a Rabl configuration unveiled the functions of nuclear membrane proteins in the clustering and tethering of centromeres at the nuclear envelope near the spindle pole body (SPB) (Fig. 1b). In this case, centromeres are associated with the linker of the nucleoskeleton and cytoskeleton (LINC) complex consisting of the inner nuclear membrane SUN (Sad1/UNC-84)-domain protein Sad1 and the outer nuclear membrane KASH (Klarsicht/ANC-1/Syne homology)-domain proteins Kms1 and Kms2, resulting in the clustering of all centromeres beneath the SPB (Fernández-Álvarez and Cooper 2017; Hou et al. 2012; Miki et al. 2004). The interaction between Sad1 and centromeres is redundantly mediated by nuclear membrane protein Lem2 and a factor involved in the centromere clustering Csi1 (Barrales et al. 2016; Fernández-Álvarez and Cooper 2017: Hou et al. 2012). Likewise, the involvement of nuclear membrane proteins in the peripheral localization and the clustering of centromeres have been shown in A. thaliana in which chromosomes show a non-Rabl configuration. In A. thaliana, the centromeric and pericentromeric regions are contained in densely compacted heterochromatic regions called chromocenters. The triple mutant sun1 sun4 sun5 shows an increased distance between the border of chromocenters and the nuclear periphery during the interphase (Poulet et al. 2017a). This suggests that the SUN proteins maintain the centromere position near the nuclear periphery, although it remains unknown whether or not the SUN proteins function as a LINC complex. In addition, the potential SUN-interacting protein CROWDED NUCLEI 1 (CRWN1) (Graumann 2014), directly binds to heterochromatic regions on chromosome arms and centromeric and pericentromeric regions, which mediates the tethering of those regions at the nuclear periphery (Hu et al. 2019). In accordance with this, chromocenters positioned more in the interior are seen in the double mutant crwn1 crwn2, as compared with the wild-type (Poulet et al. 2017a). CRWNs known as lamin-like proteins are considered to compose a meshwork layer termed nuclear lamina underneath the nuclear envelope, which mechanically supports the nuclear architecture in cooperation with other nuclear membrane proteins, such as KAKU4 and nuclear envelope-associated proteins (NEAPs) (Poulet et al. 2017b). However, KAKU4 and NEAPs do not have any role in heterochromatin-nuclear periphery tethering (Hu et al. 2019), indicating that CRWNs are the functional basis of plant nuclear lamina in chromatin positioning. Strikingly, CRWN1-bound chromatin in the pericentromeric regions highly overlap with those bound by NUCLEOPOLIN 1, a component of the nuclear pore complex (Bi et al. 2017; Hu et al. 2019). The nuclear pore complex may act in centromere positioning through interaction with the surrounding pericentromeric regions.
Recently, it has been established that CRWNs are involved not only in the tethering but also the unclustering of centromeres. The double mutant crwn1 crwn2 shows a reduced number of chromocenters (Dittmer et al. 2007; Poulet et al. 2017a; Wang et al. 2013). Consistent with this, the number of microscopically distinguishable 180-bp centromeric repeat signals is reduced in crwn1 crwn2 (3–6 signals) compared with that found in the wild-type (5–10 signals) (Wang et al. 2013). Similarly, some nuclei in crwn4 have a reduced number of chromocenters (Wang et al. 2013). In addition, chromosome conformation capture (Hi-C), which provides the genome-wide characterization of chromatin interactions, revealed that both crwn1 and crwn4 causes increased interactions between the pericentromeric regions derived from each chromosome (Grob et al. 2014). These findings suggest that at least three out of the four CRWN paralogues existing in A. thaliana contribute to the avoidance of centromere hyperclustering. In contrast, the number of chromocenters in sun1 sun4 sun5 is comparable to that found in the wild-type (Poulet et al. 2017a). Furthermore, the quintuple mutant, which lacks three KASH proteins, WPP domain-interacting proteins (WIP1–WIP3), and two KASH-interesting proteins, WPP domain-interacting tail-anchored proteins (WIT1 and WIT2) (Zhou and Meier 2014), also did not show any changes in the number of chromocenters (Poulet et al. 2017a). These suggest that the LINC complex is unlikely to function in centromere unclustering.
