Abstract
The poor prognosis of hepatocellular carcinoma (HCC) patients is mainly due to cancer metastasis. Methionine adenosyltransferase 2β (MAT2B) encodes a regulatory subunit (β) for methionine adenosyltransferase. Previous studies reveal that MAT2B provides a growth advantage for HCC, but its role in metastasis is unknown. This study showed that both in the xenograft zebra fish model and in the lung metastasis model in nude mice, the stable inhibition of MAT2B could suppress the metastasis of HCC cancer cells. Silencing of MAT2B in HCC cell lines could remarkably inhibit migration and invasion. By analysis of human phospho-kinase array membranes, we found several differentially expressed proteins, including phosphor-AKT, phospho-EGFR, phospho-Src family, phospho-FAK, phospho-STAT3 and phospho-ERK. We further confirmed the change of these EGFR pathway-related proteins was in accordance with MAT2B expression pattern through immunoblotting test. Finally, we found that MAT2B was overexpressed in HCC caner tissues and correlated with poor prognosis for HCC patients in clinical manifestation. Our study demonstrated that silencing of MAT2B could suppress liver cancer cell migration and invasion through the inhibition of EGFR signaling, which suggested that MAT2B might serve as a new prognostic marker and therapeutic target for HCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is the third leading cause of cancer death in the world [1]. Meanwhile, there are from 0.25 to 1 million newly diagnosed HCC cases each year [2,3,4]. Cancer metastasis remains the major challenge to successful management of malignant disease. The poor prognosis of HCC patients is also mainly due to cancer metastasis and recurrence. However, the underlying molecular mechanism involved in the metastasis process of HCC remains to be elusive.
Methionine adenosyl transferase (MAT) is a critical cellular enzyme responsible for generating major methyl donor, S-adenosylmethionine (SAMe), by using methionine and adenosine triphosphate [5]. Two genes (MAT1A and MAT2A) encode the catalytic subunit (α1 and α2) of different MAT isoforms, whereas a third gene (MAT2B) encodes a regulatory subunit (β) that modulates the kinetic properties of MAT2A-encoded isoenzyme [6].
MAT expression is switched from MAT2A to MAT1A during liver development, whereas it is reversed to MAT2A again during liver malignant transformation [7]. MAT1A is often silenced; instead, MAT2A and MAT2B are induced in liver cirrhosis and HCC patients [7,8,9]. Previous studies indicated that MAT2A overexpression promotes cancer cell proliferation and progression [10,11,12]. In addition, knockdown MAT2A dramatically suppresses tumor cell proliferation and induces cell cycle arrest and apoptosis [13,14,15]. Therefore, the transcriptional change from MAT1A to MAT2A is believed to play a critical role in facilitating cancer cell survival and proliferation [13, 16].
MAT2B is best known for regulation of MAT2A-encoded enzymatic activity; mounting evidence from previous studies has revealed a much broader role for MAT2B protein in cancer biology. For instance, MAT2B has been shown to offer a growth advantage in HCC [17], and most of the reports found a correlation between MAT2B and cancer cell proliferation, apoptosis and cell cycle arrest [13, 17,18,19]. However, to our knowledge, the role of MAT2B in progression and metastasis of HCC has not yet been explored.
In the current study, we tried to elucidate the functional contribution and the potential mechanisms of MAT2B in regulating the metastasis process of HCC. We revealed the correlation between MAT2B expression and clinicopathological features of HCC patients and found that MAT2B plays a key role in migration and invasion of liver cancer cells and mediated the process through EGFR signaling pathway. Our data suggested that MAT2B might serve as a potential biomarker for prognosis and a promising target for therapy in liver cancer.
Materials and methods
IHC staining of human tumor tissue array and patient follow-up
Human HCC tumor tissue array contained 80 patients with follow-up data, and IHC was stained with MAT2B-specific antibody against human MAT2B (Abcam, USA) by Shanghai Biochip (Shanghai, China), using the Dako Cytomation EnVision+ System-HRP (DAB) detection kit (Dako, USA). Briefly, the tissue array sections in 5 mm were dehydrated and subject to peroxidase blocking. MAT2B antibody was added at a dilution of 1:200 and incubated at room temperature for 30 min on the Dako Autostainer (Carpinteria, CA, USA) using the Dako Cytomation EnVisionþ System-HRP (DAB) detection kit. The slides were counterstained with hematoxylin. The stained slides were observed under microscope, and images were acquired. Tumor differentiation was defined according to the edmondson grading system. The detailed clinicopathological characteristics are given in Table 1.
