Abstract
Analysis of gene expression data generated by high-throughput microarray transcript profiling experiments coupled with cis-regulatory elements enrichment study and cluster analysis can be used to define modular gene programs and regulatory networks. Unfortunately, the high molecular weight glutenin subunits of wheat (Triticum aestivum) are more similar than microarray data alone would allow to distinguish between the three homoeologous gene pairs. However, combining complementary DNA (cDNA) expression libraries with microarray data, a co-expressional network was built that highlighted the hidden differences between these highly similar genes. Duplex clusters of cis-regulatory elements were used to focus the co-expressional network of transcription factors to the putative regulatory network of Glu-1 genes. The focused network helped to identify several transcriptional gene programs in the endosperm. Many of these programs demonstrated a conserved temporal pattern across the studied genotypes; however, few others showed variance. Based on this network, transient gene expression assays were performed with mutated promoters to inspect the control of tissue specificity. Results indicated that the interactions of the ABRE│CBF cluster with distal promoter regions may have a dual role in regulation by both recruiting the transcription complex as well as suppressing it in non-endosperm tissue. A putative model of regulation is discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat (Triticum aestivum) seed storage proteins (SSPs) serve as one of the primary sources of plant proteins in human diets and animal feed worldwide. These proteins are synthesized in the endosperm; a tissue specialized to starch and protein biosynthesis and storage.
The HMW glutenin subunits (HMW GS) are one of the main storage proteins of wheat. They are encoded by three homoeologous loci denoted as Glu-A1, Glu-B1, Glu-D1, located on the long arm of chromosome 1 of all three genomes. As a result of a tandem duplication, the three Glu-1 loci encode two paralogs of the HMW glutenin subunit, called x and y-type or Glu-1-1 or Glu-1-2, making a total of 6 Glu-1 genes present in the hexaploid wheat. Earlier studies reported that ortholog genes show higher conservation than paralogs (Anderson et al. 2002).
HMW glutenin subunits are solely expressed in the endosperm; the 3–5 active genes in the hexaploid genome account for approximately 12 % of the total seed protein content (Seilmeier et al. 1991; Halford et al. 1992). The expression levels of the homoeolog and paralog Glu-1 genes vary greatly. In general, Glu-1Bx genes have the highest transcription level followed by Glu-1Dx genes. The y-type genes are the lowest expressed. Glu-1Ax gene has a null allele in most genotypes (Payne et al. 1981). However, expression of the same gene can vary greatly across genotypes. It has been reported that the Glu-1Ay gene is always inactive in hexaploid wheat varieties (Shewry et al. 1992). However, active Glu-1Ay genes were identified from related species (Jiang et al. 2009).
The expressions of all prolamin proteins follow a well characterized, although varying, temporal pattern during seed development. Their transcription is regulated by trans-acting factors associated with cis-acting elements as well as epigenetic factors (Fauteux and Strömvik 2009; Kawakatsu and Takaiwa 2010; Juhász et al. 2011; Wen et al. 2012). Earlier studies suggested that transcription of gliadin and LMW glutenin genes (Gli-1,2 and Glu-3, respectively) were influenced by methylation and imprinting while expression of HMW GS coding Glu-1 genes is less dependent on these epigenetic factors (Wen et al. 2012). A closer analysis of the expression profiles of prolamin genes and their responses to abiotic stresses indicate the presence of different regulatory mechanisms for each prolamin protein family (DuPont et al. 2006; Hurkman et al. 2013). A model for different transcriptional regulation mechanisms have already been proposed which is based on the conserved non-coding elements of the LMW GS coding Glu-3 genes (Juhász et al. 2011).
In an earlier in silico experiment, we identified six cis-regulatory modules (CRMs) in the 1600-bp-long promoter region of Glu-1 genes. Our method was based on calculating local overrepresentation and co-occurrences of binding sites. The results are shared with the scientific community on a pre-print server (Makai et al. 2014b). Since then, the first draft genome of Triticum aestivum has been published (Mayer et al. 2014), and it gave us a new opportunity to improve the experiment by calculating the global overrepresentations of binding sites and binding site clusters. In a parallel study, Ravel and co-workers reported conserved cis-regulatory modules in the promoters of HMW-GS promoters that they identified by a simple and non-statistical method (Ravel et al. 2014).
Co-expressional network data and cis-regulatory motif information was proved to be a useful tool to identify sub-networks of genes involved in similar process (Vandepoele et al. 2009). In this study, we aimed to decipher the regulatory logic of Glu-1 genes by a co-expressional network analysis and a high-throughput cis-regulatory module detection study as well as experimental studies. The function and strength of both wild-type and mutant Glu-1x type promoters were studied in reporter gene assays using particle bombardment similarly to earlier studies (Basu et al. 2003; Ravel et al. 2014). In conclusion, a possible regulatory logic of combinatorial cis-regulation of Glu-1 genes is detailed, which was formulated by a reverse approach described in Werner and co-workers (Werner et al. 2003).
Methods and materials
Cis-regulatory module detection
All publicly available gene sequences of HMW glutenin subunits of T. aestivum were collected from NCBI’s nucleotide archive. Additional promoter sequences for Glu-1Ay2 and Glu-Dx12 were downloaded from the wheat survey sequence repository (Mayer et al. 2014). Sequences were categorized by locus, paralog types (x or y), and genotypes. Altogether, there were 156 HMW GS promoter sequences collected. One hundred and forty sequences were longer than 250 bp, 122 longer than 500 bp, and 27 were longer than 700 bp. Eighty-seven promoters belonged to x-type and 69 to y-type HMW GS genes. The promoters represented well all three loci: there were 60 from the A genome, 67 from the B genome, and 29 from the D genome. For sequence characterization, promoters longer than 700 bp were used. List of motifs generally related to prolamin gene promoters were used based on the results of Juhász and co-workers (Juhász et al. 2011). The terminology of motifs and “boxes” is the same as in the above study. Additional motifs were retrieved from PlantCare database. Conserved regulatory modules (CRMs) were identified by a locally developed perl script, Module Detective tool. The Module Detective follows the general framework of CRÈME (Sharan et al. 2003). Motifs were searched by RegEx instead of position weight matrices because RegEx allowed the use of motifs with varying length. Cluster length was set to 200 bp and maximum distance between motifs in a cluster was set to 40 bp. In addition, motif duplexes and triplexes were searched for where maximum distance of motifs were set to 50 bp. One thousand-basepair-long promoter sequences were extracted from the recently published T. aestivum survey sequences (version 22) and used as the background set (Mayer et al. 2014). Binding site (BS) distribution was calculated for both the background set and the Glu1 genes using a sliding window method (window size 100, step size 25). Significance was calculated using the hypergeometric distribution for both motif detection and local BS abundance.