In addition to nuclear membrane proteins, recently, one of the examples of the structural maintenance of chromosome (SMC) complexes, composed of two SMC proteins and three non-SMC proteins, condensin II, was identified as an important regulator of centromere distribution. In the nurse cells of Drosophila melanogaster, in which centromeres show a Rabl-type distribution, condensin II acts in the axial compaction of the chromosome, which enforces the dissociation of pericentromeric chromatin interactions. In this case, surprisingly, the increasing activity of condensin II, along with the development of nurse cells, contributes to the transition from Rabl to non-Rabl configuration (Bauer et al. 2012), which indicates that condensin II plays a critical role in establishing the interphase chromosome configuration. Condensin II is also conserved in plants (Sakamoto et al. 2011) and is indispensable for the unclustering of centromeres in A. thaliana, as shown by the drastically reduced number of microscopically distinguishable centromeric signals (the lowest number is one) in condensin II mutants (Sakamoto et al. 2019; Schubert et al. 2013). In addition, the A. thaliana meristematic nuclei with clustered centromeres, caused by the dysfunction of condensin II, seems to represent a Rabl-like distribution pattern from the viewpoint of histological analysis—polarized centromeric signals on one side of the nucleus (Schubert et al. 2013). Therefore, it is possible that condensin II is a crucial factor for ensuring the dispersed centromere distribution in plants, as is the case in D. melanogaster. However, it remains unknown if the molecular action of condensin II, reported in D. melanogaster, is also applicable to the case of A. thaliana.
Taken together, in plants, centromere tethering at the nuclear periphery would be mediated by a LINC complex and nuclear lamina consisting of CRWNs, whereas centromere unclustering would be regulated by CRWNs and condensin II. The centromere tethering at the nuclear periphery would contribute to maintaining the position of the centromeres during the interphase, regardless of the Rabl or non-Rabl configuration in plants. However, the centromere unclustering may be crucial for the establishment of dispersed centromere distribution observed in the non-Rabl configuration. Considering that both CRWNs and condensin II are highly conserved in plants (Poulet et al. 2017b; Sakamoto et al. 2011), it is interesting to determine whether the activities of those proteins are correlated with the type of centromere distribution among the plant species.
Biological significances of proper centromere distribution during interphase
Thus far, it is unclear whether or not there are advantages for each centromere distribution, Rabl or non-Rabl type, whereas the biological significance of maintaining a proper centromere distribution has been suggested. In Saccharomyces cerevisiae, which exhibits a Rabl configuration, the detachment of centromeres from the SPB on the nuclear envelope results in the increased spatial mobility of chromosomes, which would impact on the spatial organization of chromosomes during the interphase (Verdaasdonk et al. 2013). In addition, it is proposed that the formation of centromere clusters regulates the spatial organization of chromosomes in human cells that show a non-Rabl configuration (Tjong et al. 2016). In A. thaliana, each chromosome is compartmentalized into a different space within a nucleus (Pecinka et al. 2004). Hi-C analysis indicated the weakened compartmentalization of each chromosome within a nucleus and of local chromatin domains, such as a loosened chromatin region and a condensed chromatin region within a chromosome in mutants crwn1 and crwn4 (Hu et al. 2019). This suggests that heterochromatin tethering and/or centromere unclustering contributes to properly maintain the spatial and structural organization of chromosomes in plants. It is also reported that the mutation in A. thaliana condensin II subunit CAP-D3 causes the dispersed distribution of a certain chromosome arm (Schubert et al. 2013), supporting the idea that centromere unclustering is involved in the spatial organization of chromosomes. However, it should be noted that the dispersed chromosome in the condensin II mutant may not be a consequence of centromere hyperclustering. As in the case of the nurse cells of D. melanogaster (Bauer et al. 2012), it may be possible that A. thaliana condensin II promotes the compartmentalization of each chromosome by acting directly on the chromosome compaction and, consequently, induces centromere unclustering.