HCC fresh tissue sample collection
Fresh HCC tissues and their adjacent liver tissues were collected from 16 patients undergoing resection of HCCs from February 2004 to December 2008 at Huashan Hospital (Shanghai, China). HCC diagnosis was based on the World Health Organization criteria. Ethical approval was obtained from the research ethics committee of Huashan Hospital, and written informed consent was obtained from each patient.
Cell culture and reagents
Human liver cancer cell line HepG2 was obtained from the American Type Culture Collection (Manassas, VA, USA), and the highly spontaneous metastatic human HCC cell line LM6 [20, 21] was obtained from the Liver Cancer Institute, Zhongshan Hospital, Fudan University (Shanghai, China). SMMC-7721 was obtained from the Type Culture Collection of Chinese Academy of Sciences (Shanghai, China). HepG2 and LM6 were cultured in DMEM (Dulbecco’s modified Eagle’s medium) (Gibco, USA), and SMMC-7721 was cultured in RPMI-1640 (Gibco, USA), both containing 10% fetal bovine serum (Gibco, USA) and 1% penicillin–streptomycin solution (Beyotime, China), at 37 °C with 5% CO2. Stable cell lines were selected and maintained in 2 mg/ml puromycin.
siRNA silencing
Liver cancer cells were transfected with siRNA oligonucleotides using Lipofectamine RNAiMAX reagent (Invitrogen, USA). Briefly, siRNA and Lipofectamine RNAiMAX reagent were each incubated separately with Opti-MEM (Gibco, USA) for 5 min and mixed together for 20 min at room temperature, and then, the mixture was applied to the cells plated in 4 ml of medium (final concentration of siRNA is 60 nM). The sequences of siRNAs are as follows: for MAT2B, siMAT2B-1:5′-GAAUGCUGGAUCCAUCAAUTT-3′, siMAT2B-2: 5′-GUUUGAAGAAAGGAACUUUTT-3′; and for control scrambled siRNA, siControl: 5′-TTCTCCGAACGTGTCACGTTT-3′. All the above siRNA oligonucleotides were synthesized by RiboBio (Shenzhen, China).
Wound healing assay
Cells from each cell line were seeded in 6-well plates at a density of 3 × 105 cells per well in complete DMEM or RPMI-1640 medium. Cells at 100% confluence were starved in serum-free medium for more than 12 h dependent on different cell lines. Cells were then lightly and quickly scratched with a 200-μl pipette tip at the center of each well. After being removed of detached cells and washed with phosphate-buffered saline (PBS) three times, the cells were incubated with DMEM or RPMI-1640 supplemented with 10% FBS. The scratch wounds were photographed at 24 h under an inverted microscope (Leica, Germany). The moving distance was measured by ImageJ software (National Institutes of Health, USA). The formula which we used to calculate and analyze the healing index that represents the ability of cellular migration was (wound area (initial) − wound area (final))/wound area (initial) × 100%.
Kinetics of cell migration
To monitor cell migration in a label-free real-time manner, we used an impedance-based xCELLigence real-time cell analyzer (RTCA) DP instrument (ACEA Biosciences, USA) equipped with a custom RTCA CIM-Plate, which is a 16-well system, and each well is composed of upper and lower chambers. The bottom surface of the upper chamber is composed of a microporous membrane coated with gold electrodes. Migration was measured as the relative impedance change (cell index) across the upper chamber. For the experiments, the lower chambers were filled with fresh medium containing 10% FBS and the upper chambers were seeded into 5 × 104 cells per well diluted with serum-free medium. CIM plates were loaded into the RTCA station in the cell culture incubator immediately; cell index (CI) was measured periodically every 15 min. The data were analyzed using the provided RTCA software. Experiments were performed in triplicate.
