Abstract
Indian lotus (Nelumbo nucifera) is one of the dominant aquatic plants cultivated in Dal Lake, situated at 1586 m above mean sea level (MSL) in the northeast of Srinagar, Kashmir. Despite their economic and ecological role, the microbial communities associated with the lotus plant are still unexplored. In this study, we investigated the prokaryotic communities on surfaces of different lotus microhabitats (roots, rhizome, leaves, flowers, and fruits), lake water, and sediments using 16S rRNA gene amplicon sequencing. Overall, prokaryotic diversity decreased significantly on the surface of lotus microhabitats in comparison to the lake water and sediments. Among the microhabitats of lotus, roots and leaves harbored more diverse communities in comparison to rhizomes, fruits, and flowers. A total of 98 genera were shared by lotus and the Dal Lake sediments and water. However, significant differences were found in their relative abundance; for example, Pseudomonas was the most dominant genus on the majority of lotus microhabitats. On the other hand, Flavobacterium was highly abundant in the lake water, while a higher abundance of Acinetobacter was recorded in sediments. Additionally, we also noted the presence of potential human pathogenic genera including Escherichia-Shigella, Enterobacter, Pantoea, Raoultella, Serratia, and Sphingomonas on the lotus microhabitats. Predicted functions of prokaryotic communities revealed a higher abundance of genes associated with nutrient uptake in the microhabitats of the lotus. This study offered first-hand information on the prokaryotic communities harbored by lotus plants and water and sediments of the Dal Lake and demonstrated the adaptation of diverse communities to microhabitats of lotus.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Dal Lake is a shallow, open-drainage lake fed by Dachigam-Telbal Nallah (with the perennial flow) and Dara Nallah along with several minor streams. The average annual inflow of the lake is around 292 million m3, with Telbal Nallah accounting for 80% of the total and 20% contributed by other sources (Bhat 2013). Long-term studies suggested a significant reduction in lake size, while the floating aquatic vegetation on the surface increased threefold (Rashid et al. 2017). Anthropogenic activities such as the urbanization of the lake interiors, disposal of untreated household waste, and agriculture adversely affected the health of this pristine ecosystem (Rashid et al. 2017; Ahmad et al. 2020).
The phytomicrobiome is associated explicitly with plant parts and provides various services to benefit plants, like the availability of mineral nutrients and the control colonization of pathogens (Rahi 2017). In addition to these benefits, the phytomicrobiome can also provide environment-friendly solutions to current challenges, such as degradation of harmful chemicals and reducing the emission of greenhouse gasses (Hamonts et al. 2018; Ke et al. 2021). See Singh et al. (2021) for a review. Harnessing the potential of phytomicrobiome requires a detailed understanding of the composition of microbial communities (Macdonald and Singh, 2014; Hamonts et al. 2018). Multiple factors, including type of the ecosystems (terrestrial or aquatic), host plant species, microhabitats of the plant (roots, stems and leaves), and geographic location, can influence the structure and colonization of phytomicrobiome. Till now most of the phytomicrobiome studies have focused on the terrestrial plants and there is a huge knowledge gap with regard to aquatic plant microbiome, despite their crucial role in ecosystem service (García-Llorente et al. 2011).
Indian lotus (Nelumbo nucifera) is an aquatic perennial herb, which grows in temperate and tropical climates like marshes and bogs, a floating anchored macrophyte growing in shallow lakes and ponds (Bhattacharjee and Lakshminarasimhan 2018; Gowthami et al. 2021). The plant possesses an anchored rhizome that contains numerous gas canals, mostly misidentified as the lotus stem (Matthews and Seymour 2006). Almost all parts of the lotus are used for various edible preparations. The rhizome of lotus (locally known as nadru) cultivated in the Dal Lake in particular, has high demand due to its unique taste and quality, which helps the farmers fetch a reasonable price in the markets and is the main driving force in its extensive cultivation in the lake. Its leaves are used to wrap food and serve food as a plate due to their unique properties, such as super hydrophobicity and self-cleaning nature (Wang et al. 2009; Gowthami et al. 2021). Cultivating lotus is an important activity from both economic and ecological perspectives in the Dal Lake, as it is also known to remove pollutants (Mishra et al. 2009; Rasid et al. 2019).
A large majority of plant microbiome studies were focused either into the rhizosphere and/or the endosphere, considering their relevance in plant growth and health. Contrary to this, studies have suggested the microbial communities in different parts of plants are distinct and divided plant into several unique microhabitats, such as phyllosphere and anthosphere (Egamberdieva et al. 2017; Chen et al. 2021; Ke et al. 2021). Furthermore, the plants growing in the aquatic ecosystems have distinct microbial communities for above- and below-water compartments (Nielsen et al. 2001; Holmer and Laursen 2002; Shahid et al. 2020). Due to the physiological differences, each part of lotus behaves like a unique microenvironment and might harbor diverse microbial communities. Although lotus cultivation in the Dal Lake has both economic and ecological importance, like other aquatic plants a fewer microbiology studies have been performed so far on lotus microbiome, and those too also with a perspective detecting the phytopathogens (Meeboon and Takamatsu 2015; Chen and Kirschner 2017). In the present study, we deciphered the composition of prokaryotic communities in water and sediment samples and surface microbiome of various microhabitats from root-to-fruits of lotus cultivated in the Dal Lake. We also used the microbiome data to predict the plausible functions of microbial communities harbored by lotus and the Dal Lake water and sediments.