A growing body of evidence suggests that the spatial positioning of genes within a nucleus is linked to its expression and the transcriptional activity in eukaryotes (Nguyen and Bosco 2015). Therefore, it is anticipated that the abnormal centromere distribution, which is accompanied by the alternation of the spatial organization of chromosomes, has numerous effects on the control of gene expression. However, contrary to the anticipation, Hu et al. (2019) revealed that there are only 14 genes differentially expressed in mutant crwn1. In addition, they also found that the mutation in crwn1 causes a slight difference in the accessibility of chromatin in the promoter region. Similar to crwn1, A. thaliana condensin II mutant cap-d3 also has little effect on gene expression (Municio et al. 2019). These facts suggest that the centromere distribution does not have a massive impact on the local chromatin organization and transcription. Meanwhile, it is reported that the dysfunction in the LINC complex and CRWNs alters the transcription in the vicinity of the centromere in a distinct manner (Poulet et al. 2017a). Triple mutant sun1 sun4 sun5 demonstrates the release of transcriptional gene silencing at centromeric and pericentromeric repeats, whereas double mutant crwn1 crwn2 shows enhanced silencing (Poulet et al. 2017a). The release of transcriptional gene silencing requires the decondensation of the centromeric chromatin, whereas the enhancement of transcriptional gene silencing requires the further condensation of the centromeric chromatin. Therefore, the failure of the centromere-nuclear periphery tethering may cause centromere decondensation, whereas the failure of centromere unclustering, or the simultaneous failure of centromere unclustering and tethering, may cause centromere condensation. Interestingly, a similar phenomenon is also seen in other organisms that show a clustered centromere distribution. In S. pombe, the lack of MSC domain in the nuclear membrane protein Lem2 causes centromere unclustering while simultaneously facilitating the transcription of repetitive DNA at the pericentromere (Barrales et al. 2016). Furthermore, in Drosophila tissue cultured cells, whose centromere cluster is formed around the nucleolus, the failure in centromere clustering results in impaired silencing of repetitive DNA at the pericentromere (Padeken et al. 2013). Therefore, it is possible that the formation of a proper centromere distribution is required to maintain the constitutive heterochromatic state around centromeres, not only in plants but also in other organisms.
Recent studies revealed that the maintenance of the interphase centromere distribution is necessary for the proper progression of the mitotic phase in S. pombe. The failure in centromere clustering at SPB during the interphase causes severe mitotic defects, such as failure of spindle assembly and chromosome missegregation (Fernández-Álvarez et al. 2016; Hou et al. 2012), indicating that the centromere clustering prior to entry into the mitotic phase is crucial for proper cell division. In contrast, for now, no clear mitotic chromosomal defects in crwn or condensin II mutants are reported in A. thaliana, indicating that the centromere distribution during the interphase may not be linked to the progression of mitotic events in plants.
In summary, the proper centromere tethering to the nuclear periphery during the interphase would be linked to at least the maintenance of centromere condensation in eukaryotes, although a mechanism of this mutual relationship remains unclear. Meanwhile, the centromere unclustering is unlikely to involve gene expression control, despite its impact on the regulation of the spatial and structural organizations of chromosomes in plants. Thus, it is conceivable that the proper gene expression control would have to be robust, regardless of the alteration in centromere distribution, as it is a fundamental of life. This might be a reason why some plants were able to acquire a non-Rabl-type centromere distribution during their evolution.
