Abstract
Rehmannia japonica (Thunb.) Makino ex T. Yamaz. is an endangered perennial herb species in Japan. Although earlier the Japanese considered it a variety of R. glutinosa, recent Japanese taxonomists have consistently regarded it as an independent species. According to the historical literature, Rehmannia japonica seems to have been known in China and Japan in the past. However, Chinese taxonomists do not recognize R. japonica at present. In Japan, only two populations are known, and although these populations flower every year, seed reproduction has not been observed. In this study, we aimed to reveal the phylogenetic relationships and levels of genetic diversity of R. japonica. A haplotype network based on two chloroplast DNA regions (trnL-trnF and rps16) showed that the sequences of R. japonica were distinguishable by three or four sites of indels from the most closely related species, R. chingii, consistent with the separate species status of R. japonica. An analysis of genetic diversity using twelve microsatellite loci showed that all of the ramets of R. japonica collected from two geographically isolated populations had an identical multilocus genotype, including identical heterozygous genotypes at six loci. This result indicated asexual origin of all sampled ramets. This study also suggests that the absence of sexual reproduction of R. japonica is explained by self-incompatibility combined with only a single genet remaining in the R. japonica populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In Japan, many endangered plants occur in small populations in rural landscapes, and conservation activities have been conducted for them. However, some of these endangered plant species may originate from introduction by humans. For example, Loropetalum chinense (R. Br.) Oliv. (Hamamelidaceae) is a common evergreen tree species in China, but it is endangered in Japan and strictly protected by the local governments or landowners. However, genetic analysis of all wild individuals in Japan revealed only two or three genets in each population, indicating artificial propagation such as cutting and transplanting (Isagi and Kaneko 2014). Therefore, reexamination of the origin and conservation value of “endangered” species growing in rural landscapes is recommended in order to avoid spending limited conservation resources on human-introduced populations.
Although Rehmannia japonica (Thunb.) Makino ex T. Yamaz. is listed as critically endangered in the Japanese red list (Ministry of Environment of Japan 2015), its taxonomic status has been unclear. The genus Rehmannia had been included in the family Scrophulariaceae. Its current taxonomic placement is under question due to the disintegration of Scrophulariaceae (Mabberley 2008; Olmstead et al. 2001; Oxelman et al. 2005), but recent phylogenetic studies showed that a monophyletic group, including Rehmannia is sister to Orobanchaceae (Albach et al. 2007; Xia et al. 2009). In general, six species, R. elata N. E. Br., R. piasezkii Maxim., R. henryi N. E. Br., R. solanifolia P. C. Tsoong and T. L. Chin, R. glutinosa (Gaertn.) Libosch. ex Fisch. and C. A. Mey., and R. chingii H. L. Li are recognized in the genus Rehmannia (Albach et al. 2007; Chin 1998; Xia et al. 2009) and these six species are endemic to China (Chin 1998).
In addition to the six Chinese species of Rehmannia, Rehmannia japonica has been recognized in Japan. Although earlier Japanese taxonomists such as Matsuda (1918), Makino (1932), and Hara (1949) considered it as a variety of R. glutinosa, recent Japanese taxonomists have consistently regarded it as an independent species (e.g., Yamazaki 1993; Yonekura 2012). Rehmannia japonica is very similar to R. chingii in its flower size and shape (Yamazaki 1961), but it is distinguished by several morphological characters (Yamazaki 1993). Rehmannia japonica is considered to have been widely cultivated as a horticultural plant in the past in Japan (Yamazaki 1961), and historical Japanese names of R. japonica, “Senrigoma” or “Komenmou,” are mentioned in the traditional medical books of China and Japan. In China, “Komenmou” was found in the historical Chinese medical book, “Compendium of Materia Medica”, which was published in 1596 (https://books.google.co.jp/books?id=zVueCQAAQBAJ). In Japan, “Komenmou” or “Senrigoma” was illustrated in “Kai” (http://dl.ndl.go.jp/info:ndljp/pid/2555607/1) and “Honzou zufu” (http://dl.ndl.go.jp/info:ndljp/pid/1287126/1), which were published in 1765 and 1828, respectively. In the three books, “Komenmou” was distinguished from R. glutinosa, which is a famous medical plant species. Although taxonomic identification of R. japonica and “Komenmou” in the historical literature is needed, Rehmannia japonica may have been known in China and Japan in the past. However, Rehmannia japonica is not presently reported from China (Hong et al. 1998; Yamazaki 1961, 1993), and the taxonomic status and exact origin of this species is still uncertain.
