Abstract
Background
Epithelium-specific ETS 3 (ESE3) is down-regulated frequently in several malignancies and involved in carcinogenesis and progression. However, ESE3 expression pattern and its relationship with clinical features and prognosis in hepatocellular carcinoma (HCC) are still largely unknown.
Methods
ESE3 expression was analyzed by quantitative real-time PCR and western blotting in HCC cell lines, and then, it was analyzed by immunohistochemistry in HCC tissues and peritumoral normal tissues from total 94 HCC patients. The relationship between ESE3 expression and clinical features was investigated to illustrate the potential prognostic value in HCC. ESE3 roles on HCC progression were evaluated in vitro and vivo by MTT assay and mice tumor model, respectively.
Results
ESE3, mainly located in the cytoplasm, was remarkably down-regulated in HCC tissues and cell lines. Low ESE3 expression was positively associated with tumor progression and metastasis features. Kaplan–Meier analysis demonstrated that low ESE3 expression contributed to poor recurrence-free survival (RFS) and overall survival (OS) (both p < 0.01) of patients, and maintained its prognostic value in predicting poor RFS and OS of “Early-stage” HCC patients regardless of clinical features being studied. Multivariate survival analysis was also identified ESE3 as an independent prognostic factor for RFS (p = 0.05 for marginal significance) and OS (p = 0.031). ESE3 expression restoration in cells led to a significant inhibition in HepG2 cell proliferation in vitro and vivo (both p < 0.001).
Conclusions
Down-regulated ESE3 expression in HCC tissues could serve as a potential therapeutic target against HCC and appears to be as a poor prognostic indicator for prognosis, especially in “Early-stage” HCC patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC), the most frequent primary liver tumor, accounts for the third leading cause of cancer-related deaths [1]. Despite technological advances in early diagnosis and improvements in the therapy (e.g., surgical tumor resection and liver transplantation), the prognosis of HCC remains poor and the average survival time since diagnosis onset is less than 5 years [2]. Therefore, identification of a potential therapeutic target and prognostic biomarker for HCC is an urgent need for improving the therapeutic outcomes [3, 4].
The ETS family, one of the largest families of transcriptional regulators, is composed of nearly 40 distinct members [5, 6]. All ETS genes process a highly conserved sequence, the ETS domain, which recognizes and binds to a GGAA/T core sequence (ETS-Binding Site, EBS) in the promoter and enhancer region of target genes [7, 8]. ETS factors are shown to act as regulators of target genes that involved in multiple biological processes, such as cellular proliferation, differentiation, carcinogenesis, and tumor metastasis [5, 9]. One subset of ETS factors named as epithelium-specific ETS factors (ESE), including ESE1 (ERT), ESE2 (ELF5), ESE3 (EHF), and PDEF (PSE), are expressed in epithelial tissues [9, 10]. Among the ESEs’ subset, abnormal expression of ESE3 proteins has been reported to be involved in carcinogenesis and progression of many human cancers [11,12,13,14,15,16]. Recently, Zhao et al. have reported the decreasing of ESE3 expression can promote pancreatic cancer migration and invasion by downregulating E-cadherin expression [12]. Research on prostate cancer also indicated that ESE3 was down-regulated by promoter methylation and acted as tumor suppressor gene [14, 15]. Additionally, ESE3 acts as a tumor suppressor gene in the esophageal squamous cell carcinoma (ESCC) carcinogenesis through changes in subcellular localization in ESCC [13]. However, limited investigation has been carried out about characterization of ESE3 expression pattern in HCC, and its role in HCC development and progression still remains unclear.
In this regard, the aim of this study is to investigate the expression pattern of ESE3 in cell lines and liver tissues of HCC patients using immunohistochemistry stain and qRT-PCR analysis. Furthermore, the expression level of ESE3 was compared with patients’ clinical features and outcomes to examine its prognostic values. The roles of ESE3 on HCC progression are also investigated in vitro and vivo, respectively. To our knowledge, this study is the first report on the effects of ESE3 in the HCC development and progression.
Methods
Cell line culture and culture conditions
The human HCC cell lines HepG2, Huh7 and Hep3B, one normal human liver cell line L-02 and human pancreatic carcinoma cell line BxPC-3 were obtained from Cell bank of Chinese Academy of Science or were kindly provided by Prof. Bo Zhang (Shanghai Harmonious One Biotech Co., Ltd, Shanghai, China). These cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM; Gibco Laboratories) containing with 10% fetal bovine serum (FBS) (Life Technologies, Grand Island, NY) and penicillin–streptomycin, at 37 ℃ in 5% CO2 and 95% air condition.