Human phospho-kinase array test and immunoblotting
LM6 cells were treated with siMAT2B or siControl,both cell lysates were extracted with cell lysis buffer (Beyotime, China), and the protein concentration in the lysates was quantified using an Enhanced BCA Protein Assay Kit (Tiangen, China). Detection of 43 kinase phosphorylation sites was performed on human phospho-kinase array membranes (R&D Systems, USA) using 300 μg total protein lysates according to the manufacturer’s instructions. Protein samples with 30–50 μg were loaded for immunoblotting, using antibodies against phospho-EGFR (Thr669), phospho-Src family (Tyr416), phospho-FAK (Tyr397), phosphor-AKT (Ser473), phospho-STAT3 (Tyr705) (Cell Signaling Technology, USA, dilution: 1:1000), MAT2B (Abcam, USA, dilution: 1:1000), phospho-ERK (Tyr204) (Santa Cruz Biotechnology, USA, dilution: 1:500) and Tubulin (Kangwei, China, dilution: 1:2000).
Xenografts in zebrafish
The liver cancer cells which were transfected with siMAT2B-1 and siControl for 24 h stained with 5-(and-6)-carboxyfluorescein diacetate, succinimidyl ester (5(6)-CFDA, SE; CFSE) (Life Technologies, USA). After labeling, the cells were collected and resuspended in PBS before injection. Wild-type AB zebrafish embryos were dechorionated with sharp forceps at 48 h post-fertilization (dpf) and then anaesthetized with 0.02 mg/ml tricaine (Sigma-Aldrich, USA). The anesthetized embryos were emplaced for microinjection in a dish which was coated with 1% agarose. Approximately 150–200 cells were injected into the yolk sac of embryos using an electronically regulated air-pressure micro-injector. Injected zebrafish embryos were maintained in E3 medium at 33 °C. At least 15 zebrafishes were randomly selected and then photographed by stereo microscope (Leica, Germany). The tumor mass was calculated and analyzed by ImageJ software.
Tumor formation assay
Five-week-old female athymic nude mice were purchased from the Shanghai Experimental Animal Center (Shanghai, China). LM6 cells which are stable transfected with shMAT2B or shControl cultured for 24 h were trypsinized, resuspended in PBS and then subcutaneously injected into the armpit with 1 × 106 cells per injection. Tumor size was measured by a vernier caliper weekly and calculated as (length × width2)/2. All procedures were performed in accordance with the National Institutes of Health Guide for the Care and Use of Laboratory Animals.
Statistical analysis
The statistical significance for the differences between groups was examined using the Prism6 software (GraphPad Software, Inc., USA). The unpaired two-tailed t test was used for the comparison of parameters between two groups. We estimated the probabilities of overall survival rate according to the Kaplan–Meier method. For all the tests, three levels of significance (p < 0.05, p < 0.01 and p < 0.001) were used.
Results
High MAT2B expression predicted a poor prognosis in HCC patients
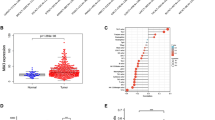
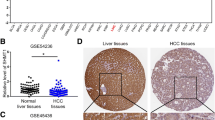
To investigate the clinical significance of MAT2B in HCC, we first examined the expression of MAT2B in tumor sites and adjacent liver tissues by immunohistochemistry (IHC) staining of human HCC tissue array containing 80 primary HCC tissues and their adjacent liver tissues. As shown in Fig. 1a, we found that MAT2B was overexpressed in the tumor tissues compared with adjacent liver tissues. Based upon the intensity of staining, we classified the samples into four groups with increasing staining intensity from weak (+) to the strongest (++++ , Fig. 1b). MAT2B expression was low (groups 1(+) and 2(++)) in most adjacent normal tissues (66.3%), but it was high (groups 3(+++) and 4(++++)) in most of the liver cancer tissues (65.0%, Fig. 1c). We found that the mRNA level of MAT2B in HCC tissues was much higher as compared with their adjacent liver tissues (a total of 16 paired cancers versus adjacent samples, Fig. 1d). Then, we further quantified the protein level of MAT2B in HCC and adjacent tissues by immunoblotting (IB) analysis. As shown in Fig. 1e, f the level of MAT2B in tumor tissues was significantly higher than that in the surrounding adjacent liver tissues from the same patient. Next, we analyzed the correlation of MAT2B expression status (low expression versus high expression) with clinicopathological features of HCC patients. We found that the MAT2B expression in the HCC tissues was negatively correlated with the survival rate of the patients (Fig. 1g, p = 0.012). Moreover, MAT2B expression seemed to be positively correlated with tumor size (Table 1, p = 0.010). However, no correlation between MAT2B expression and other clinical parameters (e.g., age and tumor location) was observed. These results indicated that MAT2B overexpression may be required for liver tumorigenesis. Thus, MAT2B overexpression has a potential to be developed as a prognosis biomarker for HCC patients.
