Abstract
Background
FoxM1 plays important regulatory roles in a variety of diseases. However, the functional role of FoxM1 and mechanisms responsible for its expression in gastrointestinal stromal tumor (GIST) is not thoroughly understood.
Methods
FoxM1 protein expression and biological function were examined in human GIST tissues and cells using immunohistochemistry, quantitative real-time PCR, western blot, CCK-8, wound-healing- and Matrigel invasion assays, respectively. The role of hypoxia-inducible factor (HIF) signaling in FoxM1 expression was investigated using chromatin immunoprecipitation and luciferase reporter and in vivo tumor growth assays.
Results
FoxM1 was highly expressed in highly proliferative and migratory/invasive GIST specimens. Upregulation of FoxM1 was positively correlated with the expression of HIF-1α and HIF-2α in GIST specimens, and hypoxia-induced FoxM1 expression in GIST cells. Functionally, ectopic expression of FoxM1 significantly promoted GIST cell proliferation, cell cycle progression, migration and invasion, whereas the knockdown of endogenous FoxM1 of hypoxic GIST cells had the opposite effects. Molecularly, FoxM1 was transcriptionally regulated by HIF-2α under normoxia, whereas it was upregulated by both HIF-1α and HIF-2α under hypoxia. The xenograft tumor data further confirmed the regulated effect of HIF-1α and HIF-2α on FoxM1, and demonstrated that the simultaneous downregulation of both HIF-1α and HIF-2α inhibited GIST tumor growth.
Conclusions
Our data demonstrated the critical role of FoxM1 in promoting GIST progression and uncovered a novel HIF-1α/HIF-2α-FoxM1 axis. These findings identify FoxM1 as a possible new molecular target for designing novel therapeutic treatments to control GIST progression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastrointestinal stromal tumor (GIST) is the most common mesenchymal neoplasm of gastrointestinal tract. Over 80% of GISTs exhibit the activating mutation of the proto-oncogene c-kit and platelet-derived growth factor receptor alpha (PDGFRA), which is critical for GIST growth and maintenance [1,2,3]. In the last few years, targeted therapies against c-kit and PDGFRA have improved survival of this disease. However, acquired resistance to c-kit/PDGFRA inhibitors eventually occurs in almost all GIST patients during treatment [4, 5].Thus, there is a need to better understand molecular mechanisms involved in GIST progression to discover new therapy strategies.
FoxM1 plays important roles in maintaining the balance between cell proliferation, differentiation and genomic stability [6,7,8]. Experimental and clinical evidence has demonstrated that FoxM1 is abnormally elevated in many human malignancies [9,10,11,12], and functions as a master activator of several cancer-related processes such as angiogenesis [13, 14], invasion, and metastasis [15,16,17]. However, the expression level of FoxM1 in GISTs is not well known. The functional significance and expression mechanism of FoxM1 in GISTs need to be further investigated.
Hypoxia is a common phenomenon in solid tumors and also is associated with aggressive phenotypes and adverse clinical outcome in many cases [18,19,20]. A major mechanism mediating oxygen-dependent transcriptional responses is hypoxia-inducible factor (HIF) that binds to cis-acting DNA hypoxia response element (HRE; 5′-RCGTG-3′) to activate downstream gene transcription. HIF is a heterodimeric protein composed of a constitutively expressed β-subunit (HIF-1β) and an oxygen-regulated α-subunit (HIF-1α, -2α and -3α). Their key regulatory subunits, HIF-1α and HIF-2α, are induced similarly by hypoxia, but the functional roles in tumor may vary, depending on their downstream effector molecules. It has been described that hypoxia contributes to the aggressive behavior of GISTs, and HIF-1α correlates with a worse prognosis of GIST patients [21, 22].These results suggest that HIF-α may act as a critical regulator of adaptive responses to hypoxia in GISTs.
In this study, we first found that FoxM1 was highly expressed in highly proliferative and migratory/invasive GIST specimens. Hypoxia-induced FoxM1 expression and FoxM1 was a target gene of both HIF-1α and HIF-2α. FoxM1 promoted GIST cell proliferation, metastasis and invasion. Our data provide a critical link among FoxM1, HIF-1α, HIF-2α and GIST progression. Thus, FoxM1 expression induced by both HIF-1α and HIF-2α may serve as a novel target for GIST therapy.
Materials and methods
Patients, specimens and gene mutation analysis
Paraffin-embedded GIST specimens were obtained from 83 constrictive patients undergoing surgery at Changhai Hospital. Studies using human specimens were approved by the institutional committee for human research of Changhai hospital. c-kit and PDGFRA mutation analysis was performed using sequencing methods. The detailed information is shown in Supplementary methods.
