Abstract
The release of high-quality chromosome-level genome sequences of members of the Triticeae tribe has greatly facilitated genetic and genomic analyses of important crops such as wheat (Triticum aestivum) and barley (Hordeum vulgare). Due to the large diploid genome size of Triticeae plants (ca. 5 Gbp), transcript analysis is an important method for identifying genetic and genomic differences among Triticeae species. In this review, we summarize our results of RNA-Seq analyses of diploid wheat accessions belonging to the genera Aegilops and Triticum. We also describe studies of the molecular relationships among these accessions and provide insight into the evolution of common hexaploid wheat. DNA markers based on polymorphisms within species can be used to map loci of interest. Even though the genome sequence of diploid Aegilops tauschii, the D-genome donor of common wheat, has been released, the diploid barley genome continues to provide key information about the physical structures of diploid wheat genomes. We describe how a series of RNA-Seq analyses of wheat relatives has helped uncover the structural and evolutionary features of genomic and genetic systems in wild and cultivated Triticeae species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat (Triticum aestivum), barley (Hordeum vulgare), rye (Secale cereale), and the wild relatives of these crops (e.g., members of the genus Aegilops) are closely related and belong to the Triticeae tribe, which evolved some 12 million years ago within the Pooideae subfamily of the Poaceae (Gaut 2002). The haploid genome of diploid Triticeae is ca. 5 Gbp, as estimated based on the genome of diploid cultivated barley (IBSC 2012); over 80% of this genome comprise repetitive elements. All three genomes (A, B, and D) of common hexaploid wheat are similar to the genome of barley in terms of genome size, gene content, and repetitive elements (IWGSC 2014; Wicker et al. 2018). Barley and wheat genes are functionally closely related. Therefore, information about cloned barley genes associated with a particular trait can be used to help identify the genes responsible for a similar trait in wheat. This was clearly demonstrated by a study involving genome editing of three wheat orthologs (Abe et al. 2019) of the barley seed dormancy gene Qsd1 (Sato et al. 2016), leading to high levels of seed dormancy in the genome-edited wheat plants.
Several years after its establishment, the International Sequencing Consortium released draft genome sequences for barley (IBSC 2012), Aegilops tauschii (Luo et al. 2013), and common wheat (IWGSC 2014). These draft genomes were based on the physical maps of BAC (bacterial artificial chromosome) clones covering entire chromosomes and on whole-genome shotgun sequencing assemblies of short reads with gene model annotations based on transcript evidence. Overlapping BAC clones were then arranged into minimum tiling paths, which were subjected to shotgun sequencing. Based on these efforts, high-quality chromosome-level assemblies were released for barley (Mascher et al. 2017), A. tauschii (Luo et al. 2017), and common wheat (IWGSC 2018). Multiple haplotypes of both barley (Jayakodi et al. 2020) and wheat (Walkowiak et al.2020) were recently de novo sequenced, revealing considerable genomic variations within each crop species.
The wheat gene pool encompasses a large number of species in the genera Triticum and Aegilops, including the donor species of hexaploid-cultivated wheat. The availability of chromosome-scale genome assemblies for wheat and barley has facilitated genetic and genomic analysis of cultivated and wild wheat species by providing the basic structures of pseudomolecules and genes. However, the sequences of wild wheat genomes other than the donor genomes of cultivated wheat remain uncharacterized. Genetic and evolutionary analyses of wild wheat species are particularly challenging due to dissimilarities in the structures and sequences of the assembled genomes.
The wheat and barley research communities have generated numerous cDNA sequences that can be used to characterize the expressed portion of the genome and provide a source of genetic markers (Close et al. 2009; Manickavelu et al. 2012). Following the development of next generation sequencers, it became possible to perform massive RNA sequencing (RNA-Seq) of cDNA libraries for expression analysis and the detection of polymorphisms among samples. Many RNA-Seq studies have been performed for expression analysis. Xiang et al. (2019) used this method to examine common and durum wheats and their three putative diploid ancestors and identified gene expression programs and the contributions of the A, B, and D subgenomes to grain development in polyploid wheats. Nevertheless, identifying polymorphisms between polyploid wheat haplotypes remains challenging because short transcript sequences from multiple (sub)genomes were assembled within gene models. Fortunately, RNA-Seq analysis can be used to efficiently detect polymorphisms even among closely related diploid germplasms, such as the breeding germplasms used for barley (Tanaka et al. 2019).
Here, we review our use of RNA-Seq analysis to detect polymorphisms in diploid wheat genomes and generate DNA markers. Analysis of the barley genome often provides useful information about the structures of diploid Triticeae genomes due to the well-annotated gene models and clear descriptions of genome structures available for barley.