MAT2B expression is elevated in human HCCs and negatively correlated with patient survival. a Representative image of IHC staining performed for human HCC tissue array using MAT2B-specific antibody, as described in “Materials and methods”. b and c Classification of samples according to the intensity of staining of MAT2B expression (n = 80). d qRT-PCR to measure relative expression levels of MAT2B in HCC tissue samples as compared with their adjacent liver tissue samples (n = 15). e IB analysis to determine expression of MAT2B in HCCs and adjacent liver tissues (n = 16). A, adjacent liver tissues; T, HCC tumor tissues. f Quantification of MAT2B expression in HCCs and adjacent liver tissues (n = 16). g Correlation analysis of MAT2B expression status and patient survival (n = 80). Data are presented as the mean ± SEM (*p < 0.05, ****p < 0.0001)
MAT2B is a key factor for migration and invasion of liver cancer cells in vitro
To evaluate the role of MAT2B in the migration and invasion of liver cancer cells, we knocked down MAT2B in HepG2, LM6 and SMMC-7721 cells using two siRNA oligos targeting two well-identified regions of MAT2B sequences (named siMAT2B-1 and siMAT2B-2, respectively, Fig. 2a). MAT2B knockdown significantly inhibited the motility compared with scrambled control in HepG2, LM6 and SMMC-7721 cells in wound healing assay in 24 h (Fig. 2b). Then, we further investigated the potential roles of MAT2B in migration ability using a real-time cell analyzer (RTCA) assay. As shown in Fig. 2c, MAT2B siRNA transfection significantly decreased the migration ability in all these three kinds of liver cancer cell lines. In addition, by matrigel-coated modified Boyden chamber assay, we also found that knockdown of MAT2B in liver cancer cells can inhibit the invasive property compared to the negative control cells in the HepG2, LM6 and SMMC-7721 cell lines (Fig. 2d).
MAT2B knockdown inhibits liver cancer cell migration and invasion in vitro. a MAT2B knockdown efficacy via two different siRNA silencing sequences (siMAT2B-1 and siMAT2B-2) assessed with IB analysis in HepG2, HCC-LM6 and SMMC-7721 cell lines. Transwell assays after incubation for 24h (b) and real-time cell analyzer (RTCA) assays (c) for assessing cell migration ability in HepG2, HCC-LM6 and SMMC-7721 cell lines. d Matrigel-coated modified Boyden chamber assays for assessing cell invasion ability in HepG2, HCC-LM6 and SMMC-7721 cell lines. Data are presented as the mean ± SEM (**p < 0.01, ***p < 0.001)
To further confirm the role of MAT2B in HCC development, we overexpressed with MAT2B into SMMC-7721 cells. Increased MAT2B was evident based on western blot analysis (Fig. 3a). Overexpression of MAT2B significantly promoted the motility compared with pcDNA3.1 control in SMMC-7721 cells in wound healing assay in 48 h (Fig. 3b), whereas RTCA assay showed that the migration ability was also elevated in MAT2B overexpression group (Fig. 3c). Thus, overexpression of MAT2B led to increased cell viability and migration ability in SMMC-7721 cells.