Rehmannia japonica is extremely rare now. Only two populations are known in Japan. These two populations are found in Shizuoka Prefecture. Because of the limited number of populations, R. japonica is classified as “critically endangered” in the Japanese Red Data List (Ministry of Environment of Japan 2015). Old stone walls of terraced fields are the main habitats of R. japonica, and renovations of these walls and the resulting habitat destruction has directly led to the decreasing numbers of populations of this species. In addition to the limited number of populations, both R. japonica populations appear to have serious problems with sexual reproduction. Despite flowering every year, Rehmannia japonica does not produce seeds (Yamazaki 1993; Kaneko personal observation). Thus, this species depends on vegetative reproduction via rhizome elongation, and continued loss of habitat by the renovation of stone walls could cause irreversible damage to the continued existence of the populations.
In this study, we aimed to reveal the phylogenetic relationships and levels of genetic diversity of R. japonica. Previous studies inferred molecular phylogenies of the six Chinese species of Rehmannia using chloroplast DNA regions but did not sample R. japonica (Albach et al. 2007; Xia et al. 2009). A molecular phylogenetic analysis including R. japonica may clarify its taxonomic identity. The results might also demand a reconsideration of the conservation status of R. japonica. If R. japonica is genetically identical to one of the other Rehmannia species and is an introduced species from China, it is not appropriate that the conservation agencies treat it as an endangered species. On the other hand, if R. japonica is genetically distinct, its populations in Japan should be conserved as the only remaining populations of this species. In addition to the molecular phylogenetic analysis, the genetic diversity of these remnant populations was evaluated using microsatellite markers developed in the present study. Detailed genetic analysis can assess the genetic diversity and number of genets in remnant populations (e.g., Isagi and Kaneko 2014). The results also allow insights into a possible reason for the lack of sexual reproduction in R. japonica and provide the basic information needed for developing an effective conservation program for this species.
Materials and methods
Study site
Only two populations are known for R. japonica. They occur in Shizuoka Prefecture, Japan (Population 1: ca. 300 flowering ramets, N35°10′, E137°53′, Population 2: ca. 100 flowering ramets, N35°08′, E137°54′). These two populations are separated by about 3 km. The habitat of this species is old stone walls of terraced fields on steep slopes. Although the age of construction of these stone walls is not known, population 2 has been known since the 1960s (Ohba personal communication). Local famers have not been made aware that this plant species is listed as critically endangered in the Red List (Ministry of Environment of Japan 2015). We also investigated a population of R. chingii, which is the putative sister taxon of R. japonica (Yamazaki 1961) at Mt. Tian-mu, Zhejiang, China (N30°19′, E119°26′).
Sampling and DNA extraction
Plant materials for molecular analysis were sampled from two populations of R. japonica and one population of R. chingii. In R. japonica, leaf samples of 39 and 19 ramets were collected from population 1 and 2, respectively, in 2009. Samples were collected across the distribution of each population, from flowering ramets and small ramets with only one or two small leaves above ground. These samples were stored at −30 °C for later analysis. Voucher was deposited in the Herbarium of the Faculty of Symbiotic Systems Science, Fukushima University (FKSE), Fukushima, Japan. In R. chingii, we collected leaf samples from 15 ramets from Mt. Tian-mu population in 2010. These samples were desiccated using silica gel and stored at room temperature before analysis. Total genomic DNA was extracted using a modified CTAB method (Milligan 1992).