DNA marker generation in A. tauschii and its application to hexaploid wheat analysis
A. tauschii is an important D-genome donor species of common wheat. The D-genome of common wheat is less polymorphic than its A- and B-genomes (Rosyara et al. 2019). Tetraploid wheat and A. tauschii can be crossed artificially to produce synthetic hexaploid wheat (Kihara and Lilienfeld 1949; Matsuoka and Nasuda 2004) (Fig. 1a). These synthetic lines can be used as intermediates to exploit the natural variation in A. tauschii for hexaploid wheat improvement (Trethowan and Mujeeb-Kazi 2008). However, compared to cultivated wheat species, the analysis of natural variation in A. tauschii has been limited by the lack of efficient genetic markers for this wild wheat species. The main PCR-based marker system that has been used until recently is simple sequence repeats (SSRs; Somers et al. 2004), but the marker resolution for this system is low, and the genomic positions of SSRs are unknown.
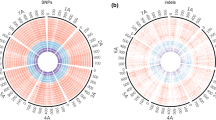
Relationships among Triticeae species. a Images of spikes of hexaploid synthetic wheat (AABBDD) and parental lines Triticum turgidum (AABB) and Aegilops tauschii (DD). b Molecular relationships among diploid Triticeae species (Tanaka et al. 2020)
By contrast, the genomic positions of transcript-based markers can be estimated. Iehisa et al. (2012) and Iehisa et al. (2014) used the early next generation sequencing (NGS) platform known as the 454 system (GS FLX Titanium, Roche Diagnostics) and RNA-Seq to detect polymorphisms among representative A. tauschii accessions (Table 1). Based on an analysis of the population structure of A. tauschii accessions and amplified fragment length polymorphism analysis of 122 accessions, Mizuno et al. (2010) identified two major phylogenetic lineages: 1 (L1) and 2 (L2). Iehisa et al. (2012) used RNA-Seq to analyze two A. tauschii accessions from each lineage. Following de novo assembly of the reads, 9435 single-nucleotide polymorphisms (SNPs) and 739 insertion/deletion polymorphisms (indels) were identified.
Iehisa et al. (2014) developed markers by deep sequencing the spike transcriptomes from the same two A. tauschii accessions and mapping the reads to the F2 population of these accessions. The authors then transferred the marker information to hexaploid wheat accessions, including synthetic hexaploid wheat lines. The synthetic wheat lines were obtained through crosses between tetraploid wheat cv. Langdon and the A. tauschii accessions, followed by chromosome doubling of the interspecific hybrids (Takumi et al. 2009; Kajimura et al. 2011). In total, 5642 out of 5808 contigs with high confidence (HC) SNPs were assigned to the A. tauschii draft genome sequence (Luo et al. 2013). Iehisa et al. (2017) detected splicing variants in the transcripts of two A. tauschii accessions and applied their splicing patterns to synthetic hexaploid wheat. They analyzed alternative splicing of 23,778 loci and identified multiple splicing variants in 4712 genes. For at least some genes, alternative splicing patterns were clearly distinct between the two A. tauschii accessions and were transmitted from the parental A. tauschii accessions to the synthetic hexaploid wheat lines.
Nishijima et al. (2016) generated additional markers using the more recent MiSeq (Illumina) NGS platform to generate 300-bp paired-end reads, which overlapped with the target fragment size of 550 bp. The number of reads ranged from 669,383 (247 Mbp) to 893,917 (386 Mbp) using the 454 platform with 500-bp single reads (Iehisa 2012; Iehisa 2014) but from 4.8 to 5.8 million using MiSeq 300-bp paired-end sequencing reads. After de novo assembly of the merged paired-end reads of ten representative accessions of A. tauschii, 33,680–65,827 transcripts were reconstructed for each accession, with N50 values of 1369–1519 bp. This analysis formed the basis for generating RNA-Seq transcripts, providing a set of unigenes for each accession, including 29,386–55,268 representative isoforms. The deduced unigenes were anchored to the chromosomes of A. tauschii and barley.
The SNPs and indels in the anchored unigenes, which covered entire chromosomes, were sufficient for linkage map construction, even when using combinations between the most closely related accessions. Interestingly, the resolution of SNP and indel distribution was higher in the barley genome than in the A. tauschii genome. Since barley chromosomes are regarded as virtual chromosomes of Triticeae species, this strategy allows genetic markers to be generated that are arranged on the chromosomes in an order based on conserved synteny. This technique is efficient for genome-wide RNA-Seq-based DNA marker generation for A. tauschii.