Overexpression of MAT2B promotes migration in SMMC-7721 cells. a MAT2B overexpression efficacy via three different plasmids (1#, 2# and 3#) assessed with IB analysis in SMMC-7721 cell lines. b Transwell assays after incubation for 48h for assessing cell migration ability in SMMC-7721 cell lines. c Real-time cell analyzer (RTCA) assays for assessing cell migration ability in SMMC-7721 cell lines. d IB analysis for indicated proteins, and MAT2B overexpression-induced p-EGFR, p-ERK and p-Src accumulation in SMMC-7721 cell lines. Data are presented as the mean ± SEM (*p < 0.05, **p < 0.01)
MAT2B knockdown depressed the migration and metastasis of liver cancer cells in vivo
Based on these in vitro findings, we further hypothesized that the ability of in vivo tumor formation and metastasis ability of MAT2B silenced cells would be significantly impaired when compared with shControl cells. To address this research inquiry, we next evaluated the effect of MAT2B knockdown on tumorigenesis using LM6 xenograft model of human HCCs with rapid growth upon transplantation into nude mice. We used lentiviral-transfected cell lines stably expressing shMAT2B and shControl in LM6 cells and then implanted the LM6 cells into the left or right flanks of nude mice, respectively. As shown in Fig. 4a, the tumor growth in the shMAT2B group was significantly inhibited by 58.5% in 8 weeks when compared with that of the shControl group (average tumor volume: 150.8 mm3 ± 49.5 mm3 vs. 362.9 mm3 ± 49.2 mm3, p = 0.009). Meanwhile, we found that the metastasis nudes in lung were also decreased in shMAT2B group (Fig. 4b). To confirm our findings, we further used zebrafish to detect the migration rate of both groups; we found that the tumor cell migration significantly decreased in the shMAT2B group than that of the shControl group (Fig. 4c).
Silencing of MAT2B inhibits invasive growth of HCC-LM6 cells in vivo. a Images for xenograft tumor on the nude mouse are shown in the left panel and tumor growth curve is shown in the right panel. (n = 7 for shControl group and n=8 for shMAT2B group). b Representative H&E images of subcutaneous tumors and lung metastasis nodes derived from shControl and shMAT2B HCC-LM6 cells. c Representative images of tumor cell migration ability in zebrafish model in both shMAT2B group and shControl groups. Data are presented as the mean ± SEM (*p < 0.05, **p < 0.01 and ***p < 0.001)
MAT2B knockdown inhibited liver cancer cell migration partially through EGFR-dependent signaling pathway
Since knockdown of MAT2B caused cells to exhibit less malignant phenotypes, we then tried to gain further insight into underlying molecular mechanism. Protein phosphorylation, especially kinase phosphorylation, is closely related to tumor metastasis. Thus, we firstly analyzed the phosphorylation profiles of kinases and their protein substrates by human phospho-kinase array. We found several epidermal growth factor receptor (EGFR) signaling pathway-related proteins were differentially expressed, including phosphor-AKT, phospho-EGFR, phospho-Src family, phospho-FAK, phospho-STAT3 and phospho-ERK (Fig. 5a). Since p38 mitogen-activated protein kinase (p38 MAPK) signaling cascade was a major downstream of EGFR pathway, we further confirmed the changes of these proteins were along with the change of MAT2B expression through immunoblotting test (Figs. 3d, 5b).
MAT2B knockdown suppresses human HCC progression through inactivation of EGFR. a Human Phospho-Kinase Array (R&D Systems, USA) for the relative expression of phosphorylation profiles of kinases and their protein substrates. b IB analysis for the indicated proteins and MAT2B silencing-induced downregulated expression of p-EGFR, p-ERK, p-FAK, p-STAT3, p-ERK and p-Src in HepG2, HCC-LM6 and SMMC-7721 cell lines
Discussion
HCC is the most common primary liver malignancy in adults [22,23,24]. The number of HCC-related deaths almost equals to the number of cases being diagnosed each year (more than 560,000), and the 5-year survival rate is below 9% [25]. The poor prognosis of HCC is mainly because of cancer metastasis within or outside the liver. In past several years, numerous molecules have been reported to be vital in recurrence and metastasis of HCC, such as ras homolog gene family, epidermal growth factor-like domain, multiple 7 (Egfl1) and micro-RNAs (miRNAs) [26,27,28,29]. However, the underlying mechanisms responsible for invasion and metastatic of HCC are still not completely understood. Therefore, further understanding of the molecular mechanism underlying the progression of metastasis and identifying novel metastatic factors are important for HCC prevention as well as treatment.
