Abstract
This study aimed to assess the performance of our in-house method for rapid direct bacterial identification (ID) and antimicrobial susceptibility testing (AST) using a positive blood culture (BC) broth. For Gram-negative bacteria, 4 mL of BC broth was aspirated and passed through a Sartorius Minisart syringe filter with a pore size of 5 µm. The filtrate was then centrifuged and washed. A small volume of the pellet was used for ID, using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry, and for AST, using automated broth microdilution. For Gram-positive cocci, 4 mL of BC broth was passed through the Minisart syringe filter. Then, 4 mL of sterile distilled water was injected in the direction opposite to that of the filtration to collect the bacterial residue trapped in the filter. Compared with the conventional method performed with pure colonies on agar plates, 94.0% (234/249) were correctly identified using the in-house method, with rates of 91.4% (127/139) and 97.3% (107/110) for Gram-positive and Gram-negative isolates, respectively. Of 234 correctly identified isolates, 230 were assessed by AST. Categorical agreement and essential agreement were 93.3% and 94.5%, respectively, with a minor error rate of 3.8%, a major error rate of 3.4%, and a very major error rate of 1.6%. Our in-house preparation method showed good performance in rapid direct ID and AST using positive BC broths compared to the conventional method. This simple method can shorten the conventional turnaround time for ID and AST by at least 1 day, potentially contributing to better patient management.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bloodstream infections (BSI) are a leading cause of morbidity and mortality worldwide [1]. Rapid identification of an infectious pathogen and accelerated determination of its antimicrobial susceptibility are key components of treating bacterial infection [2]. Conventional blood culture (BC) is the standard method for diagnosing and guiding the treatment of patients with BSI. However, the conventional method requires at least a few days because an aliquot of positive BC must be sub-cultured on solid agar plates and then subjected to a series of biochemical tests [3].
Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) is a standard method used in clinical microbiology laboratories to rapidly identify bacteria in fresh colonies [2]. Positive BCs require at least one additional day to produce pure bacterial colonies for subsequent testing and procedures, in contrast to other clinical samples [3]. Efficient antimicrobial stewardship programs and patient management depend on the rapid identification of bacterial pathogens in blood cultures and subsequent antimicrobial susceptibility testing (AST) [4, 5]. Multidrug-resistant infections increase the length of hospital stay, and mortality can result from inappropriate or suboptimal antibiotic use [6]. Therefore, it is crucial to reduce the time required for identification and AST in clinical microbiology laboratories [7]. Hence, various preparation methods have been developed to purify bacterial biomass obtained directly from a positive BC broth, including serum separator tubes [8,9,10] and chemical reagents such as saponin [11, 12], sodium dodecyl sulfate [13], ammonium chloride [14], and Triton-X [15]. In addition, molecular methods can accelerate microbial identification and predict antimicrobial susceptibility using direct blood samples or positive blood cultures [16]. In our previous study, we developed an in-house preparation method for direct bacterial identification (ID) using a positive BC broth [17], which involved using a sterile filter for Gram-negative bacteria and serial washing for Gram-positive cocci. Compared with the microfiltration method, which takes approximately 15 min, the serial washing method is time-consuming (40 min) and labor-intensive. Therefore, we established a modified procedure for Gram-positive cocci, employing a sterile filter that reduces the sample processing time from 40 to 15 min. The objective of this study was to assess the performance of our modified method for rapid direct ID and AST using positive BC broth.
Materials and methods
Test samples
This study was conducted between December 2020 and February 2021 at Korea University Anam Hospital. BD BACTEC Plus aerobic/F and anaerobic/F bottles were used to collect blood specimens, which were then incubated in a BACTEC Plus system (Becton Dickinson, Franklin Lakes, NJ, USA). Direct ID and AST were performed concurrently using conventional methods. BCs showing mixed morphotypes or yeast upon Gram staining were excluded from the study.
Sample preparation for conventional ID and AST
A small volume of BC fluid was sub-cultured on appropriate solid media and used for Gram staining. After overnight incubation, fresh colonies were transferred onto the sample spots on a steel target slide. After the spots were dried, 1.0 μL matrix solution was added and air-dried at room temperature. The target plate was identified by MALDI-TOF MS using a Microflex LT system (Bruker Daltonics, Bremen, Germany). The manufacturer’s suggested criteria were used to interpret identification scores: a score of > 2.0 indicated species-level identification, a score of ≥ 1.7 indicated genus identification, and a score of < 1.7 indicated unreliable identification.
AST was performed using the MicroScan WalkAway system (Beckman Coulter, Sacramento, CA, USA) or VITEK 2 system (bioMérieux, Marcy-l’Etoile, France) depending on the species to be identified. For Gram-negative bacilli (NM44 panel) and Gram-positive cocci (PM28 panel), except Streptococcus spp., the MicroScan system was used with the Prompt inoculation system and RENOK rehydrating inoculator (Beckman Coulter). The Prompt inoculation system utilizes a unique wand to standardize the inocula without turbidity measurements. For AST of Streptococcus spp., a colony suspension of 0.5 McFarland turbidity was used with the VITEK 2 ST03 panel.
