Abstract
Background
Tubular injury plays a crucial role in the pathogenesis of diabetic nephropathy (DN). It is well known that many microRNAs (miRNAs) exert crucial effects on tubular injury. This study intends to explore the effect of miR-142-3p on the apoptosis and oxidative stress of high glucose (HG)-treated renal tubular epithelial cells (HK-2) and its underlying mechanism.
Materials and methods
HK-2 cells were exposed to HG to mimic cell injury. MTT assays and flow cytometry analyses were conducted to measure cell viability and cell apoptosis, respectively. RT-qPCR and western blot analyses were carried out to detect RNA and protein levels, respectively. The levels of oxidative stress markers were evaluated by ELISA. The binding between miR-142-3p and biorientation of chromosomes in cell division 1 (BOD1) was validated by a luciferase reporter assay.
Result
MiR-142-3p is low-expressed in HG-stimulated HK-2 cells. Functionally, miR-142-3p overexpression attenuates the apoptosis and oxidative stress of HG-stimulated HK-2 cells. Mechanistically, BOD1 was confirmed to be targeted by miR-142-3p in HK-2 cells. Moreover, BOD1 overexpression reversed the suppressive effect of miR-142-3p overexpression on the apoptosis and oxidative stress of HK-2 cells treated with HG.
Conclusion
MiR-142-3p ameliorates HG-induced renal tubular epithelial cell injury by targeting BOD1. The finding might provide novel insight into the role of miR-142-3p/BOD1 axis in DN treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diabetic nephropathy (DN) is a common diabetic complication and a leading cause of end-stage renal diseases worldwide [1, 2]. DN is regarded as a main cause of mortality in patients diagnosed with diabetes [3]. In addition, DN is also a severe microvascular complication, and its early diagnosis is essential for DN treatment. However, there are no effective therapeutic methods for this complicated pathogenesis at present [4]. Since tubular injury is widely recognized to be associated with the pathogenesis of DN [5, 6], studies on new targets for tubular injury under high glucose condition are necessary for exploring prospective therapies for DN treatment.
MicroRNAs (miRNAs) are small noncoding RNAs with 19–24 nucleotides in length, which are crucial modulators of gene expression in many diseases [7, 8]. Specifically, miRNAs modulate gene expression by binding with 3’untranslated region (3’UTR) of messenger RNAs (mRNAs) to accelerate the degradation or limit the translation of mRNAs at the post-transcriptional level [9,10,11]. In addition, miRNAs can affect many cellular processes including cell proliferation, differentiation, apoptosis and death [12,13,14]. Growing evidence has highlighted the important role of miRNAs in the pathogenesis of DN [15,16,17], and the miRNA-mRNA-regulating network has also been verified in DN [18, 19]. For example, miR-98 directly targets 3’UTR of mRNA neural precursor cell expressed developmentally downregulated gene 4-like (Nedd4L) to promote pathological damage and fibrosis in DN rats and to facilitate the apoptosis of HG-treated renal tubular epithelial cells [20]. MiR-192 prevents renal fibrosis in DN by targeting 3’UTR of early growth response factor 1 (Egr1) [21]. MiR-30e binds with 3’UTR of GLIPR-2 to downregulate GLIPR-2 expression, thus prevents renal fibrosis in DN [22]. Previously, miR-142-3p was reported to serve as a biomarker in renal fibrosis [23]. Moreover, miR-142-3p was revealed to modulate the development of diabetes and serve as a biomarker for type II diabetes [24, 25]. Mechanistically, miR-142-3p interacts with 3’UTR of Forkhead box protein O1 (FOXO1) in gestational diabetes mellitus [26]. However, the specific role of miR-142-3p in DN remains unknown.
In our study, we intended to explore the biological function and molecular mechanism of miR-142-3p in DN by assessing the effects of miR-142-3p and its target gene on HG-induced renal tubular epithelial cell injury. This finding might provide novel insight into the role of miR-142-3p in the diagnosis and treatment of DN.
Materials and methods
Animals
Adult male Wistar rats (200–250 g) were purchased from Beijing Vital River Laboratory Animal Technology (Beijing, China) and housed under standard conditions (12 h light and dark).
Ethical statement
Animal experiments were approved by the animal ethics committee of Henan Provincial People’s Hospital (Henan, China) and conducted complying with all the relevant national regulations and institutional policies for the care and use of animals.
Rat model of diabetes
A rat model of diabetes was established by administering streptozotocin (STZ; 65 mg/kg of body weight; Sigma Aldrich, St. Louis, MO, USA) with 0.1 mol/L citrate buffer (LMAIBio, Shanghai, China) via intraperitoneal injection, according to the previous study [27]. Rats received citrate buffer were set as the normal control. On day 3 and day 5 after model establishment, blood glucose of these rats was examined. Successful modeling was defined as the concentration of blood glucose up to 16.7 mM in the two examinations.
Collection of blood samples and renal tubule
At 0, 4, 8, 12 and 16 weeks, serum was collected to determine levels of renal function indicators, i.e., serum creatinine and blood urea nitrogen (BUN), and proximal convoluted tubules were obtained at the same week interval. All rats were individually housed in metabolic cages at the above time points. Blood samples were collected from caudal vein over all time intervals, while serum was collected after 5 min of centrifugation at 4000 rpm and preserved at – 80 ℃. After the collection of blood samples, kidneys of these rats were perfused with normal saline through ascending aorta and then were removed. Proximal convoluted tubules were isolated for homogenate preparation.