Co-expression network construction
The expression analysis of HMW GS alleles and transcription factors were conducted in silico based on complementary DNA (cDNA) and microarray data of developing wheat seeds. The cDNA-based measurement was carried out as described in our earlier reports (Juhász et al. 2011; Makai et al. 2014a). All together, 31 datasets (11 cDNA libraries and 19 microarray datasets) were used. Microarray data (TA3 and TA38) were downloaded from the PlexDB (Dash et al. 2012). Libraries of cDNA used in the study are shown in Table 1. Expressions of TFs were measured by using all clusters available at the Plant Transcription Factor Database (Zhang et al. 2011). All together, 1940 sequences were used for all known TF families. Co-expression network was based by calculating the Pearson correlation of the normalized expression data and visualized by the software Gephi (Bastian et al. 2009). Focusing the network was achieved by dividing the PCC values of each edge with a p value of the corresponding TF BS duplex. Then, the 10 based logarithm was calculated and used as focused weights to build a network. The cutoff value for the modified edges was 0. Sub-networks were determined using the algorithm of Blondel and co-workers (Blondel et al. 2008).
Allelic composition of genotypes was collected using annotations supplied by their respective depositors or published results. The query sequences of each alleles used for cDNA-based expression measurements are as follows: Glu-1Ax2*—M22208 (for Glenlea, DuPont libraries); Glu-1Bx7—DQ119142 (for Glenlea lib.); and BK006773 (for Chinese Spring and DuPont libs.); Glu-1By8—JF736014 (for the Chinese Spring and the Glenlea libraries); Glu-1By9—X61026 (for the DuPont library); Glu-1Dx2—BK006460 (for the Chinese Spring lib.); Glu-1Dx5—BK006458 (for the Glenlea and the DuPont libraries); Glu-1Dy10—X12929 (for the Glenlea and DuPont libraries); and Glu-1Dy12—BK006459 (for the Chinese Spring library).
Transient reporter gene expression assay
In order to study promoter function, 1000 bp (upstream of the start codon) length wild type and mutant Glu-1Bx7 promoters were cloned to pCambia1391z binary vector (www.cambia.org). Both wild type and mutant Glu-1Bx7 promoters were custom synthesized. As for mutant promoters, the 103-bp-long ABRE|CBF|PBF motif cluster (from −288 to −177) was either deleted (bxDEL::GUS) or replaced (bxRPLC::GUS) with a non-interacting sequence of the same length (used from 16,000 bp upstream of TSS and checked negative for BSs). These were cloned between HinDIII and NcoI sites of pCambia1391z. Empty pCambia1391z with promoter-less uidA gene, and the pKPK1 plasmid, containing a maize ubiquitin constitutive promoter driven uidA gene has also been used in control experiments (Mészáros et al. 2014).
Our transient expression protocol for leaves was based on the method published by Marzin and co-workers (Marzin et al. 2008). T. aestivum cv. Chinese Spring plants served as plant material. Plants were grown in phytotron chambers (Conviron, Winnipeg, Canada) using the spring climatic program T1 (Tischner and Koszegi 1997). Second leaves of 2- or 3-week-old seedlings (two or three leaf-stage, 12–13 Zadoks’ scale, (Zadoks et al. 1974)) were cut into 2–4 cm length pieces and placed onto solid medium in Petri dishes. The medium contained 0.5 % (w/v) phytoagar, 0.4 M mannitol (osmolyte), and 10 μM thidiazuron as senescence inhibitor, respectively. Additionally, 1.65 g l−1 NH4NO3 was also included in the medium which is identical to ammonium-nitrate content of Murashige and Skoog medium. Biolistic transformation has been carried out 1 h after putting the leaves onto the medium. Particle bombardment has been accomplished by using the PDS1000/He system (Bio-Rad, USA). Gold particles of 1 μm diameter were coated according to the manufacturer’s instructions. One milligram of gold particles were coated with 750 ng of plasmid DNA, suspended in 5 μl ethanol and shot to a single Petri-dish. The particle delivery system was adjusted to 1100 psi of helium pressure and 27 mm Hg of vacuum pressure inside the chamber. The distance between the stopping screen and the target was 6 cm. After the bombardment, leaves were kept for 24 h in a climatized room [23 °C constant temperature, natural (indirect) daylight, 30 μmol m2 s−1] for 24 h. Transient expression was stopped by putting the leaves to GUS histochemical staining solution (10 mg ml−1 X-Gluc, 100 mM sodium phosphate, pH 7.0, 10 mM EDTA, 500 mM K4[Fe2(CN)6], 500 mM K3[Fe2(CN)6], 1 % Triton X-100). The staining was performed overnight at 37 °C. After it was completed, photosynthetic pigments were removed by sequential washes with 20, 50, 70, and 100 % ethanol, respectively. GUS spots were counted by using a stereomicroscope. Data are presented as mean values obtained from five independent experiments expressed in percentage of the negative control. ANOVA was performed on the data as statistical analysis.