Materials and methods
Site description and sample collection
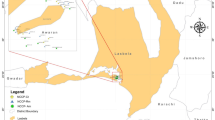
Dal Lake is a postglacial, urban water body regarded as one of the most beautiful lakes in the world located between 34°6′N and 34°10′N latitude and 74°50′E and 74°54′E longitude in the middle of Kashmir Valley (Shah et al. 2014). Water and sediments, and lotus root, rhizome, flower, fruit, and leaf samples were collected in the 50 mL falcon tube (Falcon™ 50 mL Conical Centrifuge Tubes) and sterile sample collection bags from three sites of the Dal Lake. Samples were collected in triplicate from each site and pooled to make one composite sample for each site. The samples were stored in an icebox, transported to the laboratory, and processed immediately for DNA extraction.
DNA extraction for lake water and sediments, and plant parts
Total DNA was extracted from water and sediment samples using MoBio DNA Extraction Kit (MoBio Laboratories, Inc., USA). Before DNA extraction from sediments, the samples were homogenized under aseptic conditions, and 120 mg samples were transferred into the MoBio bead tubes and used for total DNA extraction. For water DNA extraction, 250 mL water was filtered through a 33-mm-diameter sterile hydrophilic Polyethersulfone (PES) membrane filter with a 0.22 µm pore size (Merck, Millipore™, USA) using Merck laboratory filtration systems (Lab System 2). Subsequently, the filter was cut into small parts using sterile scalpel blade no. 4 (Himedia Laboratories, India). Using sterile stainless steel pointed size forceps (Himedia Laboratories, India), the filter parts were transferred into the MoBio bead tube and used for subsequent DNA extraction following the manufacturer’s instruction. To extract community DNA from plant samples, the roots were placed 50 mL centrifuge tube, while three inches of rhizome and intact fruit, flower, and leaf (compressed by folding) were placed in 500 mL sterile γ-irradiated Himedia disposable sterile wide-mouth bottles were submerged in the adequate phosphate buffer saline (PBS). The tubes and bottles were shaken vigorously by hand for 5 min and subjected to sonication for 60 s. The sonication cycles were repeated three times to separate maximum microbial cells present on the surface of plant parts into the PBS. The microbial cells containing PBS were filtered and processed as described for the Lake water samples. The yield and quality of the community DNA samples were checked on 0.8% agarose gel, and DNA concentration was measured using a NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Willingminton, USA). All the extracted DNA samples were stored at − 20 °C until further processing.
Preparation of 16S rRNA gene amplicon libraries and sequencing
The prokaryotic communities were analyzed using 16S rRNA amplicon sequencing via the Illumina-Miseq platform. PCR amplification was performed using the primer set 515F (5′ CCTACGGGNGGCWGCAG 3′) and 806R (5′ GACTACHVGGGTATCTAATCC 3′) that selectively amplified the V4 region of the 16S rRNA gene. Template and library preparation was carried out according to the manufacturer’s protocol (Illumina, USA). The sequencing of multiplexed 16S rDNA amplicon libraries was performed on the Illumina Miseq platform using paired-end 2 × 250 bp chemistry. Multiplexing of the equimolar concentration of DNA was carried out to generate at least 0.1 million (1 lakh) reads for each sample.
Bioinformatics and statistical analysis
Assembly of forward and reverse reads for each sample was carried out using FLASH (Magoč and Salzberg 2011). Microbial diversity analysis was done using a standard QIIME (v1.8.0) pipeline (Caporaso et al. 2010) on the high-quality sequences. A closed reference-based OTU picking approach was used to cluster reads into Operational Taxonomic Units (OTUs) at 97% sequence similarity using UCLUST (Edgar 2010) algorithm and SILVA database for the further downstream analysis. Representative sequences (repset) from each OTU were selected for taxonomic assignment. The chloroplast and mitochondria sequences using the qiime filter table command. Beta diversity (PCoA) was carried out using the scientific data analysis tool PAST3 (Hammer et al. 2001). Detection of the differentially abundant Phyla and genera across the samples was done using analysis of variance (ANOVA) in GraphPad Prism ver. 9.0.
The functional potentials of the prokaryotic communities were predicted using an R package, i.e., Tax4Fun (Aßhauer et al. 2015). It uses taxonomic information based on the 16S rRNA gene sequence and KEGG database to predict functional potentials of the prokaryotic communities. Analysis of differential abundance of functional potentials across different sample groups (including plant parts, lake water, and lake sediment sample) was done based on ANOVA. The genera and predicted genes with a significant difference in their relative abundance (p < 0.05) were plotted in a heatmap matrix in an R package pheatmap (version 1.0.12). Venn diagrams were constructed using an online tool developed by Van de Peer Lab (https://www.vandepeerlab.org/?q=tools/venn-diagrams).