Measurement of serum creatinine and blood urea nitrogen (BUN)
Creatinine assay kits (Abcam, Cambridge, UK) and QuantiChrom™ Urea Assay Kits (BioAssay Systems, Shanghai, China) were, respectively, employed for measuring serum creatinine and BUN following the manufacturer’s protocols.
Cell culture
Human renal tubular epithelial (HK-2) cells (Chinese Academy of Science Cell Bank, Shanghai, China) were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM; Gibco, Thermofisher Scientific, USA) containing 10% fetal bovine serum (FBS; Hyclone, South Logan, USA), 1% penicillin (100 U/mL) and streptomycin (100 mg/ml) and cultured at 37 ℃ with 5% CO2. To induce cell injury, HK-2 cells were incubated in serum-free medium with high glucose (HG, 25 mM) for 48 h. Cells in the control group were treated with normal glucose (NG, 5.5 mM) for 48 h.
Cell transfection
For cell transfection, HK-2 cells were seeded in 24-well plates (1 × 104 cells/well). MiR-142-3p mimic and its negative control (NC) mimic-NC were synthesized by Ribobio (Guangzhou, China). The full length of BOD1 was cloned into pcDNA3.1 vector (Ribobio) for overexpressing BOD1. Cell transfection was carried out using Lipofectamine 2000 (Invitrogen, USA). Cells were then kept under the condition of 5% CO2 for 48 h at 37 ℃.
Methylthiazolyldiphenyl-tetrazolium bromide (MTT) assay
The viability of HK-2 cells was evaluated by MTT assay using an MTT cell proliferation assay kit (Cayman, USA). Briefly, HK-2 cells (1 × 104 cells/mL) were grown in 96-well plates and transfected with indicated vectors for 24 h, 48 h or 72 h. Then, 10 μL MTT solution (Beyotime, Shanghai, China) was added to each well and incubated for another 4 h at 37 ℃ in a humidified atmosphere with 5% CO2. Finally, the absorbance was detected and recorded utilizing a microplate reader (Bio-Rad, USA) at 490 nm.
Cell apoptosis
The apoptosis of HK-2 cells was measured by an Annexin V-FITC/PI apoptosis detection kit (Beyotime, Shanghai, China). Briefly, cells (1 × 104 cells/well) were placed in 6-well plates. After being washed with PBS three times, HK-2 cells were resuspended in binding buffer. Afterwards, Annexin V-FITC solution (5 μl) and Propidium Iodide (PI) solution (5 μl) were added to each well for 15 min of incubation at room temperature in the dark. Cell apoptosis was examined by the FACSCalibur flow cytometry (BD, USA) and then analyzed by CellQuest software.
Reverse transcription quantitative polymerase chain reaction (RT-qPCR)
Total RNAs were isolated from HK-2 cells using TRIzol Reagent (Invitrogen, USA) and reverse transcribed into cDNA utilizing a High-Capacity RNA-to-cDNA Kit (Applied Biosystems, USA). The RT-qPCR was performed using the SYBR Green PCR Supermix kit (Bio-Rad, USA) and analyzed by the QuantStudio 7 Flex Real-Time PCR system (Applied Biosystems, USA). Gene expression levels were calculated by the 2−ΔΔCt calculation method. U6 was an internal control for miR-142-3p and GAPDH was an internal reference for candidate mRNAs.
Western blot analysis
Proteins were extracted using RIPA buffer containing cocktail. The protein concentrations were quantified using a BCA Protein Assay kit (Beyotime, Shanghai, China). Equal protein samples were subjected to 12% SDS-PAGE and then moved onto polyvinylidene difluoride (PVDF) membranes (Millipore, USA). Subsequently, the membranes were coated with 5% defatted milk powder in tris-buffered saline containing Tween®20 (TBST) for 2 h and then incubated with primary antibodies against Cleaved caspase-3 (ab2302, Abcam, Shanghai, China), BOD1 (ab122380, Abcam) and GAPDH (ab181602, Abcam) at 4 ℃ overnight. After washed with TBST, the horseradish peroxidase (HRP) conjugated secondary antibody (Zsbio, Beijing, China) was added for another 1 h of incubation at 37 ℃. Finally, the protein bands were visualized by enhanced chemiluminescence (ECL; Wanlei Biotechnology, Shanghai, China) and analyzed by ImageJ software.
Detection of malondialdehyde (MDA) content and superoxide dismutase (SOD) activity
MDA content in HK-2 cells was measured using colorimetrically thiobarbituric acid reactive substances (TBARS) assay (Cayman, USA) under manufacturer’s instructions. The SOD activity was examined with a SOD activity assay kit (Lianke, Hangzhou, China) following the manufacturer’s protocols. Enzymatic activity was presented as units per mg of protein. The samples were analyzed using a spectrophotometer (Bio-Rad, USA).