Transformation of endosperm was based on a method described by Ravel and co-workers (Ravel et al. 2014) with the modification that 1 g l−1 casein hydrolysate was included in the medium after the transformation, instead of separately adding all the 20 proteinogenic amino acids. Biolistic bombardment of endosperms was carried out identically to the leaves. The transient expression lasted for 48 h. GUS-staining was performed identically to leaves, but the tissues were simply put to 70 % ethanol after staining.
Data are presented as mean values obtained from five independent experiments. Unpaired Student’s t tests and ANOVA were performed on the data as statistical analysis.
Results
Motif composition of the promoter of Glu-1 genes
Transcription factors bound to regulatory motifs at the promoter of Glu-1 genes are the main drivers of their transcription. Identifying binding sites and characterizing differences among the six Glu-1 genes and their alleles offers a view on the underlying regulatory mechanisms. Since the allele composition of the six Glu-1 genes are genotype specific, differences across alleles may be directly correlated with the phenotypes, thus with dough making quality.
Analysis of BSs demonstrated that promoter of Glu-1 genes have a modular distribution for TBF, bZIP, PBF, ABRE, VP1, MYB, and NAC binding motifs (Fig. 1a, b). The distribution profiles demonstrated five well separated regions on the promoter (Fig. 1c). In the proximal promoter region (−150 bs), TBF and CBF (NF-YA) are overrepresented. Further down, ABRE and PBF recognition sites show distinctive peaks. MYB and VP1 recognition sites are overrepresented between position −400 and −500 bp. A high abundance regions of bZIP BS is found between position −500 and 700 where bZIP type BSs are the single most frequent motifs including the BS for SPA. Further upstream from the transcription start site (TSS), between −700 and −850 bp, recognitions sites of ABI3/VP1 and NAC type TFs are slightly overrepresented.
Surface diagrams of sliding window analysis of 1000-bp-long promoter sequences of Glu1 genes for TF BSs. The height of the surface represents the 10 based negative logarithm of significance value (p value) of the BS occurrence(s) in the region. Window size was 100 and step size was 25 bp. a Motif distribution of Glu-1x genes. b Motif distribution of Glu-1y genes. Subpanel c shows a schematic guide to help localize the regions as referred in the text. Region 1 is where TATA box is overrepresented. In the case of x-type gene, there is a peak for MYB BSs as well. Next is a region where ABRE, PBF, and CBF BSs show a peak in both genes. Region 3 is an interim region that separates region 2 from the bZIP rich region 4. Region 3 has a low peak for VP1 and MYB BSs. Region 5 show different characteristic for the x and y genes. Glu-1x promoters have VP1 (RY-elements) enriched in this region while Glu-1y show a peak for NAC BSs
The greatest difference between the analysed promoter sequences have been found in a region between the ABRE motif at −277 bp relative to transcription starting site in sense direction and the MYB BSs between -400 and -500 bps. Exact positions varied for the different alleles. In the case of Glu-1Bx genes (with the exception of Glu-1Bx13), there is a 55-bp-long insertion that resulted in a duplicated CEREAL-box and the loss of a PBF at -418 bp. In the case of Glu-1Ay genes, there is a 131-bp-long deletion resulting in the loss of CEREAL-box and the loss of PBF BS at −300 bp. In the same region at around −312 bp, Glu-1Ax genes also lack this PBF BS. Above −700 bp, the motifs are less frequent and less conserved. However, while the motif occurrences of this distal upstream region are relatively conserved, their positions are more polymorphic due to insertions and deletions described above (data not shown).
Identifying cis-regulatory modules
Determining the positions of single binding sites (BSs) is necessary but not sufficient to “decode” the regulatory mechanisms programmed in the promoters of the Glu-1 genes. Therefore, an analysis to determine local and global overrepresentation and clusters of binding sites on the studied promoters was carried out. In conclusion, during these analyses, we found that single BSs and certain motif clusters follow a highly conserved, non-overlapping distribution.
Co-occurrences of BSs are already apparent from the distribution diagram (Fig. 1), although it cannot tell how specific these co-occurrences are to the Glu-1 genes. Therefore, a motif cluster analysis was performed (Supplementary table 1). Overrepresented motif clusters, where at least two (duplex) or three (triplex) BSs or more (sliding window) are combined, were identified for Glu-1 genes. A region between position −250 and −224 was detected, where an ABRE, CBF, PBF, and HMW enhancer are in various combinations. A well conserved cluster of an antisense bZIP, sense MYB, and VP1 was detected between −450 and −550. Both the sliding window and the triplex analysis found the highly abundant bZIP region between −500 and −700 that was reported above.
Concluding the distribution analysis and motif cluster analysis, we could reconfirm putative cis-regulatory modules on the Glu1 promoters (Fig. 2) initially identified by local overrepresentation on promoters. The 177-long proximal promoter region of Glu-1 genes was named the basal promoter region. It contains a conserved composition of TATA at -92 bp and CBF at -37 bp and includes an additional cluster of MYB at -143 bp and MYBs between -174 -177 bp in x-type genes. This is followed by CRM1 that includes the HMW-enhancer, a PBF BS-rich region, the conserved ABRE at -277 bp, and the CEREAL box for x-type genes. CRM2 is a highly conserved motif cluster composed of sense MYB, VP1, and an antisense bZIP BSs. Upstream of CRM2 are regions enriched in BSs belonging to various TF families. CRM3 is a bZIP-rich region, including the BS for SPA. Their exact regions vary by the Glu-1 gene and the function of insertions and deletions. CRM4 of Glu-1x contains the invert couples of MYB BSs in the basal promoter regions as illustrated in Fig. 2. Furthermore, VP1 (RY- elements) demonstrated here a peak on the surface diagram whereas this regions of the Glu-1y genes are overrepresented by NAC BSs.