Sample preparation for rapid ID and AST
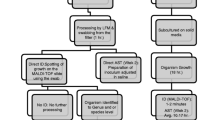
For Gram-negative bacteria, 4 mL was aspirated from a positive BC broth, filtered through a Minisart syringe filter with a pore size of 5 µm (Sartorius, Göttingen, Germany), and transferred into two 2-mL tubes. The filtrate was centrifuged at 13,000 rpm for 2 min. After decanting the supernatants, the pellets were washed twice with 1 mL sterile distilled water (DW) and centrifuged at 13,000 rpm for 2 min. An appropriate amount of the pellet from one tube was applied to a steel target and allowed to air-dry. The matrix solution (1.0 μL) was added to the sample and air-dried at room temperature. This sample was used for the mass spectrometric analysis. In addition to bacterial proteins, pellets obtained directly from positive BC broths contain blood proteins, which leads to background noise in MALDI-TOF MS [18]. Hence, cutoff scores of 1.7 and 1.5 were regarded as reliable for identifying species and genus, respectively, following previous research [19, 20]. A score < 1.5 was considered as unreliable identification [21]. For AST, a small volume of the pellet in the second tube was used in the MicroScan system. The sample processing time was ~ 15 min.
For Gram-positive cocci, 4 mL of BC broth was passed through the Minisart syringe filter membrane with a pore size of 5 µm (Sartorius). However, because of clustering, Gram-positive cocci could not be filtered; they adhered to the filter membrane as a residue. Therefore, 4 mL of sterile DW was injected in the direction opposite to that of the filtration to collect the bacterial residue trapped in the filter. The residue (in DW) was centrifuged at 13,000 rpm for 2 min. After decanting the supernatant, the pellet in each tube was washed twice, as described above. Each pellet was identified by MALDI-TOF MS and AST using the MicroScan or VITEK 2 system, depending on the genus type, as described above. The sampling processing time was approximately 15 min, much shorter than the 40 min required for bead-beating and serial washing, which we previously used for Gram-positive cocci [17].
Addition of isolates with known resistant mechanisms
The inclusion of isolates with known resistance mechanisms is important in the current study. Since testing prospective isolates obtained during the study period may result in an underrepresentation of key resistant strains, we additionally included 26 isolates of frozen stocks from clinical samples for the analysis. Four isolates of carbapenemase-producing Klebsiella pneumoniae, five isolates of multidrug-resistant (MDR) Pseudomonas aeruginosa, four isolates of MDR Acinetobacter baumannii, one isolate of carbapenemase-resistant (CR) A. baumannii, five isolates of vancomycin-resistant Enterococcus faecium, and seven isolates of methicillin-resistant Staphylococcus aureus (MRSA) were included. The bacteria used for contrived samples represented different clinical isolates, not clonal isolates from the same organism from the same patient.
Isolates that were non-susceptible to at least one agent in ≥ 3 antimicrobial categories were defined as MDR [21]. Carbapenem-resistant Enterobacterales (CRE) were classified as isolates resistant to any carbapenem. All CRE isolates were tested for detecting and differentiating the five main carbapenemase families (NDM, IMP, VIM, OXA-48-like, and KPC) using the NG Test CARBA-5 immunochromatographic assay (NG Biotech, Guipry, France). The production of extended-spectrum beta-lactamase (ESBL) was identified by interpretation of the antibiogram obtained by the MicroScan NM44 panel according to the CLSI guidelines [22]. ESBL-producing enterbacterales were defined by ≥ 3 twofold decrease in the minimum inhibitory concentration (MIC) for either antimicrobial agent (ceftazidime or cefotaxime) tested in combination with clavulanate vs. the MIC of the agent when tested alone (e.g., ceftazidime MIC = 8 μg/mL; ceftazidime-clavulanate MIC = 1 μg/mL). The MICs of ceftazidime-clavulanate and cefotaxime-clavulanate were used only for ESBL diagnosis and were not included in the AST agreement results between the conventional and rapid methods.
Simulated blood cultures were performed using 26 contrived samples stored at − 80 °C. The samples were thawed and cultured overnight on blood agar plates at 37 °C. The resulting fresh bacterial colonies were suspended in phosphate-buffered saline to achieve a turbidity of 0.5 McFarland standard. This contained approximately 1.5 × 108 colony-forming units (CFU)/mL and then serially diluted to a final 1 × 104 CFU/mL. Then, 0.1 mL of this bacterial suspension (1000 CFU) was combined with 5 mL of sterile human blood to be discarded and inoculated into a BD BACTEC Plus aerobic/F bottle [23]. These contrived samples were further processed in the same method described above for the conventional and rapid methods.
Comparison of conventional and rapid methods
In this study, the conventional MALDI-TOF MS identification method was used as the reference method, and all direct identification results were compared to those obtained using this approach. Rapid identification results were categorized as correct at the species level (same species with score of ≥ 1.7), correct at the genus level (same species with score of < 1.7 but ≥ 1.5), unreliable (score < 1.5), or misidentified (different species with score of ≥ 1.7), in accordance with the reference method.
