Abstract
Achillea is a crop with Chinese herbal characteristics and horticultural values. Its leaves and flowers contain aromatic oil, and the ripe herb can also be used as medicine to induce sweat and relieve rheumatic pains. It is seen cultivated in gardens all over China. Currently, the most comprehensive chloroplast genome sample involved in the study refers to New World clades of Achillea, which are used for marker selection and phylogenetic research. We completely sequenced the chloroplast genomes of Achillea millefolium. These sequencing results showed that the plastid genome is 149,078 bp in size and possesses a typical quadripartite structure containing one large single copy (LSC) with 82,352 bp, one small single copy (SSC) with 18,426 bp, and a pair of inverted repeat (IR) regions with 24,150 bp in Achillea millefolium. The chloroplast genome encodes a common number of genes, of which 88 are protein-coding genes, 37 transfer ribonucleic acid genes, and 8 ribosomal ribonucleic acid genes, which are highly similar in overall size, genome structure, gene content, and sequence. The exact similarity was observed when compared to other Asteraceae species. However, there were structural differences due to the restriction or extension of the inverted repeat (IR) regions—the palindromic repeats being the most prevalent form. Based on 12 whole-plastomes, 3 hypervariable regions (rpoB, rbcL, and petL-trnP-UGG) were discovered, which could be used as potential molecular markers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chloroplast genomes are highly conservative when compared to mitochondrial and nuclear genomes. They typically include 110–130 unique genes and range in size from 120 to 160 bp (Fu et al., 2017). A pair of inverted repeat (IR) regions is separated by a small single-copy (SSC) area and a long single-copy (LSC) region in the plastid genome of angiosperms, which has a standard quadripartite structure (Zhu et al., 2016; Ye et al., 2018; Wang et al., 2022). Although the plastid genome is found to have a highly conserved sequence and structure in most angiosperms, it has shown considerable variations in various taxa. Structural variations always exist along with the contraction or extension of the IR regions, large inversions or deletions (Downie & Jansen, 2015; Rather et al., 2023), and the gain or loss of genes (Sun et al., 2017). Some hot spots with single nucleotide polymorphisms could be identified, providing enough information for species identification (Zhou et al., 2018). For low nucleotide substitution rates, the lack of recombination, and the inheritance of a single parent, many chloroplast DNA sequences have been used to infer plant phylogenies and population genetic analyses (Shaw et al., 2005; Lu et al., 2016).
With the rapid development of next-generation sequencing (NGS) technology, the entire chloroplast genome is increasingly used for phylogenetic analysis and inference of phylogenetic history, which can provide numerous variable sites for phylogenetic analyses(He et al., 2021). Therefore, the whole chloroplast genomes indicate the potential to reveal evolutionary relationships and have been adopted to generate high-resolution phylogenies and genetic diversity, especially for some unknown relationships of the complex taxa or at low taxonomic levels (Huan et al., 2016; Zhang et al., 2017; Xu et al., 2019). Because different regions of the whole chloroplast genomes differ in their evolutionary rates, it might be an excellent choice for phylogenomic analyses to partition the genome by regions or genes. The concatenated coding genes have been extensively explored in plastome phylogenomics (Zhang et al., 2016; Tang et al., 2022).
To date, the plastid genomes of Achillea have not been sequenced nor obtained on GenBank. The comparative analyses of plastid genomes among different species can inform the characteristics of the plastid genome of Achillea.
Materials and methods
DNA extraction and sequencing
This study used a plant genomic DNA kit to extract total genomic DNAs from leaf material following the protocols (Tiangen Biotech, Beijing, China). We measured the DNA concentration (>50 ng μL−1) using a NanoDrop spectrophotometer and achieved fragmentation using sonication. The fragmented DNA was purified and end-repaired, and the sizes were determined by gel electrophoresis. Paired-end libraries with insert sizes of 350 bp were prepared following Illumina’s standard genomic DNA library preparation procedure, accompanied by a quality control library for sequencing. We sequenced (based on sequencing by synthesis SBS technology) the whole genome of the species using the Illumina Novaseq 6000 platform (Illumina, USA).