The cis-regulatory modules (CRMs) of the Glu-1 genes. Genes are grouped by x and y type. Sense and antisense strand is shown in grey and in light red, respectively. Gaps were inserted to the sequences between the CRM1 and CRM2 regions to align the highly conserved motif cluster of the CRM2. The insertions are marked with dashed lines and the lengths are shown. Complementary pairs of MYB BSs are encircled. Dashed and continuous circles represent separate pairs. The geometric distance and centre position of MYB BS pair are drawn for Glu-1Bx genes as example. Also for Glu-1Bx gene, a pair of DOF and bZIP BS is shown, and its distance and centre position is marked. Interestingly, both centre positions are situated at the site of CEREAL at -501 bp. Without the inserted gaps, Glu-1Ax and Glu-1Dx genes have similar structures. The picture is a modified version of the earlier published one (Makai et al. 2014b)
Constructing the co-expression network of Glu-1 genes and the interacting TFs
To further investigate the presence of different regulatory network of the paralogs, a co-expression network was constructed using expression data from microarray and cDNA libraries. Since the paralogs of Glu-1 gene are very similar, microarray data alone cannot distinguish between the two genes. Therefore, the use of cDNA libraries was necessary to obtain appropriate resolution to distinguish between the Glu-1x and Glu-1y genes. Similarly, TFs of the same family are very similar, and the use of cDNA data together with the robust measurements of microarray assays was necessary to study individual genes.
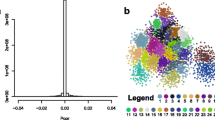
A co-expression network was built by calculating Pearson’s correlation coefficient (PCC), where a cutoff of 0.9 was applied to reduce the number of edges from 404 570 to 8 213. The network has over 1000 nodes including all known TFs of hexaploid wheat and the Glu-1 genes that are expressed in the seed.
The inferred network demonstrated all the co-expression instances that occured between all 1037 TFs used in this study. However, our aim was to study the regulatory network of Glu-1 genes; therefore, the edges and nodes had to be filtered.
The motif cluster analysis identified clusters of binding sites that were significantly overrepresented in the promoters of Glu-1 genes. The p values of the duplex clusters were used as a factor to modify the weight of the edges (see methods). This resulted in a network that was more focused on the co-expression instances most likely to be involved in Glu-1 regulation (Fig. 3). This method filtered out nodes of TFs without known binding sites on the promoter of Glu-1 genes and also decreased the weights of edges where the TF-TF interactions were too general.
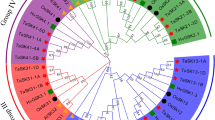
Co-expression network of wheat developing endosperm focusing on Glu-1 genes and their putative interacting partners based on cis-regulatory elements. Colors are representing the modules identified by the algorithm of Blondel and co-workers (Blondel et al. 2008). Green denotes the early expressing ABI3 (Ta058138) gene program, blue the Enhancer program (involving the SPA, PBFs, LEC1, LEC2, and an ABI3 homologue genes), red the L1L (Ta033841) and brown the late ABA-related gene cluster. Gene programs of Glu-1x (yellow) and Glu-1y (purple) are well separated. Glu-1y program is closer to the late expressing ABA-related and L1L gene programs
The hubs of the network allowed us to identify different regulatory networks (or modular gene programs) by inspecting the modularity of the network. The computationally identified sub-networks were named applying the following rules: if a gene was in the same cluster as Glu-1x or Glu-1y, then the cluster was named Glu-1x or Glu-1y, respectively. When a cluster contained the PBF and SPA genes, it was named Enhancer. In other cases, the cluster was named after the gene known to be involved in endosperm specific regulation (ABI3, L1L and ABA). Some of the programs have an absolute temporal profile, while others did not follow a conserved time course. ABI3 was early expressed while ABA and L1L programs were usually switched on late (after the Glu-1 programs) in all studied genotypes. However, the timing of the Enhancer program varied between genotypes. The Enhancer program in Chinese spring peaked late at 20 DPA, which is after the peaks of Glu-1 genes, whereas the Enhancer program in Glenlea was expressed as early as 5 DPA (Fig. 4).
Heatmaps based on expression data of (a) Glenlea and (b) Chinese spring libraries. Tree is based on hierarchical clustering of the expression data using Cluster 3.0 (de Hoon et al. 2004). The PlanTFDB IDs with and the name of gene programs are shown to right of the heatmap. Bold text highlights genes like spa (Tae003578), pbfs (Tae046928, Tae051344, and Tae021210), and abi3 (Tae058138). Genes belonging to the Enhancer program are expressing early in Glenlea as opposed to Chinese spring. The ABI3 program occurs early in both genotypes making it virtually indistinguishable from the Enhancer program in Glenlea. Similarly, the genes belonging to the L1L program are activated later during the endosperm development. Only genes that are significantly expressed in both libraries are shown
We looked at TFs participating in each program. Figure 5 shows TFs by their type except in the case of PBF and SPA where we thought it was more appropriate to mark them separately. The largest program was the one named Enhancer. In this cluster, we found all the PBFs, LECx, and the SPA TF genes. ABI3 was the second largest group, but with a less diverse TF composition. It had TFs that were expressed relatively early, and it is mostly bZIPs of the HBP-1 subfamily, VP1, NF-Y (including CBFs), and MYB genes. This composition is reflecting the highly conserved CRM2 promoter region. In Glenlea, no separate ABI3 cluster was observed and ABI3 was typically co-expressed with the Enhancer program. In the Glu-1 programs, the TF compositions were rather similar with minor differences. Glu-1x program contained three MYB TFs whereas Glu-1y had only one. JaMYB type TFs (Tae044348, Tae040197) were unique to the regulatory network of Glu-1x genes, whereas Glu-1y program has TaMYB1 type MYB (Tae025436). Slightly more stress-related TFs were in the Glu-1y subnetwork then in the Glu-1x, although the difference was not significant (data not shown). The L1L and ABA programs had distinct although lower number of TFs. L1L program contained the highest number of NAC TFs whereas the ABA program had the highest number ABA-related bZIP TFs (TaABI5, TaABF).