Western blot analysis
The protocols for western blot were well described previously [17, 18]. In brief, cells grown to ~ 60% confluence were harvested. Whole protein lysates were prepared according to the standard protocol using RIPA buffer (Cell signaling, Danvers, MA). Equal amounts of protein were located on 10% SDS-PAGE gels and transferred to nylon membranes and incubated with primary antibodies against ESE3 (Thermo Fisher Scientific, Fremont, CA, USA, Cat No. PA5-30716) and β-tubulin (Santa Cruz Biotechnology, USA). Protein bands were detected using an enhanced chemiluminescence regent (Sigma, USA). β-tubulin was used as reference proteins. Relative densities (RD) was calculated by normalizing the IOD density of each blot with that of β-tubulin.
Immunofluorescence staining
To evaluate the subcellular location of ESE3 expression, HCC cell line Hep3B and human pancreatic carcinoma cell line BxPC-3 were seeded onto glass slides, fixed with 4% paraformaldehyde for 30 min, permeabilized with 0.2% Tween-X100 for 5 min, and blocked with 3% BSA in PBS for 1 h. The rabbit polyclonal anti-ESE3 (Thermo Fisher Scientific, Fremont, CA, USA, Cat No. PA5-30716) was applied and incubated at 4℃, overnight. Slides were washed twice with PBS and incubated with FITC-conjugated anti-rabbit IgG or TRITC-conjugated anti-rabbit IgG (both from Zhongshan Biotechnology Co. Ltd., Beijing, China). Nuclei were mounted with 4,6-diamidino-2-phenylindole (DAPI). The slides were viewed under a Leica fluorescent microscope. Negative controls were treated identically with the primary antibodies omitted.
Quantitative real-time PCR (qRT-PCR)
Total RNA from HCC cell lines was extracted by TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to its protocol. cDNA was synthesized from 2 μg RNA using a Reverse Transcription Kit (Toybo Co. Ltd., Osaka, Japan). The accession number of ESE3 RNA sequence is NM_001206616. Primer pairs were as follows: ESE3: forward 5′-CATGAGTTTGCAGGAGTTCACC-3′, reverse 5′-GGTGTTCATGATGGAAGGCTC-3′; β-actin: forward 5′-TGACGTGGACATCCGCAAAG-3′, reverse 5′-CTGGAAGGTGGACAGCGAGG-3′. The qRT-PCR amplification cycling was follows: predenaturation at 95 ℃ for 10 min, followed by 40 PCR circulation of 95 ℃ for 10 s, annealing at 60 ℃ for 20 s, and extension at 72 ℃ for 15 s. A melting-cure analysis was performed at the end of amplification to verify its specificity.
mRNA levels of target genes were normalized to the internal control β-actin. The relative ESE3 mRNA expression level was determined using the comparative (2−ΔΔCT) method and normalized to the internal control β-actin. The data are representative of three independent experiments.
Patients, specimens, and follow-up
A total of 94 patients with primary HCC who underwent hepatectomy at Shandong Provincial Hospital Affiliated to Shandong University between November 2011 and September 2012 were enrolled. The criteria for curative resection were described previously [19]. All features of tumor (e.g., tumor size, Edmondson Grade, and capsular and vascular invasion, etc.) were defined independently by two pathologists (Jia-wen Xu and Ai-ju Fang). Tumor differentiation grade was based on the Edmondson grading system. All patients had not received anticancer treatment before surgery in this study. HCC was staged according the Barcelona Clinic Liver Cancer (BCLC) staging system [20]. There were 83 males and 11 females with ranging in age from 37 to 78 years (average, 54.2 years). 74 HCC patients have the cirrhosis background, and the etiology of the liver disease was hepatitis B virus (HBV) in 75 patients, hepatitis C virus (HCV) in 8 patients, and non-HBV non-HCV (NBNC) in 11 patients. The patients’ histopathologic and clinical features are summarized in Table 1.
All study participants were examined via tumor biomarkers and abdominal ultrasonography every 4 months for the first-year postoperation and every 6–7 months afterward during the follow-up periods.
A computed tomography (CT) scan of the abdomen was performed every 6 months. If recurrence was suspected, CT scan was performed immediately. Treatment modalities after recurrence, radiofrequency ablation, and selected transcatheter arterial chemoembolization were administered depending on the patients’ individual situation [21].
The mean follow-up was 40 months. The main outcomes of the study were overall survival (OS) and recurrence-free survival (RFS), which were calculated from the date of operation to the date of death or recurrence [22]. If recurrence was not diagnosed at the time of study, the survivors were censored on the date of death or the last date of follow-up.