Positional control of telomeres during the interphase and its biological significance
Although there are exceptions, such as A. thaliana, where telomeres are associated with the nucleolus throughout the interphase (Roberts et al. 2009), most telomeres are localized along the nuclear periphery, irrespective of the type of chromosome configuration—Rabl or non-Rabl—in plants (Dong and Jiang 1998). Therefore, as in the case of centromere distribution, the nuclear membrane proteins function in the telomere tethering at the nuclear periphery during the mitotic interphase; however, to our knowledge, no such protein has been yet identified in plants. In contrast, the indispensable functions of nuclear membrane proteins in the positional regulation of telomeres have been reported in yeasts whose telomeres are localized at the nuclear periphery during the interphase (Ebrahimi and Cooper 2016). In S. pombe, telomere tethering to the nuclear periphery is regulated by several inner nuclear membrane proteins, such as Bqt3–Bqt4 complex, which interacts with the telomere binding protein complex Rap1/Taz1 (Chikashige et al. 2009). Similarly, in S. cerevisiae, two redundant pathways mediated by inner nuclear membrane proteins Esc1 or SUN-domain protein yMps3, are required for the telomere tethering to the nuclear periphery (Schober et al. 2009; Taddei et al. 2004). Whereas the homologs of Bqt and Esc proteins are unlikely to be conserved in plants, Poulet et al. (2017b) have shown that the homologs of SUN domain proteins are highly conserved. In addition, a study of maize suggests the involvement of ZmSUN2 in anchoring a telomere to the nuclear periphery during meiosis, even though it remains unknown whether ZmSUN2 performs a similar function during the mitotic interphase (Murphy et al. 2014). Therefore, it is worth investigating whether the plant SUN proteins are involved in telomere tethering to the nuclear periphery during the mitotic interphase. Meanwhile, a recent study found that a nucleolar protein, NUCLEOLIN 1 (NUC1), is involved in the telomere association with the nucleolus in A. thaliana (Pontvianne et al. 2016). For instance, nuc1 shows the increased dissociation of telomeres from the nucleolus. In addition, the telomere length is shortened in all the chromosomes of nuc1. Therefore, it is proposed that the maintenance of the telomere association with the nucleolus through the NUC1 function is crucial for maintaining the telomere length (Pontvianne et al. 2016). In the case of S. pombe, the telomere association with the nuclear periphery is not involved in the telomere functions, such as telomere length control and subtelomeric gene silencing during the interphase; instead, it is required for the proper replication of the heterochromatic chromosome regions during the S phase (Chikashige et al. 2009, 2010; Ebrahimi et al. 2018). The difference in the effect of telomere positioning between A. thaliana and S. pombe might be due to the difference in the nuclear component to which the telomere is tethered.
Taken together, in comparison with centromeres, little is known regarding the factors involved in the proper positioning of telomeres during the mitotic interphase in plants. Considering the existence of protein Lem2 in S. pombe, which has roles in the spatial regulation of both centromeres and telomeres during the interphase (Barrales et al. 2016; Fernández-Álvarez and Cooper 2017), it is tempting to speculate that the proteins involved in the centromere positioning, such as CRWNs and condensin II, also function in the telomere positioning in plants. Indeed, it has been proposed that condensin II-mediated chromosome compaction drives the movement of telomeres along the nuclear periphery, which results in the transition of the Rabl to the non-Rabl-type telomere distribution in the nurse cells of D. melanogaster (Bauer et al. 2012). Meanwhile, the identification of the novel telomere binding nuclear membrane proteins and/or nucleolar proteins will expand our understanding of telomere positioning in both Rabl and non-Rabl configurations.
Conclusions
The identification of the factors involved in the maintenance of the non-Rabl-type centromere distribution during the interphase in A. thaliana enabled the exploration of the biological significance of proper centromere distribution in intranuclear events in plants. Although the studies on these factors provided some hints, no clear answer has been found. Given that the Lem2 functions in the centromere clustering prevent the loss of Rabl-type centromere distribution in S. pombe (Barrales et al. 2016; Fernández-Álvarez and Cooper 2017; Hou et al. 2012), there might be a system that maintains centromere clustering in plants exhibiting Rabl-type centromere distribution. From this point of view, the acquisition of the mutant in other species, which is defective in the maintenance of Rabl-type centromere distribution and/or the revertant of A. thaliana mutant showing a Rabl-type centromere distribution, such as mutant condensin II, will strongly contribute to a better understanding of the biological significance of proper centromere distribution. In addition, to understand the telomere positioning and its significance, further analyses of other species are necessary, particularly considering that telomere localization in A. thaliana is exceptional among plants. Finally, an integrated understanding of the positional control of centromeres and telomeres may help us explain the diversification of the chromosome configuration among organisms beyond the plant kingdom.