TF distribution on the identified gene programs during seed development. Vertical axe represents the number of TFs. The order of the programs represents a hypothetical chronology; however, the exact timing of the programs would need more data. The Enhancer program is most diverse and contains the highest number of genes. This program also shows the greatest divergence in expression dynamics. The ABA and L1L and ABI3 programs have distinct TF compositions and somewhat conserved expression dynamics. L1L has the highest number of NAC TFs suggesting a greater role in Glu-1y type genes based on cis-regulatory elements during the mid-late term of the endosperm development. ABA program contains the likes of abi5 and abf genes suggesting a role during the end of the development. On the other hand, genes belonging to the ABI3 program seem to be an early expressing gene module (cluster) with a TF composition reminiscent to the motif composition of CRM2. Colors of binding sites are as in Fig. 2
Inspecting the roles of modules
In order to study how the highly conserved ABRE|CBF cluster affects tissue specificity, endosperm and a non-target tissue (leaf) were selected for transient gene expression assays. Experiments were based on the presumption that tissue specificity is maintained by negative regulation (i.e., suppression in non-target tissues). To test this hypothesis, two modification of the promoter were designed to study both the effect of presence/absence of the cluster and the necessity of distance between elements upstream and downstream of the implied promoter region. Modified promoters driving the expression of a reporter gene and the results are presented in Fig. 6a, b. As a negative control, an empty pCam1391z construct (GUS without promoter) was used. In leaves, both bxWT::GUS and bxRPLC::GUS demonstrated significant differences compared to the negative control whereas the bxDEL::GUS presented no significant difference in this comparison. The bxRPLC::GUS construct was the highest expressing in all the repeated experiments while the bxDEL::GUS was the least active. The promoter bxRPLC was found to be significantly stronger than both bxDEL (at 99 % level) and bxWT (at 95 % level). As expected, bombardment of leaves with the constitutive ubiquitin promoter driven uidA gene resulted in the most spots (174 spots on average). All bxWT::GUS, bxRPLC::GUS and bxDEL::GUS presented a slight “leak” in wheat leaves exhibiting less than 5 % activity compared to the constitutive promoter (Suppl. figure 1).
To inspect the role of the ABRE|CBF motif cluster, transient gene expression assays were carried out. a Promoter strength expressed in normalized spot counts in biolistic transformation-based transient reporter gene expression assay with Glu-1Bx promoters on wheat leaves (green) and starchy endosperm (brown). b The schematics of the constructs are as follows, bxRPLC stands for the mutated promoter and bxDEL is the line where the CRM1 was removed (abbreviation bxWT stands for the wild type promoter)
Transformation of endosperms presented contrasting results compared to leaves (Fig. 6a and suppl. figure 2). The highest activity was exhibited by bxWT::GUS, followed by the two mutant promoters, bxRPLC and bxDEL. The difference was, in both cases, significant (at 95 and 99 % level, respectively). Only the bxDEL::GUS construct ranked similarly to leaves and demonstrated the lowest activity.
Discussion
Co-expression network and cis-regulatory elements together are excellent tools to determine putative gene interactions related to target genes. In our study, a focused co-expressional network of transcription factors and the Glu-1 genes were constructed. Since co-expression does not necessarily mean interaction, focusing was achieved by (a) filtering out nodes of TFs that has no binding sites (BS) on the promoter of the Glu-1 genes and (b) amending the weight of the edges by the significance of BS co-occurrence of the two TFs linked by the edge.
We previously reported that the motif clusters of cis-regulatory modules CRM4, CRM6, and the basal promoter demonstrated the greatest variance between x- or y-type Glu-1 genes (Fig. 2) (Makai et al. 2014b). Now, the analysis was repeated calculating global overrepresentation using the complete genome of hexaploid wheat as a background. Glu-1y genes have NAC BSs in their CRM4 while x-type genes do not have these BSs at all. It was already suggested that a NAC transcription factor, ENAC1, can be involved in seed development and abiotic stress response in rice (Sun et al. 2012). As for Glu-1x genes, a highly conserved pattern of MYB BSs in the CRM4 and in the basal promoter region were identified. The importance of the interaction of MYB TFs is well known for plants (Dubos et al. 2010; Zhang et al. 2012). Our network analysis further highlighted the importance of these MYB BS pairs genes because of the higher number of MYB TFs in the sub-network of Glu-1x genes.
The co-expression network demonstrated that the Glu-1 paralogs belong to different regulatory networks during grain development. Network analysis was used previously to gain insights to Arabidopsis seed protein regulation (Peng and Weselake 2011). In the case of Arabidopsis, the BSs belonging to B3 and bZIP type TFs are overrepresented in the gene network of seed storage proteins. In case of rice, it has been proposed that multiple regulatory mechanisms may be involved in the endosperm-specific expression of glutelin genes (Qu et al. 2008). Earlier studies already reported that homoeolog wheat Glu-1 genes are controlled by the same regulatory systems (Wanous et al. 2003). Results presented here are in harmony with the general view that duplication is usually followed by divergence of expression and/or sub-functionalization (Li et al. 2005). In addition, differences in promoter profiles of the Glu-1 paralogs suggest two distinct regulatory mechanisms at action.
Beside these gene programs, gene modules related to ABI3, ABA related and L1L genes were identified that demonstrated conserved temporal dynamics across the genotypes. It is an interesting finding of our study that the Enhancer gene program showed temporal variety across genotypes. This can be concluded that it has an influence on the varying HMW-GS contents of the endosperm between cultivars. The Enhancer program of Chinese spring turned on late, after the Glu-1x and y programs, while in the case in Glenlea, Enhancer program started as early as 5 DPA.