Tissue microarray construction, immunohistochemistry, and assessment
Tissue microarrays were prepared as described previously [19]. Ninety-four pathology-proven HCC tissues and 51 peritumoral normal tissues (at least 5 cm from the tumor edge, only matched HCC tissues of Edmondson Grade II and III) were collected. After examining HE-stained slides for the location and differentiation of HCC tissue and normal tissues, tissue microarray (TMA) slides (collaborated with Shanghai Biochip Company, Ltd., Shanghai, China) were constructed.
Immunohistochemistry (IHC) stain was performed using a DAB substrate kit (Maxin) as described previously [19]. Signal was detected by a standard streptavidin immunoperoxidase reaction, followed by chromagen detection with diaminobenzidine. Negative control sections were incubated with normal rabbit serum in place of primary antibody (both from Zhongshan Biotechnology Co. Ltd., Beijing, China). Two pathologists evaluated the immunoreactivity independently without knowledge of clinical data. Immunohistochemical staining was also evaluated using Image-Pro v6.2 software (Media Cybernetics Inc., Bethesda, MD, USA) as previously described [19]. Integrated optical density (IOD) was measured in each field, and ESE3 density was formulated as ratio IOD/the total area of each field. The results were log transformed before statistical analysis. Sections were assigned a score by a first observer that was confirmed by a second observer.
Construction of recombinant ESE3 over-expressed lentiviruses
The full-coding sequence of human ESE3 gene was synthesized from Haro Biothech (Shanghai, China) and directionally cloned into pLVX-IRES-Puromycin vector (pLVXpuro-ESE3) by Xho I and Bamh I restriction endonuclease enzymes, and the recombinant ESE3 over-expressed lentivirus was constructed successfully. The mock vector (pLVXpuro) used as control.
3-(4,5-Dimethyl-2-thiazolyl)-2,5-diphenyl-2-H-tetrazolium bromide (MTT) Assay
The HepG2 cells, which stably transfected with these two types of lentivirus, were prepared. MTT assay was used to evaluate the cell proliferation. Briefly, HepG2 cells were seeded in 96-well plates at a rate of 2.6 × 103 cells/well and cultured for 24, 48, 72, 96, and 120 h. Then, 10 μl MTT (Solarbio, Beijing, China) was added to each well to dissolve the formazan crystals. The absorbance at 490 nm was detected using a spectrophotometer (TECAN Nanoquant, Switzerland).
Xenograft nude mice tumor model
All animal experiments were performed in the animal laboratory center, and in accordance with guidelines of the institutional Animal care and Use Committee and approved by the Medical Ethics Committee of Shandong Province Hospital Affiliated to Shandong University. The 5-week-old nude mice were purchased from Shanghai Laboratory Animal Center (Shanghai, China). HepG2 cells (1 × 107) stably transfected with the recombinant ESE3 over-expressed lentivirus or mock control were subcutaneously injected into the flanks of 5-week-old male BALB/c nude mice (n = 6 mice/group) The tumor volumes were measured every 3 days for 30 days. Following resection, the tumor weights were detected.
Statistical analysis
For ESE3, The cut-off value for definition of subgroups was the median value. All data were expressed as Mean ± SD. Analyses were performed using software package SPSS V.15.0. Chi-square test was applied to compare qualitative variable. Quantitative variables were analyzed using Student’s t test and one-way analysis ANOVAs for comparison between groups. Survival curves as analyzed by Kaplan–Meier method and comparison between subgroups were examined by the log-rank test. Multivariable analysis was performed with the Cox regression model. A value of p < 0.05 was taken as being significant.
Results
ESE3 is mainly located on the cytoplasm and down-regulated in human HCC
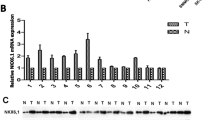
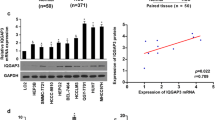
As shown in Fig. 1a, ESE3, a protein band of approximately 34 kDa, was detected in L-02 cell line, with human pancreatic carcinoma cell line BxPC-3 used as positive control [12]. ESE3 was also detected in three HCC cell lines at various levels. The results of relative densities (RD) by normalizing with β-tubulin showed that the ESE3 expression level in these HCC cell lines was remarkably lower than that of L-02 (*p < 0.05, **p < 0.001 vs. L-02, Fig. 1b). Consistent with the ESE3 protein expression pattern, the qRT-PCR results also confirmed the low expression of ESE3 mRNA in HCC cell lines studied (*p < 0.05, **p < 0.001 vs. L-02, Fig. 1c). In addition, immunofluorescence analysis was performed to characterize the subcellular localization of ESE3 in HCC cell lines. These results showed that ESE3 was mainly located on the cytoplasm of HCC cell line Hep3B and the nucleus of pancreatic carcinoma cell line BxPC-3 respectively, whereas the negative control showed the absence of specific fluorescence signal (Fig. 1d).