Results
Sequencing results and diversity analyses
Overall, a total of 16,51,507 sequences were generated in the study. Quality filtered sequences (n = 12,93,594) were assigned to 6876 OTUs, belonging to 58 phyla. Values for goods coverage were more than 0.97% for all the samples. A significant decline in all alpha-diversity indices except Simpson was noted for lotus plants in comparison to the water and sediments of the Dal Lake (Fig. 1). No such differences were observed in alpha-diversity indices except observed ASVs among the microhabitats of the lotus plant (Fig. S1). Beta diversity analysis using Bray NMDS plot showed unique clustering of the Dal Lake water and sediments and lotus microhabitats (Fig. 2). The sediments and water of the Dal Lake are placed on the positive side of axis 1, while samples of the lotus microhabitats roots are placed on the negative side of axis 1 with sediments and water of the Dal Lake.
Prokaryotic community structure and differential abundance analysis
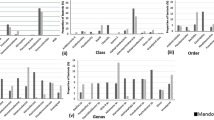
Proteobacteria, Firmicutes, Bacteroidetes, Actinobacteria, Verrucomicrobia, Acidobacteria, Cyanobacteria, Chloroflexi, Nitrospirae, Thaumarchaeota, and Planctomycetes represented more than 95% of the prokaryotic communities across the sample groups (Fig. 3 and Table S1). Proteobacteria was the most dominant phylum across all the samples except lotus leaves and represent more than 90% of the total prokaryotes in the fruit and flower samples. Members of the phylum Firmicutes dominated the prokaryotic communities on the leaves and represent more than 10% of the total prokaryotes in the roots and rhizomes of lotus and sediments of the Dal Lake. Likewise, Actinobacteria makes approximately 20% of the prokaryotic communities in the waters and ≈8% in the sediments of the Dal Lake, while less than 0.5% in lotus plants. Additionally, more than 15% proportion of the prokaryotic communities in the water and sediments of the Dal Lake consists of Bacteroidetes. The proportion of Lokiarchaeota, Deferribacteres, Hadesarchaea, Cyanobacteria, and Bathyarchaeota was less than 1.0% for all the samples, and sediments harbored the highest abundance of these phyla.
Genera level community analysis revealed a high abundance of Pseudomonas, Acinetobacter, Clostridium, Pantoea, Pectobacterium, Flavobacterium, Sphingomonas, Weissella, Rhizobium, Serratia, Polynucleobacter, Bacillus, Alpinimonas, Enterobacter, Pelosinus, Leuconostoc, Polaromonas, and Dickeya across the samples (Fig. 4; Table S2). A significant difference was recorded in the abundance of 71 prokaryotic genera (ANOVA, p < 0.05). The relative abundance of Pseudomonas was consistently higher in lotus microhabitats, whereas water and sediments of the Dal Lake have increased Flavobacterium. Similarly, a high relative abundance of Clostridium was recorded in leaves, rhizomes, and roots of lotus and sediments. A high relative abundance of Bacillus, Massilia, Geobacter, and Arthrobacter was noted in the sediments of Dal Lake, in comparison to water and all microhabitats of lotus. The Dal Lake water exhibited a higher abundance of Polynucleobacter, Alpinimonas, Pseudarcicella, Luteolibacter, and Opitutus. On the other hand, Pantoea, Pectobacterium, Serratia, Rhizobium, Sphingomonas, Weissella, Escherichia-Shigella, Enterobacter, Chryseobacterium, Xanthomonas, Leuconostoc, Lactococcus, and Dickeya were present in higher proportions on one or more microhabitats of the lotus (Fig. 5). These genera were either absent or present in low proportions in the Dal Lake sediments and water.
Shared and unique microbial taxa
Overall, 98 genera were present across the lotus plant, and sediment and water of the Dal Lake (Fig. 6; Table S3). These core members of the communities belonging to genera like Acinetobacter, Flavobacterium, Bacillus, Pseudomonas, Clostridium, Rhizobium, Sphingomonas, Polaromonas, Enterobacter, Pelosinus, Caulobacter, Brevundimonas, Novosphingobium, Hyphomicrobium, Weissella, Cryobacterium, Ruminococcaceae, Stenotrophomonas, Serratia, Hymenobacter, Dialister, Aeromonas, Pantoea, Escherichia-Shigella, Pectobacterium, Methylobacterium, Paracoccus, Psychrobacter, Klebsiella, Aquabacterium, Thauera, and Cronobacter. Sediments of the Dal Lake exhibited the presence of 154 unique genera, comprising several uncultured bacteria and archaea, and genera such as Nitrosospira, Bryobacter, Arcticibacter, Gracilibacter, Oxobacter, Syntrophomonas, Anaerobacillus, Methanocella, Isosphaera, Lacibacter, Anaerovorax, Schlesneria, Singulisphaera, Steroidobacter, Symbiobacterium, Flavitalea, and Methanosaeta. On the other hand, the Dal Lake water has 86 genera unique to sediments and lotus plants, while the lotus plant has only 34 unique genera. Among the microhabitats of the lotus plant, 18 genera including Pseudomonas, Acinetobacter, Clostridium, Sphingomonas, Methylobacterium, Rhizobium, and the members of Enterobacteriaceae such as Klebsiella, Escherichia-Shigella, and Pantoea were found in all microhabitats making them the core of the lotus microbiome. Additionally, 48, 28, 20, 13, and 11 genera were exclusively present in the leaf, root, rhizome, flower, and fruit, respectively.