To compare the AST results of the direct and conventional methods, MIC was categorized as susceptible, intermediate, or resistant, following the CLSI guidelines [22]. Accuracy of the direct AST is measured in categorical agreement (CA) and essential agreement (EA), essential agreement of evaluable results (EAER) [24, 25]. CA is the proportion of isolates that yielded the same susceptibility category (i.e., susceptible, intermediate, or resistant) by the conventional method and the direct method. EA is the percentage of isolates with a MIC value within a single doubling dilution of the corresponding conventional method. Acceptance criteria for CA and EA should be ≥ 90% [24, 26]. EAER is the proportion of isolates with on-scale MIC results within a 1log2 dilution for the direct and conventional methods. CA discrepancies are classified into three types of error. Minor error (mE) indicated that the result was resistant or susceptible by the conventional method and intermediate by the direct method or vice versa. Major error (ME) indicated that the isolate was categorized susceptible by the conventional method and resistant by the direct method (false resistance). Very major error (VME) indicated that the conventional method result was resistant and the direct method result was susceptible (false susceptibility). For mE error rate calculations, the total number of tested isolates were used as the denominator. For ME, the number of susceptible isolates were used as the denominator. VME was calculated using the number resistant isolates as the denominator. ME and VME rates that are acceptable are typically < 3% of susceptible and resistant isolates tested, respectively [24, 26].
Results
Comparison of conventional and rapid ID results
A total of 249 nonduplicate monomicrobial positive BCs, including 26 contrived samples—139 Gram-positive isolates and 110 Gram-negative isolates—were used in this study (Table 1). Compared to the conventional method, 94.0% (234/249) were correctly identified at the species level using the rapid direct method, with rates of 91.4% (127/139) and 97.3% (107/110) for Gram-positive and Gram-negative isolates, respectively. The unreliable identification rate was greater for Gram-positive bacteria (6.5%, 9/139) than that for Gram-negative bacteria (0%, 0/110). The misidentification rates for Gram-positive and Gram-negative bacteria were 0% (0/139) and 2.7% (3/110), respectively.
Comparison of conventional and rapid AST results
Of the 234 correctly identified isolates, four Bacteroides spp. were excluded, and the remaining 230 (127 Gram-positive and 103 Gram-negative) were assessed for antimicrobial susceptibility (Table 2). In total, 4052 isolate/antimicrobial agent combinations were analyzed. The CA, EA, and EAER between the direct and reference methods were 93.3% (3782/4052), 94.5% (3752/3972), and 91.8% (270/294), respectively, with an mE rate of 3.8% (156/4052), an ME rate of 3.4% (88/2350), and a VME rate of 1.6% (26/1582). CA, EA, and EAER among the Gram-positive isolates were 93.3% (1703/1826), 94.1% (1643/1746), and 91.0% (181/199), respectively. The mE, ME, and VME rates were 4.0% (73/1826), 3.1% (31/1007), and 2.4% (19/785), respectively. The Gram-negative isolates showed a CA of 93.4% (2079/2226), EA of 94.7% (2109/2226), and EAER of 93.7% (89/95), with an mE rate of 3.7% (83/2226), ME rate of 4.2% (57/1343), and VME rate of 0.9% (7/797).
The overall AST agreement for S. aureus was high (> 90%) for both CA and EA (Supplementary Table S1). Fifteen isolates of MRSA were successfully detected based on cefoxitin screening and oxacillin agreement. For staphylococci other than S. aureus (SOSA), AST agreement was high for most antibiotics except gentamicin, erythromycin, and trimethoprim-sulfamethoxazole (Supplementary Table S2 and S3). Among the six isolates of Streptococcus spp., only two mEs were found: benzylpenicillin in S. salivarius, and tetracycline in S. dysgalactiae (Supplementary Table S4). The AST agreement in Enterococcus spp. was high for most antibiotics except teicoplanin and rifampin due to the high numbers of mE (12.5% and 34.4%, respectively) (Supplementary Table S5). The rapid AST method successfully detected 16 isolates of vancomycin-resistant enterococci (VRE) by vancomycin agreement results. For E. coli, the CA/EA/EAER were 92.0%, 93.5%, and 90.0%, respectively (Supplementary Table S6). The ME was high (> 3%) for E. coli, mainly due to the large number of ME for cefoxitin (14.6%) and cefotaxime (13.3%). AST agreement rates in K. pneumoniae were high in CA (93.5%), EA (94.7%), and EAER (94.7%), with no VME (0%) (Supplementary Table S7). Enterobacterales other than E. coli and K. pneumoniae showed good AST agreement (Supplementary Table S8). Of the 12 Acinetobacter spp., 11 were A. baumannii, seven were MDR A. baumannii, and four were CR A. baumannii. The AST agreement for Acinetobacter spp. was 96.1%, 96.1%, and 100% for CA/EA/EAER (Supplementary Table S9). For all the antimicrobial agents tested, the correlation was high (> 91%), except tetracycline, for which the agreement was 75% due to a high mE. In the case of P. aeruginosa, the AST agreement was very high (greater than 97.7%) without ME or VME (Supplementary Table S10). Other non-Enterobacterales showed a high AST agreement between the conventional and rapid methods (Supplementary Table S11).