Chloroplast DNA analysis
We amplified and sequenced the trnL intron, trnL exon, and trnL–trnF spacer region (referred to here as the trnL–F region) and rps16 region of chloroplast DNA for four individuals of R. japonica and two individuals of R. chingii. The trnL–F region was amplified with primers “c” and “f” of Taberlet et al. (1991). The rps16 intron was amplified with primers rps16-2F and rps16-R3 (Bremer et al. 2002). The PCR amplification was performed in a thermal cycler GeneAmp PCR System 2700 (Applied Biosystems, Foster City, California, USA). A High Pure PCR Product Purification Kit (Roche Diagnostics, Mannheim, Germany) was used, and the purified products were sequenced directly with an ABI BigDye Terminator Cycle Sequencing Kit ver. 3.1 (Applied Biosystems) on the ABI PRISM 3100 Genetic Analyzer (Applied Biosystems). Both strands of the amplified PCR products were sequenced. Electropherograms were assembled with FinchTV (http://www.geospiza.com/finchtv/). The assembled sequences of each locus were aligned with those obtained from DDBJ/EMBL/GenBank (Tables 1, 2) for R. elata, R. piasezkii, R. henryi, R. solanifolia, R. glutinosa, and R. chingii using CLUSTAL X (Thompson et al. 1997). All consecutive indels were treated as one-point mutations. Based on these aligned sequences, a parsimony network of chloroplast DNA haplotypes was drawn using the program TCS1.06 (Clement et al. 2000).
Microsatellite marker development and analysis
Microsatellite markers were developed using the improved technique for isolating codominant compound microsatellite markers of Lian and Hogetsu (2002) and Lian et al. (2006). An adaptor-ligated, restricted DNA library for R. japonica was constructed as follows: genomic DNA was digested with the blunt-end restriction enzyme SspI. The restriction fragments were then ligated with a specific blunt adaptor (consisting of the 48-mer: 5′-GTAATACGACTCACTATAGGGCACGCGTGGTCGACGGCCCGGGCTGGT-3′ and an 8-mer with the 3′-end capped with an amino residue: 5′-ACCAGCCC-NH2-3′) using the Takara DNA ligation kit (Takara, Otsu, Japan). Fragments from the SspI DNA library were amplified by PCR using a compound SSR primer, (AC)6(AG)7, (AC)6(TC)7, or (TC)6(AC)7, and an adaptor primer (5′-CTATAGGGCACGCGTGGT-3′). The amplified fragments, ranging from 400 to 800 bp, were then separated on a 1.5% LO3 agarose gel (Takara) and purified using the QIAquick Gel Extraction Kit (Qiagen, Valencia, California, USA). The purified DNA fragments were cloned using the Qiagen PCR Cloning plus Kit (Qiagen), following the manufacturer’s instructions. The cloned fragments were amplified using the M13 forward and reverse primers. Amplified fragments were sequenced using the BigDye Terminator Cycle Sequencing Kit (Applied Biosystems). For each fragment containing a compound SSR sequence at one end, a specific primer was designed from the sequence flanking the compound SSR using Primer 3 (version 0.4.0; Rozen and Skaletsky 2000).
Polymorphism of the developed microsatellite markers was evaluated for 58 individuals of R. japonica and 15 individuals of R. chingii. Amplifications followed the standard protocol of the Qiagen Multiplex PCR Kit (Qiagen) in a final volume of 10 µL, which contained 5 ng of extracted DNA, 5 µL of 2×Multiplex PCR Master Mix, and 0.2 µM of each multiplexed primer. Compound SSR primers ((AC)6(AG)7, (AC)6(TC)7, or (TC)6(AC)7) were labeled with fluorochromes FAM, PET, or VIC (Applied Biosystems), respectively. Amplifications were performed with the GeneAmp PCR System 2700 thermal cycler using the following conditions: initial denaturation at 95 °C for 15 min, followed by 30 cycles of denaturation at 94 °C for 30 s, annealing 57 °C for 1 min 30 s, extension at 72 °C for 1 min, and a final extension at 60 °C for 30 min. The sizes of the PCR products were measured using the ABI PRISM 3100 Genetic Analyzer (Applied Biosystems) and GENESCAN and GENOTYPER software (Applied Biosystems). The loci showing polymorphism were tested for deviation from Hardy–Weinberg equilibrium and linkage disequilibrium between loci using FSTAT (version 2.9.3; Goudet 1995). Significance levels were adjusted using Bonferroni correction for multiple testing.