Detection of intracellular reactive oxygen species (ROS) and extracellular nitric oxide (NO)
The ROS production in HK-2 cells was evaluated by a ROS Assay kit (Beyotime). Briefly, HK-2 cells were grown in 96-well plates. After exposure to HG or transfection with different plasmids, 10 μM of 2’-7’-dichlorofluorescein diacetate (DCFH-DA) was added to each well for 1 h at 37 ℃ in the dark. After 30 min of incubation, each well was washed with PBS for three times and detected by a microplate reader (Biotek, USA) with an excitation wavelength of 488 nm and an emission wavelength of 535 nm. Nitric oxide (NO) release was detected using the NO Assay Kit (Beyotime). HK-2 cells were seeded in 96-well plates. Griess Reagent I (50 μL/well) and Griess Reagent II (50 μL/well) were added to each well. Finally, the absorbance at 540 nm wavelength was detected by a microplate reader (Biotek).
Luciferase reporter assay
The fragment from BOD1 3’UTR containing the predicted binding site of miR-142-3p was amplified by RT-qPCR and was cloned into a pmirGLO Dual-luciferase miRNA Target Expression Vector (Promega, USA) to construct the reporter vector BOD1-wild type (BOD1-WT). The reporter vector BOD1-mutant type (BOD1-WT) was also constructed. Then, the vectors were co-transfected with miR-142-3p mimic or mimic-NC into HK-2 cells. The luciferase activity was then detected by the dual-luciferase reporter assay system (Promega, USA) after 48 h of transfection.
Statistical analysis
All data were analyzed using SPSS 20.0 software (IBM SPSS Statistics, USA). Data are shown as the mean ± standard deviation (SD). Student’s t test was utilized for comparison between 2 groups, and one-way analysis of variance (ANOVA) followed by Tukey’s post hoc analysis was utilized for multiple comparisons more than 2 groups. The value of p < 0.05 was considered statistically significant.
Results
MiR-142-3p exhibits low expression in diabetic rats and is negatively correlated with serum creatinine and blood urea nitrogen (BUN) level
After the establishment of diabetic rat model and the collection of blood samples and renal tubules, RT-qPCR was conducted to analyze miR-142-3p expression in these rats. Low miR-142-3p expression was examined in renal tubules of diabetic rats than that in normal rats (Fig. 1A). In addition, miR-142-3p expression was decreased in a time-dependent manner and maintained at a stable level approximately at week 12 (Fig. 1A). Moreover, the levels of indicators for renal function (serum creatinine and BUN) were detected at indicated time points after model establishment. Spearman correlation analysis was used to identify the correlation between miR-142-3p expression and serum creatinine (or BUN) level. The results suggested that both serum creatinine and BUN levels were negatively correlated with miR-142-3p expression in tissues of diabetic rats (Fig. 1B, C).
MiR-142-3p exhibits low expression in renal tubules of diabetic rats and is negatively correlated with serum creatinine and BUN. A The expression of miR-142-3p in renal tubules and blood samples of normal rats and diabetic rats was examined by RT-qPCR at indicated time points. B The correlation between miR-142-3p expression and serum creatinine level was identified by Spearman correlation coefficient analysis. C MiR-142-3p expression was negatively correlated with blood urea nitrogen (BUN) level in tissues of diabetic rats, as suggested by Spearman correlation coefficient analysis. *p < 0.05, **p < 0.01
MiR-142-3p inhibits HG-induced HK-2 cell apoptosis
Previously, miR-142-3p was extensively reported as a biomarker for type II diabetes [24, 25], whereas the specific function of miR-142-3p in DN still needs to be explored. In our study, we found that miR-142-3p expression was decreased in a glucose-dependent manner. High concentration of glucose resulted in low expression of miR-142-3p (Fig. 2A). Similarly, cell viability was gradually suppressed by high concentration of glucose, as shown in MTT assays (Fig. 2B). HK-2 cells were stimulated via normal glucose (NG; 5.5 mM) or high glucose (HG; 45 mM), respectively. We discovered that miR-142-3p presented evident downregulation in HK-2 cells under HG stimulation (Fig. 2C). Next, we decided to probe the effect of miR-142-3p on the apoptosis of HG-induced HK-2 cells. RT-qPCR revealed that miR-142-3p expression was significantly elevated after the transfection of miR-142-3p mimic (Fig. 2C). Subsequently, MTT assay showed that HG inhibited the viability of HK-2 cells while miR-142-3p overexpression significantly rescued HG-induced inhibitory effect (Fig. 2D). As presented in flow cytometry analysis, the apoptosis of HK-2 cells was enhanced in HG group compared with that in NG group, while the promotion of cell apoptosis was reversed by overexpressing miR-142-3p (Fig. 2E, F). Western blot analysis was performed to examine the protein level of cleaved caspase-3, one of the apoptotic markers. We found that miR-142-3p overexpression reversed HG-mediated increase in cleaved caspase-3 protein level in HK-2 cells (Fig. 2G). These data indicated that miR-142-3p alleviates HG-induced injury on HK-2 cells.