Applications of DNA markers to the genetic analysis of A. tauschii
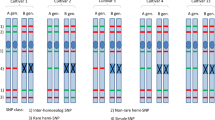
The ability to generate markers by RNA-Seq analysis allowed us to use these markers for genetic analysis. Bulked segregant analysis (BSA) based on RNA-Seq is a powerful method for narrowing the search for candidate causal genes of target phenotypes in polyploid wheat and its wild relatives (Trick et al. 2012; Ramirez-Gonzalez et al. 2015; Edae and Rouse 2019; Li et al. 2019). Nishijima et al. (2018) performed BSA of a bi-parental mapping population of two synthetic hexaploid wheat lines that shared identical A and B genomes but contained D genomes of distinct origins with different Net2 alleles (Fig. 2). An F2 mapping population was generated via a cross between these hexaploid lines. Heterozygotes in subsequent generations were selected and used to generate two F5 lines: one with ten plants homozygous for Net2 and the other with ten plants homozygous for net2. Using the platform described by Nishijima et al. (2016), pooled RNA samples were analyzed by RNA-Seq-based BSA, and D-genome-specific polymorphisms around the Net2 gene were identified. The resulting SNPs were classified into homoeologous polymorphisms of the A, B, and D genomes and D-genome allelic variations based on the RNA-Seq results from a parental tetraploid and two A. tauschii accessions. The difference in allele frequency at the D-genome-specific SNP sites between the contrasting bulks (the ∆SNP-index) was higher for the target chromosome than for the other chromosomes. Several SNPs with the highest ∆SNP-indices were converted into PCR-based markers and assigned to the Net2 chromosomal region, which will subsequently be narrowed down to a single gene.

modified from Nishijima et al. (2018)
An example of RNA-Seq based bulk segregant analysis for locus mapping in wheat. RNA-seq reads generated by Illumina MiSeq were aligned to A. tauschii cDNAs, and SNP calling was conducted for SNP-index calculation to identify SNP markers linked to the locus. This figure was
RNA-Seq-based markers are also being used to detect segments introgressed from A. tauschii to a synthetic hexaploid wheat line showing early heading as a novel source of variation. Takumi et al. (2020) backcrossed two early heading lines of a synthetic hexaploid wheat line derived from a cross between durum wheat and A. tauschii with four Japanese elite cultivars to develop early heading lines of bread wheat. For this technique, it is important that the introgressed lines continue to perform as well as the repeated (original) parent, since A. tauschii may have wild traits that are not suitable for cultivation. RNA-Seq-based genotyping was performed to detect SNPs between the selected lines and their parental wheat cultivars, which successfully revealed the chromosomal regions that were transmitted from the parental synthetic wheat line to the selected lines. These results demonstrate that this technique is efficient for the introgression and identification of traits from A. tauschii in the D-genome.
RNA-Seq-based DNA markers can be used to map genic regions, but not non-genic repeated regions, in the genome. The two examples of this technique being used described above demonstrate that RNA-Seq-based DNA markers can provide sufficient resolution for genetic distance-based analysis of A. tauschii and synthetic hexaploid wheat.
RNA-Seq based DNA marker analysis of other Triticeae species
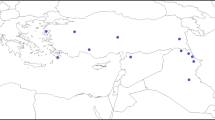
Wild species of the genera Triticum and Aegilops are useful genetic resources for wheat breeding, as they can be used for interspecific crosses among cultivated and wild wheat species (Fig. 1). Tetraploids and hexaploids occur not only in Triticum but also in Aegilops and are thought to be derived from intercrosses between diploid species of Triticum and Aegilops (Tsunewaki 2009 and Chen et al. 2020). However, reference genome sequences are not yet available for most of these species. To evaluate intragenic DNA polymorphisms within species and nucleotide substitutions between species, we used RNA-Seq to analyze 12 species: T. urartu (AA genome), T. monococcum ssp. monococcum (AmAm genome), T. monococcum ssp. aegilopoides (AmAm genome) (Michikawa et al. 2019), A. umbellulata (UU genome) (Okada et al. 2018), A. speltoides (SS genome), A. bicornis (SbSb genome), A. searsii (SsSs genome), A. longissima (SlSl genome), A. sharonensis (SlSl genome) (Miki et al. 2019), A. caudata (CC genome), A. comosa (MM genome), and A. uniaristata (NN genome) (Tanaka et al. 2020). RNA-Seq of these diploid species allowed us to detect genome-wide SNPs and indels across chromosomes. These studies demonstrate the usefulness of RNA-Seq for detecting nucleotide polymorphisms, generating markers that distinguish each genome, and exploring the molecular evolution in these species. A summary diagram is shown in Fig. 1b.