Results
Chloroplast DNA analysis
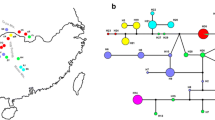
The aligned sequences consisted of 1,520 bp from the two regions of chloroplast DNA: trnL–F (840 bp) and rps16 (680 bp). In trnL–F, seven nucleotide substitutions and four indels were observed across Rehmannia (Table 1). In rps16, ten nucleotide substitutions and five indels were observed (Table 2). Sequences for these two DNA regions were identical for the four samples of R. japonica. The sequences obtained in this study were deposited in the DDBJ (accession numbers LC152768-LC152779). A haplotype network based on the nucleotide substitutions and indels of the two chloroplast DNA regions showed that the sequence of R. japonica was different from the other six Rehmannia species (Fig. 1). The most similar sequences to R. japonica belong to R. chingii (No. 2 and 4), but the sequences of R. chingii were distinguished from R. japonica by three or four sites of indels (Tables 1, 2). The sequences of the other five species (R. elata, R. piasezkii, R. henryi, R. solanifolia, and R. glutinosa) were clearly different from that of R. japonica.
Microsatellite analysis
Twelve loci were identified that showed a clear, strong, single band for each allele (Table 3). Levels of polymorphism were evaluated for 58 ramets from the two populations of R. japonica. The 58 samples collected from two geographically-isolated populations showed same multilocus genotype at the twelve loci (Table 4). Only the single alleles were observed for the six loci (Reja052, Reja075, Reja087, Reja152, Reja171, and Reja185), while two alleles were observed for the other six loci (Reja042, Reja057, Reja083, Reja156, Reja163, and Reja167). In R. chingii, five out of the twelve loci were amplified, and the 15 samples showed 15 multilocus genotypes at these five loci (Table 4). The number of alleles per locus ranged from 4 to 12 with an average of 8.0. The observed and expected heterozygosities (H O and H E) ranged from 0.25 to 0.73 and from 0.52 to 0.88, respectively, with averages of 0.73 for both. A significant deviation from Hardy–Weinberg equilibrium (P < 0.05), which is likely due to the existence of null alleles, was observed at four loci (Reja057, Reja075, Reja087, and Reja156). There was no evidence of significant linkage disequilibrium (P < 0.05).
Discussion
Phylogenetic identity and origin of R. japonica
The present results indicate that R. japonica may be genetically distinct from the other six Rehmannia species, as is also suggested by their morphology. Yamazaki (1961) suggested that the most closely-related species to R. japonica is R. chingii based on morphological characteristics, and R. japonica and R. chingii are distinguished morphologically by the length of petiole, density of pilose hairs on leaf, and short granular of filament (Chin 1998; Yamazaki 1961, 1993). These taxonomic conclusions based on morphological comparisons were supported by the present analysis of chloroplast DNA sequences. The sister taxon of R. japonica was R. chingii and these two species were distinguishable by the chloroplast DNA haplotypes. In the sequences of the trnL–F and rps16 regions, a number of substitutions and indels were variable among the seven species of Rehmannia. However, taxonomically-distinct R. glutinosa and R. solanifolia shared an identical haplotype, and thus, the three indels that distinguish R. japonica and R. chingii could potentially confirm their identities as different species.
The microsatellite analysis also suggested genetic differentiation between R. japonica and R. chingii. Seven of twelve microsatellite markers developed from R. japonica did not amplify in R. chingii, and significant deficiency of observed heterozygosity was observed at four loci in R. chingii. These unamplified loci and samples probably contain null alleles, likely due to nucleotide substitutions, insertions and deletions in the primer annealing sites between R. japonica and R. chingii. Although transferability of microsatellite markers among different species varies among taxa and loci, it decreases with increasing genetic distance among taxa in general. Microsatellite markers developed for various plant species using the same protocol as in the present study showed clearly higher transferability among related species than that seen between R. japonica and R. chingii (e.g., Izuno et al. 2011; Kaneko et al. 2011; Mori et al. 2008). This fact is consistent with the assertion that R. japonica and R. chingii are genetically distinct enough to treat as different species.
The present findings also suggested the need for extensive morphological and genetic analysis about Rehmannia species in China. In order to discuss the origin of R. japonica, an understanding of the genetic variation of each Rehmannia species is needed. Although we could not determine the origin of R. japonica based on the present and previous studies, background genetic information of Chinese Rehmannia is quite limited at present. In addition, Chinese and Japanese taxonomists are separately investigating the Rehmannia species, and most Chinese taxonomists probably have not recognized the possibility that R. japonica could exist in China. Therefore, the extensive analysis of Rehmannia species, especially R. chingii, may help identify the original populations of R. japonica.