Bioinformatics analysis
The Oncomine database (http://www.oncomine.org) was used to examine the transcriptional profiles of FoxM1 in human tumors as well as GISTs.
Cell culture and hypoxic condition
GIST-T1 and GIST 882 cells were cultured at 37 °C in a 5% CO2 atmosphere. For experiments involving hypoxia, cells were incubated in a hypoxia incubator in an atmosphere consisting of 94% N2, 5% CO2, and 1% O2.
Plasmid construction and transfection
Plasmids and oligonucleotides used in this study are listed in Supplementary Tables 1 and 2, respectively. The detailed information about plasmid construction and transfection is shown in Supplementary methods.
RNA isolation and quantitative real-time PCR
RNA was isolated from GIST cells, reverse transcribed, and used as a template for the real-time quantitative polymerase chain reaction (qPCR) as described in Supplementary Methods.
Western blot
GIST cells were lysed, and protein expression was evaluated via western blot. The detailed procedure is described in Supplementary Methods.
Chromatin immunoprecipitation (ChIP) assay
ChIP assay was performed using the Pierce agarose ChIP kit (ThermoScientific, Rock- ford, IL). The detailed information for the promoters analyzed is provided in Supplementary Table 3.
Luciferase reporter assay
GIST-T1 cells were co-transfected with the expression plasmids, the promoter reporter plasmids and the pRL-TK (Renila luciferase) plasmids. Luciferase reporter assay was performed using the dual-luciferase reporter assay (Promega, Madison, WI, USA).
Assays of cell proliferation, cell cycle, wound-healing- and Matrigel invasion
GIST cells were transfected with the indicated constructs and exposed to normoxia or hypoxia. After the optimum time of incubation, cell proliferation, cell cycle, wound-healing- and Matrigel invasion assays were performed. The detailed procedure is described in Supplementary Methods.
Tumor growth assay
The animal protocol was approved by the Ethical Review Committee for Animal Experimentation of Changhai Hospital. The tumor growth assay was performed as described in Supplementary methods.
HE and immunohistochemical staining
HE staining was performed according to the standard protocol. Immunohistochemical staining was carried out using a peroxidase Envision method (DAKO, Carpinteria, CA, USA) according to standard methodology. The detailed information about antibodies and immunohistochemical grading is shown in Supplementary methods.
Statistical analysis
Data are mean ± standard error (SD) for continuous variables and as percentages for categorical variables. The qualitative variables were compared using the Pearson’s χ2 test or Fisher’s exact test, quantitative variables were analyzed by the Student’s t test, Wilcoxon signed-rank test or Spearman’s rank correlation test. P value < 0.05 was considered statistically significant for all tests. Statistical analyses were performed using GraphPad Prism 6.0 (GraphPad Software, Inc., La Jolla, CA, USA).
Results
Patient characteristics and demographics
The clinical and pathological characteristics of the 83 patients with GISTs are summarized in Supplementary Table 4. The patients were 42 males (50.6%) and 41 females (49.4%), ranging in age from 32 to 82 years old. Primary sites included stomach (n = 38; 45.78%), small (n = 33; 39.76%) and large bowel (n = 9; 10.84%), as well as extra-gastrointestinal sites (n = 3; 3.61%). Mutational analysis showed that 63 GISTs had mutations in c-kit, including 6 mutations in exon 9 (7.23%), 55 mutations in exon 11 (66.27%) and 2 mutations in exon 13 (2.41%). PDGFRA mutation was found in 5 (6.02%) patients. Representative c-kit/PDGFRA gene mutations are shown in Supplementary Fig. 1.
Increased expression of FoxM1 is associated with tumor progression in GIST patients
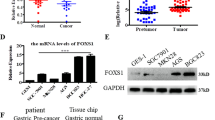
We searched the public datasets on Oncomine (http://www.oncomine.org), and found that the expression of FoxM1 was upregulated in many tumors (Supplementary Fig. 2a). Furthermore, analysis of the public datasets confirmed the frequent FoxM1 amplification in GIST tissues (Supplementary Fig. 2b). Then, we examined the expression levels of FoxM1 in GISTs and adjacent non-tumor tissues by immunohistochemistry. Consistent with an important role of FoxM1 in cell cycle progression, it was expressed in highly proliferative cells of glandular epithelium in the adjacent non-tumor tissues (Fig. 1a). Whereas in GIST specimens, the expression of FoxM1 showed a wide variability in terms of proportion with nuclear staining, from 0 to 100% (Fig. 1a).