A. umbellulata (UU genome) can be crossed with tetraploid wheat (T. turgidum AABB genome), which allows synthetic hexaploids (AABBUU genome) to be generated via ABU triploids. We performed RNA-Seq analysis of 12 representative accessions of A. umbellulata and reconstructed the transcripts of reads for each accession via de novo assembly (Okada et al. 2018). We anchored the deduced transcripts to the pseudomolecules of A. tauschii and barley, both of which are regarded as virtual chromosomes of A. umbellulata, and determined the distribution of SNPs and indels across the entire chromosomes. Genetic diversity in A. umbellulata was high despite its narrow habitat. No clear lineages were differentiated, and lower-frequency alleles were predominantly detected in A. umbellulata. These rare alleles might be the main source of the high genetic diversity of A. umbellulata.
Two wild diploid wheat species, Triticum monococcum ssp. aegilopoides and T. urartu, are closely related and harbor the Am and A genomes, respectively. T. urartu is the A-genome donor of tetraploid and common wheat, and T. monococcum ssp. monococcum is the cultivated form derived from the wild Am genome wheat subspecies aegilopoides. Since the Am and A genomes are genetically close, identifying a large number of markers that can discriminate between these genomes is challenging. Michikawa et al. (2019) detected genome-wide SNPs and indels from RNA-Seq data from the leaf transcripts of 15 accessions of these two diploid wheat species. The SNPs between the Am and A genomes, which were detected using the A-genome of common wheat as the reference genome, covered all of the chromosomes, facilitating the construction of PCR-based cleaved amplified polymorphic sequence (CAPS) markers that discriminate between the Am and A genomes. These markers effectively confirmed the addition of aegilopoides chromosomes to tetraploid wheat in nascent allohexaploid lines with AABBAmAm genomes. In addition, the markers allowed linkage maps to be constructed for mapping populations of aegilopoides accessions.
The efficient generation of markers in several Triticeae species prompted us to analyze genome differentiation in diploid wild wheat by applying RNA-Seq to wild wheat species with more diverse diploid genomes. We estimated genome differentiation based on an RNA-Seq-based survey of genome-wide polymorphisms throughout homoeologous loci in Triticum and Aegilops (Tanaka et al. 2020). The genome nomenclatures in these species were defined based on their affinity for chromosomal pairing. However, few studies have evaluated genome differentiation based on genome-wide nucleotide variations, especially in the three genomes of the genus Aegilops: A. caudata L. (CC genome), A. comosa Sibth. et Sm. (MM genome), and A. uniaristata Vis. (NN genome). Genetic divergence of the exon regions throughout all of the chromosomes was larger between the M and N genomes vs. the A and Am genomes. A. caudata had the second highest genetic diversity after A. speltoides, the putative B-genome donor of common wheat. In phylogenetic trees derived from nuclear and chloroplast genome-wide polymorphism data, the species with C, D, M, N, U, and S genomes were connected by short internal branches, suggesting that these diploid species emerged during a relatively short evolutionary period. The highly consistent nuclear and chloroplast phylogenetic topologies indicate that the nuclear and chloroplast genomes of the diploid Triticum and Aegilops species coevolved after their diversification into each genome, accounting for most of the genome differentiation among the diploid species. RNA-Seq-based analyses successfully revealed genome differentiation among the diploid Triticum and Aegilops species and supported the chromosome-pairing-based genome nomenclature system, except for the position of A. speltoides. Phylogenomic and epigenetic analyses of the intergenic and centromeric regions are needed to help clarify the basis of this inconsistency.
The S-genome of five species of section Sitopsis of the genus Aegilops is considered to be the origin of the B-genome in cultivated tetraploid and hexaploid wheat species, although the actual donor is unclear. Miki et al. (2019) attempted to elucidate the phylogenetic relationships among Sitopsis species by performing RNA-Seq of the coding regions of each chromosome. They extensively analyzed genome-wide polymorphisms in 19 accessions of the Sitopsis species in reference to the tetraploid and hexaploid wheat B genome sequences. Consequently, these polymorphisms were efficiently anchored to the B genome chromosomes. The results of genome-wide exon sequencing and subsequent phylogenetic analysis indicate that A. speltoides is likely the direct donor of all chromosomes of the wheat B genome. Only three chromosomal regions contradicted this phylogenetic relationship, and these exceptions could be explained by the higher recombination rates in distal regions of wheat chromosomes. The rate of genome differentiation during wheat allopolyploidization from S to B is not constant but varies along the chromosomes; recombination could affect this differentiation rate. This observation could potentially be generalized to genome differentiation during allopolyploidization in other plants.