Chloroplast genome assembly and annotation analyses
First, the pair-end Illumina raw reads were cleared from adaptors and barcodes and filtered quality using Trimmomatic (Bolger et al., 2014). We trimmed the reads from both ends, removing individual bases with a Phred quality score lower than 20bp and over three consecutive uncalled bases. After trimming, we discarded the entire read with a median quality score lower than 21 or a length less than 40 bp. After the quality filter, we mapped the reads to the chloroplast genome of the closest species with a chloroplast genome available (NCBI download), using Bowtie2 v.2.2.6 to exclude the reads of nuclear and mitochondrial origins (Langmead et al., 2009). Subsequently, we used all putative chloroplast reads mapped to the reference sequence above for de novo assembly to reconstruct the chloroplast genomes using GetOrganelle 1.7.5 (Jin et al., 2020). Finally, we again mapped the clean reads to the complete plastome to examine and correct misassemblies. CpGAVAS2 generated the automatic annotations of the chloroplast genomes and then manually corrected regarding previously published plastomes using the Geneious (Shi et al., 2019; Drummond, 2012). A circular representation of both sequences was drawn using the online tool OGDRAW (Lohse et al., 2013) (https://chlorobox.mpimp-golm.mpg.de/OGDraw.html).
Codon usage bias and Ka/Ks analysis
To identify codon usage patterns, we used all coding sequences (CDSs) to estimate codon usage using CodonW with a translational table = 11. CODEML in PAML v4.973 was used to calculate the nonsynonymous (Ka) and synonymous substitutions (Ks), as well as the ratio of nonsynonymous to synonymous nucleotide substitutions (Ka/Ks) for each gene (Yang & Evolution, 2007).
Genome comparative analysis
For the comparative analyses, the chloroplast genome sequences of Achillea millefolium and 11 Asteraceae species, including Dendrosenecio cheranganiensis (NC_037956.1), Ligularia veitchiana (NC_039385.1), Saussurea tsoongii (NC_044737.1), Marshallia graminifolia (NC_042605.1), Aster hersileoides (NC_042944.1), Erigeron canadensis (NC_046789.1), Leucanthemum maximum (NC_046827.1), Tanacetum coccineum (NC_047308.1), Stilpnolepis centiflora (NC_052884.1), Artemisia fukudo (NC_044156.1), and Ajania pacifica (NC_050690.1), were retrieved from the GenBank. We then compared the chloroplast genome borders of LSC, SSC, and IRs according to their annotations. We rearranged the genome with the Mauve Genome Alignment v2.3.1 Plugin and the progressive Mauve algorithm (Darling et al., 2004). We used the mVISTA program in the Shuffle-LAGAN mode to compare the whole chloroplast genome of these species (Frazer et al., 2004; Mayor et al., 2000).
Identification of hypervariable regions
We first aligned the sequences using MAFFT v7 and manually adjusted them using BioEdit software (Katoh et al., 2005). Subsequently, we implemented a sliding window analysis to evaluate the nucleotide variability (Pi) of the cp genome using DnaSP version 5.1 software (Hall, 1999; Librado and R.J. Bioinformatics, 2009). The window range was set at 600 base pairs, and the step size was 200 base pairs.
Results
Chloroplast genome content and organization of Achillea millefolium
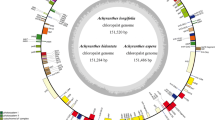
Like most land plants, the whole chloroplast genome of Achillea millefolium features a typical quadripartite genome organization with a size of 149,078 bp, including an LSC region of 82,352 bp and an SSC region of 18,426 bp, which are separated by two IR (IRa and IRb) regions of 24,150 bp. The gene map of the Achillea millefolium plastid genome is presented in Fig. 1 as a representative. The genome encodes an identical set of 133 genes (Table 1). The 133 genes consist of 8 ribosomal RNA (rRNA) genes, 37 transfer RNA (tRNA) genes, and 88 protein-coding genes.