FoxM1 was overexpressed and related to tumor proliferation and invasion/metastasis in primary human GIST specimens. a Immunohistochemical staining showed the expression of FoxM1 in human primary GIST tissues and adjacent non-tumor tissues obtained from the same patient. b Immunohistochemical staining of FoxM1 and Ki-67 in GIST tissues obtained from different patients, showed that FoxM1-overexpressing tumor tissue displayed strong Ki-67 staining, vice versa in tumor tissue of low FoxM1 expression. Positive correlation of FoxM1 expression and Ki-67 staining was further confirmed by a Spearman’s analysis. c Immunohistochemical staining of FoxM1 in GIST invasion cases
We further investigated the clinical significance of FoxM1 expression in GIST tissues. All 83 GIST patients were divided into two groups based on immunohistochemical results: high-FoxM1 expression group (n = 21) and low-FoxM1 expression group (n = 62). As shown in Table 1, a high-FoxM1 expression was significantly correlated with tumor size (P < 0.01), mitotic count (P < 0.01), invasion/metastasis (P < 0.01) and risk grading (P < 0.01). We also quantitated the number of FoxM1- and Ki-67-positive nuclei and found that a positive correlation between FoxM1 and the proliferation marker Ki-67 expression levels (P < 0.01, Fig. 1b), an association that was further confirmed by a Spearman’s analysis (r = 0.6640, P < 0.01, Fig. 1a). Moreover, most invasion/metastasis GISTs (13/17, 76%) were found to express high level of FoxM1 (Fig. 1c). These results suggest that increased FoxM1 expression may contribute to GIST progression by accelerating cell proliferation and promoting invasion/metastasis.
FoxM1 expression is upregulated under hypoxia in GIST cells
It has been reported that GIST is commonly characterized by the hypoxia pathway, which leads to constitutive activation of HIF-1α [21, 22]. Therefore, we examined the expression of several endogenous markers of tumor hypoxia in GIST tissues, and found that HIF-1α, HIF-2α, carbonic anhydrase 9 (CA 9) and glucose transporter 1 (GLUT1) were upregulated in most of the GISTs (Fig. 2a). It is well known that dysfunction of succinate dehydrogenase (SDH) complex leads to HIF-α activation [23]. We, therefore, examined SDH subunit B (SDHB) expression in GISTs to verify whether HIF-α was activated via a non-hypoxia pathway. Results showed that only one tumor was negative for SDHB (Supplementary Fig. 3), suggesting that hypoxia is the main factor responsible for the upregulation of HIF-α in GISTs.
The expression of HIF-1α and HIF-2α was upregulated in primary human GIST specimens and related to FoxM1 expression. a Immunohistochemical staining showed the expression of endogenous markers of tumor hypoxia, including HIF-1α, HIF-2α, CA9 and GLUT1, in human primary GIST tissues. b Immunohistochemical staining of HIF-1α, HIF-2α and FoxM1 in GIST tissues obtained from the different patients, showed that the expression of HIF-1α/HIF-2α was related to the expression of FoxM1. Positive correlation of HIF-1α/HIF-2α expression and FoxM1 staining was further confirmed by a Spearman’s analysis. c The expression of HIF-1α, HIF-2α and FoxM1 was examined by western blot and qPCR assays under hypoxia (1% O2) from 0 to 48 h, respectively
Based on immunohistochemistry, we observed a close correlation of FoxM1 expression and the nuclear localization of HIF-1α and HIF-2α in 83 GIST samples (Fig. 2b). The clinicopathological significance and expression pattern of HIF-1α and HIF-2α are summarized in Supplementary Tables 5 to 7.
Next, we sought to investigate whether FoxM1 could be induced by hypoxia in GIST cells. When GIST cell lines, GIST-T1 and GIST 882, were cultured under hypoxic conditions (1% O2) for up to 48 h, the expression of FoxM1 was significantly increased at the mRNA and protein levels in a time-dependent manner (Fig. 2c). Meanwhile, the protein levels of HIF-1α and HIF-2α increased rapidly in these cell lines (Fig. 2c). These results suggest that FoxM1 expression can be upregulated by hypoxia in GISTs, and both HIF-1α and HIF-2α may involve in this process.