As part of an IWGSC project, RNA-Seq-derived transcript sequences were used to generate BAC contigs for genome sequencing in wheat. Kobayashi et al. (2015) assembled 689 informative BAC contigs (hereafter referred to as contigs) representing 91% of the entire physical length of wheat chromosome 6B. The authors integrated the contigs into a radiation hybrid (RH) map of chromosome 6B, with one linkage group consisting of 448 loci with 653 markers. They then determined the order and direction of 480 contigs, corresponding to 87% of the total length of chromosome 6B. The authors also characterized contigs that contained part of the nucleolus organizer region or the centromere based on their positions on the RH map and the assembled BAC clone sequences. Analysis of the virtual gene order along chromosome 6B using information collected for the integrated map revealed the presence of several chromosomal rearrangements, representing evolutionary events that occurred on chromosome 6B. The physical map provided a high-quality, map-based reference sequence (IWGSC 2018).
As described above, RNA-Seq-based DNA marker generation could be performed for most diploid wild wheat species of Triticum and Aegilops. These species share orthologous exon sequences, allowing the sequences among species to be compared to estimate their evolutionary relationships. The finding that A. speltoides is likely the B genome donor of common wheat represents one of the most significant contributions of RNA-Seq-based DNA marker generation in wild wheat species. The use of markers for genome sequencing of wheat chromosome 6B is another important byproduct of RNA-Seq-based DNA marker generation.
Current status and future prospects for RNA-Seq-based DNA marker analysis in Triticeae
In conclusion, we reviewed information about (1) the establishment and application of an RNA-Seq-based DNA marker generation system for A. tauschii and (2) the use of RNA-Seq-based DNA markers for other diploid wild wheat species for molecular evolutionary studies.
Due to technical advancements in the ability to assemble large plant genomes (e.g., Triticeae species), chromosome-scale assembly can now be performed quickly and at a reasonable cost (Monat et al. 2019). Even a polyploid genome can be assembled using well-organized whole-genome shotgun reads and scaffold ordering techniques (e.g., Hi-C). These techniques can be used to reveal the variation across genome structures in a crop and related species in pan-genome studies (Sato 2020; Jayakodi et al. 2021). However, balancing economic importance and sequencing costs for the analysis of wild wheat species is an ongoing challenge. RNA-Seq has provided a valuable source of information for identifying polymorphisms in wild wheat accessions for use in genetic and evolutionary studies (Glémin et al. 2019; Miki et al. 2019; Tanaka et al. 2020), even when reference genomes for these species were not available and A. tauschii or barley diploid genomes were used to determine the genomes’ physical structures. As sequencing of each species in Triticeae is currently underway in the research community, the eventual availability of reference genomes for wild wheat species will further increase the efficiency of RNA-Seq-based DNA marker generation. cDNA sequencing has also been improved due to the development of single-molecule sequencing techniques provided by the PacBio and Nanopore platforms. If the gene models of each wild wheat genome are established with high-quality chromosome-scale assembly and mature transcript sequences, it might be possible to map RNA-Seq data onto the reference genome more efficiently.
The main advantage of using RNA-Seq to analyze multiple accessions is that the sequences are generated de novo, in contrast to analyses using pre-fixed polymorphism detection systems such as microarrays. Thus, RNA-Seq analysis has the flexibility to use reference sequences even if they are not from the same species as the reads. As a platform for analyzing the expressed portion of the genome and exons, RNA-Seq is a rapid and cost-effective technique that is particularly useful for analyzing the large genomes of Triticeae species.
References