Plastid genome map of Achillea millefolium. Note: The genes inside the circle are transcribed clockwise, and those outside are transcribed counterclockwise. The darker gray in the inner circle shows the GC content, while the lighter gray shows the AT content. The genes of different functions are color-coded
Codon usage bias
Based on the coding sequence (CDS) of 88 protein-coding genes, the relative synonymous codon usage (RSCU) value is estimated, with a total length of 78,336 bp, accounting for 52.55% of the complete plastid genome of Achillea millefolium. A total of 26,112 codons were found in these coding regions, with the most common amino acid being Leucine (2827 codons, approximately 10.83%), followed by Isoleucine (2207 codons, about 8.45%), and Serine (1997 codons, about 7.65%), the rarest one being Cysteine (291 codons, about 1.11%) (Supplementary Table S1). Moreover, almost all A/U-ending codons show RSCU values greater than 1, while the same values of the C/G-ending codons are less than 1. For example, the corresponding RSCU values of the four codons encoding valine in synonymous codons GUU, GUC, GUA, and GUG are 1.46, 0.49, 1.52, and 0.52, respectively, which expected the low GC content of CDS (Fig. 2).
SSR and repeat sequences analysis
Simple sequence repeats (SSRs) are DNA stretches comprising 1–6 bp short tandem repeat motifs, which have been widely applied as molecular markers in population genetics and evolutionary biology (Chmielewski et al., 2015). This study for Achillea millefolium identified 60 SSRs from chloroplasts with ranges from 10 to 21 bp (Fig. 3; Supplementary Table S2). These SSRs could be used to study the genetic diversity, the genetic structure of natural populations, and the cultivars of this species.
Long repeats can promote the plastid genome rearrangement and increase the population’s genetic diversity. Using REPuter (https://bibiserv.cebitec.uni-bielefeld.de/reputer), a total of 44 long repeats were identified across the Achillea millefolium plastome, ranging from 30 to 24,150 bp (Supplementary Table S3), among which 17 are forward repeats, 24 palindromic repeats, 3 reverse repeats, and 0 complement repeats (Fig. 4).
IRscope expansion and contraction analysis of asteraceae
The expansion and contraction of IR often give rise to variations in genome size among different plant lineages, making the study of the phylogenetic classification and the genome evolution among plant lineages. We compared 12 species in Asteraceae, indicating that the gene rps19 crosses the LSC/IRB region, and the gene ycf1 crosses the SSC/IRA region. The genes rpl2, trnN, ndhF, and trnH are close to the junction of IR/SC (Fig. 5). Overall, the IR boundary regions vary slightly in 12 chloroplast genomes.
Comparison of the large single-copy (LSC), short single-copy (SSC), and inverted repeat (IR) regions in the plastomes of twelve Asteraceae species. Note: The colored boxes denote the genes. The gaps between the genes and the boundaries are indicated by the base lengths (bp). The thin lines represent each area connection points, and the figures give information about the genes near the connection points
Genome comparison
We examined the rate of nonsynonymous (Ka) and synonymous (Ks) substitutions and the Ka/Ks ratio to detect the selective pressure on the genes. Only 80 protein-coding genes have Ka or Ks values (Fig. 6; Supplementary Table S4). We could not determine the Ka/Ks values of the additional protein-coding genes because either Ka or Ks equals 0, which indicates that these sequences are preserved and do not contain any nonsynonymous or synonymous nucleotide substitutions. The Ka/Ks ratios vary from 0.00 to 0.70, indicative of purifying selective constraint acting on the chloroplast protein-coding genes. Genes with Ka/Ks values equal to 0 include atpH, petD, petG, petN, psaC, psaI, psaJ, psbA, psbE, psbI, psbJ, psbM, rpl32, and rpl36, indicating that they are under solid purifying selection.