HIF-1α and HIF-2α mediates FoxM1 expression in GIST cells
To elucidate the underlying role of HIF-1α and HIF-2α in regulating FoxM1 expression, we knocked down the expression of HIF-1α or HIF-2α by specific short hairpin RNA (shRNA) in GIST-T1 and GIST 882 cells. Under hypoxic conditions, silencing of either HIF-1α or HIF-2α significantly downregulated FoxM1 at the mRNA (Fig. 3a) and protein levels (Supplementary Fig. 4). Although regulated by hypoxia, GIST cells expressed FoxM1under normoxia, and HIF-2α protein could also be detected under normoxic condition (Fig. 2c). We, therefore, determine whether normoxic HIF-2α may drive FoxM1 expression. Results showed that HIF-2α suppression could partially downregulate FoxM1 expression under normoxia (Fig. 3a, Supplementary Fig. 4).
The expression of FoxM1 is transcriptionally regulated by both HIF-1α and HIF-2α in GIST cells. a qPCR analysis for the expression of FoxM1, HIF-1α and HIF-2α in GIST-T1 and GIST 882 cells, respectively, after silencing of HIF-1α or HIF-2α with shRNA under normoxia or hypoxia (1% O2) for 24 h. b ChIP assay was performed with antibody against HIF-1α, HIF-2α or control IgG in GIST-T1 cells exposed to normoxia or hypoxia for 24 h. The immunoprecipitated DNA was analyzed by PCR followed by agarose gel electrophoresis. Genomic DNA input was 1%. P: prime. c The HRE1-mutated FoxM1 promoter construct or the wild-type FoxM1 promoter were co-transfected with pCDNA3.1/HIF-1α or pCDNA3.1 and pRL-TK in GIST-T1 cells. After 24 h, dual-luciferase reporter assay was performed to detect the promoter activity. Relative luciferase activity was described in comparison with the control samples co-transfected with pCDNA3.1/HIF-1α and wild-type FoxM1 promoter. d Mutated FoxM1 promoter were co-transfected with pCDNA3.1-HIF-2α or pCDNA3.1 and pRL-TK in GIST-T1 cells. After 24 h, dual-luciferase reporter assay was performed to detect the promoter activity. Relative luciferase activity was described in comparison with the control samples co-transfected with pCDNA3.1/HIF-2α and wild-type FoxM1 promoter. *P < 0.05
Subsequently, we explored whether FoxM1 was a transcriptional target of HIF-1α and HIF-2α. Sequence analysis identified five putative HREs, located at − 25 (HRE1), − 297 (HRE2), − 309 (HRE3), − 761 (HRE4) and − 2324 (HRE5) base pairs (bp) relative to the transcriptional start site of FoxM1. Chromatin immunoprecipitation (ChIP) demonstrated (Fig. 3b) that HIF-1α directly interacted with the FoxM1 promoter between − 266 and 26 under hypoxia. In addition, HIF-2α bound the FoxM1 promoter at two distinct sites under normoxia, whereas under hypoxia, HIF-2α bound at three sites (Fig. 3b).
To determine which HRE was responsive to HIF-α-mediated transcriptional activation of FoxM1 promoter, we cloned the FoxM1 promoter into the luciferase reporter plasmid pGL4.0-Basic (WT), and we co-transfected this WT reporter together either with an expression plasmid of HIF-1α (Fig. 3c) or HIF-2α (Fig. 3d) in GIST-T1 cells. We found that FoxM1 promoter activity could be strongly induced by both HIF-1α and HIF-2α, indicating the presence of a functional HIF-α-binding site in FoxM1 promoter. Based on the ChIP results, we also performed HRE site-directed mutagenesis, transfected the mutant luciferase reporter construct (Mut) into the GIST-T1 cells and compared their activity with that of WT reporter plasmid. Luciferase assay showed that HIF-1α directly bound to HRE1 site, whereas HRE2, HRE3 and HRE5 were functional HIF-2α-binding site. Altogether, these data suggest that FoxM1 is a target gene of both HIF-1α and HIF-2α. HIF-2α may be an important mediator of FoxM1 expression under normoxia, whereas both HIF-1α and HIF-2α may be involved in the expression of FoxM1 under hypoxia.