Prediction of metabolic potentials of the prokaryotic communities
Predictive functional potential analysis showed that the abundance of different functional traits was significantly different among the lotus, and sediments and water of the Dal Lake (Fig. S2; Table S4). The communities of lotus plant microhabitats had functional features unique to the sediment and water of the Dal Lake and are clustered together in the heat map (Fig. S2). The relative abundance of several genes such as glutamine synthetase [EC:6.3.1.2] and glutamate synthase [EC:1.4.1.13], cobalt/nickel transport system permease protein, indolepyruvate ferredoxin oxidoreductase [EC:1.2.7.8], ferrous iron transport protein B and carbamoyl-phosphate synthase [EC:6.3.5.5] was significantly high in the lotus plant microhabitats. On the other hand, a significant increase was observed in the relative abundance of genes encoding isoleucyl-tRNA synthetase [EC:6.1.1.5], penicillin-binding protein 1A [EC:2.4.1. − 3.4. − . −], phosphoribosylformylglycinamidine synthase [EC:6.3.5.3], heterodisulfide reductase subunit A [EC:1.8.98.1], and DNA polymerase III subunit alpha [EC:2.7.7.7] in the Dal Lake water.
Discussion
Diversity analysis of prokaryotic communities
Dal Lake is an urban valley lake in Srinagar, India, known for its beauty and cultivation of the Indian lotus (Shah et al. 2014). The present study deciphered the prokaryotic community structure associated with the lake samples and the root, rhizome, leaf, flower, and fruits of lotus cultivated in the Dal Lake. The diversity and species richness were significantly high in the lake water and sediments, and a significant decrease was observed on the plant surfaces. The decrease in the prokaryotic diversity on the plant microhabitats can be attributed to the availability of nutrient and influence of plant physiological processes such as photosynthesis, respiration, and nutrient uptake. On the other hand, Dal Lake receives large amount of organic matter due to primary production, lotus cultivation, sewage disposal, and fertilizer runoff, making it rich in organic matter and nutrients, which support a much diverse prokaryotic communities in comparison to the lotus microbiome. Similar decline in the bacterial diversity and richness from the soil to the roots and other parts of the plant has been noted in previous studies (Yamamoto et al. 2018).
Built-up of unique and specific microbial communities in different microhabitats of lotus plants was evident from the beta diversity plots. Several factors including ecological drivers like dispersal, diversification, ecological drift, and selection could be responsible for this specific community assemblage on plant surfaces (Sivakumar et al. 2020). Complex interactions between microbe-environment, microbe-microbe, and microbe-plants play an important role in shaping specific prokaryotic communities and making them niche-specific (Rodriguez et al. 2019). In addition to this, difference in the local environment (fluctuations of temperature, moisture, and radiation) and physiology contributes in shaping unique microbial communities specific to the microhabitats. Such specific compartmentation of microbial communities was also observed for A. wallichii (Chen et al. 2021).
Relative abundance of prokaryotic communities in the microhabitats of lotus, and water and sediments of the Dal Lake
The majority of members of microbial communities in the host plant are horizontally acquired from the surrounding environments, and lake prokaryotic communities are the primary reservoir for the communities associated with plants growing in the aquatic ecosystems (Mitter et al. 2017). However, plants tend to shape their microbiome, to meet their requirements for nutrient uptake and stress amelioration (Chaudhari et al. 2020a). In the present study, we noted that Proteobacteria was the most abundant phylum across all microhabitats of lotus and its relative abundance was also high in the Dal Lake water and sediments. The members of phylum Proteobacteria perform diverse metabolic functions and have been found in higher proportions in various environments (Moon et al. 2018). Additionally, it appears that lotus plant provides much suitable colonizing environment for Proteobacteria contrary to the Dal Lake water and sediments, which led to the higher relative abundance of Proteobacteria in lotus microhabitats and thereby lotus can benefit from the presence of the plant growth–promoting Proteobacteria (Compant et al. 2019; Ren et al. 2019; Chaudhari et al. 2020a; Rahi et al. 2020). Furthermore, a higher relative abundance of phyla like Lokiarchaeota, Deferribacteres, Hadesarchaea, Cyanobacteria, and Bathyarchaeota in the lake sediments, in comparison to lake water and lotus parts, indicates their possible role in nutrient cycling in the Dal Lake as reported for the anoxic sediments of other urban water bodies including Aohai Lake and Kunming Lake in Beijing (Wang and Wang 2020). Significant differences recorded in the prokaryotic community structure of water and sediments of the Dal Lake suggested that the lake water is not a mere suspension of sediments as reported previously for the high-altitude Pangong Lake (Chaudhari et al. 2020b). The higher abundance of Actinobacteria in water could be attributed to the presence of high amounts of organic matter suspended in the Dal Lake, and indicates their role to decompose the dead and decaying organisms. Similarly, a higher relative abundance of prokaryotic genera including Geobacter, Thiobacillus, Desulfitobacterium, Desulfatiglans, Thermoanaerobaculum, Thermincola, Ferruginibacter, Lutispora, and Candidatus Nitrosotalea, which have been known for various biogeochemical processes, was noted in sediments of the Dal Lake. Microbial communities are involved in the decomposition of organic and inorganic matter and play a key role in determining the structure and functions of aquatic ecosystems (Wang et al. 2022).