Expression level and subcellular localization of ESE3 in cell lines. a Western blot was performed to detect the ESE3 protein expression in three HCC cell lines, one human hepatocyte cell line L-02 and pancreatic carcinoma cell line BxPC-3. b Distribution of ESE3 relative densities (RD) in these cell lines studied. c The ESE3 mRNA expression in panel of HCC cell lines and L-02 was analyzed by qRT-PCR. d Immunofluorescence signal of ESE3 protein was observed in Hep3B cell (red arrow), but not in negative control, while BxPC-3 was used as positive control (yellow arrow). Data are representative of three independent experiments (*p < 0.05,**p < 0.001 vs. L-02).
To further confirm the subcellular localization and expression level of ESE3 in HCC tissues, we next performed IHC analysis in HCC TMA (including 51 matched pairs of HCC tissues and corresponding peritumoral normal tissues, see “Methods” for detail). In Fig. 2a, b, in accordance with the previous studies [12, 13], the ESE3 protein was mainly located in nucleus of pancreatic cancer and normal pancreas tissue. On the contrary, except for weakly positive staining in cytoplasm of HCC cells in HCC parenchyma (Fig. 2c), positive staining for ESE3 was located predominately in the cytoplasm of hepatocytes in peritumoral normal tissues, while specific immunoreactivity was absent in all infiltrating lymphocytes and neutrophils (Fig. 2d).
Immunohistochemistry for ESE3 in HCC tissues and peritumoral normal tissues. Positive staining of ESE3 in the nucleus of pancratic cancer tissues (blue arrow) (a) and normal pancreas tissue (black arrow) (b). Negative staining of ESE3 in the nucleus of in HCC tissues (red arrow) (c) but positive in the cytoplasm of peritumoral normal tissue (green arrow) (d). Analysis of ESE3 expression in the TMA specimens according to the intergral optical density (IOD) value (*p < 0.05 vs. peritumoral normal tissues) (e). Immunohistochemistry for ESE3 in TMAs including three Edmondson grades tissue and normal tissues (f). Western blot was performed to detect the ESE3 expression in 5 paired human HCC tissues (T) and matched peritumoral normal tissues (N), *p<0.05 vs. peritumoral normal tissues (g).
Image Pro Plus v6.2 software was used to analyze immunoreactivity of ESE3 in the TMAs. The scores for ESE3 expression was significantly lower than those in corresponding peritumoral tissues (mean score 4.723 ± 0.243 vs. 4.829 ± 0.114; p = 0.004, Fig. 2e). Using 4.758 as cut-off value, we found that the proportion of low ESE3 expression in the HCC tissues (39.2%, 20/51) was significantly higher than that in corresponding peritumoral normal tissues (19.6%, 10/51) (p = 0.03, Table 2). The scores of ESE3 expression of Edmondson grade I HCC tissues were significantly higher than those of Edmondson grade II or III (4.735 ± 0.166 vs. 4.854 ± 0.031; p = 0.028; 4.625 ± 0.306 vs. 4.854 ± 0.031; p < 0.05). Some representative immunohistochemistry images are displayed in Fig. 2f.
In addition, using western blot, expression of ESE3 protein was further demonstrated in five paired fresh samples tested. Despite some interindividual variations, the expression of ESE3 protein was significantly lower in HCC than that in normal liver tissue (Fig. 2g) and suggested that down-regulated expression of ESE3 was involved in HCC progression.
Low ESE3 expression is correlated with aggressive clinical features of HCC patients
To better evaluate the clinical relevance of ESE3 expression in HCC, we investigated the relationship between ESE3 expression and clinicopathologic features in HCC TMA, and the results are summarized in Table 1. We found that patients with low ESE3 expression in HCC was positively correlated with the aggressive clinical feature, such as presence of vascular invasion (p = 0.046), capsulation invasion (p = 0.067 for marginal significance), multiple tumor number (p = 0.003), low BCLC stage (p = 0.001), and high Edmondson grade (p = 0.004). No statistical relationship was found between ESE3 expression and patient’s age, gender, liver cirrhosis, or tumor size (all p > 0.05).