Downregulation of FoxM1 inhibits GIST cell proliferation, migration and invasion under hypoxia
Given that the expression level of FoxM1 was strongly increased under hypoxia, we first examined the effects of FoxM1 suppression on proliferation, migration and invasion of GIST cells under hypoxia. As shown in Supplementary Fig. 6, the expression of FoxM1 in GIST-T1 and GIST 882 cells was knocked down using specific shRNA. And then, FoxM1 shRNA-treated GIST cells were cultured under hypoxic conditions. As shown in Fig. 4a, the CCK-8 assay showed that FoxM1 suppression in GIST-T1 and GIST 882 cell lines significantly inhibited cell proliferation. To determine whether the decrease in GIST cell proliferation was related to cell cycle regulation, we investigated the effects of FoxM1 suppression on cell cycle progression. Remarkably, FoxM1 shRNA-treated cells displayed cell cycle arrest (Fig. 4b, d). Cell migration and invasion were further evaluated using a wound-healing method and a Matrigel-coated Transwell system. After excluding the influence of cell proliferation, migration and matrigel invasion assays showed that FoxM1 suppression partially inhibited the migration and invasion of GIST-T1 and GIST 882 cells under hypoxia (Fig. 4c, d). Western blot exhibited that transfection of FoxM1 shRNA reduced the expression levels of the cyclin-dependent kinase 2 (CdK2), proliferating cell nuclear antigen (PCNA) and matrix metalloproteinase-2 (MMP-2) (Supplementary Fig. 5).
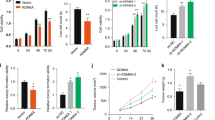
FoxM1 knockdown decreases proliferation, migration and invasion of GIST cells under hypoxia. a GIST cells were transfected with shFoxM1 or control, and cultured under hypoxia (1% O2) for up to 4 days. The growth rates were determined by CCK-8 assay at the indicated times. b shFoxM1- or shCon-transfected cells were cultured under hypoxia (1% O2) for 48 h, and cell cycle was analyzed by flow cytometry. c Wound-healing assays and transwell invasion assays were performed to examine the migration ability of GIST-T1 and GIST 882 cells after transfected with shFoxM1 or shCon under hypoxia. d Statistical analyses of cell cycle distribution, wound-healing assays and Matrigel invasion assays. *P < 0.05
Upregulation of FoxM1 promotes GIST cell proliferation, migration and invasion under normoxia
Given that GIST cells expressed FoxM1under normoxia, we also examined the effects of FoxM1 overexpression on proliferation, migration and invasion of GIST cells under normoxia. As shown in Supplementary Fig. 5, we ectopically expressed FoxM1 in GIST-T1 and GIST 882 cell lines. The CCK-8 assay and cell cycle analysis showed that FoxM1 overexpression markedly enhanced cell proliferation and cell cycle progression (Fig. 5a, b, d). Wound-healing- and transwell invasion assay further showed that FoxM1 overexpression markedly enhanced the mobility and invasiveness of GIST cells, compared with the control cells (Fig. 5c, d). Consistent with the above results, FoxM1 overexpression resulted in the increased expression of the CdK2, PCNA and MMP-2 (Supplementary Fig. 6). The above findings indicate that FoxM1 promotes the proliferation, migration and invasion of GIST cells in vitro.
FoxM1 overexpression increases proliferation, migration and invasion of GIST cells under normoxia. a GIST cells were transfected with pcFoxM1 or control, and cultured under normoxia (21% O2) for up to 4 days. The growth rates were determined by CCK-8 assay at the indicated times. b pcFoxM1- or pcEGFP-transfected cells were cultured for 48 h, and cell cycle was analyzed by flow cytometry. c Wound-healing assays and transwell invasion assays were performed to examine the migration ability of GIST-T1 and GIST 882 cells after transfected with pcFoxM1 or pcEGFP. d Statistical analyses of cell cycle distribution, wound-healing assays and Matrigel invasion assays. *P < 0.05
HIF-1α/ HIF-2α-FoxM1 axis promotes GIST xenograft growth
For in vivo tumor growth study, GIST-T1 was subcutaneously injected into athymic nude mice. Mice were randomized into four groups after implantation and administered with PT-2385 (PT, HIF-1α-specific inhibitor), PX-478 2HCl (PX, HIF-2α-specific inhibitor) or phosphate-buffered saline (PBS), or co-administered with PT + PX, respectively, in drinking water. As shown in Fig. 6a, all mice injected with GIST-T1 cells formed tumors after 2 weeks. The PT + PX-treated mice showed a significantly smaller tumor size than PT-, PX-, and PBS-treated cases (Fig. 6a, b). Immunohistochemistry demonstrated that all xenograft tumors were positive for CD117, DOG-1 and CD34 (Supplementary Fig. 7). Furthermore, the treatment of PT + PX caused significant downregulation of FoxM1 and Ki-67 expression (Fig. 6b, c).