As we know, MAT2B, which regulates MAT2A-encoded isoenzyme, is overexpressed in human cirrhosis and HCC and has been shown to confer a growth advantage [17]. In hepatocellular carcinoma (HCC), MAT1A is often silenced, whereas MAT2A and MAT2B are induced [7,8,9]. Although MATβ is best known for regulation of MATII enzymatic activity, many recent studies revealed a much broader role for MAT2B protein in cancer biology. Thus, our hypothesis was that MAT2B also serves a vital function in metastasis of HCC.
Firstly, by analyzing in wound healing assay and RTCA, we revealed that silencing of MAT2B could lead to the inhibition of the migration and invasion ability in vitro and this simulation could be reversed when MAT2B was overexpressed in liver cancer cell lines. Secondly, we aimed to confirm whether downregulation of MAT2B can also inhibit the metastasis of liver cancer cells in vivo. After constructing lentiviral-transfected cell lines stably expressing shMAT2B and shControl in LM6 cells, we applied them into zebrafish and node mice model and found that silencing of MAT2B could lead to the suppression of the ability of tumor formation and metastasis ability in vivo.
Although the above results suggested that MAT2B was important in the metastasis process of HCC because we can partially inhibit liver cancer cell migration and metastasis through knockdown of MAT2B, the underlying mechanism remains unclear. Considering that protein phosphorylation is closely related to tumor metastasis, we hope to find some breakthroughs in protein phosphorylation chips. After unscrambling the phosphorylation profiles of kinases and their protein substrates by human phospho-kinase array, we found several EGFR signaling pathway-related proteins were differentially expressed in MAT2B knockdown group.
According to previous reports, EGFR system plays an essential role in cell proliferation, survival and migration and its alteration has been implicated in the development and growth of many tumors including HCC [30,31,32,33].Accordingly, the overexpression of EGFR and some of its ligands have been correlated with more aggressive liver tumors and poor survival [34, 35]. The interactions between EGFR and its ligands would trigger intracellular signaling pathways resulting in the activation of extracellular signal-regulated kinase (ERK), c-jun NH2-terminal kinase (JNK) and p38 mitogen-activated protein kinase (p38 MAPK) through the ras/raf/MEK/MAPK cascade, the protein kinase C (PKC) pathway, the PI3 K/Akt pathway (which can lead to NF-κB activation) and the STAT pathway [33, 36, 37]. Researchers also have found that MAT2B/GIT1 interplay is essential to MEK/ERK activation and could promote both hepatocellular carcinoma cell and colon cancer cell proliferation [38]. As mentioned above, the continuous activation of EGFR signaling is a hallmark of HCC and contributes to the proliferation, resistance to apoptosis and invasive behavior of HCC cells. Our results also showed that MAT2B silencing suppressed the phosphor-AKT, phospho-EGFR, phospho-Src family, phospho-FAK, phospho-STAT3 and phospho-ERK in HCC cancer cells. These results suggested that the molecular mechanism by which MAT2B inhibits the migration and invasion of HCC was partially through the EGFR signaling pathway.
Moreover, we found that MAT2B was overexpressed in human liver cancer tissues and the group of HCC patients with higher expression of MAT2B had shorter survival time. Our data also indicated that MAT2B expression may serve as an independent predictor for prognosis of liver cancer patients, although this needs to be confirmed with a larger cohort study.
In summary, our current study has provided mechanistic insights into how MAT2B promotes liver cancer progression. EGFR pathway was identified as an important pathway involved in regulating liver cancer cell migration and metastasis. MAT2B also serves as a potential biomarker for liver cancer prognosis.
References
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–86.
Llovet JM, Bruix J. Molecular targeted therapies in hepatocellular carcinoma. Hepatology. 2008;48:1312–27.
Feo F, Pascale RM, Simile MM, De Miglio MR, Muroni MR, Calvisi D. Genetic alterations in liver carcinogenesis: implications for new preventive and therapeutic strategies. Crit Rev Oncog. 2000;11:19–62.
Bosch FX, Ribes J, Díaz M, Cléries R. Primary liver cancer: worldwide incidence and trends. Gastroenterology. 2004;127:S5–16.
Lu SC, Mato JM. S-Adenosylmethionine in cell growth, apoptosis and liver cancer. J Gastroenterol Hepatol. 2008;23(Suppl 1):S73–7.