Single genet remaining in R. japonica
Detailed genotyping of the two remnant populations using microsatellite markers showed that all sampled ramets of R. japonica are an identical genet that presumably originated from vegetative reproduction. The microsatellite analysis indicated remarkable differences in genetic diversity for R. japonica and R. chingii. All 58 ramets of R. japonica showed an identical multilocus genotype at the twelve loci, whereas the 15 ramets of R. chingii showed 15 different multilocus genotypes with high allelic polymorphism at the five amplified loci (Table 4). These results suggest that the microsatellite markers developed here will be valid for estimating the genetic diversity of R. japonica. In addition, all samples of R. japonica showed the identical heterozygous genotype for six loci. This result indicates that not only is the genetic diversity of R. japonica populations very low but also that all sampled ramets were produced without the recombination of sexual reproduction.
With the assumption of self-incompatibility, absence of sexual reproduction of R. japonica (Yamazaki 1993; Kaneko personal observation) may be explained by the only genet remaining in the R. japonica populations. Although there is no report of self-incompatibility in R. japonica, self-incompatibility was reported in R. glutinosa (Zhou et al. 2010). If R. japonica were a self-incompatible plant like R. glutinosa, seeds would not be produced in populations composed of a single genet. In the habitat of R. japonica, small ramets with only one or two small leaves were found that looked like seedlings. However, we found that these small ramets were physically connected by rhizome with flowering ramets, and the multilocus genotypes of all sampled ramets (small or flowering) were identical. Therefore, the probability of the existence of ramets having different genotypes is small. Most likely, only a single genet survives in the R. japonica populations in Japan.
Implications for conservation
The present findings suggest that R. japonica has high conservation value due to morphological and genetic differentiation from the other six species of Rehmannia and its extremely limited number of the populations. Rehmannia japonica is likely to remain as a single genet, and we should consider the genetic and ecological circumstances of this species when planning conservation measures. In general, it is thought that clonal plant without genetic variation can be particularly vulnerable to permanent and sudden environmental changes (Callaghan et al. 1992), and R. japonica in Japan may be unable to adapt against environment change due to its limited number of ramets and lack of genetic variation. The fact that only two populations are remaining also leads to high vulnerability to the catastrophic disturbance of habitat. The stone walls which have been the habitat of R. japonica are old and likely not sustainable for a long time. These habitats are located on steep slopes and landslide-prone terrain. Each population could disappear easily with just one landslide. Therefore, monitoring and conservation measures such as maintenance of these stone walls are required to prevent the extinction of R. japonica in the in situ environment, and the construction of ex situ populations is needed to prevent unexpected extinction by catastrophic events.
References
Albach DC, Li HQ, Zhao N, Jensen SR (2007) Molecular systematics and phytochemistry of Rehmannia (Scrophulariaceae). Biochem Syst Ecol 35:293–300
Bremer B, Bremer K, Heidari N, Erixon P, Olmstead RG, Anderberg AA, Källersjö M, Barkhordarian E (2002) Phylogenetics of asterids based on 3 coding and 3 non-coding chloroplast DNA markers and the utility of non-coding DNA at higher taxonomic levels. Mol Phylogenet Evol 24:274–301
Callaghan TV, Carlsson BÅ, Jónsdóttir IS, Svensson BM, Jonasson S (1992) Clonal plants and environmental change: introduction to the proceedings and summary. Oikos 63:341–347
Chin CL (1998) Rehmannia. In: Wu ZY, Raven PH (eds) Scrophulariaceae through Gesneriaceae, Flora of China, vol 18. Science Press, Beijing and Missouri Botanical Garden Press, St. Louis, pp 53–55