Concomitant inhibition of HIF-1α and HIF-2α reduces the growth of human GIST tumor xenografts in nude mice. a At the end of the experiments, the mice were humanely killed, and the tumors were excised and weighed to determine the average tumor volume of the animals in this experiment. b Representative FoxM1 and Ki-67 expression in tissues from the same mice. c Statistical analyses of tumor volume, FoxM1 expression and Ki-67 expression. d Proposed HIF-1α/HIF-2α-FoxM1 axis in GIST progression. *P < 0.05
Discussion
In this study, we first found that the elevated expression of FoxM1 was correlated with cell proliferation, metastasis/invasion and a higher tumor stage by analyzing the clinical and pathological characteristics of GIST patients. Furthermore, ectopic expression of FoxM1 significantly promoted GIST cell proliferation, cell cycle progression, migration and invasion, whereas the knockdown of endogenous FoxM1 of hypoxic GIST cells had the opposite effects. These data were further confirmed in vivo where HIF-1α/HIF-2α-FoxM1 axis promoted GIST tumor growth by increasing cell proliferation using human GIST xenograft model. To study potential molecular mechanism of FoxM1 in contributing to GIST progression, we examined the effect of FoxM1 on the expression of several GIST progression-related proteins, such as PCNA [24], CdK2 [25] and MMP-2 [26]. Consistent with our expectations, FoxM1 overexpression upregulated the expression of these proteins in vitro. These findings indicate that FoxM1 enhances GIST progression by driving the proliferation, invasion and metastasis of GIST cells.
The modulation of FoxM1 by hypoxia had been reported by Xia et al. and they found that hypoxia upregulated FoxM1 expression through HIF-1α in HepG2, MCF-7 and Hela cells [27]. However, some studies reported that hypoxia induced FoxM1 in hypoxic lungs through HIF-2α [28, 29]. Nevertheless, the exact function of HIF-α in FoxM1 expression, and the molecular relationship among HIF-1α, HIF-2α and FoxM1 in GIST cells, have not been clearly elucidated. In this study, we found that the expression levels of FoxM1 were significantly increased under hypoxic conditions in GIST cells, along with the upregulation of both HIF-1α and HIF-2α. Specifically, we found that HIF-1α was only stabilized in GIST cells under hypoxic conditions, whereas HIF-2α was stabilized at a wider range of oxygen tensions (from normoxia to hypoxia), which is consistent with previous reports [30,31,32]. In accordance with the expression of HIF-2α, GIST cells also expressed FoxM1 under normoxia. In mRNA and protein expression assays, we showed that FoxM1 was transcriptionally regulated by HIF-2α under normoxia, whereas it was upregulated by both HIF-1α and HIF-2α under hypoxia. Using chromatin immunoprecipitation and luciferase reporter assays, we further identified that HIF-1α directly interacted with FoxM1 promoter under hypoxia, whereas HIF-2α bound the FoxM1 promoter under either normoxia or hypoxia. The above results indicate that HIF-1α cooperated with HIF-2α to regulate the transactivation activity of FoxM1, and their effects are depending on oxygen levels (Fig. 6d). The existence of such an interaction was further supported by the observation that the concomitant inhibition of HIF-1α and HIF-2α blocked FoxM1 expression and markedly inhibited GIST cell growth in vivo.
There were two limitations to this study. First, we identified mutations in c-kit in 75.9% and in PDGFRA in 6% of patients, which were lower than the 85–90% frequency of c-kit/PDGFRA mutations in GISTs reported by others [3, 33]. The variations in experimental protocols and sequencing methods may affect the mutational results [34]. In addition, during a mean follow-up period of 11.7 ± 7.6 months, no patients died of disease and only two patients had evidence of recurrence. Therefore, we did not perform prognostic analysis regarding whether the FoxM1 expression was associated with patient outcome because of the short follow-up period and the small number of recurrent cases, which was another limitation to the current study.
In summary, we first revealed that FoxM1 significantly promoted the proliferation, invasion and metastasis of GIST cells. It’s expression was tightly regulated by both HIF-1α and HIF-2α depending on oxygen levels. These findings suggest that HIF-1α/HIF-2α-FoxM1 axis may be a potential candidate in the prevention and treatment of GISTs.
Abbreviations
- CdK2:
-
Cyclin-dependent kinase 2
- ChIP:
-
Chromatin immunoprecipitation
- GIST:
-
Gastrointestinal stromal tumor
- HIF:
-
Hypoxia-inducible factor
- HRE:
-
Hypoxia response element
- MMP-2:
-
Matrix metalloproteinase-2
- PDGFRA:
-
Platelet-derived growth factor receptor alpha
- PCNA:
-
Proliferating cell nuclear antigen
- PBS:
-
Phosphate-buffered saline
- PT:
-
PT-2385
- PX:
-
PX-478 2HCl
- qPCR:
-
Quantitative polymerase chain reaction
- SD:
-
Standard error
- shRNA:
-
Short hairpin RNA
References
Hirota S, Ohashi A, Nishida T, Isozaki K, Kinoshita K, Shinomura Y, et al. Gain-of-function mutations of platelet-derived growth factor receptor alpha gene in gastrointestinal stromal tumors. Gastroenterology. 2003;125:660–7.