Discussion
While the conventional method for bacterial ID and AST takes 48–96 h from the time a blood culture is flagged as positive [27], various direct testing methods provide results within 6–24 h [22]. The clinical benefits of these rapid methods include a shortened time to optimal therapy and improved antibiotic stewardship in patients with sepsis [28]. In addition, rapid diagnostic testing was associated with significant reductions in mortality risk and length of hospital stay in the presence of an antimicrobial stewardship program [29]. This study describes a simple, straightforward, and cheap method to obtain a rapid ID and AST directly from positive blood cultures. The modified in-house method provided reliable ID and AST results for bacteria causing BSI.
The correct identification rate at the species level in earlier reports varied between investigations (90–100% for Gram-negative isolates and 73–92% for Gram-positive isolates) [2, 8,9,10,11,12,13,14,15, 17]. This variation is most likely the result of differences in preparation techniques, proportions of different bacteria recovered, and criteria defined for correct identification [30]. However, direct identification of Gram-positive bacteria is less accurate than that of Gram-negative bacteria, regardless of the preparation method used. In our study, the correct identification rates of Gram-positive and Gram-negative bacteria were 91.4% and 97.3%, respectively. This trend was also observed in previous reports using commercially available and validated kits such as Sepsityper Kit (Bruker Daltonics GmbH, Bremen, Germany) [19, 31, 32]. In our study, the relatively low identification rate for Gram-positive isolates was mainly due to SOSA, such as Staphylococcus epidermidis (40/46, 87.0%). SOSA are generally recognized as contaminants when recovered from BC broth. The accuracy of our method was improved when these strains were excluded. The correct species identification rates for Staphylococcus aureus, Enterococcus spp., and Streptococcus spp. reached a maximum of 94.7% (54/57). Although the reason for the relatively poor identification of Gram-positive cocci is not apparent, several possible reasons have been proposed [33]. First, owing to their thick cell walls, less protein is extracted from Gram-positive bacteria than that from Gram-negative bacteria. Hence, higher amounts of Gram-positive bacteria may be necessary relative to the amount of Gram-negative bacteria needed. Second, misidentification is possible because some Gram-positive cocci share highly similar protein structures. Regardless of the cause, the results of the direct approach should be carefully scrutinized if Gram-positive cocci are detected in BC bottles.
Among the Gram-negative bacilli, one Acinetobacter nosocomialis isolate was misidentified as Acinetobacter baumannii, and one A. baumannii isolate was misidentified as A. nosocomialis. Because the clinical outcomes and antibiotic sensitivity of members of closely related and clinically important Acinetobacter spp. vary [34], their direct identification warrants careful evaluation. Otherwise, inadequate clinical care and inappropriate antibiotic use are unavoidable. In addition, one Klebsiella pneumoniae isolate was misidentified as Klebsiella variicola, which corroborated the difficulty in differentiating between the two species using protein spectra [35]. Consequently, at present, direct ID and MALDI-TOF MS employing any preparation procedure cannot completely replace conventional techniques for bacterial identification.
For AST, the overall CA and EA values between the rapid and conventional methods were high (93.3% and 94.5%, respectively). Generally, the acceptable rates for CA and EA are ≥ 90% [24, 26]. The CA/EA for Gram-positive and Gram-negative isolates were 93.3%/94.1% and 93.4%/94.7%, respectively, indicating that performance of the rapid method was acceptable. The differences in sample processing techniques, commercial AST panels, and antibacterial agents make it challenging to compare our results with those of other recent investigations. However, the agreement was comparable to those of earlier studies [3, 23, 32, 36].
Resistance to antimicrobials is a primary global health concern. MRSA is associated with worse clinical outcomes than those produced by methicillin-sensitive S. aureus [37]. Accurate and timely laboratory diagnosis and susceptibility testing are critical for treating, controlling, and preventing MRSA infections [38]. Our study showed that 100% CA/EA agreement was observed with oxacillin, suggesting that our method is suitable for rapidly detecting MRSA. VRE is a major cause of healthcare-associated infections, especially in patients with weakened immune systems. VRE infection results in higher mortality and extended hospital stays [39]. In our study, Enterococcus spp. isolates showed 100% agreement with vancomycin, indicating that our method effectively identifies VRE. Additionally, ESBL and carbapenemases contribute to a significant burden of morbidity and mortality globally, and their prompt identification from positive blood cultures is critical for the timely administration of effective antimicrobial therapy for BSI caused by Enterobacterales [40]. In our study, among 22 ESBL-producing Enterobacterales (Supplementary Table S6, S7, and S8), 95.5% (21/22) of ESBL was correctly identified by the direct method based on the MicroScan NM44 panel antibiogram (data not shown); one ESBL E. coli isolate was identified as a CR strain. In addition, all five carbapenemase-producing Klebsiella pneumoniae strains (all KPC types) were correctly detected, given that 100% CA/EA agreements with carbapenem antimicrobial agents were accomplished (Supplementary Table S7), which was confirmed by the NG-test CARBA-5 for carbapenemase production. Among the 77 Enterobacterales, only one VME was found for imipenem susceptibility in one E. coli isolate (Supplementary Table S6, S7, and S8). This false susceptibility may result in the possible misdiagnosis of carbapenemase-producing Enterobacterales, resulting in the administration of inappropriate antibiotics.