Abe F, Haque E, Hisano H, Tanaka T, Kamiya Y, Mikami M, Kawaura K, Endo M, Onishi K, Hayashi T, Sato K (2019) Genome-edited triple-recessive mutation alters seed dormancy in wheat. Cell Rep 28:1362–1369
Chen N, Chen W, Yan H, Wang Y, Kang H, Zhang H, Zhou Y, Sun G, Sha L, Fan X (2020) Evolutionary patterns of plastome uncover diploid-polyploid maternal relationships in Triticeae. Mol Phylogenet Evol 149:106838
Close TJ, Bhat PR, Lonardi S, Wu Y, Rostoks N, Ramsay L, Druka A, Stein N, Svensson JT, Wanamaker S, Bozdag S, Roose ML, Moscou MJ, Chao S, Varshney RK, Szűcs P, Sato K, Hayes PM, Matthews DE, Kleinhofs A, Muehlbauer GJ, DeYoung J, Marshall DF, Madishetty K, Fenton RD, Condamine P, Graner A, Waugh R (2009) Development and implementation of high-throughput SNP genotyping in barley. BMC Genomics 10:582–594
Edae EA, Rouse MN (2019) Bulked segregant analysis RNA-seq (BSR-Seq) validated a stem resistance locus in Aegilops umbellulata, a wild relative of wheat. Plos One 14:e0215492
Gaut BS (2002) Evolutionary dynamics of grass genomes. New Phytol 154:15–28
Glémin S, Scornavacca C, Dainat J, Burgarella C, Viader V, Ardisson M, Sarah G, Santoni S, David J, Ranwez V (2019) Pervasive hybridizations in the history of wheat relatives. Sci Adv 5:eaav9188
Iehisa M, Shimizu A, Sato K, Nasuda S, Takumi S (2012) Discovery of high-confidence SNPs from large-scale de novo analysis of leaf transcripts of Aegilops tauschii, a wild wheat progenitor. DNA Res 19:487–497
Iehisa M, Shimizu A, Sato K, Nishijima R, Sakaguchi K, Matsuda R, Nasuda S, Takumi S (2014) Genome-wide marker development for the wheat D-genome based on single nucleotide polymorphisms identified from transcripts in the wild wheat progenitor Aegilops tauschii. Theor Appl Genet 127:261–271
Iehisa JCM, Okada M, Sato K, Takumi S (2017) Detection of splicing variants in the leaf and spike transcripts of wild diploid wheat Aegilops tauschii and transmission of the splicing patterns to synthetic hexaploid wheat. Plant Gene 9:6–12
Jayakodi M, Padmarasu S, Haberer G, Bonthala VS, Gundlach H, Monat C, Lux T, Kamal N, Lang D, Himmelbach A, Ens J, Zhang XQ, Angessa TT, Zhou G, Tan C, Hill C, Wang P, Schreiber M, Boston LB, Plott C, Jenkins J, Guo Y, Fiebig A, Budak H, Xu D, Zhang J, Wang C, Grimwood J, Schmutz J, Guo G, Zhang G, Mochida K, Hirayama T, Sato K, Chalmers KJ, Langridge P, Waugh R, Pozniak CJ, Scholz U, Mayer KFX, Spannagel M, Li C, Mascher M, Stein N (2020) The barley pan-genome reveals the hidden legacy of mutation breeding. Nature 588:284–289
Jayakodi M, Schreiber M, Stein N and Mascher M (2021) Building pan-genome infrastructures for crop plants and their use in association genetics. DNA Res 28:dsaa030. https://doi.org/10.1093/dnares/dsaa030
Kajimura T, Murai K, Takumi S (2011) Distinct genetic regulation of flowering time and grain-filling period based on empirical study of D-genome diversity in synthetic hexaploid wheat lines. Breed Sci 61:130–141
Kihara H, Lilienfeld F (1949) A new-synthesized 6x wheat. Hereditas 35:307–319
Kobayashi F, Wu J, Kanamori H, Tanaka T, Katagiri S, Karasawa W, Kaneko S, Watanabe S, Sakaguchi T, Hanawa Y, Fujisawa H, Kurita K, Abe C, Iehisa JCM, Ohno R, Safár J, Simková H, Mukai Y, Hamada M, Saito M, Ishikawa G, Katayose Y, Endo TR, Takumi S, Nakamura T, Sato K, Ogihara Y, Hayakawa K, Dolezel J, Nasuda S, Matsumoto T, Handa H (2015) A high-resolution physical map integrating an anchored chromosome with the BAC physical maps of wheat chromosome 6B. BMC Genomics 16:595
Li H, Dong Z, Ma C, Tian X, Xiang Z, Xia Q, Ma P, Liu W (2019) Discovery of powdery mildew resistance gene candidates from Aegilops biuncialis chromosome 2Mb based on transcriptome sequencing. PLoS One 14(11):e0220089
Luo MC, Gu YQ, You FM, Deal KR, Ma Y, Hu Y, Huo N, Wang Y, Wang J, Chen S, Jorgensen CM, Zhang Y, McGuire PE, Pasternak S, Stein JC, Ware D, Kramer M, McCombie WR, Kianian SF, Martis MM, Mayer KF, Sehgal SK, Li W, Gill BS, Bevan MW, Simková H, Dolezel J, Weining S, Lazo GR, Anderson OD, Dvořák J (2013) A 4-gigabase physical map unlocks the structure and evolution of the complex genome of Aegilops tauschii, the wheat D-genome progenitor. Proc Natl Acad Sci USA 110:7940–7945