Rubin BP, Singer S, Tsao C, Duensing A, Lux ML, Ruiz R, et al. KIT activation is a ubiquitous feature of gastrointestinal stromal tumors. Cancer Res. 2001;61:8118–21.
Lasota J, Miettinen M. KIT and PDGFRA mutations in gastrointestinal stromal tumors (GISTs). Semin Diagn Pathol. 2006;23:91–102.
Antonescu CR, Besmer P, Guo T, Arkun K, Hom G, Koryotowski B, et al. Acquired resistance to imatinib in gastrointestinal stromal tumor occurs through secondary gene mutation. Clin Cancer Res. 2005;11:4182–90.
Heinrich MC, Corless CL, Blanke CD, Demetri GD, Joensuu H, Roberts PJ, et al. Molecular correlates of imatinib resistance in gastrointestinal stromal tumors. J Clin Oncol. 2006;24:4764–74.
Xie Z, Tan G, Ding M, Dong D, Chen T, Meng X, et al. Foxm1 transcription factor is required for maintenance of pluripotency of P19 embryonal carcinoma cells. Nucleic Acids Res. 2010;38:8027–38.
Teh MT, Gemenetzidis E, Chaplin T, Young BD, Philpott MP. Upregulation of FOXM1 induces genomic instability in human epidermal keratinocytes. Mol Cancer. 2010;9:45.
Koo CY, Muir KW, Lam EW. FOXM1: From cancer initiation to progression and treatment. Biochim Biophys Acta. 2012;1819:28–37.
Huang C, Du J, Xie K. FOXM1 and its oncogenic signaling in pancreatic cancer pathogenesis. Biochim Biophys Acta. 2014;1845:104–16.
Cai Y, Balli D, Ustiyan V, Fulford L, Hiller A, Misetic V, et al. Foxm1 expression in prostate epithelial cells is essential for prostate carcinogenesis. J Biol Chem. 2013;288:22527–41.
Gemenetzidis E, Bose A, Riaz AM, Chaplin T, Young BD, Ali M, et al. FOXM1 upregulation is an early event in human squamous cell carcinoma and it is enhanced by nicotine during malignant transformation. PLoS One. 2009;4:e4849.
Uddin S, Ahmed M, Hussain A, Abubaker J, Al-Sanea N, AbdulJabbar A, et al. Genome-wide expression analysis of Middle Eastern colorectal cancer reveals FOXM1 as a novel target for cancer therapy. Am J Pathol. 2011;178:537–47.
Li Q, Zhang N, Jia Z, Le X, Dai B, Wei D, et al. Critical role and regulation of transcription factor FoxM1 in human gastric cancer angiogenesis and progression. Cancer Res. 2009;69:3501–9.
Wang Z, Banerjee S, Kong D, Li Y, Sarkar FH. Down-regulation of Forkhead Box M1 transcription factor leads to the inhibition of invasion and angiogenesis of pancreatic cancer cells. Cancer Res. 2007;67:8293–300.
Huang C, Qiu Z, Wang L, Peng Z, Jia Z, Logsdon CD, et al. A novel FoxM1-caveolin signaling pathway promotes pancreatic cancer invasion and metastasis. Cancer Res. 2012;72:655–65.
Li D, Wei P, Peng Z, Huang C, Tang H, Jia Z, et al. The critical role of dysregulated FOXM1-PLAUR signaling in human colon cancer progression and metastasis. Clin Cancer Res. 2013;19:62–72.
Xia L, Huang W, Tian D, Zhu H, Zhang Y, Hu H, et al. Upregulated FoxM1 expression induced by hepatitis B virus X protein promotes tumor metastasis and indicates poor prognosis in hepatitis B virus-related hepatocellular carcinoma. J Hepatol. 2012;57:600–12.
Keith B, Johnson RS, Simon MC. HIF1α and HIF2α: sibling rivalry in hypoxic tumour growth and progression. Nat Rev Cancer. 2011;12:9–22.
Martin SK, Diamond P, Gronthos S, Peet DJ, Zannettino AC. The emerging role of hypoxia, HIF-1 and HIF-2 in multiple myeloma. Leukemia. 2011;25:1533–42.