MiR-142-3p inhibits HG-induced HK-2 cell apoptosis. A The expression of miR-142-3p in HK-2 cells treated with mannitol, different dose of glucose (15, 30, 45 mM) or without any treatment was measured by RT-qPCR. B The viability of HK-2 cells under the above treatment was evaluated by MTT assays. C RT-qPCR was performed to measure miR-142-3p level in HK-2 cells under NG or HG, and the transfection efficacy of miR-142-3p mimic was also confirmed. D MTT assay was conducted to measure the viability of HK-2 cells under NG or HG conditions and transfected with miR-142-3p mimic or mimic-NC. E, F Flow cytometry was employed to assess the apoptosis of HK-2 cells with the above treatment or transfection. G Western blot was conducted to evaluate the effect of miR-142-3p on cleaved caspase-3 protein level in HG-treated HK-2 cells. *p < 0.05, **p < 0.01
MiR-142-3p inhibits HG-induced oxidative stress in HK-2 cells
To investigate the effect of miR-142-3p on oxidative stress in HG-induced HK-2 cells, we first measured malondialdehyde (MDA) content. As shown in Fig. 3A, miR-142-3p overexpression reversed HG-induced increase in MDA content in HK-2 cells. Furthermore, overexpressing miR-142-3p rescued the suppressive effect of HG on SOD activity in the cells (Fig. 3B). In addition, we assessed intracellular ROS production and NO in the extracellular supernatant of HK-2 cells. The results suggested that both intracellular ROS and extracellular NO levels were remarkably upregulated under HG conditions and then reversed by miR-142-3p overexpression (Fig. 3C, D). In summary, miR-142-3p inhibits HG-stimulated oxidative stress in HK-2 cells.
MiR-142-3p inhibits HG-induced oxidative stress in HK-2 cells. A–D The effect of miR-142-3p mimic on levels of oxidative-related factors (MDA content, SOD activity, intracellular ROS level and extracellular NO release) in HG-stimulated HK-2 cells was measured by corresponding kits. *p < 0.05, **p < 0.01
BOD1 is targeted by miR-142-3p
We continued to explore the underlying mechanism of miR-142-3p. The Venn diagram showed that there are 10 overlapped mRNAs from the website of miRmap (https://mirmap.ezlab.org/) [28], PITA (https://genie.weizmann.ac.il/pubs/mir07/mir07_dyn_data.html) [29] and RNA22 (https://cm.jefferson.edu/data-tools-downloads/rna22-full-sets-of-predictions/) [30] (Fig. 4A). To identify the specific mRNA for study, we conducted RT-qPCR to explore the effect of miR-142-3p overexpression on expression levels of candidate mRNAs. Compared with the expression of other mRNAs, BOD1 expression was markedly downregulated by overexpressing miR-142-3p (Fig. 4B). Thus, we selected BOD1 for further exploration. Specific binding area between miR-142-3p and BOD1 3’UTR were obtained from TargetScan (http://www.targetscan.org/vert_72/) [31] (Fig. 4C). Dual-luciferase reporter experiment revealed that the luciferase activity of BOD1-WT was markedly decreased by miR-142-3p overexpression, while that of BOD1-Mut group was not significantly affected by miR-142-3p mimic (Fig. 4D). According to western blot analysis, miR-142-3p overexpression reversed HG-stimulated upregulation of BOD1 protein level in HK-2 cells (Fig. 4E). Overall, BOD1 is negatively modulated by miR-142-3p in HK-2 cells.
BOD1 is the target of miR-142-3p. A Ten potential mRNAs identified from miRmap, PITA and RNA22 databases were illustrated in the Venn diagram. B The expression levels of 10 potential mRNAs in HK-2 cells transfected with miR-142-3p mimic or NC mimic were examined by RT-qPCR. C The specific binding region of miR-142-3p on BOD1 3’UTR was predicted from TargetScan. D Dual-luciferase reporter experiment was performed to explore the binding ability between miR-142-3p and BOD1. E Western blot analysis was conducted to assess the effect of miR-142-3p on BOD1 protein level in HK-2 cells treated with HG. **p < 0.01
MiR-142-3p suppresses HK-2 cell apoptosis and oxidative stress by downregulating BOD1 expression
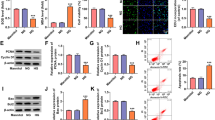
Rescue assays were conducted to figure out whether miR-142-3p suppressed HG-mediated HK-2 cell apoptosis and oxidative stress by targeting BOD1. As exhibited in western blot analysis, the protein level of BOD1 was dramatically overexpressed by transfection of pcDNA3.1/BOD1, and BOD1 overexpression reversed the suppressive effect of miR-142-3p on it in HG-treated HK-2 cells (Fig. 5A). As suggested by MTT assay, overexpressing BOD1 considerably reversed the promotion of cell viability induced by miR-142-3p mimic (Fig. 5B). Moreover, BOD1 overexpression rescued the suppressive effect of miR-142-3p overexpression on the apoptosis of HK-2 cells (Fig. 5C). Similarly, the downregulation of cleaved caspase-3 protein level mediated by miR-142-3p overexpression was rescued by overexpressing BOD1 (Fig. 5D). The decrease in MDA content and the increase in SOD activity induced by miR-142-3p mimic were counteracted by its co-transfection with pcDNA3.1/BOD1 (Fig. 5E, F). Moreover, BOD1 upregulation rescued the inhibitory effect of miR-142-3p overexpression on ROS and NO levels (Fig. 5G, H). These findings implied that miR-142-3p inhibits HK-2 cell apoptosis and oxidative stress by targeting BOD1.