Low ESE3 expression in HCC tissues indicated poor prognosis in HCC patients
At the final investigation, the follow-up rate was 95.8% (4 patients were lost to follow-up). 77 (81.9%) patients had recurrence and 55 (58.5%) patients died, including 2 patients died from liver failure without tumor recurrence and 4 patients died from the massive upper gastrointestinal hemorrhage.
Univariate survival analysis showed that ESE3 expression, tumor size, BCLC stage, and vascular invasion were significantly associated with RFS and OS. Low Edmondson grade has prognostic significance for poor RFS (Table 3). Above all, Kaplan–Meier analysis showed that HCC patients in the low ESE3 expression group have worse RFS and OS than did those in the high ESE3 expression group. The 3-year RFS and OS rate in the low ESE3 expression were only 13.5% and 59.4% compared to RFS of 45.8% and OS of 85.8% in the high ESE3 expression group, respectively (both p < 0.05; Fig. 3a).
Prognostic significance of ESE3 expression and representative images in two patients’ subgroups. Kaplan–Meier curves analysis for RFS and OS according to the ESE3 expression level of HCC tissues in all patients (a). Representative immunohistochemistry images of ESE3 loss group (case19) and gain/retain group (case 1 and 6) in HCC tissues and corresponding peritumoral mormal tissus (b). Kaplan–Meier curves for RFS and OS according to ESE3 loss and gain/retain in HCC tissues compared to peritumoral normal tissues (c).
Additionally, according to the intensity of ESE3 staining in HCC tissues and corresponding peritumoral normal tissues, the patients were classified two groups: the ESE3 loss group (ESE3T < N, n = 27) and ESE3 gain/retain group (ESE3T ≥ N, n = 24), to further evaluate the significance of ESE3 loss and gain in HCC tissues compared to peritumoral normal tissues (Fig. 3b). As expected, ESE3 loss group exhibited the shorter RFS (median, 22 months) and OS (median, 37 months) time, whereas the ESE3 gain/retain group showed the longer survival time (RFS median, 31 months vs. OS median, 47 months, both p < 0.05, Fig. 3c).
Moreover, patients were stratified by primary clinical features, including BCLC stage, tumor size, capsular invasion, tumor number, vascular invasion, as well as Edmondson grade, to fully evaluate the prognostic value of ESE3 in different clinical features being studied in this study. The patients with one or more these features such as BCLC stage (0 + A), tumor size < 5 cm, no vascular/capsular invasion, single tumor, or Edmondson grade (I + II) were recognized as “Early-stage” subgroup, or else be recognized as “Late-stage” subgroup. In the “Late-stage” subgroup, ESE3 expression level in HCC tissues did not display a satisfying predictive value for prediction of RFS and OS (Fig. 4a–f, all p > 0.05, except for RFS in the tumor size ≥ 5 cm, multiple tumors, and Edmondson grade III group, respectively, for which p < 0.05). However, in “Early-stage” subgroup, patients with low ESE3 expression exhibited a significantly poorer OS and RFS than did those with high ESE3 expression regardless of clinical features (Fig. 5a–f, all p < 0.05). These results suggested that ESE3 expression was a potential prognostic indicator in “Early-stage” HCC patients.
Low ESE3 expression in HCC tissues was an independent prognostic factor
To investigate whether the low ESE3 expression in HCC tissues was an independent prognostic factor for outcomes of HCC, a multivariable analysis using Cox regression model was performed. The tumor size (p = 0.004), Edmondson grade (p = 0.042), and low ESE3 expression (p = 0.05 for marginal significance) were identified as independent risk factors for RFS, while the low ESE3 expression (p = 0.031) and tumor size (p = 0.011) were recognized as independent prognostic factors for OS. The hazard ratios for low ESE3 expression for RFS and OS were 0.595 (95% CI 0.354–1.000) and 0.528 (95% CI 0.296–0.942), respectively (Table 3).
Restoration of the ESE3 expression reversed the proliferation of HCC cells and shows antitumor activity in xenograft nude mice tumor model
We have demonstrated that ESE3 expression was remarkably down-regulated in both HCC tissues and cell lines, and low expression of ESE3 associated with HCC progression and metastasis features. To further verify the role of ESE3 in the HCC progression, we restored the ESE3 expression by infection with the recombinant ESE3 over-expressed lentivirus in HepG2 cell lines and measured the proliferation of cells in vitro. Significantly elevated expression of ESE3 in the infected HepG2 cells was confirmed by western blotting (Fig. 5a). Results obtained from the MTT assay indicated that restored ESE3 expression in HepG2 cells significantly inhibited the proliferation (Fig. 6b).