Rouault-Pierre K, Hamilton A, Bonnet D. Effect of hypoxia-inducible factors in normal and leukemic stem cell regulation and their potential therapeutic impact. Expert Opin Biol Ther. 2016;16:463–76.
Takahashi R, Tanaka S, Hiyama T, Ito M, Kitadai Y, Sumii M, et al. Hypoxia-inducible factor-1alpha expression and angiogenesis in gastrointestinal stromal tumor of the stomach. Oncol Rep. 2003;10:797–802.
Chen WT, Huang CJ, Wu MT, Yang SF, Su YC, Chai CY. Hypoxia-inducible factor-1alpha is associated with risk of aggressive behavior and tumor angiogenesis in gastrointestinal stromal tumor. Jpn J Clin Oncol. 2005;35:207–13.
Pollard PJ, El-Bahrawy M, Poulsom R, Elia G, Killick P, Kelly G, et al. Expression of HIF-1alpha, HIF-2alpha (EPAS1), and their target genes in paraganglioma and pheochromocytoma with VHL and SDH mutations. J Clin Endocrinol Metab. 2006;91:4593–8.
Ray R, Tahan SR, Andrews C, Goldman H. Stromal tumors of the stomach: prognostic value of the PCNA index. Mod Pathol. 1994;7:26–30.
Nemoto Y, Mikami T, Hana K, Kikuchi S, Kobayashi N, Watanabe M, et al. Correlation of enhanced cell turnover with prognosis of gastrointestinal stromal tumors of the stomach: relevance of cellularity and p27kip1. Pathol Int. 2006;56:724–31.
Sun B, Qie S, Zhang S, Sun T, Zhao X, Gao S, et al. Role and mechanism of vasculogenic mimicry in gastrointestinal stromal tumors. Hum Pathol. 2008;39:444–51.
Xia LM, Huang WJ, Wang B, Liu M, Zhang Q, Yan W, et al. Transcriptional up-regulation of FoxM1 in response to hypoxia is mediated by HIF-1. J Cell Biochem. 2009;106:247–56.
Torres-Capelli M, Marsboom G, Li QO, Tello D, Rodriguez FM, Alonso T, et al. Role Of Hif2α Oxygen Sensing Pathway In Bronchial Epithelial Club Cell Proliferation. Sci Rep. 2016;6:25357.
Raghavan A, Zhou G, Zhou Q, Ibe JC, Ramchandran R, Yang Q, et al. Hypoxia-induced pulmonary arterial smooth muscle cell proliferation is controlled by forkhead box M1. Am J Respir Cell Mol Biol. 2012;46:431–6.
Nilsson H, Jögi A, Beckman S, Harris AL, Poellinger L, Påhlman S. HIF-2alpha expression in human fetal paraganglia and neuroblastoma: relation to sympathetic differentiation, glucose deficiency, and hypoxia. Exp Cell Res. 2005;303:447–56.
Holmquist-Mengelbier L, Fredlund E, Löfstedt T, Noguera R, Navarro S, Nilsson H, et al. Recruitment of HIF-1alpha and HIF-2alpha to common target genes is differentially regulated in neuroblastoma: HIF-2alpha promotes an aggressive phenotype. Cancer Cell. 2006;10:413–23.
Li Z, Bao S, Wu Q, Wang H, Eyler C, Sathornsumetee S, et al. Hypoxia-inducible factors regulate tumorigenic capacity of glioma stem cells. Cancer Cell. 2009;15: 501–13.
Lasota J, Miettinen M. Clinical significance of oncogenic KIT and PDGFRA mutations in gastrointestinal stromal tumours. Histopathology. 2008;53:245–66.
Hostein I, Debiec-Rychter M, Olschwang S, Bringuier PP, Toffolati L, Gonzalez D, et al. A quality control program for mutation detection in KIT and PDGFRA in gastrointestinal stromal tumours. J Gastroenterol. 2011;46:586–94.
Acknowledgements
This work was supported by National Natural Science Foundation of China (Grants 81472276) and Shanghai Municipal Commission of Health and Family planning, Key Developing Disciplines (No. 2015ZB0202).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1964 and later versions. Informed consent was obtained from all patients for the use of their samples.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bai, C., Liu, X., Qiu, C. et al. FoxM1 is regulated by both HIF-1α and HIF-2α and contributes to gastrointestinal stromal tumor progression. Gastric Cancer 22, 91–103 (2019). https://doi.org/10.1007/s10120-018-0846-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-018-0846-6