Halim AB, LeGros L, Geller A, Kotb M. Expression and functional interaction of the catalytic and regulatory subunits of human methionine adenosyltransferase in mammalian cells. J Biol Chem. 1999;274:29720–5.
Lu SC, Mato JM. S-adenosylmethionine in liver health, injury, and cancer. Physiol Rev. 2012;92:1515–42.
Yang H, Cho ME, Li TWH, Peng H, Ko KS, Mato JM, et al. MicroRNAs regulate methionine adenosyltransferase 1A expression in hepatocellular carcinoma. J Clin Investig. 2013;123:285–98.
Ramani K, Mato JM, Lu SC. Role of methionine adenosyltransferase genes in hepatocarcinogenesis. Cancers (Basel). 2011;3:1480–97.
Kotb M, Mudd SH, Mato JM, Geller AM, Kredich NM, Chou JY, et al. Consensus nomenclature for the mammalian methionine adenosyltransferase genes and gene products. Trends Genet. 1997;13:51–2.
Halim AB, LeGros L, Chamberlin ME, Geller A, Kotb M. Regulation of the human MAT2A gene encoding the catalytic alpha 2 subunit of methionine adenosyltransferase, MAT II: gene organization, promoter characterization, and identification of a site in the proximal promoter that is essential for its activity. J Biol Chem. 2001;276:9784–91.
LeGros L, Halim AB, Chamberlin ME, Geller A, Kotb M. Regulation of the human MAT2B gene encoding the regulatory beta subunit of methionine adenosyltransferase. MAT II. J Biol Chem. 2001;276:24918–24.
Wang Q, Liu Q, Liu Z-S, Qian Q, Sun Q, Pan D. Inhibition of hepatocelluar carcinoma MAT2A and MAT2beta gene expressions by single and dual small interfering RNA. J Exp Clin Cancer Res. 2008;27:72.
Zhang T, Zheng Z, Liu Y, Zhang J, Zhao Y, Liu Y, et al. Overexpression of methionine adenosyltransferase II alpha (MAT2A) in gastric cancer and induction of cell cycle arrest and apoptosis in SGC-7901 cells by shRNA-mediated silencing of MAT2A gene. Acta Histochem. 2013;115:48–55.
Liu Q, Wu K, Zhu Y, He Y, Wu J, Liu Z. Silencing MAT2A gene by RNA interference inhibited cell growth and induced apoptosis in human hepatoma cells. Hepatol Res. 2007;37:376–88.
Ramani K, Yang H, Kuhlenkamp J, Tomasi L, Tsukamoto H, Mato JM, et al. Changes in the expression of methionine adenosyltransferase genes and S-adenosylmethionine homeostasis during hepatic stellate cell activation. Hepatology. 2010;51:986–95.
Martínez-Chantar ML, García-Trevijano ER, Latasa MU, Martín-Duce A, Fortes P, Caballería J, et al. Methionine adenosyltransferase II beta subunit gene expression provides a proliferative advantage in human hepatoma. Gastroenterology. 2003;124:940–8.
Yang H, Ara AI, Magilnick N, Xia M, Ramani K, Chen H, et al. Expression pattern, regulation, and functions of methionine adenosyltransferase 2beta splicing variants in hepatoma cells. Gastroenterology. 2008;134:281–91.
Ramani K, Yang H, Xia M, Ara AI, Mato JM, Lu SC. Leptin’s mitogenic effect in human liver cancer cells requires induction of both methionine adenosyltransferase 2A and 2beta. Hepatology. 2008;47:521–31.
Mikol YB, Hoover KL, Creasia D, Poirier LA. Hepatocarcinogenesis in rats fed methyl-deficient, amino acid-defined diets. Carcinogenesis. 1983;4:1619–29.
Ghoshal AK, Farber E. The induction of liver cancer by dietary deficiency of choline and methionine without added carcinogens. Carcinogenesis. 1984;5:1367–70.
El-Serag HB, Rudolph KL. Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology. 2007;132:2557–76.
Perry JF, Poustchi H, George J, Farrell GC, McCaughan GW, Strasser SI. Current approaches to the diagnosis and management of hepatocellular carcinoma. Clin Exp Med. 2005;5:1–13.