Prognostic significance of ESE3 expression level in “Early-stage” subgroup of HCC patients. Kaplan–Meier curves for RFS and OS according to BCLC stage (0 + A) (a), tumor size (< 5 cm) (b), capsular invasion (no) (c), tumor number (single) (d), vascular invasion (no) (e) and Edmondson grade (I + II) (f)
Restored ESE3 expresssion inhibited the tumor growth in vitro and vivo by MTT assay and xenograft nude mice tumor model. The stably expression of ESE3 was confirmed by western blotting in HepG2 cells (a). MTT assay was used to detecte cell proliferation in vitro (b). Images were presented to show the isolated tumors from ESE3 over-expressed lentivirus infected group and mock control group (c). The tumor size was monitored every 3 days to draw the growth curve of two group (d). The average tumor volume (e) and weight (f) of the ESE3 over-expressed lentivirus infected group compared with mock control group were statistically analyzed. *p< 0.05;**p< 0.01;***p<0.001.
Additionally, the xenograft nude mice tumor model was also constructed. The mice were randomly divided into the treatment group which received subcutaneous injection of HepG2 stably transfected with the recombinant ESE3 over-expressed lentivirus and the control group which received subcutaneous injection of HepG2 transfected with mock control (Fig. 6c). Growth curve of the formed tumor showed that ESE3 over-expressed tumor has lower growth rate than the mock control group (Fig. 6d). Analysis of the excised tumors showed that the average weights and average volumes of the ESE3 over-expressed tumor were lower than those of the mock control group (Fig. 6e–f). These data suggested that ESE3 shows antitumor effects by inhibiting tumor growth and have potential to become a therapeutic target.
Discussion
Epithelium-specific ETS factors (ESEs), a member of the ETS family of DNA-binding proteins, play critical role in differentiation and development programs for many epithelial tissues [9]. Among the ESEs, increasing evidences implicate that ESE3 can be directly involved in epithelial-to-mesenchymal transition (EMT) [8], stem-like features [14, 15], and tumor progression [16, 23]. In the present study, we certified that ESE3 expression, main located on the cytoplasm of cells, was downregulated in HCC tissues and involved with HCC progression as well as poor prognosis in HCC patients. Additionally, restoration of the ESE3 expression was verified to inhibit cell proliferation and tumor grow in vitro and vivo experiments. These data indicated that ESE3 may be used as a potential predictive indicator and therapeutic target of HCC treatment.
As a nuclear transcription factor, ESE3 was found in the nuclei of normal pancreatic cells, and was down-regulated and involved in pancreatic cancers’ invasion and metastasis processes [12]. Wang et al. reported that comparing with its expression in the nuclei of normal esophageal epithelial cells, ESE3 was also mainly localized in the cytoplasm of ESCC cells [13]. In this study, we present a detailed investigation on the ESE3 expression pattern in both HCC cell lines and tissues from HCC patients. Using pancreatic cancer and corresponding peritumoral tissues as positive control, the results of immunohistochemistry showed that ESE3 protein is mainly located in the cytoplasm but not the nucleus of HCC cell lines and tissues (Fig. 2c). This expression pattern was consistent with that observed in ESCC [13]. Meanwhile, when we were concerned with the ESE3 expression pattern in peritumoral normal liver tissues, cytoplasmic expression pattern were also verified in hepatocyte (Fig. 2d). However, the subcellular localization of ESE3 in esophageal epithelial cells was found in the nucleus distribution and these changes in localization from nucleus to cytoplasm may be critical role in carcinogenesis of ESCC [13]. This difference in localization of ESE3 in these cancer cells may derive from tissue-specific change, and suggest that a different mechanism could be involved in the hepatocytes carcinogenesis. Further studies are needed to confirm the mechanism.
We further try to study the association between the ESE3 expression and HCC progression and prognosis in HCC patients. Our data confirmed that ESE3 was significantly down-regulated in HCC tissue comparing with the adjacent normal tissue. In addition, HCC patients with low expression of ESE3 were prone to have poor cancer differentiation and more advanced clinicopathologic features. These results indicated that downregulation of ESE3 was certainly involved in HCC carcinogenesis and progression. ESE3 protein plays a key role in maintaining the epithelial phenotype [5, 24]. Aberrant promoter methylation has been certified as the cause of downregulation of ESE3 gene expression, and further lead to loss of the epithelial properties, promotion of aggressive behavior in some epithelial cell-derived cancers carcinogenesis [12]. It was reported that ESE3 controlled prostate epithelial cell differentiation and its loss led to prostate tumors with mesenchymal and stem-like features [11]. According to the data of HCC in this study, we, therefore, inferred that the above mechanism might also resident in carcinogenesis and progression of HCC, and ESE3 might be a new potential therapeutic target for HCC treatment. This opinion was also supported by the findings of tumor growth in xenograft nude mice tumor model. The findings showed that restored ESE3 expression by infected with recombinant ESE3 over-expressed lentivirus can significantly inhibit the tumor growth comparing with the mock control group. Nevertheless, the details of the mechanisms require further study.