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659
Goudet J (1995) FSTAT (version 1.2): a computer program to calculate F-statistics. J Hered 86:485–486
Hara H (1949) Enumeratio Spermatophytarum Japonicarum 1. Iwanami Shoten, Tokyo
Hong DY, Yang HB, Jin CL, Holmgren NH (1998) Scrophulariaceae. In: Wu ZY, Raven PH (eds) Flora of China 18. Science Press, Beijing, pp 1–212
Isagi Y, Kaneko S (2014) Ubiquitous genotyping for conservation of endangered plant species. In: Nakano S et al (eds) Integrative observations and assessments. Springer, Tokyo, pp 311–326
Izuno A, Takamiya M, Kaneko S, Isagi Y (2011) Microsatellite loci in an endangered fern species, Athyrium viridescentipes (Woodsiaceae), and cross-species amplification. Am J Bot 98:e339–e341
Kaneko S, Kondo T, Isagi Y (2011) Development of microsatellite markers for the northern Australian endemic fan palm Livistona rigida (Arecaceae), with cross-amplification in the five related species. Conserv Genet Resour 3:697–699
Lian C, Hogetsu T (2002) Development of microsatellite markers in black locust (Robinia pseudoacasia) using a dual-suppression-PCR technique. Mol Ecol Notes 2:211–213
Lian C, Wadud MA, Geng Q, Shimatani K, Hougetsu T (2006) An improved technique for isolating codominant compound microsatellite markers. J Plant Res 119:415–417
Mabberley DJ (2008) Mabberley’s plant-book: a portable dictionary of plants, their classification and uses. Cambridge University Press, Cambridge
Makino T (1932) A contribution to the knowledge of the flora of Nippon. J Jpn Bot 8:24
Matsuda S (1918) Dziwau-zoku-oyobi-hanadziau-ni-tsuite (On Rehmannia and R. glutinosa Libosch. var. makinoi Matsuda, var. nov.). Bot Mag Tokyo 32:140–142 (in Japanese)
Milligan B (1992) Plant DNA isolation. In: Hoelzel AR (ed) Molecular genetic analysis of populations: a practical approach. IRL Press, Oxford, pp 59–88
Ministry of Environment of Japan (2015) Red Data Book 2014-Threatened Wildlife of Japan-Vol. 8, Vascular Plants. GYOSEI, Tokyo (in Japanese)
Mori K, Kaneko S, Isagi Y, Murakami N, Kato H (2008) Isolation and characterization of 10 microsatellite loci in Callicarpa subpubescens (Verbenaceae), an endemic species of the Bonin Islands. Mol Ecol Resour 8:1423–1425
Olmstead RG, Wolfe AD, Young ND, Elisons W, Reeves PA (2001) Disintegration of the Scrophulariaceae. Am J Bot 88:348–361
Oxelman B, Kornhall P, Olmstead RG, Bremer B (2005) Further disintegration of Scrophulariaceae. Taxon 54:411–425
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics Methods and Protocols: Methods in Molecular Biology. Humana Press, Totowa, pp 365–386
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Xia Z, Wang YZ, Smith JF (2009) Familial placement and relations of Rehmannia and Triaenophora (Scrophulariaceae sl) inferred from five gene regions. Am J Bot 96:519–530
Yamazaki T (1961) Meeting with Rehmannia japonica Makino. J Jpn Bot 36:351–352 (in Japanese)
Yamazaki T (1993) Scrophulariaceae 17. Rehmannia. In: Iwatsuki K, Yamazaki T, Goufford DE, Ohba H (eds) Flora of Japan IIIa. Kodansha, Tokyo, pp 343–344
Yonekura K (2012) An enumeration of the vascular plants of Japan. Hokuryukan, Tokyo
Zhou Y, Gu F, Zhou C, Chu S (2010) Genetic diversity of Rehmannia glutinosa cultivars based on sequence-related amplified polymorphism markers. Sci Hortic 125:789–794
Acknowledgements
The authors wish to acknowledge T. Kurosawa for manuscript preparation. We also thank S. Ohba, J. Fukui, and S. Sakaguchi for sampling. This work was supported by the Global Environment Research Fund (D-0903) of the Ministry of the Environment, Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kaneko, S., Matsuki, Y., Qiu, YX. et al. Chloroplast DNA sequencing and detailed microsatellite genotyping of all remnant populations suggests that only a single genet survives of the critically endangered plant Rehmannia japonica . J Plant Res 130, 117–124 (2017). https://doi.org/10.1007/s10265-016-0873-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-016-0873-5