To characterize the structure and collinearity of these species, we aligned their plastid genome sequences by Mauve (Fig. 7). The results of the alignments show the similarities of gene clusters, called local collinear blocks (LCBs). The LCBs also visualize major rearrangements because they are connected to lines in the alignment and indicate inverted regions. The synteny analysis identified no rearrangement events.
Mauve alignment of plastid genomes of 12 species of Asteraceae. The Achillea millefolium genome is put at the top as the reference genome. Within each of the alignments, local collinear blocks are represented by blocks of the same color connected by lines. Note: The rectangular blocks represent the similarity between genomes. The lines between the rectangular blocks represent the collinear relationship. The short squares represent the gene locations of the genomes. Among them, white represents CDs, green represents tRNA, and red represents rRNA
We calculated the nucleotide diversity (Pi) values by sliding the window to determine the hotspots of divergence. These values range from 0.001 to 0.158, and high levels of genetic variation are detected in 3 regions (Fig. 8). In addition, rbcL was found to be more variable than others. The results indicated that the rbcL region might undergo rapid nucleotide substitution in the Achillea species and may contain more important variable information for species authentication and phylogenetic classification.
Discussion
The current study disclosed the first complete plastid genome sequence for A. millefolium, genus Achillea, and family Asteraceae. We compared the plastid genome with eleven plastid genomes of the related species from the Asteraceae family. The chloroplast genomes vary in size from 149 bp to 153 kb, comprising four core parts of the chloroplast genome structure. All the plastid genomes are conserved and fall in the same range, with the genome sizes ranging from 149,078 bp in A. millefolium to 152,721 bp in E. canadensis, which encode 129–134 genes (129 in T. coccineum, 133 in A. millefolium, and 134 in D. cheranganiensis). As the plastid genome length of typical plant angiosperms ranges from 20 to 28 kb, the IRs of these species are 24–26 kb. However, we also observed some variations in these plastid genomes, mainly because of variations in the LSC regions rather than the recently discovered contraction and expansion of the IR region.
We determined 21 genes with introns in these genomes, including 8 tRNA genes and 13 protein-coding genes. Among these introns-containing genes, ycf3 and rps12 genes revealed two introns. In coordination with the previously reported plastid genomes, angiosperms rps12 divide unevenly. The A. millefolium LSC, SSC, and IR regions exhibited 35.62%, 30.71%, and 43.28% GC contents, respectively. High levels of GC content were discovered in IRs because of the presence of 8 rRNAs, which is typical of other angiosperm plastid genomes.
We compared the complete A. millefolium plastid genome with the 11 related plant plastid genomes. The analysis of plastid genes with known functions showed 74 protein-coding genes in A. millefolium and associated species. In addition, the gene contents and organization of A. millefolium are like those of other Asteraceae plastid genomes.
Although conservative gene order has been reported in most plants, some significant changes have occurred at the boundary between IRs, SSCs, and LSCs, such as sequence inversion, gene loss, and contraction and expansion. Because of the contraction and development of the IR region, similar length variations were observed in plastid genomes previously. In terms of SC boundary positions, size, structure, and IR, the plastid genome of A. millefolium is extremely constrained and conservative. However, due to the contraction and expansion of IR regions, it is common to see some diversions in most land plants.
Besides, we performed detailed analyses of JSA, JSB, JLA, and JLB between the IRs and LSCs regions of A. millefolium and with 11 related species. Despite the similar lengths of the IR regions of A. millefolium and the associated species, we determined some contractions and extensions, with the IR regions ranging from 24,150 bp in A. millefolium to 25,423 bp in M. graminifolia. Despite the four conserved junctions in these cp genomes, we also observed some variations with S. centiflora and M. graminifolia cp genomes. The ycf1 gene is 759 bp away in the IRa region in S. centiflora. Simultaneously, we detected in other genomes some partially duplicated genes in the IRs, including 904 bp of ycf1 in M. graminifolia.