References
Adams MD, Celniker SE, Holt RA, Evans CA, Gocayne JD, Amanatides PG, Scherer SE, Li PW, Hoskins RA, Galle RF et al (2000) The genome sequence of Drosophila melanogaster. Science 287:2185–2195
Ananiev EV, Riera-Lizarazu O, Rines HW, Phillips RL (1997) Oat-maize chromosome addition lines: a new system for mapping the maize genome. Proc Nat Acad Sci USA 94:3524–3529
Barrales RR, Forn M, Georgescu PR, Sarkadi Z, Braun S (2016) Control of heterochromatin localization and silencing by the nuclear membrane protein Lem2. Genes Dev 30:133–148
Bauer CR, Hartl TA, Bosco G (2012) Condensin II promotes the formation of chromosome territories by inducing axial compaction of polyploid interphase chromosomes. PLos Genet 8:12
Bi XL, Cheng YJ, Hu B, Ma XL, Wu R, Wang JW, Liu C (2017) Nonrandom domain organization of the Arabidopsis genome at the nuclear periphery. Genome Res 27:1162–1173
Brenchley R, Spannagl M, Pfeifer M, Barker GL, D’Amore R, Allen AM, McKenzie N, Kramer M, Kerhornou A, Bolser D et al (2012) Analysis of the bread wheat genome using whole-genome shotgun sequencing. Nature 491:705–710
Chikashige Y, Yamane M, Okamasa K, Tsutsumi C, Kojidani T, Sato M, Haraguchi T, Hiraoka Y (2009) Membrane proteins Bqt3 and -4 anchor telomeres to the nuclear envelope to ensure chromosomal bouquet formation. J Cell Biol 187:413–427
Chikashige Y, Haraguchi T, Hiraoka Y (2010) Nuclear envelope attachment is not necessary for telomere function in fission yeast. Nucleus 1:481–486
Choulet F, Wicker T, Rustenholz C, Paux E, Salse J, Leroy P, Schlub S, Le Paslier MC, Magdelenat G, Gonthier C (2010) Megabase level sequencing reveals contrasted organization and evolution patterns of the wheat gene and transposable element spaces. Plant Cell 22:1686–1701
Dittmer TA, Stacey NJ, Sugimoto-Shirasu K, Richards EJ (2007) LITTLE NUCLEI genes affecting nuclear morphology in Arabidopsis thaliana. Plant Cell 19:2793–2803
Dong F, Jiang J (1998) Non-Rabl patterns of centromere and telomere distribution in the interphase nuclei of plant cells. Chromosome Res 6:551–558
Ebrahimi H, Cooper JP (2016) Finding a place in the SUN: telomere maintenance in a diverse nuclear landscape. Curr Opin Cell Biol 40:145–152
Ebrahimi H, Masuda H, Jain D, Cooper JP (2018) Distinct ‘safe zones’ at the nuclear envelope ensure robust replication of heterochromatic chromosome regions. eLIFE 7:e3291
Fang Y, Spector DL (2005) Centromere positioning and dynamics in living Arabidopsis plants. Mol Biol Cell 16:5710–5718
Fernández-Álvarez A, Cooper JP (2017) The functionally elusive Rabl chromosome configuration directly regulates nuclear membrane remodeling at mitotic onset. Cell Cycle 16:1392–1396
Fernández-Álvarez A, Bez C, O’Toole ET, Morphew M, Cooper JP (2016) Mitotic nuclear envelope breakdown and spindle nucleation are controlled by interphase contacts between centromeres and the nuclear envelope. Dev Cell 39:544–559
Fransz P, de Jong JH, Lysak M, Castiglione MR, Schubert I (2002) Interphase chromosomes in Arabidopsis are organized as well defined chromocenters from which euchromatin loops emanate. Proc Nat Acad Sci USA 99:14584–14589
Fujimoto S, Ito M, Matsunaga S, Fukui K (2005) An upper limit of the ratio of DNA volume to nuclear volume exists in plants. Genes Genet Syst 80:345–350
Funabiki H, Hagan I, Uzawa S, Yanagida M (1993) Cell cycle dependent specific positioning and clustering of centromeres and telomeres in fission yeast. J Cell Biol 121:961–976
Goffeau A, Barrell BG, Bussey H, Davis RW, Dujon B, Feldmann H, Galibert F, Hoheisel JD, Jacq C, Johnston M et al (1996) Life with 6000 genes. Science 274:546–567
Graumann K (2014) Evidence for LINC1-SUN Associations at the plant nuclear periphery. PLos One 9:7