Among the microhabitats of lotus, leaves are responsible for photosynthesis and exchange of gasses, and contary to other lotus microhabitates, leaves showed the predominace of Firmicuties. This difference could be due to the direct exposure of leaves to several abiotic factors like light, humidity, and temperature. Additionally, a higher relative abundance of Clostridium, Deinococcus, and Exiguobacterium was recorded in the leaves; members of these genera are tolerant to a wide range of biotic stress like desiccation and radiations and are reported as phyllosphere colonizers (Wang et al. 2019). The release of carbon-rich exudates by the roots can be attributed to the dominance of Bacteroidetes in the lotus roots, but not in other microhabitats including the adjoining rhiozomes. The higher relative abundance of genera like Polaromonas, Lactococcus, Paenibacillus, Chryseobacterium, Pelosinus, and Sphingobacterium in roots indicated their possible role in plant growth–promoting activities and their preference to utilize organic exudates released by the lotus through its roots. Rhizosphere exudates like organic acids are involved in the bio-stimulation and induction of specific bacterial groups (Macias-Benitez et al., 2020).
A higher proportion of Flavobacterium in the prokaryotic communities was observed in sediment and water samples of the Dal Lake. Flavobacterium has been reported as one of the most abundant genera in freshwater lakes (Eiler and Bertilsson 2007; Michaud et al. 2012), and an increase in the abundance of Flavobacterium was noted in response to the increase in nitrogen inputs, and Flavobacterium are also known to play an important role in the denitrifying process (Li et al 2019). On the other hand, the microhabitats of the lotus plant were dominated by the members of genera, which are known for their beneficial plant growth–promoting activities like Pseudomonas (Vyas et al. 2009; Chen et al. 2021).
A majority of the studies on plant-associated microbial communities remained restricted to the plant rhizosphere, as it is considered the most active site for plant–microbe interactions (Yurgel et al. 2018; Chen et al. 2019; Chaudhari et al. 2020a). In the present study, we found that a deeper investigation of the microbial community structure of different plant parts exhibited a differential relative abundance of prokaryotic taxa across the plant microhabitats. Unique to other microhabitats of lotus, flowers are thermoregulatory (Seymour and Schultze-Motel 1996) and harbored a higher relative abundance of bacterial genera, such as Erwinia, Sphingomonas, Dickeya, Escherichia-Shigella, Pantoea, Serratia, Raoultella and Tolumonasi and Xanthomonas. Several factors including the frequent visit by the insects and the presence of sugar-rich nectar in the flower could provide an explanation to the development of unique flower microbiome (Massoni et al. 2020). Likewise, a higher relative abundance of Pseudoclavibacter was recorded in lotus fruit; this observation was in corroboration to the previous reports on the high relative abundance of members of Pseudoclavibacter in the unwashed citrus fruits (Gomba et al. 2017). Further, the lotus leaves and rhizomes exhibited a high relative abundance of lactic acid bacteria (LAB) like Weissella and Leuconostoc. These results agree with earlier reports on the higher proportions of LAB in the leaves of various plants like Matricaria chamomilla and Calendula officinalis (Köberl et al. 2019). Furthermore, we noted the presence of several potential human pathogenic bacteria, such as Escherichia-Shigella, Enterobacter, Pantoea, Raoultella, Serratia, and Sphingomonas in different parts of lotus, indicating the potential hazard associated with usage of lotus leaves and rhizome. Almost all parts of lotus are edible, especially rhizome, and the leaves are also used to warp and serve food; the presence of pathogenic bacteria indicated that proper hygiene practices should be used to reduce the chances of pathogen exposure.
Differentially abundant potential function of prokaryotic communities
A trend similar to the community structure was observed for the predicted functional profiles of communities, indicating a clear distinction among the lotus microhabitats and lake sediment and water. The genes encoding function such as nutrient and mineral acquisition were more abundant for the communities on the lotus microhabitats. The role of microorganisms in maintaining the flow of nutrients to the host plant is crucial for overall plant growth and productivity, as the direct uptake of nutrients by plants via roots can lead to a nutrient depletion zone (Sun et al. 2021).