Various former researchers have found that with their essential role in phylogenetic studies, repeat sequences can contribute significantly to genome rearrangement. We noted a total of 60 repeats in the A. millefolium cp genome altogether. The comparative analysis found higher repeats (94) in L. maximum while the lowest in S. tsoongii (44) cp genome, respectively. SSRs are useful molecular markers to determine the high variation of similar species, so they have been used to delve into population genetics and polymorphisms. They have been mainly used to analyze gene flow, estimate genetic variation, and demystify the evolutionary history of flora and fauna. In this study, we have detected 60 microsatellites in the plastid genome of A. millefolium and about 33 microsatellites in the noncoding regions. They cooperated with angiosperm plastid genomes. Mono SSRs were primarily detected in the A. millefolium plastid genome.
Additionally, these studies aided in the identification of about 55% of SSRs in noncoding areas. Other research has revealed the disparity of SSRs, which may enable it to choose molecular markers for inter- and intra-specific polymorphisms. Our results have confirmed other reports regarding the family Asteraceae that SSRs with ‘A’ or ‘T’ mononucleotide repeats dominate the plastid genomes. Mononucleotide, penta-nucleotide, and hexa-nucleotide repeats contain higher amounts of ‘A’ or ‘T’, suggesting a biased base composition with an overall ‘AT’ richness in the plastid genomes.
Conclusions
In this study, we have elucidated the complete chloroplast genome of Achillea millefolium for the first time. We conducted an in-depth analysis of the boundary regions, long repeats, and SSRs in the plastid genomes of 12 species of Asteraceae. Simple sequence repeats (SSRs) and DNA barcodes are two examples of long-sequence repeats that can be used as innovative molecular markers. We found that the gene order and plastid genome rearrangement of Achillea millefolium are similar to other related species. However, we also detected some divergence in the intergenic regions of genes and some protein-coding genes. Our study has provided a valuable set of information, which could hopefully boost species identification and solve taxonomic questions.
Data availability
The complete plastid genome sequence can be found in GenBank with the accession number ON320384. The supplementary information files include the data used to support the results. The identified long repeat regions (Kurtz et al., 2001) are deposited in GenBank (Achillea millefolium ON320384). The following supplementary files are available online: Table S1: The length of exons and introns in genes with introns in the Achillea millefolium plastid genome; Table S2: Simple sequence repeats (SSRs) in the Achillea millefolium plastid genome; Table S3: Long repeat sequences in the Achillea millefolium plastid genome.
Code availability
Not applicable.
References
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120
Chmielewski M et al (2015) Chloroplast microsatellites as a tool for phylogeographic studies: the case of white oaks in Poland. iforest-Biogeosci Forest 8
Darling ACE et al (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res 14(7):1394–1403
Downie SR, Jansen RKJSB (2015) A comparative analysis of whole plastid genomes from the apiales: expansion and contraction of the inverted repeat, mitochondrial to plastid transfer of DNA, and identification of highly divergent noncoding regions. Syst Bot 40(1):336–351
Drummond AJB (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28(12):1647–1649
Frazer KA et al (2004) VISTA: computational tools for comparative genomics. Nucleic Acids Res 32(Web Server issue):W273–W279
Fu CN et al (2017) Comparative analyses of plastid genomes from fourteen Cornales species: inferences for phylogenetic relationships and genome evolution. BMC Genomics 18(1):956
Hall TAJNAS (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis. Nucleic Acids Symp Ser 41
He T et al (2021) Comparative analysis of mitochondrial genomes of soybean cytoplasmic male-sterile lines and their maintainer lines. Funct Integr Genomics 21(1):43–57
Huan H et al (2016) Species delimitation and interspecific relationships of the genus Orychophragmus (Brassicaceae) inferred from whole chloroplast genomes. Front Plant Sci 7