Various different kinds of aberrant gene expression are indeed involved in individual carcinogenesis and progression. As showed in Table 2, in all 51 pairs of HCC tissues and self-matched peritumoral normal tissues, we observed that ESE3 expression down-regulated in only 20 out 51 HCC tissues and in 10 out 51 matched peritumoral normal tissues. It was noted that the pattern of ESE3 low expression in HCC tissues was not consistent in all pairs of samples. Meanwhile, ESE3 expression was retained or even upregulated in some HCC tissues compared with its expression in matched peritumoral normal tissues. Therefore, based on the results of immunohistochemistry stain scores in these both tissues, we classified these 51 patients as “ESE3 loss group”, whose ESE3 expression in HCC tissues was lower than that in peritumoral normal tissue (ESE3T < N group) and “ESE3 gain/retain group”, whose ESE3 expression in HCC tissues was equal or higher than that in peritumoral normal tissues (ESE3T ≥ N group). The prognosis analysis showed that ESE3T < N group displayed poorer prognosis in both RFS and OS. We can inferred that the ESE3 expression change was likely to largely account for the HCC progression independently, and ESE3 might be a novel potential therapeutic target.
Clinical stage is one of the key determining factors for the prognosis of HCC patients [25]. The “Early-stage” subgroup patients usually have one or more clinical features such as early BCLC stage, single tumor, no vascular/capsular invasion, or high Edmondson grade et al. [26]. These patients commonly have opportunity to receive surgical tumor resection and better prognosis. Hence, it is important to identify reliable and valid measures for predicting prognosis. In “Early-stage” subgroup of our study, down-regulated ESE3 expression in HCC tissues exhibited a significant predictive value in discriminating between patients with and without poor RFS and OS regardless of features being studied (Fig. 4 all p < 0.05), and was proved to be an independent prognostic indicator for prognosis of HCC in the multivariable analysis (Table 3). These data indicated that down-regulated ESE3 in HCC tissues could be a potential predictive indicator and can be applied to indicating a worse prognosis in “Early-stage” HCC patients, which would influence treatment decisions regarding individual clinical therapy.
In conclusion, although the relationship between low ESE3 expression and carcinogenesis in many types of cancers has been previously reported [11, 12, 14, 15], the significant aberrance of ESE3 in malignantly transformed human hepatocyte was investigated for the first time in this study. Our observations have provided evidence for the role of ESE3 in the progression of HCC and the prognostic significance in patients with HCC. Although the mechanism of ESE3 function needs further experimental study, targeting the ESE3 and its associated pathway may hold promise in the treatment of HCC.
References
Forner A, Reig M, Bruix J (2018) Hepatocellular carcinoma. Lancet 391(10127):1301–1314
Kang KJ, Ahn KS (2017) Anatomical resection of hepatocellular carcinoma: a critical review of the procedure and its benefits on survival. World J Gastroenterol 23(7):1139–1146
Juarez-Hernandez E, Motola-Kuba D, Chavez-Tapia NC et al (2017) Biomarkers in hepatocellular carcinoma: an overview. Expert Rev Gastroenterol Hepatol 11(6):549–558
Di Tommaso L, Roncalli M (2017) Tissue biomarkers in hepatocellular tumors: which, when, and how. Front Med (Lausanne) 4:10
Feldman RJ, Sementchenko VI, Watson DK (2003) The epithelial-specific Ets factors occupy a unique position in defining epithelial proliferation, differentiation and carcinogenesis. Anticancer Res 23(3A):2125–2131
Sharrocks AD (2001) The ETS-domain transcription factor family. Nat Rev Mol Cell Biol 2(11):827–837
Oikawa T, Yamada T (2003) Molecular biology of the Ets family of transcription factors. Gene 303:11–34
Luk IY, Reehorst CM, Mariadason JM (2018) ELF3, ELF5, EHF and SPDEF transcription factors in tissue homeostasis and cancer. Molecules 23(9):2191
Seth A, Watson DK (2005) ETS transcription factors and their emerging roles in human cancer. Eur J Cancer 41(16):2462–2478