MiR-142-3p attenuates HK-2 cell injury by downregulating BOD1 expression. A The overexpression efficiency of BOD1 in HG-treated HK-2 cells was detected by western blot analysis. B The viability of HK-2 cells in NC mimics, miR-142-3p mimic, miR-142-3p mimic + empty pcDNA3.1, and miR-142-3p mimic + BOD1 groups was analyzed by MTT assays. C Flow cytometry analysis was performed to assess the apoptosis rate of HK-2 cells in the above groups. D The protein level of cleaved caspase-3 in indicated groups was determined by western blot analysis. E, H The levels of oxidative-related factors (MDA content, SOD activity, intracellular ROS level and extracellular NO release) in the above groups were evaluated. *p < 0.05, **p < 0.01. ***p < 0.001
Discussion
DN, a common microvascular complication, resulted from chronic exposure to HG in patients with diabetes mellitus. Clinically, DN patients are generally diagnosed with glomerular hypertrophy, glomerulosclerosis, extracellular matrix accumulation and/or renal tubular epithelial cell injury [32, 33]. MicroRNAs (miRNAs) are small noncoding molecules containing approximately 22 nucleotides, which have the potential to become diagnostic markers and therapeutic targets for DN [34, 35]. Herein, miR-142-3p was found to be low-expressed in renal tubules and blood samples of diabetic rats, and its expression was negatively correlated with serum creatinine and BUN levels in tissues of diabetic rats. Since serum creatinine and BUN are indicators for renal functions, we explored the relationship between aberrant expression of miR-142-3p and diabetes-mediated renal tubule injury. MiR-142-3p expression was downregulated by HG, and miR-142-3p overexpression rescued HG-induced inhibitory effect on HK-2 cell viability and reversed the enhancement of HG on cell apoptosis.
Furthermore, accumulating evidences have demonstrated that oxidative stress is a major pathophysiological change in DN [36, 37]. In addition, oxidative stress may contribute to renal tubular epithelial cell apoptosis in DN [38], and therapies inhibiting oxidative stress might effectively maintain normal renal function and delay DN progression [39,40,41]. Thus, it is necessary to probe function of oxidative stress in renal tubular epithelial cell injury, which may provide an effective direction for DN treatment. In the present study, miR-142-3p reversed HG-mediated upregulation of MDA content, intracellular ROS production as well as extracellular NO release and rescued the decrease in SOD activity, suggesting that miR-142-3p ameliorated HG-induced HK-2 cell injury by inhibiting oxidative stress.
Despite HG-induced dysfunction of oxidative stress, autophagic dysfunction after HG treatment is also an important pathogenic process that might result in the initiation and progression of diabetic renal injury [42]. As a part of the catabolic process that degrades the injured proteins and organelles, autophagy contributes to the integrity of cell structure and functions under different stress conditions [42, 43]. Previously, the activation of autophagy was revealed to protect the proximal tubule from degeneration [44] and to delay the progression of diabetic renal injury [45]. MiR-142-3p suppresses the autophagy of 3T2-L1 cells in vitro and inhibits fat cell autophagy in obese mice [46]. MiR-142-3p overexpression inhibits the autophagy of cardiomyocytes [47]. In our future studies, we will further explore the association between miR-142-3p and HG-induced autophagy.
Mechanistically, abundant studies have indicated that miRNA can bind to 3’UTRs of their target genes, thereby regulating mRNA expression at the post-transcriptional level [41, 48]. Moreover, the molecular function of miRNA-mRNA has also been demonstrated in DN [49, 50], and the mRNA-regulating role of miR-142-3p at the post-transcriptional level has been validated in diabetes mellitus and many other diseases [26, 51, 52]. Hence, we hypothesized that miR-142-3p might act in the same pattern in DN. After bioinformatics prediction and experimental screening, mRNA biorientation of chromosomes in cell division 1 (BOD1) was found to be targeted by miR-142-3p in HK-2 cells. A previous study claimed that miR-142-3p interacted with BOD1 in carcinoma cells [53]. In addition, BOD1 was reported to regulate the chromosome biorientation and associated with adult height [54, 55]. In our study, BOD1 expression was suppressed by miR-142-3p overexpression in HK-2 cells under HG condition, and BOD1 overexpression reversed the suppressive effect of miR-142-3p on it. Moreover, BOD1 overexpression reversed the promotion of miR-142-3p on HK-2 cell viability and rescued the inhibitory effects of miR-142-3p on cell apoptosis and oxidative stress under HG conditions. The results suggested that miR-142-3p alleviates HG-induced HK-2 cell injury by targeting BOD1.
Furthermore, the mechanism underlying HG-induced decrease in miR-142-3p expression is unclear, which will be further explored in our future studies. In addition, we will focus on the mechanism of HG-induced miR-142-3p downregulation by exploring the upstream long noncoding RNAs (lncRNAs) of miR-142-3p or transcriptional factors involving miR-142-3p. Many lncRNAs were reported to serve as competing endogenous RNAs to interact with miRNAs and downregulate the expression of miRNAs. For example, lncRNA MCM3AP-AS1 was reported to interact with miR-142-3p, thereby promoting cell apoptosis [56].
In conclusion, our present study illustrated that miR-142-3p alleviates HG-mediated HK-2 injury by targeting BOD1. The study might offer a better understanding on the pathophysiology of DN and provide novel insight into the role of the miR-142-3p/BOD1 axis in DN diagnosis and treatment.