Mazzoccoli G, Tarquini R, Valoriani A, Oben J, Vinciguerra M, Marra F. Management strategies for hepatocellular carcinoma: old certainties and new realities. Clin Exp Med. 2016;16:243–56.
Nordenstedt H, White DL, El-Serag HB. The changing pattern of epidemiology in hepatocellular carcinoma. Dig Liver Dis. 2010;42(Suppl 3):S206–14.
Wu F, Yang L-Y, Li Y-F, Ou D-P, Chen D-P, Fan C. Novel role for epidermal growth factor-like domain 7 in metastasis of human hepatocellular carcinoma. Hepatology. 2009;50:1839–50.
Ye Q-H, Qin L-X, Forgues M, He P, Kim JW, Peng AC, et al. Predicting hepatitis B virus-positive metastatic hepatocellular carcinomas using gene expression profiling and supervised machine learning. Nat Med. 2003;9:416–23.
Luedde T. MicroRNA-151 and its hosting gene FAK (focal adhesion kinase) regulate tumor cell migration and spreading of hepatocellular carcinoma. Hepatology. 2010;52:1164–6.
Yang F, Yin Y, Wang F, Wang Y, Zhang L, Tang Y, et al. miR-17-5p Promotes migration of human hepatocellular carcinoma cells through the p38 mitogen-activated protein kinase-heat shock protein 27 pathway. Hepatology. 2010;51:1614–23.
Zandi R, Larsen AB, Andersen P, Stockhausen M-T, Poulsen HS. Mechanisms for oncogenic activation of the epidermal growth factor receptor. Cell Signal. 2007;19:2013–23.
Berasain C, Castillo J, Prieto J, Avila MA. New molecular targets for hepatocellular carcinoma: the ErbB1 signaling system. Liver Int. 2007;27:174–85.
Breuhahn K, Longerich T, Schirmacher P. Dysregulation of growth factor signaling in human hepatocellular carcinoma. Oncogene. 2006;25:3787–800.
Citri A, Yarden Y. EGF-ERBB signalling: towards the systems level. Nat Rev Mol Cell Biol. 2006;7:505–16.
Kira S, Nakanishi T, Suemori S, Kitamoto M, Watanabe Y, Kajiyama G. Expression of transforming growth factor alpha and epidermal growth factor receptor in human hepatocellular carcinoma. Liver. 1997;17:177–82.
Daveau M, Scotte M, François A, Coulouarn C, Ros G, Tallet Y, et al. Hepatocyte growth factor, transforming growth factor alpha, and their receptors as combined markers of prognosis in hepatocellular carcinoma. Mol Carcinog. 2003;36:130–41.
Liebmann C. EGF receptor activation by GPCRs: an universal pathway reveals different versions. Mol Cell Endocrinol. 2011;331:222–31.
Jorissen RN, Walker F, Pouliot N, Garrett TPJ, Ward CW, Burgess AW. Epidermal growth factor receptor: mechanisms of activation and signalling. Exp Cell Res. 2003;284:31–53.
Peng H, Dara L, Li TWH, Zheng Y, Yang H, Tomasi ML, et al. MAT2B-GIT1 interplay activates MEK1/ERK 1 and 2 to induce growth in human liver and colon cancer. Hepatology. 2013;57:2299–313.
Funding
This work was supported by grant from the National Natural Science Foundation of China ‘(81401980)’ (to Lijun Wu) and the Development Fund for Shanghai Talents ‘(201660)’ (to Dongqin Yang) and the Natural Science Foundation and Major Basic Research Program of Shanghai ‘(16JC1420104)’ (to Jie Liu).
Author information
Authors and Affiliations
Contributions
Lijun Wu was responsible for manuscript preparation and contributed to study design and experiment process. Ping Chen contributed to the data analysis and experiment process. Dongqin Yang contributed to study design. All co-authors contributed to the data interpretation and manuscript revision. All authors approved the final version of this manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wu, L., Chen, P., Ying, J. et al. MAT2B mediates invasion and metastasis by regulating EGFR signaling pathway in hepatocellular carcinoma. Clin Exp Med 19, 535–546 (2019). https://doi.org/10.1007/s10238-019-00579-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10238-019-00579-2