Kar A, Gutierrez-Hartmann A (2013) Molecular mechanisms of ETS transcription factor-mediated tumorigenesis. Crit Rev Biochem Mol Biol 48(6):522–543
Albino D, Civenni G, Rossi S et al (2016) The ETS factor ESE3/EHF represses IL-6 preventing STAT3 activation and expansion of the prostate cancer stem-like compartment. Oncotarget 7(47):76756–76768
Zhao T, Jiang W, Wang X et al (2017) ESE3 Inhibits pancreatic cancer metastasis by upregulating E-cadherin. Cancer Res 77(4):874–885
Wang L, Xing J, Cheng R et al (2015) Abnormal localization and tumor suppressor function of epithelial tissue-specific transcription factor ese3 in esophageal squamous cell carcinoma. PLoS ONE 10(5):e0126319
Albino D, Longoni N, Curti L et al (2012) ESE3/EHF controls epithelial cell differentiation and its loss leads to prostate tumors with mesenchymal and stem-like features. Cancer Res 72(11):2889–2900
Albino D, Civenni G, Dallavalle C et al (2016) Activation of the Lin28/let-7 axis by Loss of ESE3/EHF promotes a tumorigenic and stem-like phenotype in prostate cancer. Cancer Res 76(12):3629–3643
Cangemi R, Mensah A, Albertini V et al (2008) Reduced expression and tumor suppressor function of the ETS transcription factor ESE-3 in prostate cancer. Oncogene 27(20):2877–2885
Li T, Zhu Y, Han L et al (2015) VEGFR-1 activation-induced MMP-9-dependent invasion in hepatocellular carcinoma. Future Oncol 11(23):3143–3157
Liu J, Han G, Liu H et al (2013) Suppression of cholangiocarcinoma cell growth by human umbilical cord mesenchymal stem cells: a possible role of Wnt and Akt signaling. PLoS ONE 8(4):e62844
Li T, Zhu Y, Ren W et al (2012) High co-expression of vascular endothelial growth factor receptor-1 and Snail is associated with poor prognosis after curative resection of hepatocellular carcinoma. Med Oncol 29(4):2750–2761
Bruix J, Llovet JM (2002) Prognostic prediction and treatment strategy in hepatocellular carcinoma. Hepatology 35(3):519–524
Livraghi T (2003) Radiofrequency ablation, PEIT, and TACE for hepatocellular carcinoma. J Hepatobiliary Pancreat Surg 10(1):67–76
Llover JMDBA, Bruix J (2008) Design and end-points of clinical trials in hepatocellular carcinoma. J Natl Cancer Inst 100:698–711
Longoni N, Kunderfranco P, Pellini S et al (2013) Aberrant expression of the neuronal-specific protein DCDC2 promotes malignant phenotypes and is associated with prostate cancer progression. Oncogene 32(18):2315–2324
Galang CK, Muller WJ, Foos G et al (2004) Changes in the expression of many Ets family transcription factors and of potential target genes in normal mammary tissue and tumors. J Biol Chem 279(12):11281–11292
Fujiwara N, Friedman SL, Goossens N et al (2018) Risk factors and prevention of hepatocellular carcinoma in the era of precision medicine. J Hepatol 68:526–549
Choi SB, Lee JG, Kim KS et al (2008) The prognosis and survival analysis according to seven staging systems of hepatocellular carcinoma following curative resection. Hepatogastroenterology 55(88):2140–2145
Acknowledgements
We are grateful to Prof. Bo Zhang (Shanghai Harmonious One Biotech Co., Ltd, Shanghai, China) for kindly providing HCC cell lines, technical assistance, and helpful discussion regarding this research area. This study was funded by grants from National Natural Science Foundation of China (No. 81602550 and No. 81871700), Science and Technology Development Project of Shandong Province (No. 2018GSF118072), and Natural Science Foundation of Shandong Province (No. ZR2018BH003).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Research involving human participants and animal
Ethical approval for using human samples and animals was obtained from the Ethics Committee of Provincial Hospital affiliated to Shandong University.
Informed consent
Patients had to give written informed consent for the use of their tumor tissues for further investigations at time of the initial diagnosis.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
About this article
Cite this article
Lyu, Z., Ma, M., Xu, Y. et al. Expression and prognostic significance of epithelial tissue-specific transcription factor ESE3 in hepatocellular carcinoma. Int J Clin Oncol 25, 1334–1345 (2020). https://doi.org/10.1007/s10147-020-01675-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10147-020-01675-0