References
Lin YC, Chang YH, Yang SY, Wu KD, Chu TS. Update of pathophysiology and management of diabetic kidney disease. J Formos Med Assoc. 2018;117(8):662–75. https://doi.org/10.1016/j.jfma.2018.02.007.
Qi C, Mao X, Zhang Z, Wu H. Classification and differential diagnosis of diabetic nephropathy. J Diabetes Res. 2017;2017:8637138. https://doi.org/10.1155/2017/8637138.
Papadopoulou-Marketou N, Chrousos GP, Kanaka-Gantenbein C. Diabetic nephropathy in type 1 diabetes: a review of early natural history, pathogenesis, and diagnosis. Diabetes Metab Res Rev. 2017. https://doi.org/10.1002/dmrr.2841.
Papadopoulou-Marketou N, Paschou SA, Marketos N, Adamidi S, Adamidis S, Kanaka-Gantenbein C. Diabetic nephropathy in type 1 diabetes. Minerva Med. 2018;109(3):218–28. https://doi.org/10.23736/s0026-4806.17.05496-9.
Ioannou K. Diabetic nephropathy: is it always there? Assumptions, weaknesses and pitfalls in the diagnosis. Hormones (Athens). 2017;16(4):351–61. https://doi.org/10.14310/horm.2002.1755.
Meza Letelier CE, San Martín Ojeda CA, Ruiz Provoste JJ, Frugone Zaror CJ. Pathophysiology of diabetic nephropathy: a literature review. Medwave. 2017;17(1):e6839. https://doi.org/10.5867/medwave.2017.01.6839.
Lu TX, Rothenberg ME. MicroRNA. J Allergy Clin Immunol. 2018;141(4):1202–7. https://doi.org/10.1016/j.jaci.2017.08.034.
Mohr AM, Mott JL. Overview of microRNA biology. Semin Liver Dis. 2015;35(1):3–11. https://doi.org/10.1055/s-0034-1397344.
Regazzi R. MicroRNAs as therapeutic targets for the treatment of diabetes mellitus and its complications. Expert Opin Ther Targets. 2018;22(2):153–60. https://doi.org/10.1080/14728222.2018.1420168.
Rosado JA, Diez-Bello R, Salido GM, Jardin I. Fine-tuning of microRNAs in type 2 diabetes mellitus. Curr Med Chem. 2019;26(22):4102–18. https://doi.org/10.2174/0929867325666171205163944.
Kumar M, Nath S, Prasad HK, Sharma GD, Li Y. MicroRNAs: a new ray of hope for diabetes mellitus. Protein Cell. 2012;3(10):726–38. https://doi.org/10.1007/s13238-012-2055-0.
Simonson B, Das S. MicroRNA therapeutics: the next magic bullet? Mini Rev Med Chem. 2015;15(6):467–74. https://doi.org/10.2174/1389557515666150324123208.
Yu J, Yu C, Feng B, Zhan X, Luo N, Yu X, et al. Intrarenal microRNA signature related to the fibrosis process in chronic kidney disease: identification and functional validation of key miRNAs. BMC Nephrol. 2019;20(1):336. https://doi.org/10.1186/s12882-019-1512-x.
Zanchi C, Macconi D, Trionfini P, Tomasoni S, Rottoli D, Locatelli M, et al. MicroRNA-184 is a downstream effector of albuminuria driving renal fibrosis in rats with diabetic nephropathy. Diabetologia. 2017;60(6):1114–25. https://doi.org/10.1007/s00125-017-4248-9.
Zhao Y, Yin Z, Li H, Fan J, Yang S, Chen C, et al. MiR-30c protects diabetic nephropathy by suppressing epithelial-to-mesenchymal transition in db/db mice. Aging Cell. 2017;16(2):387–400. https://doi.org/10.1111/acel.12563.
Sun J, Zhao F, Zhang W, Lv J, Lv J, Yin A. BMSCs and miR-124a ameliorated diabetic nephropathy via inhibiting notch signalling pathway. J Cell Mol Med. 2018;22(10):4840–55. https://doi.org/10.1111/jcmm.13747.
Zhou L, Xu DY, Sha WG, Shen L, Lu GY, Yin X, et al. High glucose induces renal tubular epithelial injury via Sirt1/NF-kappaB/microR-29/Keap1 signal pathway. J Transl Med. 2015;13:352. https://doi.org/10.1186/s12967-015-0710-y.
Kantharidis P, Hagiwara S, Brennan E, McClelland AD. Study of microRNA in diabetic nephropathy: isolation, quantification and biological function. Nephrology (Carlton). 2015;20(3):132–9. https://doi.org/10.1111/nep.12374.
Rudnicki M, Beckers A, Neuwirt H, Vandesompele J. RNA expression signatures and posttranscriptional regulation in diabetic nephropathy. Nephrol Dial Transplant. 2015;30(Suppl 4):35–42. https://doi.org/10.1093/ndt/gfv079.
Zeng Y, Feng Z, Liao Y, Yang M, Bai Y, He Z. Diminution of microRNA-98 alleviates renal fibrosis in diabetic nephropathy by elevating Nedd4L and inactivating TGF-β/Smad2/3 pathway. Cell Cycle. 2020;19(24):3406–18. https://doi.org/10.1080/15384101.2020.1838780.