A. baumannii and P. aeruginosa are innately resistant to many antibiotics and, more crucially, are capable of developing resistance to almost any antimicrobial treatment [41]. These organisms are nosocomial pathogens of clinical concern and are of public health significance, mainly because of their exceptional adaptability to the clinical setting [42, 43]. Treatment complications, adverse effects on clinical outcomes, and increased treatment costs are associated with nosocomial isolates of MDR A. baumannii and P. aeruginosa [44]. MDR A. baumannii and P. aeruginosa can spread easily from patient to patient on healthcare personnel's hands in the hospital setting [45]. Hence, accurate and rapid identification of MDR A. baumannii and P. aeruginosa is crucial for proper patient management and appropriate infection control in the clinical setting. Given that no VME occurred for either A. baumannii or P. aeruginosa in our study (Supplementary Table S9 and 10), the direct method successfully identified all MDR and CR strains. Overall, our preparation method resulted in good agreement between direct and conventional AST; therefore, the adoption of the direct preparation method in a clinical setting could improve patient management. However, it is advisable to correlate direct AST results with conventional AST because of the potential for false susceptibility, as observed in the current and previous studies [3, 46].
VME (false susceptibility) are the greatest concern, as they may result in the administration of inappropriate antibiotics, which can be fatal, particularly in critically ill patients where these antibiotics are commonly used as first-line therapy [47]. The data obtained in this study included 1.6% VME, which was below the acceptable criteria (< 3%). However, the overall ME rate of 3.4% of our in-house filter method was beyond the CLSI criteria (> 3%). Discordant AST results between the direct and conventional methods might result from differences in inoculum size. We used the Prompt inoculation Wand (Beckman Coulter) to transfer bacterial biomass to MicroScan AST cards in this study, except for Streptococcus spp. Compared to the three colonies on solid agar plates picked up by the Prompt inoculation Wand and using the conventional AST method, for which the manufacturer’s instructions were followed, “appropriate amounts” of bacterial sediment were picked up by the wand and utilized for the direct AST method. Hence, it is possible that the initial inoculum density of the bacterial isolates may differ between the two methods. The inoculum effect is the phenomenon of diminished antibacterial activity at inocula higher than those used for susceptibility testing [48]. In our study, the direct AST method showed the tendency to read high, in comparison to the reference method, based on higher ME than VME (Table 2). The inoculum effect can compromise the accuracy of technologies that do not use a uniform inoculum size because even small variations in the inoculum can have a significant impact on the determined MIC [49, 50]. Previous study demonstrated that the inoculum effect was associated with beta-lactam/beta-lactamase inhibitor combinations against E. coli [48], which was corroborated by our findings of high ME for beta-lactam antibiotics in E. coli (Supplementary Table S6). ME may prevent the patients from receiving a proper antibiotic and result in the use of more expensive second-line antibiotics, contributing to higher healthcare costs [47]. Therefore, it is advised to apply the same inoculum density for our direct AST method as for the conventional AST by measuring McFarland turbidity of bacterial suspension in future studies or in a real clinical setting. Other confounding factors of our direct preparation method, such as the presence of red blood cells, cellular debris, constituents of BC media, and technical errors, may also have contributed to the AST discrepancy.
This study had several limitations. First, given the small sample size drawn from a single clinical microbiology laboratory, only a restricted range of bacterial species was examined. Another limitation of this study was the omission of polymicrobial bacteremia. Therefore, direct ID and AST are only practical for blood specimens considered monomicrobial and free of the skin or mucosal contaminants. Third, the discrepant identification of closely related species was not resolved using a more accurate approach, such as whole-genome sequencing. In addition, although we used the term “rapid” for this study, we did not measure the turnaround time for the filter method compared to the conventional method. Thus, future studies are necessary to evaluate the turnaround time of the filter method for ID and AST. Lastly, as previously stated, a high ME could be related to the difference in inoculum density for AST between the conventional and direct methods. Nevertheless, the present study showed that the in-house filter method is promising, given that it was able to provide > 90% CA and EA with the conventional method on the clinical samples. Agreement rates could have been higher with the same inoculum density between the methods.
Conclusion
Our method has the major benefit of providing accurate results for ID and AST 1 day earlier than the conventional ID/AST methods, which are considered the gold standard. Our in-house method is straightforward, quick, and affordable for rapid direct ID and AST using positive BC broths. Additional studies are required to further assess its potential role in clinical management and antimicrobial stewardship program.
Data availability
In the article or as supplementary information, all data pertinent to the study are included.
Code availability
Not applicable.