Earlier studies showed that promoter region upto −277 nucleotides of Glu-1Dx5 is enough to secure tissue specific expression of the transgene uidA but at a low level (Halford et al. 1989; Robert et al. 1989; Anderson et al. 1998). Other experiment with chimeric promoters with an act1 intron downstream of the basal promoter region of Glu-1 genes increased the activity of reporter genes in rice with an even shorter promoter (251 bp length) (Oszvald et al. 2008). This suggests a basic role of the 277/251 long promoter regions of Glu-1 genes. Indeed, the CCAAT-box of HMW enhancer binds NF-YA TF which in combination with NF-YB and NF-YC TFs have an important role of combinatory regulation in plants (Laloum et al. 2013). The transient expression assay demonstrated in vivo that in the case of Glu-1Bx7 promoter, both the presence of the cluster ABRE at -277 bp & CBF at -230 bp (ABRE|CBF) and the distance of all other distal cis-regulatory elements compared to the transcription start site have important roles in the regulation. These results suggest that the ABRE|CBF cluster has a dual role in recruiting and inhibiting the transcription complex recruited by TATA at -91 bp. When the cluster is deleted, the gene is dysfunctional, and the transcription activity of the reporter gene was reduced significantly compared to the wild type promoter both in leaves and endosperm. However, when the distance constraints were kept and only the cluster was replaced with non-binding sequence, the reporter gene showed higher activity than the wild type promoter but only in leaves. As expected, this construct demonstrated weaker activity in the endosperm than the wild type. This highlights the fact that the implied region may have a principal role in the tissue specificity of its target. To explain all these results, we propose a spatial rearrangement mechanism of the promoter region that mediates interaction between the CRM4 and the basal region that together can drive transcription. In leaves, this mediation is suppressed by factors bound to the ABRE|CBF cluster, while in the endosperm, this suppression is not applied. By mutating (but not deleting) the ABRE|CBF cluster the suppression was eased in non-target tissues. Deletion, on the other hand, appeared to ab ovo inhibit transcription. This distal mediation can be attributed to either MYB-MYB and/or bZIP-PBF interactions (Fig. 7). This extends the earlier hypothesis that assigned tissue specificity solely to the basal promoter (Makai et al. 2014b). Additionally, PBF and bZIP binding sites in close proximity were known to assure tissue specific expression; however, TFs bound to these BSs are not necessarily present in the early phase of seed development and are more characteristic in the Enhancer program.
The schematic representation of the suggested mechanism of the Glu-1x promoter when ABRE|CBF was replaced (bxRPLC; upper) and when it was deleted (bxDEL; lower). When the distance constraints between the CRMs are shortened, distal interaction partners do not overlay and no transcription initiation occurs. When distance is kept, the Enhancer complex of either the MYB-MYB or bZIP-PBF pairs drove transcription. The transcription activity of the bxRPLC was stronger in leaves then that of the wild type promoter that suggests that ABRE|CBF cluster has a role in transcription inhibition
DOF and bZIP interactions are specific to plant seeds (Agarwal et al. 2011); therefore, it indicates that the DOF abundant CRM1 and bZIP abundant CRM3 modules interact. Their communication is likely to be mediated by the highly conserved CRM2 module that contains a tripartite element and requires the right combination of MYB, bZIP, and VP1 TFs. The co-expression network presented in this study demonstrated a tight co-ordinated presence of TFs of these families in the ABI3 and Enhancer gene programs. Bioinformatic analysis have previously demonstrated that tripartite elements of bZIP, MYB, and VP1 BSs are evolutionary conserved and appear to synergistically contribute to auxin-inducible expression (Berendzen et al. 2012). The abi3 (a VP1 type TF) appears early in all studied genotypes in the ABI3 program. Besides, CRM2 is the most conserved module and is present in all Glu-1 genes showing a conserved order and orientation of BSs. A DNA loop may be formed by a TF complex at CRM2 and that brings CRM3 and CRM1 in proximity. This is supported by earlier studies reporting that interaction between MYB and bZIP TFs can form DNA loops at a relatively short (>150 bp) distance (Tahirov et al. 2002). Once, PBF emerge in the endosperm tissue during the Enhancer program and it binds to its cognate BSs in CRM1, it forms a complex with bZIPs bound to CRM3. Subsequently, the CRM1-CRM3 modules take over the transcriptional control of Glu-1 genes from the basal promoter.
The results of the transient gene expression assay let us conclude that the ABRE|CBF cluster and MYB/MYB and/or PBF/SPA enhancer complex (Makai et al. 2014b) works against each other in young wheat leaves. This explains why a minimal level of Glu-1 gene activity can be observed in non-endosperm tissues (Fig. 7).
Furtado and his co-workers reported that a 425-bp-long Glu-1 promoter was leaky and showed expression in the stomata, midrib, and veins of leaf tissue and in root tissue in transgenic rice (Furtado et al. 2008). In one hand, this may prove that tissue specificity is controlled differently in rice and wheat; on the other hand, however, in the light of the modular structure of the promoter, the interaction of CRM3 with CRM1 is missing in this construct that could abolish tissue specificity. In addition, a 425-bp-long Glu-1Dy promoter was also expressed in the aleurone (Furtado et al. 2009). This may indicate that cis-elements upstream of 425 bp may have a role in controlling repression in this tissue. Another study reported that a truncated promoter of 700 bp length of Glu1-Bx7 drove less active transcription than a full length 2000 bp long (Wang et al. 2013). This finding hints possible further evidence of an interaction between the conserved, complementary pairs of MYB BSs in the basal promoter region and CRM4 region that are at equal distance from the CEREAL box (Fig. 2). Since MYB of CRM4 is missing in this construct, no interaction can be formed between the MYB TFs that reduces transcription activity.