Luo MC, Gu YQ, Puiu D, Wang H, Twardziok SO, Deal KR, Huo N, Zhu T, Wang L, Wang Y, McGuire PE, Liu S, Long H, Ramasamy RK, Rodriguez JC, Van SL, Yuan L, Wang Z, Xia Z, Xiao L, Anderson OD, Ouyang S, Liang Y, Zimin AV, Pertea G, Qi P, Bennetzen JL, Dai X, Dawson MW, Müller HG, Kugler K, Rivarola-Duarte L, Spannagl M, Mayer KFX, Lu FH, Bevan MW, Leroy P, Li P, You FM, Sun Q, Liu Z, Lyons E, Wicker T, Salzberg SL, Devos KM, Dvořák J (2017) Genome sequence of the progenitor of the wheat D-genome Aegilops tauschii. Nature 551:498–502
Manickavelu A, Kawaura K, Oishi K, Shin-I T, Kohara Y, Yahiaoui N, Keller B, Abe R, Suzuki A, Nagayama T, Yano K, Ogihara Y (2012) Comprehensive functional analyses of expressed sequence tags in common wheat (Triticum aestivum). DNA Res 19:165–177
Matsuoka Y, Nasuda S (2004) Durum wheat as a candidate for the unknown female progenitor of bread wheat: an empirical study with a highly fertile F1 hybrid with Aegilops tauschii Coss. Theor Appl Genet 109:1710–1717
Michikawa A, Yoshida K, Okada M, Sato K, Takumi S (2019) Genome-wide polymorphisms from RNA sequencing assembly of leaf transcripts facilitate phylogenetic analysis and molecular marker development in wild einkorn wheat. Mol Genet Genomics 294:1327–1341
Middleton CP, Senerchia N, Stein N. Akhunov, ED, Keller, B, Wicker T (2014) Sequencing of chloroplast genomes from wheat, barley, rye and their relatives provides a detailed insight into the evolution of the Triticeae tribe. PLoS One 9:e85761
Miki Y, Yoshida K, Mizuno N, Nasuda S, Sato K, Takumi S (2019) Origin of wheat B-genome chromosomes inferred from RNA sequencing analysis of leaf transcripts from section Sitopsis species of Aegilops. DNA Res 26:171–182
Mizuno N, Yamasaki M, Matsuoka Y, Kawahara T, Takumi S (2010) Population structure of wild wheat D-genome progenitor Aegilops tauschii Coss.: implications for intraspecific lineage diversification and evolution of common wheat. Mol Ecol 19:999–1013
Monat C, Padmarasu S, Lux T, Wicker T, Gundlach H, Himmelbach A, Ens J, Li C, Muehlbauer GJ, Schulman AH, Waugh R, Braumann I, Pozniak C, Scholz U, Mayer KFX, Spannagl M, Stein N, Mascher M (2019) TRITEX: chromosome-scale sequence assembly of Triticeae genomes with open-source tools. Genome Biol 20:284
Nishijima R, Yoshida K, Motoi Y, Sato K, Takumi S (2016) Genome-wide identification of novel genetic markers from RNA sequencing assembly of diverse Aegilops tauschii accessions. Mol Genet Genomics 291:1681–1694
Nishijima R, Yoshida K, Sakaguchi K, Yoshimura S, Sato K, Takumi S (2018) RNA sequencing-based bulked segregant analysis facilitates efficient D-genome marker development for a specific chromosomal region of synthetic hexaploid wheat. Int J Mol Sci 19:3749
Okada M, Yoshida K, Nishijima R, Michikawa A, Motoi Y, Sato K, Takumi S (2018) RNA-seq analysis reveals considerable genetic diversity and provides genetic markers saturating all chromosomes in the diploid wild wheat relative Aegilops umbellulata. BMC Plant Biol 18:271
Ramirez-Gonzalez RH, Segovia V, Bird N, Fenwick P, Holdgate S, Bery S, Jack P, Caccamo M, Uauy C (2015) RNA-Seq bulked segregant analysis enables the identification of high-resolution genetic markers for breeding in hexaploid wheat. Plant Biotechnol J 13:613–624
Rosyara U, Kishii M, Payne T, Sansaloni CP, Singh RP, Braun HJ, Dreisigacker S (2019) Genetic contribution of synthetic hexaploid wheat to CIMMYT’s spring bread wheat breeding germplasm. Sci Rep 9:12355
Sato K (2020) History and future perspectives of barley genomics. DNA Res 27(4):dsaa023. https://doi.org/10.1093/dnares/dsaa023
Sato K, Yamane M, Yamaji N, Kanamori H, Tagiri A, Schwerdt JG, Fincher JB, Matsumoto T, Takeda K, Komatsuda T (2016) Alanine aminotransferase controls seed dormancy in barley. Nat Commun 7:11625
Somers DJ, Isaac P, Edwards K (2004) A high-density wheat microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Takumi S, Naka Y, Morihiro H, Matsuoka Y (2009) Expression of morphological and flowering time variation through allopolyploidization: an empirical study with 27 wheat synthetics and their parental Aegilops tauschii accessions. Plant Breed 128:585–590