References
McNamara JF, Righi E, Wright H, Hartel GF, Harris PNA, Paterson DL (2018) Long-term morbidity and mortality following bloodstream infection: a systematic literature review. J Infect 77:1–8. https://doi.org/10.1016/j.jinf.2018.03.005
Prod’hom G, Bizzini A, Durussel C, Bille J, Greub G (2010) Matrix-assisted laser desorption ionization-time of flight mass spectrometry for direct bacterial identification from positive blood culture pellets. J Clin Microbiol 48:1481–1483. https://doi.org/10.1128/JCM.01780-09
Infante A, Ortiz de la Tabla V, Martín C, Gázquez G, Buñuel F (2021) Rapid identification and antimicrobial susceptibility testing of Gram-negative rod on positive blood cultures using MicroScan panels. Eur J Clin Microbiol Infect Dis 40:151–157. https://doi.org/10.1007/s10096-020-04014-3
Juttukonda LJ, Katz S, Gillon J, Schmitz J, Banerjee R (2020) Impact of a rapid blood culture diagnostic test in a children’s hospital depends on Gram-positive versus Gram-negative organism and day versus night shift. J Clin Microbiol 58:e01400-e1419. https://doi.org/10.1128/JCM.01400-19
Banerjee R, Komarow L, Virk A, Rajapakse N, Schuetz AN, Dylla B, Earley M, Lok J, Kohner P, Ihde S, Cole N, Hines L, Reed K, Garner OB, Chandrasekaran S, de St MA, Kanatani M, Curello J, Arias R, Swearingen W, Doernberg SB, Patel R (2021) Randomized trial evaluating clinical impact of RAPid IDentification and susceptibility testing for Gram-negative bacteremia: RAPIDS-GN. Clin Infect Dis 73:e39–e46. https://doi.org/10.1093/cid/ciaa528
Campion M, Scully G (2018) Antibiotic use in the intensive care unit: optimization and de-escalation. J Intensive Care Med 33:647–655. https://doi.org/10.1177/0885066618762747
Munson EL, Diekema DJ, Beekmann SE, Chapin KC, Doern GV (2003) Detection and treatment of bloodstream infection: laboratory reporting and antimicrobial management. J Clin Microbiol 41:495–497. https://doi.org/10.1128/JCM.41.1.495-497.2003
Barnini S, Ghelardi E, Brucculeri V, Morici P, Lupetti A (2015) Rapid and reliable identification of Gram-negative bacteria and Gram-positive cocci by deposition of bacteria harvested from blood cultures onto the MALDI-TOF plate. BMC Microbiol 15:124. https://doi.org/10.1186/s12866-015-0459-8
Azrad M, Keness Y, Nitzan O, Pastukh N, Tkhawkho L, Freidus V, Peretz A (2019) Cheap and rapid in-house method for direct identification of positive blood cultures by MALDI-TOF MS technology. BMC Infect Dis 19:72. https://doi.org/10.1186/s12879-019-3709-9
ZenginCanalp H, Bayraktar B (2021) Direct rapid identification from positive blood cultures by MALDI-TOF MS: specific focus on turnaround times. Microbiol Spectr 9:e0110321. https://doi.org/10.1128/spectrum.01103-21
Jakovljev A, Bergh K (2015) Development of a rapid and simplified protocol for direct bacterial identification from positive blood cultures by using matrix-assisted laser desorption ionization time-of-flight mass spectrometry. BMC Microbiol 15:258. https://doi.org/10.1186/s12866-015-0594-2
Huang YL, Sun QL, Li JP, Hu YY, Zhou HW, Zhang R (2019) Evaluation of an in-house MALDI-TOF MS rapid diagnostic method for direct identification of micro-organisms from blood cultures. J Med Microbiol 68:41–47. https://doi.org/10.1099/jmm.0.000866
López-Pintor JM, Navarro-San Francisco C, Sánchez-López J, García-Caballero A, Fernández L, de Bobadilla E, Morosini MI, Cantón R (2019) Direct antimicrobial susceptibility testing from the blood culture pellet obtained for MALDI-TOF identification of Enterobacterales and Pseudomonas aeruginosa. Eur J Clin Microbiol Infect Dis 38:1095–1104. https://doi.org/10.1007/s10096-019-03498-y
Sakarikou C, Altieri A, Bossa MC, Minelli S, Dolfa C, Piperno M, Favalli C (2018) Rapid and cost-effective identification and antimicrobial susceptibility testing in patients with Gram-negative bacteremia directly from blood-culture fluid. J Microbiol Methods 146:7–12. https://doi.org/10.1016/j.mimet.2018.01.012
Simon L, Ughetto E, Gaudart A, Degand N, Lotte R, Ruimy R (2019) Direct identification of 80 percent of bacteria from blood culture bottles by matrix-assisted laser desorption ionization-time of flight mass spectrometry using a 10-minute extraction protocol. J Clin Microbiol 57:e01278-e1318. https://doi.org/10.1128/JCM.01278-18
Peker N, Couto N, Sinha B, Rossen JW (2018) Diagnosis of bloodstream infections from positive blood cultures and directly from blood samples: recent developments in molecular approaches. Clin Microbiol Infect 24:944–955. https://doi.org/10.1016/j.cmi.2018.05.007
Ko YJ, Kook JK, Lee CK (2021) In-house method for direct bacterial identification in positive blood culture broths using microfiltration, bead beating, and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. J Microbiol Methods 180:106065. https://doi.org/10.1016/j.mimet.2020.106065