Grob S, Schmid MW, Grossniklaus U (2014) Hi-C analysis in Arabidopsis identifies the KNOT, a structure with similarities to the flamenco locus of Drosophila. Mol Cell 55:678–693
Hou HT, Zhou Z, Wang Y, Wang JY, Kallgren SP, Kurchuk T, Miller EA, Chang F, Jia ST (2012) Csi1 links centromeres to the nuclear envelope for centromere clustering. J Cell Biol 199:735–744
Hu B, Wang N, Bi XL, Karaaslan ES, Weber AL, Zhu WS, Berendzen KW, Liu C (2019) Plant lamin-like proteins mediate chromatin tethering at the nuclear periphery. Genome Biol 20:18
Jin QW, Trelles-Sticken E, Scherthan H, Loidl J (1998) Yeast nuclei display prominent centromere clustering that is reduced in nondividing cells and in meiotic prophase. J Cell Biol 141:22–29
Macas J, Neumann P, Navrátilová A (2007) Repetitive DNA in the pea (Pisum sativum L.) genome: comprehensive characterization using 454 sequencing and comparison to soybean and Medicago truncatula. BMC Genom 8:427
Manuelidis L (1984) Different central nervous system cell types display distinct and nonrandom arrangements of satellite DNA sequences. Proc Nat Acad Sci USA 81:3123–3127
Matsunaga S, Katagiri Y, Nagashima Y, Sugiyama T, Hasegawa J, Hayashi K, Sakamoto T (2013) New insights into the dynamics of plant cell nuclei and chromosomes. In: Jeon KW (ed) International review of cell and molecular biology. Elsevier Academic Press Inc, San Diego, pp 253–301
Miki F, Kurabayashi A, Tange Y, Okazaki K, Shimanuki Y, Niwa O (2004) Two-hybrid search for proteins that interact with Sad1 and Kms1, two membrane-bound components of the spindle pole body in fission yeast. Mol Genet Genom 270:449–461
Muller H, Gil J, Drinnenberg IA (2019) The impact of centromeres on spatial genome architecture. Trends Genet 35:565–578
Municio C, Antosz W, Grasser KD, Kornobis E, Van Bel M, Eguinoa I, Coppens F, Bräutigam A, Lermontova I, Bruckmann A, Houben A, Schubert V (2019) The Arabidopsis condensin CAP-D subunits arrange interphase chromatin. bioRxiv. https://doi.org/10.1101/2019.12.12.873885
Murphy SP, Gumber HK, Mao Y, Bass HW (2014) A dynamic meiotic SUN belt includes the zygotene-stage telomere bouquet and is disrupted in chromosome segregation mutants of maize (Zea mays L.). Front Plant Sci 5:314
Nguyen HQ, Bosco G (2015) Gene positioning effects on expression in eukaryotes. In: Bassler BL (ed) Annual review of genetics. Annual Reviews, Palo Alto, pp 627–646
Padeken J, Mendiburo MJ, Chlamydas S, Schwarz HJ, Kremmer E, Heun P (2013) The nucleoplasmin homolog NLP mediates centromere clustering and anchoring to the nucleolus. Mol Cell 50:236–249
Paterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J, Gundlach H, Haberer G, Hellsten U, Mitros T, Poliakov A et al (2009) The Sorghum bicolor genome and the diversification of grasses. Nature 457:551–556
Pecinka A, Schubert V, Meister A, Kreth G, Klatte M, Lysak MA, Fuchs J, Schubert I (2004) Chromosome territory arrangement and homologous pairing in nuclei of Arabidopsis thaliana are predominantly random except for NOR-bearing chromosomes. Chromosoma 113:258–269
Pontvianne F, Carpentier MC, Durut N, Pavlištová V, Jaške K, Schořová S, Parrinello H, Rohmer M, Pikaard CS, Fojtová M et al (2016) Identification of nucleolus-associated chromatin domains reveals a role for the nucleolus in 3D organization of the A. thaliana genome. Cell Rep 16:1574–1587
Poulet A, Duc C, Voisin M, Desset S, Tutois S, Vanrobays E, Benoit M, Evans DE, Probst AV, Tatout C (2017a) The LINC complex contributes to heterochromatin organisation and transcriptional gene silencing in plants. J Cell Sci 130:590–601
Poulet A, Probst AV, Graumann K, Tatout C, Evans DE (2017b) Exploring the evolution of the proteins of the plant nuclear envelope. Nucleus 8:46–59
Rabl C (1885) Über Zelltheilung. Morphol Jahrb 10:214–330
Rawlins DJ, Shaw PJ (1990) Three-dimensional organization of ribosomalDNA in interphase nuclei of Pisum sativum by in situ hybridization and optical tomography. Chromosoma 99:143–151