Liu F, Zhang ZP, Xin GD, Guo LH, Jiang Q, Wang ZX. miR-192 prevents renal tubulointerstitial fibrosis in diabetic nephropathy by targeting Egr1. Eur Rev Med Pharmacol Sci. 2018;22(13):4252–60. https://doi.org/10.26355/eurrev_201807_15420.
Zhao D, Jia J, Shao H. miR-30e targets GLIPR-2 to modulate diabetic nephropathy: in vitro and in vivo experiments. J Mol Endocrinol. 2017;59(2):181–90. https://doi.org/10.1530/jme-17-0083.
Gholaminejad A, Abdul Tehrani H, Gholami FM. Identification of candidate microRNA biomarkers in renal fibrosis: a meta-analysis of profiling studies. Biomarkers. 2018;23(8):713–24. https://doi.org/10.1080/1354750x.2018.1488275.
Liang YZ, Li JJ, Xiao HB, He Y, Zhang L, Yan YX. Identification of stress-related microRNA biomarkers in type 2 diabetes mellitus: a systematic review and meta-analysis. J Diabetes. 2020;12(9):633–44. https://doi.org/10.1111/1753-0407.12643.
Zhu H, Leung SW. Identification of microRNA biomarkers in type 2 diabetes: a meta-analysis of controlled profiling studies. Diabetologia. 2015;58(5):900–11. https://doi.org/10.1007/s00125-015-3510-2.
Zhang T, Ji C, Shi R. miR-142-3p promotes pancreatic β cell survival through targeting FOXO1 in gestational diabetes mellitus. Int J Clin Exp Pathol. 2019;12(5):1529–38.
Zhou L, Xu DY, Sha WG, Shen L, Lu GY, Yin X. Long non-coding MIAT mediates high glucose-induced renal tubular epithelial injury. Biochem Biophys Res Commun. 2015;468(4):726–32. https://doi.org/10.1016/j.bbrc.2015.11.023.
Vejnar CE, Zdobnov EM. MiRmap: comprehensive prediction of microRNA target repression strength. Nucleic Acids Res. 2012;40(22):11673–83. https://doi.org/10.1093/nar/gks901.
Kertesz M, Iovino N, Unnerstall U, Gaul U, Segal E. The role of site accessibility in microRNA target recognition. Nat Genet. 2007;39(10):1278–84. https://doi.org/10.1038/ng2135.
Miranda KC, Huynh T, Tay Y, Ang YS, Tam WL, Thomson AM, et al. A pattern-based method for the identification of MicroRNA binding sites and their corresponding heteroduplexes. Cell. 2006;126(6):1203–17. https://doi.org/10.1016/j.cell.2006.07.031.
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120(1):15–20. https://doi.org/10.1016/j.cell.2004.12.035.
Qi C, Mao X. Classification and Differential Diagnosis of Diabetic. Nephropathy. 2017;2017:8637138. https://doi.org/10.1155/2017/8637138.
Flyvbjerg A. The role of the complement system in diabetic nephropathy. Nat Rev Nephrol. 2017;13(5):311–8. https://doi.org/10.1038/nrneph.2017.31.
Tang J, Yao D, Yan H, Chen X, Wang L, Zhan H. The role of MicroRNAs in the pathogenesis of diabetic nephropathy. Int J Endocrinol. 2019;2019:8719060. https://doi.org/10.1155/2019/8719060.
Lu Y, Liu D, Feng Q, Liu Z. Diabetic nephropathy: perspective on extracellular vesicles. Front Immunol. 2020;11:943. https://doi.org/10.3389/fimmu.2020.00943.
Sagoo MK, Gnudi L. Diabetic nephropathy: is there a role for oxidative stress? Free Radical Biol Med. 2018;116:50–63. https://doi.org/10.1016/j.freeradbiomed.2017.12.040.
Magee C, Grieve DJ, Watson CJ, Brazil DP. Diabetic Nephropathy: a tangled web to unweave. Cardiovasc Drugs Ther. 2017;31(5–6):579–92. https://doi.org/10.1007/s10557-017-6755-9.
Jha JC, Banal C, Chow BS, Cooper ME, Jandeleit-Dahm K. Diabetes and kidney disease: role of oxidative stress. Antioxid Redox Signal. 2016;25(12):657–84. https://doi.org/10.1089/ars.2016.6664.
Bao L, Li J, Zha D, Zhang L, Gao P, Yao T, et al. Chlorogenic acid prevents diabetic nephropathy by inhibiting oxidative stress and inflammation through modulation of the Nrf2/HO-1 and NF-ĸB pathways. Int Immunopharmacol. 2018;54:245–53. https://doi.org/10.1016/j.intimp.2017.11.021.
Hou Y, Shi Y, Han B, Liu X, Qiao X, Qi Y, et al. The antioxidant peptide SS31 prevents oxidative stress, downregulates CD36 and improves renal function in diabetic nephropathy. Nephrol Dial Transplant. 2018;33(11):1908–18. https://doi.org/10.1093/ndt/gfy021.