Klein S, Zimmermann S, Köhler C, Mischnik A, Alle W, Bode KA (2012) Integration of matrix-assisted laser desorption/ionization time-of-flight mass spectrometry in blood culture diagnostics: a fast and effective approach. J Med Microbiol 61(Pt 3):323–331. https://doi.org/10.1099/jmm.0.035550-0
Morgenthaler NG, Kostrzewa M (2015) Rapid identification of pathogens in positive blood culture of patients with sepsis: review and meta-analysis of the performance of the sepsityper kit. Int J Microbiol 2015:827416. https://doi.org/10.1155/2015/827416
Saffert RT, Cunningham SA, Mandrekar J, Patel R (2012) Comparison of three preparatory methods for detection of bacteremia by MALDI-TOF mass spectrometry. Diagn Microbiol Infect Dis 73:21–26. https://doi.org/10.1016/j.diagmicrobio.2012.01.010
Magiorakos AP, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, Giske CG, Harbarth S, Hindler JF, Kahlmeter G, Olsson-Liljequist B, Paterson DL, Rice LB, Stelling J, Struelens MJ, Vatopoulos A, Weber JT, Monnet DL (2012) Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect 18:268–281. https://doi.org/10.1111/j.1469-0691.2011.03570.x
CLSI (2022) Performance standards for antimicrobial susceptibility testing, 32nd ed. CLSI supplement M100. Clinical and Laboratory Standards Institute, Wayne, PA
Wong AYW, Johnsson ATA, Özenci V (2022) Performance of dRAST on prospective clinical blood culture samples in a simulated clinical setting and on multidrug-resistant bacteria. Microbiol Spectr 10:e0210721. https://doi.org/10.1128/spectrum.02107-21
CLSI (2015) Verification of commercial microbial identification and antimicrobial susceptibility testing systems, 1st ed. CLSI Guideline M52. Clinical and Laboratory Standards Institute, Wayne, PA
U.S. Food and Drug Administration (2018) Antimicrobial susceptibility test (AST) systems - Class II special controls guidance for industry and FDA. FDA. https://www.fda.gov/medicaldevices/guidance-documents-medical-devices-and-radiation-emitting-products/antimicrobial-susceptibility-test-astsystems-class-ii-special-controls-guidance-industry-and-fda. Accessed 19 Dec 2022
Humphries RM, Ambler J, Mitchell SL, Castanheira M, Dingle T, Hindler JA, Koeth L, Sei K (2018) CLSI Methods development and standardization working group best practices for evaluation of antimicrobial susceptibility tests. J Clin Microbiol 56:e01934-e2017. https://doi.org/10.1128/JCM.01934-17
Tabak YP, Vankeepuram L, Ye G, Jeffers K, Gupta V, Murray PR (2018) Blood culture turnaround time in U.S. acute care hospitals and implications for laboratory process optimization. J Clin Microbiol 56:e00500-e518. https://doi.org/10.1128/JCM.00500-18
Banerjee R, Humphries R (2021) Rapid antimicrobial susceptibility testing methods for blood cultures and their clinical impact. Front Med (Lausanne) 8:635831. https://doi.org/10.3389/fmed.2021.635831
Timbrook TT, Morton JB, McConeghy KW, Caffrey AR, Mylonakis E, LaPlante KL (2017) The Effect of molecular rapid diagnostic testing on clinical outcomes in bloodstream infections: a systematic review and meta-analysis. Clin Infect Dis 64:15–23. https://doi.org/10.1093/cid/ciw649
Meex C, Neuville F, Descy J, Huynen P, Hayette MP, De Mol P, Melin P (2012) Direct identification of bacteria from BacT/ALERT anaerobic positive blood cultures by MALDI-TOF MS: MALDI Sepsityper kit versus an in-house saponin method for bacterial extraction. J Med Microbiol 61:1511–1516. https://doi.org/10.1099/jmm.0.044750-0
Ponderand L, Pavese P, Maubon D, Giraudon E, Girard T, Landelle C, Maurin M, Caspar Y (2020) Evaluation of Rapid Sepsityper® protocol and specific MBT-Sepsityper module (Bruker Daltonics) for the rapid diagnosis of bacteremia and fungemia by MALDI-TOF-MS. Ann Clin Microbiol Antimicrob 19:60. https://doi.org/10.1186/s12941-020-00403-w
Watanabe N, Koyama S, Taji Y, Mitsutake K, Ebihara Y (2022) Direct microorganism species identification and antimicrobial susceptibility tests from positive blood culture bottles using rapid Sepsityper Kit. J Infect Chemother 28(4):563–568. https://doi.org/10.1016/j.jiac.2021.12.030
Yonetani S, Ohnishi H, Ohkusu K, Matsumoto T, Watanabe T (2016) Direct identification of microorganisms from positive blood cultures by MALDI-TOF MS using an in-house saponin method. Int J Infect Dis 52:37–42. https://doi.org/10.1016/j.ijid.2016.09.014
Marí-Almirall M, Cosgaya C, Higgins PG, Van Assche A, Telli M, Huys G, Lievens B, Seifert H, Dijkshoorn L, Roca I, Vila J (2017) MALDI-TOF/MS identification of species from the Acinetobacter baumannii (Ab) group revisited: inclusion of the novel A. seifertii and A. dijkshoorniae species. Clin Microbiol Infect 23:210.e211-210.e219. https://doi.org/10.1016/j.cmi.2016.11.020
Rodrigues C, Passet V, Rakotondrasoa A, Brisse S (2018) Identification of Klebsiella pneumoniae, Klebsiellaquasipneumoniae, Klebsiellavariicola and related phylogroups by MALDI-TOF mass spectrometry. Front Microbiol 9:3000. https://doi.org/10.3389/fmicb.2018.03000