In contrast, a higher relative abundance of acetyl-CoA C-acetyltransferase was recorded in lake sediments, which plays an important role in various biosynthesis and degradation pathways (Goudarzi, 2019). Dal Lake water samples showed a higher relative abundance of several genes encoding key functions that are necessary for cellular processes. Though the function assignments of prokaryotic communities are based on prediction, it still offers a fast and reliable alternative to understanding the potential functions of communities. The relevance of such predictions has been found promising in providing information on ecosystem functioning (Djemiel et al. 2022).
Conclusions
In conclusion, the study reveals that parts of the lotus plant have specific prokaryotic communities and site-specific functions. Furthermore, differences were observed in the community structure of sediment and water samples, with enrichment of genera with the potential to degrade organic and inorganic compounds in sediments of the Dal Lake. Despite the variations, a few prokaryotic genera were found constantly on all microhabitats of the lotus plant, lake water, and sediment samples, forming the core microbiome, while the impact of local environment and plant physiology was also noticed shaping microhabitat-specific microbiome. Pseudomonas was explicitly enriched in all plant parts, except the flowers, which have a higher relative abundance of Sphigomonas. These two genera appear to be promising candidates for developing microbial bio-inoculants to promote the growth and health of lotus.
Data availability
The sequence data is available at NCBI SRA submission with SRA no. SRP335402.
Abbreviations
- PCoA :
-
Principal co-ordinate analysis
- NMDS :
-
Non-metric multidimensional scaling
- OTUs :
-
Operational taxonomic unit
- ANOVA :
-
Analysis of variance
References
Ahmad T, Gupta G, Sharma A, Kaur B, Alsahli AA, Ahmad P (2020) Multivariate statistical approach to study spatiotemporal variations in water quality of a Himalayan urban fresh water lake. Water 12(9):2365. https://doi.org/10.3390/w12092365
Aßhauer KP, Wemheuer B, Daniel R, Meinicke P (2015) Tax4Fun: predicting functional profiles from metagenomic 16S rRNA data. Bioinform 31(17):2882–2884. https://doi.org/10.1093/bioinformatics/btv287
Bhat T (2013) Physicochemical characteristics of Dal Lake under temperate conditions of Kashmir (Jammu and Kashmir), India. Int J Curr Sci 9:133–136
Bhattacharjee B, Lakshminarasimhan P (2018) A description of N.nucifera; Central National Herbarium: Howrah, 2018.
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Method 7:335–336. https://doi.org/10.1038/nmeth.f.303
Chaudhari D, Rangappa K, Das A et al (2020) Pea (Pisum sativum l.) plant shapes its rhizosphere microbiome for nutrient uptake and stress amelioration in acidic soils of the north-east region of India. Fron Microbiol 11:968. https://doi.org/10.3389/fmicb.2020a.00968
Chaudhari DS, Dhotre DP, Jani K et al (2020b) Bacterial communities associated with the biofilms formed in high-altitude brackish water Pangong Tso located in the Himalayan Plateau. Curr Microbiol 77:4072–4084. https://doi.org/10.1007/s00284-020-02244-4
Chen K-L, Kirschner R (2017) Fungi from leaves of lotus (Nelumbo nucifera). Mycol Prog 17:275–293. https://doi.org/10.1007/s11557-017-1324-y
Chen S, Waghmode TR, Sun R et al (2019) Root-associated microbiomes of wheat under the combined effect of plant development and nitrogen fertilization. Microbiome 7 https://doi.org/10.1186/s40168-019-0750-2
Chen X, Krug L, Yang M et al (2021) The Himalayan onion (Allium wallichii Kunth) harbors unique spatially organized bacterial communities. Microb Ecol https://doi.org/10.1007/s00248-021-01728-5
Compant S, Samad A, Faist H, Sessitsch A (2019) A review on the plant microbiome: ecology, functions, and emerging trends in microbial application. J Adv Res 19:29–37. https://doi.org/10.1016/j.jare.2019.03.004
Djemiel C, Maron PA, Terrat S et al (2022) Inferring microbiota functions from taxonomic genes: a review. GigaScience 11:giab090. https://doi.org/10.1093/gigascience/giab090
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinform 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Egamberdieva D, Wirth SJ, Alqarawi AA et al (2017) Phytohormones and beneficial microbes: essential components for plants to balance stress and fitness. Front Microbiol 8:2104. https://doi.org/10.3389/fmicb.2017.02104