Roberts NY, Osman K, Armstrong SJ (2009) Telomere distribution and dynamics in somatic and meiotic nuclei of Arabidopsis thaliana. Cytogenet Genome Res 124:193–201
Sakamoto T, Inui YT, Uraguchi S, Yoshizumi T, Matsunaga S, Mastui M, Umeda M, Fukui K, Fujiwara T (2011) Condensin II alleviates DNA damage and is essential for tolerance of boron overload stress in Arabidopsis. Plant Cell 23:3533–3546
Sakamoto T, Sugiyama T, Yamashita T, Matsunaga S (2019) Plant condensin II is required for the correct spatial relationship between centromeres and rDNA arrays. Nucleus 10:116–125
Santos AP, Shaw P (2004) Interphase chromosomes and the Rabl configuration: does genome size matter? J Microsc 241:201–206
Schnable PS, Ware D, Fulton RS, Stein JC, Wei F, Pasternak S, Liang C, Zhang J, Fulton L, Graves TA (2009) The B73 maize genome: complexity, diversity, and dynamics. Science 326:1112–1115
Schober H, Ferreira H, Kalck V, Gehlen LR, Gasser SM (2009) Yeast telomerase and the SUN domain protein Mps3 anchor telomeres and repress subtelomeric recombination. Genes Dev 23:928–938
Schubert I, Shaw P (2011) Organization and dynamics of plant interphase chromosomes. Trends Plant Sci 16:273–281
Schubert V, Lermontova I, Schubert I (2013) The Arabidopsis CAP-D proteins are required for correct chromatin organisation, growth and fertility. Chromosoma 122:517–533
Taddei A, Hediger F, Neumann FR, Bauer C, Gasser SM (2004) Separation of silencing from perinuclear anchoring functions in yeast Ku80, Sir4 and Esc1 proteins. EMBO J 23:1301–1312
Tjong H, Li WY, Kalhor R, Dai C, Hao SL, Gong K, Zhou YG, Li HC, Zhou XJ, Le Gros MA et al (2016) Population-based 3D genome structure analysis reveals driving forces in spatial genome organization. Proc Nat Acad Sci USA 113:E1663–E1672
Vanrobay E, Thomas M, Tatout C (2013) Heterochromatin positioning and nuclear architecture (In: Evans DE, Graumann K, Bryant JA (eds) Plant nuclear structure, genome architecture and gene regulation. Wiley-Backwell). Annu Plant Rev 46:157–190
Verdaasdonk JS, Vasquez PA, Barry RM, Barry T, Goodwin S, Forest MG, Bloom K (2013) Centromere tethering confines chromosome domains. Mol Cell 52:819–831
Wang HY, Dittmer TA, Richards EJ (2013) Arabidopsis CROWDED NUCLEI (CRWN) proteins are required for nuclear size control and heterochromatin organization. BMC Plant Biol 13:13
Wood V, Gwilliam R, Rajandream MA, Lyne M, Lyne R, Stewart A, Sgouros J, Peat N, Hayles J, Baker S et al (2002) The genome sequence of Schizosaccharomyces pombe. Nature 415:871–880
Zhou X, Meier I (2014) Efficient plant male fertility depends on vegetative nuclear movement mediated by two families of plant outer nuclear membrane proteins. Proc Nat Acad Sci USA 111:11900–11905
Arabidopsis-Genome-Initiative (2000) Analysis of the genome sequence of the floweringplant Arabidopsis thaliana. Nature 408:796–815
International Human Genome Sequencing Consortium (2004) Finishing the euchromatic sequence of the human genome. Nature 431:931–945
International-Rice-Genome-Sequencing-Project (2005) The map-based sequence of the rice genome. Nature 436:793–800
International-Barley-Genome-Sequencing-Consortium (2012) A physical, genetic and functional sequence assembly of the barley genome. Nature 491:711–716
Acknowledgements
This work was supported by MXT/JSPS KAKENHI (19K06748) to TS. We would like to thank Editage (www.editage.com) for English language editing.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Oko, Y., Ito, N. & Sakamoto, T. The mechanisms and significance of the positional control of centromeres and telomeres in plants. J Plant Res 133, 471–478 (2020). https://doi.org/10.1007/s10265-020-01202-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-020-01202-2