Pan HW, Li W, Li RG, Li Y, Zhang Y, Sun EH (2018) Simple sample preparation method for direct microbial identification and susceptibility testing from positive blood cultures. Front Microbiol 9:481. https://doi.org/10.3389/fmicb.2018.00481
van Hal SJ, Jensen SO, Vaska VL, Espedido BA, Paterson DL, Gosbell IB (2012) Predictors of mortality in Staphylococcus aureus bacteremia. Clin Microbiol Rev 25:362–386. https://doi.org/10.1128/CMR.05022-11
Brown DF, Edwards DI, Hawkey PM, Morrison D, Ridgway GL, Towner KJ, Wren MW (2005) Guidelines for the laboratory diagnosis and susceptibility testing of methicillin-resistant Staphylococcus aureus (MRSA). J Antimicrob Chemother 56:1000–1018. https://doi.org/10.1093/jac/dki372
Xie O, Slavin MA, Teh BW, Bajel A, Douglas AP, Worth LJ (2020) Epidemiology, treatment and outcomes of bloodstream infection due to vancomycin-resistant enterococci in cancer patients in a vanB endemic setting. BMC Infect Dis 20:228. https://doi.org/10.1186/s12879-020-04952-5
Keshta AS, Elamin N, Hasan MR, Pérez-López A, Roscoe D, Tang P, Suleiman M (2021) Evaluation of rapid immunochromatographic tests for the direct detection of extended spectrum beta-lactamases and carbapenemases in enterobacterales isolated from positive blood cultures. Microbiol Spectr 9:e0078521. https://doi.org/10.1128/Spectrum.00785-21
Zavascki AP, Carvalhaes CG, Picão RC, Gales AC (2010) Multidrug-resistant Pseudomonas aeruginosa and Acinetobacter baumannii: resistance mechanisms and implications for therapy. Expert Rev Anti Infect Ther 8:71–93. https://doi.org/10.1586/eri.09.108
Paterson DL (2006) The epidemiological profile of infections with multidrug-resistant Pseudomonas aeruginosa and Acinetobacter species. Clin Infect Dis 43(Suppl 2):S43-48. https://doi.org/10.1086/504476
Peleg AY, Seifert H, Paterson DL (2008) Acinetobacter baumannii: emergence of a successful pathogen. Clin Microbiol Rev 21:538–582. https://doi.org/10.1128/CMR.00058-07
Motbainor H, Bereded F, Mulu W (2020) Multi-drug resistance of blood stream, urinary tract and surgical site nosocomial infections of Acinetobacter baumannii and Pseudomonas aeruginosa among patients hospitalized at Felegehiwot referral hospital, Northwest Ethiopia: a cross-sectional study. BMC Infect Dis 20:9235. https://doi.org/10.1186/s12879-020-4811-8
Exner M, Bhattacharya S, Christiansen B, Gebel J, Goroncy-Bermes P, Hartemann P, Heeg P, Ilschner C, Kramer A, Larson E, Merkens W, Mielke M, Oltmanns P, Ross B, Rotter M, Schmithausen RM, Sonntag HG, Trautmann M (2017) Antibiotic resistance: What is so special about multidrug-resistant Gram-negative bacteria? GMS Hyg Infect Control 12:05. https://doi.org/10.3205/dgkh000290
Waites KB, Brookings ES, Moser SA, Zimmer BL (1998) Direct susceptibility testing with positive BacT/Alert blood cultures by using MicroScan overnight and rapid panels. J Clin Microbiol 36:2052–2056. https://doi.org/10.1128/JCM.36.7.2052-2056.1998
Nassar MSM, Hazzah WA, Bakr WMK (2019) Evaluation of antibiotic susceptibility test results: how guilty a laboratory could be? J Egypt Public Health Assoc 94:4. https://doi.org/10.1186/s42506-018-0006-1
Lenhard JR, Bulman ZP (2019) Inoculum effect of β-lactam antibiotics. J Antimicrob Chemother 74:2825–2843. https://doi.org/10.1093/jac/dkz226
Idelevich EA, Becker K (2019) How to accelerate antimicrobial susceptibility testing. Clin Microbiol Infect 25:1347–1355. https://doi.org/10.1016/j.cmi.2019.04.025
Smith KP, Kirby JE (2018) The inoculum effect in the era of multidrug resistance: minor differences in inoculum have dramatic effect on MIC determination. Antimicrob Agents Chemother 62:e00433-e518. https://doi.org/10.1128/AAC.00433-18
Funding
The authors declare that no funds, grants, or other support was received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to the conception and design of this study. Material preparation, data collection, and analysis were performed by all authors. The first draft of the manuscript was written by Keun Ju Kim, and all the authors commented on the previous versions of the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This study was approved by the Institutional Review Board (IRB) of the Korea University Anam Hospital (2018AN0057).
Consent to participate
A waiver of informed consent was granted by the IRB owing to the use of remnant and de-identified samples for the study.
Consent for publication
Not applicable
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kim, K.J., Yun, S.G., Cho, Y. et al. Evaluation of a sterile, filter-based, in-house method for rapid direct bacterial identification and antimicrobial susceptibility testing using positive blood culture. Eur J Clin Microbiol Infect Dis 42, 691–700 (2023). https://doi.org/10.1007/s10096-023-04592-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-023-04592-y