Eiler A, Bertilsson S (2007) Flavobacteria blooms in four eutrophic lakes: linking population dynamics of freshwater bacterioplankton to resource availability. Appl Environ Microbiol 73:3511–3518. https://doi.org/10.1128/AEM.02534-06
García-Llorente M, Martín-López B, Díaz S, Montes C (2011) Can ecosystem properties be fully translated into service values? An economic valuation of aquatic plant services. Ecol Appl 21:3083–3103
Gomba A, Chidamba L, Korsten L (2017) Effect of postharvest practices including degreening on citrus carpoplane microbial biomes. J Appl Microbiol 122:1057–1070. https://doi.org/10.1111/jam.13396
Gowthami R, Sharma N, Pandey R et al (2021) A model for integrated approach to germplasm conservation of Asian lotus (Nelumbo nucifera Gaertn.). Genet Resour Crop Evol 68:1269–1282. https://doi.org/10.1007/s10722-021-01111-w
Goudarzi A (2019) The recent insights into the function of ACAT1: a possible anti-cancer therapeutic target. Life Sci 232:116592
Hammer Ø, Harper DA, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electr 4:9
Hamonts K, Trivedi P, Garg A et al (2018) Field study reveals core plant microbiota and relative importance of their drivers. Environ Microbiol 20(1):124–140. https://doi.org/10.1111/1462-2920.14031
Holmer M, Laursen L (2002) Effect of shading of Zostera marina (eelgrass) on sulfur cycling in sediments with contrasting organic matter and sulfide pools. J Exp Mar Biol Ecol 270:25–37. https://doi.org/10.1016/s0022-0981(02)00015-1
Ke J, Wang B, Yoshikuni Y (2021) Microbiome engineering: synthetic biology of plant-associated microbiomes in sustainable agriculture. Trend Biotechnol 39(3):244–261. https://doi.org/10.1016/j.tibtech.2020.07.008
Köberl M, Erschen S, Etemadi M et al (2019) Deciphering the microbiome shift during fermentation of medicinal plants. Sci Rep 9:49799. https://doi.org/10.1038/s41598-019-49799-2
Li Y, Sun Y, Zhang H et al (2019) The responses of bacterial community and N2O emission to nitrogen input in lake sediment: estrogen as a co-pollutant. Environ Res 179:108769
Macdonald C, Singh B (2014) Harnessing plant-microbe interactions for enhancing farm productivity. Bioengineered 5(1):5–9. https://doi.org/10.4161/bioe.25320
Macias-Benitez S, Garcia-Martinez AM, Caballero Jimenez P, Gonzalez JM, Tejada Moral T, Parrado Rubio J (2020) Rhizospheric organic acids as biostimulants: monitoring feedbacks on soil microorganisms and biochemical properties. Front Plant Sci 11:633. https://doi.org/10.3389/fpls.2020.00633
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27(21):2957–2963. https://doi.org/10.1093/bioinformatics/btr507
Matthews PGD, Seymour RS (2006) Anatomy of the gas canal system of Nelumbo nucifera. Aqua Bot 85:147–154. https://doi.org/10.1016/j.aquabot.2006.03.002
Meeboon J, Takamatsu S (2015) Erysiphe takamatsui, a powdery mildew of lotus: rediscovery of teleomorph after 40 years, morphology and phylogeny. Mycosci 56:159–167. https://doi.org/10.1016/j.myc.2014.05.002
Michaud L, Caruso C, Mangano S, Interdonato F, Bruni V, Giudice AL (2012) Predominance of Flavobacterium, Pseudomonas, and Polaromonas within the prokaryotic community of freshwater shallow lakes in the northern Victoria Land, East Antarctica. FEMS Microbiol Ecol 82:391–404. https://doi.org/10.1111/j.1574-6941.2012.01394.x
Mishra V, Pathak V, Tripathi B (2009) Accumulation of cadmium and copper from aqueous solutions using Indian lotus (Nelumbo nucifera). AMBIO J Human Environ 38:110–112
Mitter EK, de Freitas JR, Germida JJ (2017) Bacterial root microbiome of plants growing in oil sands reclamation covers. Front Microbiol 8:849. https://doi.org/10.3389/fmicb.2017.00849
Moon CD, Young W, Maclean PH et al (2018) Metagenomic insights into the roles of Proteobacteria in the gastrointestinal microbiomes of healthy dogs and cats. Microbiol Open 7:e00677. https://doi.org/10.1002/mbo3.677
Massoni J, Bortfeld-Miller M, Jardillier L et al (2020) Consistent host and organ occupancy of phyllosphere bacteria in a community of wild herbaceous plant species. ISME J 14:245–258. https://doi.org/10.1038/s41396-019-0531-8
Nielsen LB, Finster K, Welsh DT et al (2001) Sulphate reduction and nitrogen fixation rates associated with roots, rhizomes and sediments from Zostera noltii and Spartina maritima meadows. Environ Microbiol 3:63–71. https://doi.org/10.1046/j.1462-2920.2001.00160.x
Rahi P (2017) Phytomicrobiome: a reservoir for sustainable agriculture. In: Kalia VC, Shouche Y, Purohit HJ, Rahi P (eds) Mining of Microbial Wealth and MetaGenomics. Springer, Singapore, pp 117–132