References
Agarwal P, Kapoor S, Tyagi AK (2011) Transcription factors regulating the progression of monocot and dicot seed development. Bioessays 33:189–202. doi:10.1002/bies.201000107
Anderson OD, Abraham-Pierce FA, Tam A (1998) Conservation in wheat high-molecular-weight glutenin gene promoter sequences: comparisons among loci and among alleles of the GLU-B1-1 locus. TAG Theor Appl Genet 96:568–576. doi:10.1007/s001220050775
Anderson OD, Larka L, Christoffers MJ et al (2002) Comparison of orthologous and paralogous DNA flanking the wheat high molecular weight glutenin genes: sequence conservation and divergence, transposon distribution, and matrix-attachment regions. Genome 380:367–380. doi:10.1139/G01-137
Bastian M, Heymann S, Jacomy M (2009) Gephi: An Open Source Software for Exploring and Manipulating Networks
Basu C, Kausch AP, Luo H, Chandlee JM (2003) Promoter analysis in transient assays using a GUS reporter gene construct in creeping bentgrass (Agrostis palustris). J Plant Physiol 160:1233–1239. doi:10.1078/0176-1617-01104
Berendzen KW, Weiste C, Wanke D et al (2012) Bioinformatic cis-element analyses performed in Arabidopsis and rice disclose bZIP- and MYB-related binding sites as potential AuxRE-coupling elements in auxin-mediated transcription. BMC Plant Biol 12:125. doi:10.1186/1471-2229-12-125
Blondel VD, Guillaume J-L, Lambiotte R, Lefebvre E (2008) Fast unfolding of communities in large networks. J Stat Mech Theory Exp 2008:P10008. doi:10.1088/1742-5468/2008/10/P10008
Dash S, Van Hemert J, Hong L et al (2012) PLEXdb: gene expression resources for plants and plant pathogens. Nucleic Acids Res 40:D1194–D1201. doi:10.1093/nar/gkr938
Dubos C, Stracke R, Grotewold E et al (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15:573–581. doi:10.1016/j.tplants.2010.06.005
DuPont FM, Hurkman WJ, Vensel WH et al (2006) Differential accumulation of sulfur-rich and sulfur-poor wheat flour proteins is affected by temperature and mineral nutrition during grain development. J Cereal Sci 44:101–112. doi:10.1016/j.jcs.2006.04.003
Fauteux F, Strömvik MV (2009) Seed storage protein gene promoters contain conserved DNA motifs in Brassicaceae, Fabaceae and Poaceae. BMC Plant Biol 9:126. doi:10.1186/1471-2229-9-126
Furtado A, Henry RJ, Pellegrineschi A (2009) Analysis of promoters in transgenic barley and wheat. Plant Biotechnol J 7:240–253. doi:10.1111/j.1467-7652.2008.00394.x
Furtado A, Henry RJ, Takaiwa F (2008) Comparison of promoters in transgenic rice. Plant Biotechnol J 6:679–693. doi:10.1111/j.1467-7652.2008.00352.x
Halford N, Forde J, Shewry P, Kreis M (1989) Functional analysis of the upstream regions of a silent and an expressed member of a family of wheat seed protein genes in transgenic tobacco. Plant Sci 62:207–216
Halford NG, Field JM, Blair H et al (1992) Analysis of HMW glutenin subunits encoded by chromosome 1A of bread wheat (Triticum aestivum L.) indicates quantitative effects on grain quality. Theor Appl Genet. doi:10.1007/BF00224285
De Hoon MJL, Imoto S, Nolan J, Miyano S (2004) Open source clustering software. Bioinformatics 20:1453–1454. doi:10.1093/bioinformatics/bth078
Hurkman WJ, Tanaka CK, Vensel WH et al (2013) Comparative proteomic analysis of the effect of temperature and fertilizer on gliadin and glutenin accumulation in the developing endosperm and flour from Triticum aestivum L. cv. Butte 86. Proteome Sci 11:8. doi:10.1186/1477-5956-11-8
Jiang Q-T, Wei Y-M, Wang F et al (2009) Characterization and comparative analysis of HMW glutenin 1Ay alleles with differential expressions. BMC Plant Biol 9:16. doi:10.1186/1471-2229-9-16
Juhász A, Makai S, Sebestyén E et al (2011) Role of conserved non-coding regulatory elements in LMW-glutenin gene expression. PLoS One 6:e29501. doi:10.1371/journal.pone.0029501
Kawakatsu T, Takaiwa F (2010) Cereal seed storage protein synthesis: fundamental processes for recombinant protein production in cereal grains. Plant Biotechnol J 8:1–15. doi:10.1111/j.1467-7652.2010.00559.x
Laloum T, De Mita S, Gamas P et al (2013) CCAAT-box binding transcription factors in plants: Y so many? Trends Plant Sci 18:157–166. doi:10.1016/j.tplants.2012.07.004
Li W-H, Yang J, Gu X (2005) Expression divergence between duplicate genes. Trends Genet 21:602–607. doi:10.1016/j.tig.2005.08.006
Makai S, Juhász A, Balázs E, Tamás L (2014) A simple and efficient way to in silico study expression of highly similar genes highly. doi:10.6084/m9.figshare.1254162
Makai S, Tamas L, Juhasz A (2014) Distinct regulatory modules identified in the promoters of wheat Glu-1 genes suggest different regulatory mechanisms. bioRxiv 011635. doi:10.1101/011635
Marzin S, Mihaly R, Pauk J, Schweizer P (2008) A transient assay system for the assessment of cell-autonomous gene function in dehydration-stressed barley. J Exp Bot 59:3359–3369. doi:10.1093/jxb/ern186
Mayer KFX, Rogers J, Dole el J et al (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788–1251788. doi:10.1126/science.1251788 (80-)
Mészáros K, Éva C, Kiss T et al (2014) Generating marker-free transgenic wheat using minimal gene cassette and cold-inducible Cre/Lox System. Plant Mol Biol Report. doi:10.1007/s11105-014-0830-1