Takumi S, Mitta S, Komura S, Ikeda TM, Matsunaka H, Sato K, Yoshida K, Murai K (2020) Introgression of chromosomal segments conferring early heading date from wheat diploid progenitor, Aegilops tauschii Coss., into Japanese elite wheat cultivars. PLOS One 15(1):e0228397
Tanaka S, Yoshida K, Sato K, Takumi S (2020) Diploid genome differentiation conferred by RNA sequencing-based survey of genome-wide polymorphisms throughout homoeologous loci in Triticum and Aegilops. BMC Genomics 21:246
Tanaka T, Ishikawa G, Ogiso-Tanaka E, Yanagisawa T, Sato K (2019) Development of genome-wide SNP markers for barley via reference-based RNA-Seq analysis. Frontiers in Plant Sci. https://doi.org/10.3389/fpls.2019.00577
The International Barley Genome Sequencing Consortium (IBSC) (2012) A physical, genetic and functional sequence assembly of the barley genome. Nature 491:711–716
The International Wheat Genome Sequencing Consortium (IWGSC) (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788
The International Wheat Genome Sequencing Consortium (IWGSC) (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361:6403
Trick M, Adamski NM, Mugford SG, Jiang CC, Febrer M, Uauy C (2012) Combining SNP discovery from next-generation sequencing data with bulked segregant analysis (BSA) to fine-map genes in polyploid wheat. BMC Plant Biol 12:14
Trethowan RM, Mujeeb-Kazi A (2008) Novel germplasm resources for improving environmental stress tolerance of hexaploid wheat. Crop Sci 48:1255–1265
Tsunewaki K (2009) Plasmon analysis in the Triticum-Aegilops complex. Breed Sci 59:455–470
Walkowiak S, Gao L, Monat C et al (2020) Multiple wheat genomes reveal global variation in modern breeding. Nature 588:277–283
Wicker T, Gundlach H, Spannagl M et al (2018) Impact of transposable elements on genome structure and evolution in bread wheat. Genome Biol 19:103
Xiang D, Quilichini TD, Liu Z, Gao P, Pan Y, Li Q, Nilsen KT, Venglat P, Esteban E, Pasha A, Wang Y, Wen R, Zhang Z, Hao Z, Wang E, Wei Y, Cuthbert R, Kochian LV, Sharpe A, Provart N, Dolf Weijers C, Gillmor S, Pozniak C, Datla R (2019) The transcriptional landscape of polyploid wheats and their diploid ancestors during embryogenesis and grain development. Plant Cell 31:2888–2911
Mascher M, Gundlach H, Himmelbach A, Beier S, Twardziok SO, Wicker T, Radchuk V, Dockter C, Hedley PE, Russell J, Bayer M, Ramsay L, Liu H, Haberer G, Zhang XQ, Zhang Q, Barrero RA, Li L, Taudien S, Groth M, Felder M, Hastie A, Šimková H, Staňková H, Vrána J, Chan S, Muñoz-Amatriaín M, Ounit R, Wanamaker S, Bolser D, Colmsee C, Schmutzer T, Aliyeva-Schnorr L, Grasso S, Tanskanen J, Chailyan A, Sampath D, Heavens D, Clissold L, Cao S, Chapman B, Dai F, Han Y, Li H, Li X, Lin C, McCooke JK, Tan C, Wang P, Wang S, Yin S, Zhou G, Poland JA, Bellgard MI, Borisjuk L, Houben A, Doležel J, Ayling S, Lonardi S, Kersey P, Langridge P, Muehlbauer GJ, Clark MD, Caccamo M, Schulman AH, Mayer KFX, Platzer M, Close TJ, Scholz U, Hansson M, Zhang G, Braumann I, Spannagl M, Li C, Waugh R, Stein N (2017) A chromosome conformation capture ordered sequence of the barley genome. Nature 544:427–433
Acknowledgements
The wheat and barley seed samples used in these studies were provided by the National Bioresource Project. Computations for RNA sequence assembly of reads were performed on the NIG supercomputer at the ROIS National Institute of Genetics, Japan. This work was supported by the MEXT as part of a Joint Research Program implemented at the Institute of Plant Science and Resources, Okayama University, Japan. This work was also supported by Grant-in-Aid for Scientific Research (B) No. 16H04862 from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan to ST and Grant-in-Aid for Scientific Research on Innovative Areas No. 19H04863 from MEXT to ST and KY. KY was supported by JST, PRESTO (No. JPMJPR15QB).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Shigeo Takumi: April 28, 1968 (date of birth)–June 4, 2020 (date of death)
Rights and permissions
About this article
Cite this article
Sato, K., Yoshida, K. & Takumi, S. RNA-Seq-based DNA marker analysis of the genetics and molecular evolution of Triticeae species. Funct Integr Genomics 21, 535–542 (2021). https://doi.org/10.1007/s10142-021-00799-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-021-00799-4