Jin JJ et al (2020) GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol 21(1):241
Katoh K et al (2005) Improvement in the accuracy of multiple sequence alignment program MAFFT. Genome Inform 16(1):22–33
Kurtz S et al (2001) REPuter: the manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res 29(22):4633–4642
Langmead B et al (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10(3):R25
Librado and R.J. Bioinformatics (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25(11):1451–1452
Lohse M et al (2013) OrganellarGenomeDRAW – a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res 41(Web Server issue):W575–W581
Lu RS, Pan L, Qiu YXJFIPS (2016) The complete chloroplast genomes of three cardiocrinum (Liliaceae) species: comparative genomic and phylogenetic analyses. Front Plant Sci 7
Mayor C et al (2000) VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 16(11):1046–1047
Rather SA et al (2023) Characterization of the complete chloroplast genome of Dalbergia congesta (Fabaceae), an endangered legume endemic to the Nilgiri Hills of Tamil Nadu, India. Funct Integr Genomics 23(2):126
Shaw J et al (2005) The tortoise and the hare II: relative utility of 21 noncoding chloroplast DNA sequences for phylogenetic analysis. Am J Bot 92(1):142–166
Shi L et al (2019) CPGAVAS2, an integrated plastome sequence annotator and analyzer. Nucleic acids Res 47(W1):W1
Sun Y et al (2017) Complete plastome sequencing of both living species of Circaeasteraceae (Ranunculales) reveals unusual rearrangements and the loss of the ndh gene family. BMC Genom 18(1):592
Tang D et al (2022) Characteristics and comparative analysis of Mesona chinensis Benth chloroplast genome reveals DNA barcode regions for species identification. Funct Integr Genomics 22(4):467–479
Wang R et al (2022) Comparative analysis of mitochondrial genomes provides insights into the mechanisms underlying an S-type cytoplasmic male sterility (CMS) system in wheat (Triticum aestivum L.). Funct Integr Genomics 22(5):951–964
Xu WQ et al (2019) Comparative genomics of figworts (Scrophularia,Scrophulariaceae), with implications for the evolution of Scrophularia and Lamiales. J Syst Evol 57(1):11
Yang ZJMB, Evolution YZH (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24(8):1586–1591
Ye W-Q et al (2018) Plastome organization, genome-based phylogeny and evolution of plastid genes in Podophylloideae (Berberidaceae). Mol Phylogenet Evol 127:978–987
Zhang N et al (2016) Another look at the phylogenetic position of the grape order Vitales: chloroplast phylogenomics with an expanded sampling of key lineages. Mol Phylogenet Evol 101:216–223
Zhang S et al (2017) Diversification of Rosaceae since the late Cretaceous based on plastid phylogenomics. New Phytol 214:3
Zhou T et al (2018) Comparative chloroplast genome analyses of species in Gentiana section Cruciata (Gentianaceae) and the development of authentication markers. Int J Mol Sci 19(7)
Zhu A et al (2016) Evolutionary dynamics of the plastid inverted repeat: the effects of expansion, contraction, and loss on substitution rates. New Phytol 209(4):1747–1756
Funding
The study was supported by the Start-up Foundation of Introducing Talents for Scientific Research, Nanjing Vocational University of Industry Technology (No.201050619YK701).
Author information
Authors and Affiliations
Contributions
FL: conceptualization, software, formal analysis, writing – original draft, visualization, and project administration; AM: writing – review and editing, visualization, supervision, and funding acquisition; WY: validation, writing – review and editing, visualization, supervision, and funding acquisition; DX: review and editing, and data curation; CJ: review and editing, data curation.
Corresponding authors
Ethics declarations
Ethical approval
This article contains no studies with human participants or animals performed by authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key message
The analysis of the whole chloroplast genome of Achillea millefolium revealed 149,078 bp, including 82,352 bp of LSC, 18,426 bp of SSC, and 24,150 bp of IR.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, F., Movahedi, A., Yang, W. et al. The complete plastid genome and characteristics analysis of Achillea millefolium. Funct Integr Genomics 23, 192 (2023). https://doi.org/10.1007/s10142-023-01121-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-023-01121-0