Oszvald M, Gardonyi M, Tamas C, et al. (2008) Development and characterization of a chimaeric tissue-specific promoter in wheat and rice endosperm. In Vitro 1–7. doi:10.1007/s11627-007-9082-1
Payne PI, Holt LM, Law CN (1981) Structural and genetical studies on the high-molecular-weight subunits of wheat glutenin. Theor Appl Genet 60:229–236. doi:10.1007/BF02342544
Peng FY, Weselake RJ (2011) Gene coexpression clusters and putative regulatory elements underlying seed storage reserve accumulation in Arabidopsis. BMC Genomics 12:286. doi:10.1186/1471-2164-12-286
Qu LQ, Xing YP, Liu WX et al (2008) Expression pattern and activity of six glutelin gene promoters in transgenic rice. J Exp Bot 59:2417–2424. doi:10.1093/jxb/ern110
Ravel C, Fiquet S, Boudet J et al (2014) Conserved cis-regulatory modules in promoters of genes encoding wheat high-molecular-weight glutenin subunits. Front Plant Sci 5:1–17. doi:10.3389/fpls.2014.00621
Robert LS, Thompson RD, Flavell RB (1989) Tissue-specific expression of a wheat high molecular weight glutenin gene in transgenic tobacco. Plant Cell 1:569–578. doi:10.1105/tpc.1.6.569
Seilmeier W, Belitz H-D, Wieser H (1991) Separation and quantitative determination of high-molecular-weight subunits of glutenin from different wheat varieties and genetic variants of the variety Sicco. Z Lebensm Unters Forsch 192:124–129. doi:10.1007/BF01202625
Sharan R, Ovcharenko I, Ben-Hur A, Karp RM (2003) CREME: a framework for identifying cis-regulatory modules in human-mouse conserved segments. Bioinformatics 19:i283–i291. doi:10.1093/bioinformatics/btg1039
Shewry PR, Halford NG, Tatham AS (1992) High molecular weight subunits of wheat glutenin. J Cereal Sci 15:105–120. doi:10.1016/S0733-5210(09)80062-3
Sun H, Huang X, Xu X et al (2012) ENAC1, a NAC transcription factor, is an early and transient response regulator induced by abiotic stress in rice (Oryza sativa L.). Mol Biotechnol 52:101–110. doi:10.1007/s12033-011-9477-4
Tahirov T, Sato K, Ichikawa-Iwata E (2002) Mechanism of c-Myb–C/EBPβ cooperation from separated sites on a promoter. Cell 108:57–70
Tischner T, Koszegi B (1997) Climatic programms used in the Martonvasar phytotron most frequently in recent years. Acta Agron. Hungarica
Vandepoele K, Quimbaya M, Casneuf T et al (2009) Unraveling transcriptional control in Arabidopsis using cis-regulatory elements and coexpression networks. Plant Physiol 150:535–546. doi:10.1104/pp. 109.136028
Wang K, Zhang X, Zhao Y et al (2013) Structure, variation and expression analysis of glutenin gene promoters from Triticum aestivum cultivar Chinese Spring shows the distal region of promoter 1Bx7 is key regulatory sequence. Gene. doi:10.1016/j.gene.2013.06.068
Wanous MK, Munkvold JD, Kruse JD et al (2003) Identification of chromosome arms influencing expression of the HMW glutenins in wheat. Theor Appl Genet 106:213–220. doi:10.1007/s00122-002-1098-7
Wen S, Wen N, Pang J et al (2012) Structural genes of wheat and barley 5-methylcytosine DNA glycosylases and their potential applications for human health. Proc Natl Acad Sci. doi:10.1073/pnas.1217927109
Werner T, Fessele S, Maier H, Nelson PJ (2003) Computer modeling of promoter organization as a tool to study transcriptional coregulation. FASEB J 17:1228–1237. doi:10.1096/fj.02-0955rev
Zadoks JC, CHANG TT, KONZAK CF (1974) A decimal code for the growth stages of cereals. Weed Res 14:415–421. doi:10.1111/j.1365-3180.1974.tb01084.x
Zhang H, Jin J, Tang L et al (2011) PlantTFDB 2.0: update and improvement of the comprehensive plant transcription factor database. Nucleic Acids Res 39:D1114–D1117. doi:10.1093/nar/gkq1141
Zhang L, Zhao G, Jia J et al (2012) Molecular characterization of 60 isolated wheat MYB genes and analysis of their expression during abiotic stress. J Exp Bot 63:203–214. doi:10.1093/jxb/err264
Acknowledgments
The authors are grateful to László Tora and Bob Anderssen for their careful reading, valuable discussions, and their lot of useful advice. In addition, we would like to thank the International Wheat Genome Sequencing Consortium (IWGSC, www.wheatgenome.org) for pre-publication access to the IWGSC Chromosome Survey Sequences hosted at http://wheat-urgi.versailles.inra.fr/Seq-Repository.
Funding
This work was supported by the Hungarian Scientific Research Fund [grant agreement no. OTKA-K100881] and the European Union together with the European Social Fund [grant agreement no. TAMOP 4.2.2/A-11/1/KONV-2012-0008].
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 50 kb)
Supplementary fig. 1
Representative close-up images of GUS histochemical staining of wheat leaves after 24 hours of transient expression following bombardment with (a) bxWT::GUS, (b) bxRPLC::GUS and (c) bxDEL::GUS. (GIF 241 kb)
Supplementary fig. 2
Representative images of GUS histochemical staining of wheat starchy endosperm, after 24 hours of transient expression following bombardment with (a) bxWT::GUS, (b) bxRPLC::GUS and (c) bxDEL::GUS. (GIF 219 kb)
Rights and permissions
About this article
Cite this article
Makai, S., Éva, C., Tamás, L. et al. Multiple elements controlling the expression of wheat high molecular weight glutenin paralogs. Funct Integr Genomics 15, 661–672 (2015). https://doi.org/10.1007/s10142-015-0441-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-015-0441-4