Al-Rasheed NM, Al-Rasheed NM, Bassiouni YA, Hasan IH, Al-Amin MA, Al-Ajmi HN, et al. Simvastatin ameliorates diabetic nephropathy by attenuating oxidative stress and apoptosis in a rat model of streptozotocin-induced type 1 diabetes. Biomed Pharmacother. 2018;105:290–8. https://doi.org/10.1016/j.biopha.2018.05.130.
Wang Y, Li Y, Yang Z, Wang Z, Chang J, Zhang T, et al. Pyridoxamine treatment of HK-2 human proximal tubular epithelial cells reduces oxidative stress and the inhibition of autophagy induced by high glucose levels. Med Sci Monit. 2019;25:1480–8. https://doi.org/10.12659/msm.914799.
Chen F, Sun Z, Zhu X, Ma Y. Astilbin inhibits high glucose-induced autophagy and apoptosis through the PI3K/Akt pathway in human proximal tubular epithelial cells. Biomed Pharmacother. 2018;106:1175–81. https://doi.org/10.1016/j.biopha.2018.07.072.
Kimura T, Takabatake Y, Takahashi A, Kaimori JY, Matsui I, Namba T, et al. Autophagy protects the proximal tubule from degeneration and acute ischemic injury. J Am Soc Nephrol. 2011;22(5):902–13. https://doi.org/10.1681/asn.2010070705.
Kitada M, Ogura Y, Monno I, Koya D. Regulating autophagy as a therapeutic target for diabetic nephropathy. Curr Diab Rep. 2017;17(7):53. https://doi.org/10.1007/s11892-017-0879-y.
Wei Z, Qin X, Kang X, Zhou H, Wang S, Wei D. MiR-142-3p inhibits adipogenic differentiation and autophagy in obesity through targeting KLF9. Mol Cell Endocrinol. 2020;518: 111028. https://doi.org/10.1016/j.mce.2020.111028.
Su Q, Liu Y, Lv XW, Ye ZL, Sun YH, Kong BH, et al. Inhibition of lncRNA TUG1 upregulates miR-142-3p to ameliorate myocardial injury during ischemia and reperfusion via targeting HMGB1- and Rac1-induced autophagy. J Mol Cell Cardiol. 2019;133:12–25. https://doi.org/10.1016/j.yjmcc.2019.05.021.
Plotnikova OM, Skoblov MY. Efficiency of the miRNA- mRNA interaction prediction programs. Mol Biol. 2018;52(3):543–54. https://doi.org/10.7868/s0026898418030187.
Yang Z, Guo Z, Dong J, Sheng S, Wang Y, Yu L, et al. miR-374a regulates inflammatory response in diabetic nephropathy by targeting MCP-1 expression. Front Pharmacol. 2018;9:900. https://doi.org/10.3389/fphar.2018.00900.
Wang J, Duan L, Guo T, Gao Y, Tian L, Liu J, et al. Downregulation of miR-30c promotes renal fibrosis by target CTGF in diabetic nephropathy. J Diabetes Complicat. 2016;30(3):406–14. https://doi.org/10.1016/j.jdiacomp.2015.12.011.
Qin B, Shu Y, Long L, Li H, Men X, Feng L, et al. MicroRNA-142-3p induces atherosclerosis-associated endothelial cell apoptosis by directly targeting rictor. Cell Physiol Biochem. 2018;47(4):1589–603. https://doi.org/10.1159/000490932.
Sukma Dewi I, Celik S, Karlsson A, Hollander Z, Lam K, McManus JW, et al. Exosomal miR-142-3p is increased during cardiac allograft rejection and augments vascular permeability through down-regulation of endothelial RAB11FIP2 expression. Cardiovasc Res. 2017;113(5):440–52. https://doi.org/10.1093/cvr/cvw244.
Pan D, Du Y, Ren Z, Chen Y, Li X, Wang J, et al. Radiation induces premature chromatid separation via the miR-142-3p/Bod1 pathway in carcinoma cells. Oncotarget. 2016;7(37):60432–45. https://doi.org/10.18632/oncotarget.11080.
Di J, Rutherford S, Chu C. Review of the cervical cancer burden and population-based cervical cancer screening in China. Asian Pac J Cancer Prev. 2015;16(17):7401–7. https://doi.org/10.7314/apjcp.2015.16.17.7401.
Porter IM, Schleicher K, Porter M, Swedlow JR. Bod1 regulates protein phosphatase 2A at mitotic kinetochores. Nat Commun. 2013;4:2677. https://doi.org/10.1038/ncomms3677.
Gao Y, Zhao H, Li Y. LncRNA MCM3AP-AS1 regulates miR-142-3p/HMGB1 to promote LPS-induced chondrocyte apoptosis. BMC Musculoskelet Disord. 2019;20(1):605. https://doi.org/10.1186/s12891-019-2967-4.
Funding
This research was supported by Henan Science and Technology Research Project (No. 162102310287), Henan health and Family Planning Commission Provincial Ministry Co Construction Project (No.201701022), Henan Medical Science and Technology Research Project (No.2018020395).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors report no declarations of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
About this article
Cite this article
Zhao, N., Luo, Q., Lin, R. et al. MiR-142-3p ameliorates high glucose-induced renal tubular epithelial cell injury by targeting BOD1. Clin Exp Nephrol 25, 1182–1192 (2021). https://doi.org/10.1007/s10157-021-02102-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10157-021-02102-y