Rahi P, Giram P, Chaudhari D et al (2020) Rhizobium indicum sp nov, isolated from root nodules of pea (Pisum sativum) cultivated in the Indian trans-Himalayas. Sys App Microbiol 43:126127. https://doi.org/10.1016/j.syapm.2020.126127
Rashid I, Romshoo SA, Amin M, Khanday SA, Chauhan P (2017) Linking human-biophysical interactions with the trophic status of Dal Lake, Kashmir Himalaya, India. Limnologica 62:84–96. https://doi.org/10.1016/j.limno.2016.11.008
Rasid NA, Naim MN, Man HC, Bakar NA, Mokhtar MN (2019) Evaluation of surface water treated with lotus plant Nelumbo nucifera. J Environ Chem Engineer 7(3):103048. https://doi.org/10.1016/J.JECE.2019.103048
Ren X-M, Guo S-J, Tian W et al. (2019) Effects of plant growth-promoting bacteria (PGPB) inoculation on the growth, antioxidant activity, cu uptake, and bacterial community structure of rape (Brassica napus L.) grown in Cu-contaminated agricultural soil. Front Microbiol 10: 1455. https://doi.org/10.3389/fmicb.2019.01455
Rodriguez PA, Rothballer M, Chowdhury SP et al (2019) Systems biology of plant-microbiome interactions. Mol Plant 12:804–821. https://doi.org/10.1016/j.molp.2019.05.006
Seymour RS, Schultze-Motel P (1996) Thermoregulating Lotus Flowers Nat 383:305
Shah JA, Pandit AK, Farooq S (2014) Periphytic algal community of Dal Lake in Kashmir valley, India. Res J Environ Sci 8:391–398. https://doi.org/10.3923/rjes.2014.391.398
Shahid MJ, AL-surhanee AA, Kouadri F et al (2020) Role of microorganisms in the remediation of wastewater in floating treatment wetlands: a review. Sustain 12:5559. https://doi.org/10.3390/su12145559
Singh T, Awasthi G, Tiwari Y (2021) Recruiting endophytic bacteria of wetland plants to phytoremediate organic pollutants. Int J Environ Sci Technol. https://doi.org/10.1007/s13762-021-03476-y
Sivakumar N, Sathishkumar R, Selvakumar G et al (2020) Phyllospheric microbiomes: diversity, ecological significance, and biotechnological applications. in: Yadav, A., Singh, J., Rastegari, A., Yadav, N., (Eds) Plant microbiomes for sustainable agriculture. Sustainable Development and Biodiversity, 25. Springer, Cham. https://doi.org/10.1007/978-3-030-38453-1_5
Sun Y, Wang M, Mur LAJ et al (2021) The cross-kingdom roles of mineral nutrient transporters in plant-microbe relations. Physiol Plant 171(4):771–784
Szymańska S, Sikora M, Hrynkiewicz K et al (2021) Choosing source of microorganisms and processing technology for next generation beet bioinoculant. Sci Rep 11. https://doi.org/10.1038/s41598-021-82436-5
Vyas P, Rahi P, Gulati A (2009) Stress tolerance and genetic variability of phosphate-solubilizing fluorescent Pseudomonas from the cold desert of the trans-Himalayas. Microb Ecol 58:425–434
Wang Y, Guo M, Li X et al (2022) Shifts in microbial communities in shallow lakes depending on trophic states: Feasibility as an evaluation index for eutrophication. Ecol Indicators 136:108691. https://doi.org/10.1016/j.ecolind.2022.108691
Wang J, Chen H, Sui T, Li A, Chen D (2009) Investigation on hydrophobicity of lotus leaf: experiment and theory. Plant Sci 176:687–695. https://doi.org/10.1016/j.plantsci.2009.02.013
Wang W, Ma Y, He J et al (2019) Gene regulation for the extreme resistance to ionizing radiation of Deinococcus radiodurans. Gene 715:144008. https://doi.org/10.1016/j.gene.2019.144008
Wang Y, Wang C (2020) Comparative study on archaeal diversity in the sediments of two urban landscape water bodies. PLoS One 15:e0229097. https://doi.org/10.1371/journal.pone.0229097
Yamamoto K, Shiwa Y, Ishige T et al (2018) Bacterial diversity associated with the rhizosphere and endosphere of two halophytes: Glaux maritima and Salicornia europaea. Front Microbiol 9:2878. https://doi.org/10.3389/fmicb.2018.02878
Yurgel SN, Douglas GM, Dusault A et al (2018) Dissecting community structure in wild blueberry root and soil microbiome. Front Microbiol 9:1187. https://doi.org/10.3389/fmicb.2018.01187
Acknowledgements
The authors acknowledge the Director, NCCS, Pune.
Funding
This work was supported by funding from the Department of Biotechnology (BT/Coord. II/01/03/2016), Government of India. B.A.M. also acknowledges the funding under Energy Bioscience Overseas Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chaudhari, D., Kiran, S., Choudhary, A. et al. Prokaryotic communities adapted to microhabitats on the Indian lotus (Nelumbo nucifera) growing in the high-altitude urban Dal Lake. Int Microbiol 26, 257–267 (2023). https://doi.org/10.1007/s10123-022-00297-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10123-022-00297-y